

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146341.8 + phase: 0
(268 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
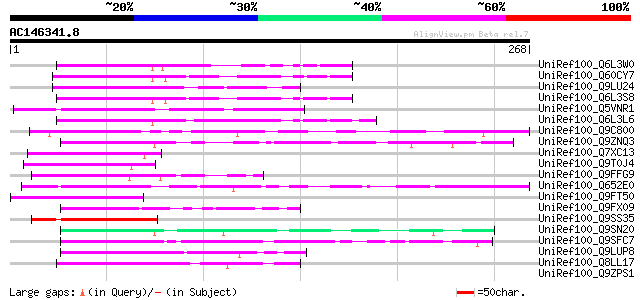
Sequences producing significant alignments: (bits) Value
UniRef100_Q6L3W0 Putative F-box protein [Solanum demissum] 58 2e-07
UniRef100_Q60CY7 Putative F-box protein [Solanum demissum] 57 5e-07
UniRef100_Q9LU24 Gb|AAF25964.1 [Arabidopsis thaliana] 57 6e-07
UniRef100_Q6L3S8 Putative F-Box protein [Solanum demissum] 55 1e-06
UniRef100_Q5VNR1 Hypothetical protein B1150E06.18-1 [Oryza sativa] 55 2e-06
UniRef100_Q6L3L6 Putative F-box-like protein [Solanum demissum] 54 3e-06
UniRef100_Q9C800 Hypothetical protein F10C21.17 [Arabidopsis tha... 52 1e-05
UniRef100_Q9ZNQ3 Hypothetical protein At2g27520 [Arabidopsis tha... 50 5e-05
UniRef100_Q7XC13 Hypothetical protein [Oryza sativa] 50 8e-05
UniRef100_Q9T0J4 Hypothetical protein AT4g38870 [Arabidopsis tha... 49 1e-04
UniRef100_Q9FFG9 Arabidopsis thaliana genomic DNA, chromosome 5,... 49 2e-04
UniRef100_Q652E0 Hypothetical protein P0463G11.10 [Oryza sativa] 48 3e-04
UniRef100_Q9FT50 Hypothetical protein T25B15_90 [Arabidopsis tha... 47 4e-04
UniRef100_Q9FX09 T3F24.1 protein [Arabidopsis thaliana] 47 5e-04
UniRef100_Q9SS35 F14P13.16 protein [Arabidopsis thaliana] 47 7e-04
UniRef100_Q9SN20 Hypothetical protein F3A4.60 [Arabidopsis thali... 46 9e-04
UniRef100_Q9SFC7 F17A17.21 protein [Arabidopsis thaliana] 46 9e-04
UniRef100_Q9LUP8 Gb|AAD25583.1 [Arabidopsis thaliana] 46 9e-04
UniRef100_Q8LL17 Suppressor of nim1-1 [Arabidopsis thaliana] 46 9e-04
UniRef100_Q9ZPS1 Hypothetical protein At2g02030 [Arabidopsis tha... 46 0.001
>UniRef100_Q6L3W0 Putative F-box protein [Solanum demissum]
Length = 427
Score = 58.2 bits (139), Expect = 2e-07
Identities = 51/172 (29%), Positives = 78/172 (44%), Gaps = 43/172 (25%)
Query: 25 MFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRPP--------RLIL 76
+ LP+ELI +IL L VK++ + +C+SKSW LIS PTFV+ +K RLI
Sbjct: 51 LLLPDELITEILLKLPVKSLSKFMCVSKSWLQLISSPTFVKNHIKLTADDKGYIHHRLIF 110
Query: 77 K-----------PPCWVYPMRTIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCKVVISC 125
+ PP + T + I ++ +++ +V S
Sbjct: 111 RNIDGNFKFCSLPPLFTKQQHTEELFHIDSPIERSTLST---------------HIVGSV 155
Query: 126 NGLLCFIFCSDNKEYHNYWFRIWNPATGTRSKALGTNHDYNLQLRSLRFNFG 177
NGL+C + KE + IWNP T T+SK L N+ S+++ FG
Sbjct: 156 NGLICVV--HGQKEAY-----IWNP-TITKSKEL-PKFTSNMCSSSIKYGFG 198
>UniRef100_Q60CY7 Putative F-box protein [Solanum demissum]
Length = 383
Score = 57.0 bits (136), Expect = 5e-07
Identities = 53/169 (31%), Positives = 77/169 (45%), Gaps = 32/169 (18%)
Query: 23 LSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRPP--------RL 74
+ + LP+ELI +IL L +K++ + +C+SKSW LIS P FV+K +K RL
Sbjct: 7 IPLLLPDELITEILLRLPIKSLSKFMCVSKSWLQLISSPAFVKKHIKLTANDKGYIYHRL 66
Query: 75 ILKPP------CWVYPMRTIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCKVVISCNGL 128
I + C + P+ T Q L I +L S + +V S NGL
Sbjct: 67 IFRNTNNDFKFCPLPPLFTNQQL-IEEILHIDSPIERTTL---------STHIVGSVNGL 116
Query: 129 LCFIFCSDNKEYHNYWFRIWNPATGTRSKALGTNHDYNLQLRSLRFNFG 177
+C + Y IWNPA T+SK L + NL ++ FG
Sbjct: 117 ICVAHVRQREAY------IWNPAI-TKSKELPKSTS-NLCSDGIKCGFG 157
>UniRef100_Q9LU24 Gb|AAF25964.1 [Arabidopsis thaliana]
Length = 360
Score = 56.6 bits (135), Expect = 6e-07
Identities = 39/128 (30%), Positives = 57/128 (44%), Gaps = 15/128 (11%)
Query: 23 LSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRPPRLILKPPCWV 82
+S FLPEEL +IL L +K + R C+ K+W LI+DP F + P +
Sbjct: 1 MSKFLPEELAIEILVRLSMKDLARFRCVCKTWRDLINDPGFTETYRDMSPAKFVSFYDKN 60
Query: 83 YPMRTIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCKVVISCNGLLCFIFCSDNKEYHN 142
+ M ++ K+P IT DF D + D V+ C+G LC N
Sbjct: 61 FYMLDVEG-------KHPVITNKLDFPLD-QSMIDESTCVLHCDGTLCVTL-------KN 105
Query: 143 YWFRIWNP 150
+ +WNP
Sbjct: 106 HTLMVWNP 113
>UniRef100_Q6L3S8 Putative F-Box protein [Solanum demissum]
Length = 327
Score = 55.5 bits (132), Expect = 1e-06
Identities = 53/167 (31%), Positives = 75/167 (44%), Gaps = 32/167 (19%)
Query: 25 MFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRPP--------RLIL 76
+ LP+ELI +IL L +K++L+ +C+SKSW LIS P FV+ +K RLI
Sbjct: 10 LLLPDELITEILLKLPIKSLLKFMCVSKSWLQLISSPAFVKNHIKLTADDKGYIYHRLIF 69
Query: 77 KPP------CWVYPMRTIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCKVVISCNGLLC 130
+ C + P+ T Q L I L S + +V S NGL+C
Sbjct: 70 RNTNDDFKFCPLPPLFTQQQL-IKELYHIDSPIERTTL---------STHIVGSVNGLIC 119
Query: 131 FIFCSDNKEYHNYWFRIWNPATGTRSKALGTNHDYNLQLRSLRFNFG 177
+ Y IWNP T T+SK L + NL ++ FG
Sbjct: 120 AAHVRQREAY------IWNP-TITKSKELPKSRS-NLCSDGIKCGFG 158
>UniRef100_Q5VNR1 Hypothetical protein B1150E06.18-1 [Oryza sativa]
Length = 433
Score = 54.7 bits (130), Expect = 2e-06
Identities = 46/151 (30%), Positives = 70/151 (45%), Gaps = 14/151 (9%)
Query: 3 YSKLMSIKKKKKDKKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPT 62
Y+ + S K + + K K T + L ++L+ ILS L +K++ C+ KSW +L SD
Sbjct: 41 YTAINSTLKGEMNSKKSKCTCT--LTDDLVVDILSRLPLKSVCCFKCVCKSWASLFSDQY 98
Query: 63 FVQKQLKRPPRLILKPPCWVYPMRTIQSLPITRLLK-NPSITVSGDFCYDGGGLNDNCKV 121
F K +RP L+ Y S+ I +L N I S F ++N K+
Sbjct: 99 FCTKLPRRPAGLL-------YQDSNNSSIQIAKLPSGNSEIGTSLSFMPH----HENLKL 147
Query: 122 VISCNGLLCFIFCSDNKEYHNYWFRIWNPAT 152
V NGL+ F S + + F + NPAT
Sbjct: 148 VDCSNGLILFTHGSKSDSPDSSHFIVCNPAT 178
>UniRef100_Q6L3L6 Putative F-box-like protein [Solanum demissum]
Length = 384
Score = 54.3 bits (129), Expect = 3e-06
Identities = 51/173 (29%), Positives = 76/173 (43%), Gaps = 23/173 (13%)
Query: 25 MFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRPP--------RLIL 76
+ LP+ELI +IL L +K++ + +C+SKSW LIS P FV+ +K RLI
Sbjct: 10 LLLPDELITEILLRLPIKSLSKFMCVSKSWLQLISSPAFVKNHIKLTANGKGYIYHRLIF 69
Query: 77 KPPCWVYPMRTIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCKVVISCNGLLCFIFCSD 136
+ + + SL K I D + +V S NGL+C
Sbjct: 70 RNTNDDFKFCPLPSL----FTKQQLIEELFDIVSPIERTTLSTHIVGSVNGLICAAHVRQ 125
Query: 137 NKEYHNYWFRIWNPATGTRSKALGTNHDYNLQLRSLRFNFGSEAFGSCYYDYR 189
+ Y IWNP T T+SK L + NL ++ FG + + DY+
Sbjct: 126 REAY------IWNP-TITKSKELPKSRS-NLCSDGIKCGFG---YDESHDDYK 167
>UniRef100_Q9C800 Hypothetical protein F10C21.17 [Arabidopsis thaliana]
Length = 441
Score = 52.4 bits (124), Expect = 1e-05
Identities = 65/274 (23%), Positives = 113/274 (40%), Gaps = 74/274 (27%)
Query: 11 KKKKDKKNK-----KLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQ 65
+KK+++ K + TL++ LP+ L+ +IL L VK ++RL +SK W +LI +
Sbjct: 76 RKKQERSGKSPSCSETTLAVELPDVLVEEILQRLPVKYLVRLKSISKGWKSLIESDHLAE 135
Query: 66 KQLKRPPRLILKPPCWVYPMRTIQSLPITRLLKNPSITVSGDFCYDGGGLN-------DN 118
K L RL+ K Y ++ I+ + ++ S ++ F G+N D
Sbjct: 136 KHL----RLLEKK----YGLKEIK----ITVERSTSKSICIKFFSRRSGMNAINSDSDDL 183
Query: 119 CKVVISCNGLLCFIFCSDNKEYHNYWFRIWNPATGTRSKALGTNHDYNLQLRSLRFNFGS 178
+V SCNGL+C E + + + NP TG R+L G+
Sbjct: 184 LRVPGSCNGLVCVY------ELDSVYIYLLNPMTGV--------------TRTLTPPRGT 223
Query: 179 EAFGSCYYDYRRRYLLGCLKFTFGCGILSGTYKTVEFRAKGDEENKYGPWRSQVRILSLS 238
+ L FG +++GTYK + YG R + L
Sbjct: 224 K-----------------LSVGFGIDVVTGTYKVMVL---------YGFDRVGTVVFDLD 257
Query: 239 DDCWR----NIDSFPVIPLISPFNNGVYLSGTIY 268
+ WR P+ + +P N V+++G+++
Sbjct: 258 TNKWRQRYKTAGPMPLSCIPTPERNPVFVNGSLF 291
>UniRef100_Q9ZNQ3 Hypothetical protein At2g27520 [Arabidopsis thaliana]
Length = 347
Score = 50.4 bits (119), Expect = 5e-05
Identities = 63/252 (25%), Positives = 103/252 (40%), Gaps = 54/252 (21%)
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRPPR---LILKPPCWVY 83
LP +L+ +ILS L ++ RL K WN L DP F+ KQ + + +++ VY
Sbjct: 6 LPWDLVDEILSRLPATSLGRLRFTCKRWNALFKDPEFITKQFHKAAKQDLVLMLSNFGVY 65
Query: 84 PMRTIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCKVVISCNGLLCFIFCSDNKEYHNY 143
M T L + P+ N V CNGLL CS +E +
Sbjct: 66 SMS-------TNLKEIPN--------------NIEIAQVFHCNGLL---LCS-TEEGNKT 100
Query: 144 WFRIWNPATG-TRSKALGTNHDYNLQLRSLRFNFGSEAFGSCYYDYRRRYLLGCLKFTFG 202
+ NP TG TR T+++YN + +G+ + Y Y+ L+ T+G
Sbjct: 101 KLVVVNPCTGQTRWIEPRTDYNYN---HDIALGYGNNSTKKSYDSYK------ILRITYG 151
Query: 203 CGIL------SGTYKTVEFRAKGDEENKYGP--------WRSQVRILSLSDDCWRNIDSF 248
C ++ S +++ + E++ YG W S + LS D ++F
Sbjct: 152 CKLVEIFELKSNSWRVLSKVHPNVEKHYYGGVSFKGNTYWLSYTKFNILSFDF--TTETF 209
Query: 249 PVIPLISPFNNG 260
+PL + +G
Sbjct: 210 RSVPLPFLYQDG 221
>UniRef100_Q7XC13 Hypothetical protein [Oryza sativa]
Length = 445
Score = 49.7 bits (117), Expect = 8e-05
Identities = 24/74 (32%), Positives = 43/74 (57%), Gaps = 5/74 (6%)
Query: 10 KKKKKDKKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQL- 68
K+K+ T LPEE++ +IL+ L VK++LR + + W +IS+P+F++ QL
Sbjct: 38 KRKRTVVPPAAATFPDLLPEEIVVEILARLPVKSLLRFKSVCRGWRAIISEPSFIRTQLQ 97
Query: 69 ----KRPPRLILKP 78
K+ P +++ P
Sbjct: 98 CSASKQEPSILISP 111
>UniRef100_Q9T0J4 Hypothetical protein AT4g38870 [Arabidopsis thaliana]
Length = 426
Score = 48.9 bits (115), Expect = 1e-04
Identities = 26/72 (36%), Positives = 41/72 (56%), Gaps = 4/72 (5%)
Query: 8 SIKKKKKDKKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDP----TF 63
S + ++ K K S LP +LI +IL L +K ++R +C+SK W ++I DP F
Sbjct: 34 STMETQRKKFTKVSVNSELLPVDLIMEILKKLSLKPLIRFLCVSKLWASIIRDPYFMKLF 93
Query: 64 VQKQLKRPPRLI 75
+ + LKRP L+
Sbjct: 94 LNESLKRPKSLV 105
>UniRef100_Q9FFG9 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MLN1
[Arabidopsis thaliana]
Length = 295
Score = 48.5 bits (114), Expect = 2e-04
Identities = 37/128 (28%), Positives = 60/128 (45%), Gaps = 24/128 (18%)
Query: 12 KKKDKKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISD----PTFVQKQ 67
+KK +++ T S LP +LI +IL L KT+ R +C+SK W+++I F+ +
Sbjct: 47 RKKVSDDRRSTNSDLLPMDLIKEILKRLPAKTLARFLCVSKLWSSIIRSRDLMKLFLTES 106
Query: 68 LKRPPRLIL----KPPCWVYPMRTIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCKVVI 123
RP RL K C+++ +S T L P+ G +NC V
Sbjct: 107 SARPGRLFFTFRRKDDCFLFSSE--ESSVATYLFTIPA-----------SGYTNNCSFV- 152
Query: 124 SCNGLLCF 131
+GL+C+
Sbjct: 153 --HGLICY 158
>UniRef100_Q652E0 Hypothetical protein P0463G11.10 [Oryza sativa]
Length = 503
Score = 47.8 bits (112), Expect = 3e-04
Identities = 66/268 (24%), Positives = 111/268 (40%), Gaps = 52/268 (19%)
Query: 7 MSIKKKKKDKKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQK 66
M+ KK + +KK+ +++ LP++++ IL L V T+LR + K W+ +I DP F
Sbjct: 1 MASKKALAMQPSKKICMTL-LPQDIVELILLRLPVSTLLRCRGVCKQWDGIIRDPQFAMV 59
Query: 67 QLKRPPRLILKPPCWVYPMRTIQSLPITRLLKNPSITVSGDFCYDGGG-----LNDNCKV 121
++R PR P+ Q + LL PS + D + + + +
Sbjct: 60 HIQRAPR---------RPLFFFQRENLVHLL-YPSEAILFDEAWSPSKWVVPVIEPDDFL 109
Query: 122 VISCNGLLCFIFCSDNKEYHNYWFRIWNPATGTRSKALGTNHDYNLQLRSLRFNFGSEAF 181
SCNGL+C SD +I N ATG + L N ++ F
Sbjct: 110 CASCNGLIC--LHSDKST-----IKIANLATGE-----------CMHLVKPVRNSKTDHF 151
Query: 182 GSCYYDYRRRYLLGCLKFTFGCGILSGTYKTVEFRAKGDEENKYGPWRSQVRILSLSDDC 241
YY +FG ++ YK + F DE G S +++ +L D+
Sbjct: 152 S--YY-------------SFGFHPVTKQYKVMHFLR--DEHLHVGTSFSIIQVYTLGDEK 194
Query: 242 WRNIDSFPVIPLISPFNNGVY-LSGTIY 268
WR++ + + L +GV + G +Y
Sbjct: 195 WRDVRTPQALSLRCVERSGVVNVGGAMY 222
>UniRef100_Q9FT50 Hypothetical protein T25B15_90 [Arabidopsis thaliana]
Length = 390
Score = 47.4 bits (111), Expect = 4e-04
Identities = 23/69 (33%), Positives = 40/69 (57%)
Query: 1 MRYSKLMSIKKKKKDKKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISD 60
M S ++++KK+K K+ K + +PEE++ IL L K+++R C+SK W +LI+
Sbjct: 1 MGSSLCVAVRKKEKQKQKKTVVFLPEIPEEMLIDILIRLPAKSLMRFKCVSKLWLSLITS 60
Query: 61 PTFVQKQLK 69
F + K
Sbjct: 61 RYFTNRFFK 69
>UniRef100_Q9FX09 T3F24.1 protein [Arabidopsis thaliana]
Length = 370
Score = 47.0 bits (110), Expect = 5e-04
Identities = 41/124 (33%), Positives = 56/124 (45%), Gaps = 20/124 (16%)
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRPPRLILKPPCWVYPMR 86
LP ELI +ILS + ++++R +SK WN L D F+ K R IL Y
Sbjct: 7 LPCELIEEILSRVPPESLVRFRTVSKKWNALFDDKMFINNH-KMTFRFILATESKFY--- 62
Query: 87 TIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCKVVISCNGLLCFIFCSDNKEYHNYWFR 146
S+ +T P I V + D GL K++I CNG F+ C KE
Sbjct: 63 ---SVSMT-----PKIEVR-ELSLDIPGLELKPKILIDCNG---FLLCGMEKE----GIV 106
Query: 147 IWNP 150
+WNP
Sbjct: 107 VWNP 110
>UniRef100_Q9SS35 F14P13.16 protein [Arabidopsis thaliana]
Length = 389
Score = 46.6 bits (109), Expect = 7e-04
Identities = 22/65 (33%), Positives = 42/65 (63%), Gaps = 2/65 (3%)
Query: 12 KKKDKKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRP 71
++K K++K + S +P +L+++IL L K++ R C+SK W+++ ++P F+ R
Sbjct: 14 QRKSKRSKSGSSS--IPLDLVSEILLRLPEKSVARFRCVSKPWSSITTEPYFINLLTTRS 71
Query: 72 PRLIL 76
PRL+L
Sbjct: 72 PRLLL 76
>UniRef100_Q9SN20 Hypothetical protein F3A4.60 [Arabidopsis thaliana]
Length = 382
Score = 46.2 bits (108), Expect = 9e-04
Identities = 53/234 (22%), Positives = 85/234 (35%), Gaps = 56/234 (23%)
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRPPR----LILKPPCWV 82
LP+EL+ +IL + ++ +L K WN L +D TF +K + P+ +L+ C
Sbjct: 5 LPQELLEEILCRVPATSLKQLRLTCKEWNRLFNDRTFSRKHFDKAPKQFLITVLEERC-- 62
Query: 83 YPMRTIQSLPITRLLKNPSITVSGDFC---YDGGGLNDNCKVVISCNGLLCFIFCSDNKE 139
+ SL I PS +G+ Y + C+GL D +
Sbjct: 63 ----RLSSLSINLHSGFPSEEFTGELSPIDYHSNSSQVIIMKIFHCDGLFVCTILKDTR- 117
Query: 140 YHNYWFRIWNPATGTRS-KALGTNHDYNLQLRSLRFNFGSEAFGSCYYDYRRRYLLGCLK 198
+WNP TG + G N D N G + Y D +
Sbjct: 118 -----IVVWNPCTGQKKWIQTGENLDEN----------GQDFVLGYYQDNKSS------- 155
Query: 199 FTFGCGILSGTYKTVEFRA--KGDEENKYGPWRSQVRILSLSDDCWRNIDSFPV 250
+YK + ++ GD+E K I + + WRN+D P+
Sbjct: 156 --------DKSYKILSYKGYNYGDQEFK---------IYDIKSNTWRNLDVTPI 192
>UniRef100_Q9SFC7 F17A17.21 protein [Arabidopsis thaliana]
Length = 417
Score = 46.2 bits (108), Expect = 9e-04
Identities = 57/229 (24%), Positives = 97/229 (41%), Gaps = 31/229 (13%)
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRPPRLILKPPCWVYPMR 86
LPE++IA I S L + +I RL+ + +SW ++++ + P + L C P+R
Sbjct: 28 LPEDIIADIFSRLPISSIARLMFVCRSWRSVLTQHGRLSSSSSSPTKPCLLLHC-DSPIR 86
Query: 87 TIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCKVVISCNGLLCFIFCSDNKEYHNYWFR 146
L L + + F VV SCNGLLC + +N
Sbjct: 87 --NGLHFLDLSEEEKRIKTKKFTLRFASSMPEFDVVGSCNGLLCL-----SDSLYNDSLY 139
Query: 147 IWNPATGTRSKALGTNHDYNLQLRSLRFNFGSEAFGSCYYDYRRRYLLGCLKFTFGCGIL 206
++NP T + ++ Y+ Q L F FG +++ + Y + LK + G
Sbjct: 140 LYNPFTTNSLELPECSNKYHDQ--ELVFGFG-------FHEMTKEYKV--LKIVYFRG-- 186
Query: 207 SGTYKTVEFRAKGDEENKYGPWRSQVRILSLSDD------CWRNIDSFP 249
S + +R +G + K +S+V+IL+LS WR++ P
Sbjct: 187 SSSNNNGIYRGRGRIQYK----QSEVQILTLSSKTTDQSLSWRSLGKAP 231
>UniRef100_Q9LUP8 Gb|AAD25583.1 [Arabidopsis thaliana]
Length = 388
Score = 46.2 bits (108), Expect = 9e-04
Identities = 38/131 (29%), Positives = 60/131 (45%), Gaps = 11/131 (8%)
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRPPRLILKPPCWVYPMR 86
L E+L+ +ILS + ++ RL K WN L +D F +K + P+ L + +
Sbjct: 6 LSEDLVEEILSRVPAISLKRLRYTCKQWNALFNDQRFSKKHRDKAPKTYLGLTLKDFRIY 65
Query: 87 TIQSLPITRLLKNPSITVSGDFCYDGGGLND----NCKVVISCNGLLCFIFCSDNKEYHN 142
++ S + LL N +I + +F LND + C+GL I CS + N
Sbjct: 66 SMSS-NLHGLLHNNNIDLLMEFKGKLSSLNDLNDFEISQIYPCDGL---ILCSTKR---N 118
Query: 143 YWFRIWNPATG 153
+WNP TG
Sbjct: 119 TRLVVWNPCTG 129
>UniRef100_Q8LL17 Suppressor of nim1-1 [Arabidopsis thaliana]
Length = 370
Score = 46.2 bits (108), Expect = 9e-04
Identities = 38/129 (29%), Positives = 59/129 (45%), Gaps = 13/129 (10%)
Query: 25 MFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRP-PRLILKPPCWVY 83
M LP EL ILS L +++R +SK WN+L +D TF+ L R P I+ +Y
Sbjct: 1 MALPWELEEDILSRLPPISLVRFRTVSKHWNSLFNDKTFINNHLSRSRPEFIILTNSKIY 60
Query: 84 PMRTIQSLPITRLLKNPSITVSGDFCYD--GGGLNDNCKVVISCNGLLCFIFCSDNKEYH 141
+ I I +P+I + YD G + N ++ +C+ L + N Y
Sbjct: 61 SVDIIDHNNI-----DPTIRLHEIPTYDIHSRGTDLNRTIINTCDEFLFY-----NYRYW 110
Query: 142 NYWFRIWNP 150
+ +WNP
Sbjct: 111 DNMTALWNP 119
>UniRef100_Q9ZPS1 Hypothetical protein At2g02030 [Arabidopsis thaliana]
Length = 334
Score = 45.8 bits (107), Expect = 0.001
Identities = 38/143 (26%), Positives = 68/143 (46%), Gaps = 10/143 (6%)
Query: 13 KKDKKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQ--LKR 70
K+ K+ +K + +P E++ +IL L VK++ R +SK W TLI+ F ++ L++
Sbjct: 25 KEIKRRRKEFEKIHIPNEIVEEILVRLPVKSLTRFQTVSKHWRTLITSKYFGKRHMALEK 84
Query: 71 PPRLILKPPCWVYPMRTIQSLPI-TRLLKNPSITVSGDFCYDGGGLNDNCKVVISCNGLL 129
L C + R +L + T L+ S++ + ++ G + SC+GL+
Sbjct: 85 SKGCKLLFVCDDFVDRAEDTLFLKTVALEKTSVSEGDEQAFEFEGYKGFLDISESCDGLV 144
Query: 130 CFIFCSDNKEYHNYWFRIWNPAT 152
CF + E + NPAT
Sbjct: 145 CFYDTTRAVE-------VMNPAT 160
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.324 0.142 0.455
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 490,962,109
Number of Sequences: 2790947
Number of extensions: 21038521
Number of successful extensions: 45982
Number of sequences better than 10.0: 283
Number of HSP's better than 10.0 without gapping: 236
Number of HSP's successfully gapped in prelim test: 47
Number of HSP's that attempted gapping in prelim test: 45699
Number of HSP's gapped (non-prelim): 307
length of query: 268
length of database: 848,049,833
effective HSP length: 125
effective length of query: 143
effective length of database: 499,181,458
effective search space: 71382948494
effective search space used: 71382948494
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 74 (33.1 bits)
Medicago: description of AC146341.8


