

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146329.8 + phase: 0
(402 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
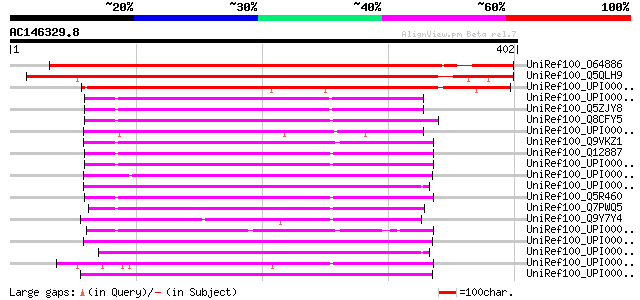
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O64886 Putative heme A:farnesyltransferase [Arabidopsi... 482 e-135
UniRef100_Q5QLH9 Putative heme A:farnesyltransferase [Oryza sativa] 464 e-129
UniRef100_UPI0000259A1C UPI0000259A1C UniRef100 entry 274 4e-72
UniRef100_UPI00003AB46C UPI00003AB46C UniRef100 entry 200 7e-50
UniRef100_Q5ZJY8 Hypothetical protein [Gallus gallus] 200 7e-50
UniRef100_Q8CFY5 COX10 homolog, cytochrome c oxidase assembly pr... 200 7e-50
UniRef100_UPI00003C25B8 UPI00003C25B8 UniRef100 entry 197 6e-49
UniRef100_Q9VKZ1 CG5037-PA [Drosophila melanogaster] 196 1e-48
UniRef100_Q12887 Protoheme IX farnesyltransferase, mitochondrial... 194 3e-48
UniRef100_UPI000036B006 UPI000036B006 UniRef100 entry 193 8e-48
UniRef100_UPI00002CA06A UPI00002CA06A UniRef100 entry 192 1e-47
UniRef100_UPI000028260F UPI000028260F UniRef100 entry 192 2e-47
UniRef100_Q5R460 Hypothetical protein DKFZp459I052 [Pongo pygmaeus] 190 5e-47
UniRef100_Q7PWQ5 ENSANGP00000013941 [Anopheles gambiae str. PEST] 187 3e-46
UniRef100_Q9Y7Y4 SPBC365.02c protein [Schizosaccharomyces pombe] 187 5e-46
UniRef100_UPI0000433F63 UPI0000433F63 UniRef100 entry 186 8e-46
UniRef100_UPI0000310A37 UPI0000310A37 UniRef100 entry 186 8e-46
UniRef100_UPI00002AFF7B UPI00002AFF7B UniRef100 entry 184 4e-45
UniRef100_UPI00002362EA UPI00002362EA UniRef100 entry 182 1e-44
UniRef100_UPI0000274407 UPI0000274407 UniRef100 entry 181 3e-44
>UniRef100_O64886 Putative heme A:farnesyltransferase [Arabidopsis thaliana]
Length = 431
Score = 482 bits (1240), Expect = e-135
Identities = 242/370 (65%), Positives = 291/370 (78%), Gaps = 14/370 (3%)
Query: 32 STFSSSPSVSASKSNDLISLARHYGNCYSQLSKARLSLLVVATSGTGFVLGSGGA-VDLS 90
+T +++ +S+ L L HY CY +LSKA+LS+LVVATSGTG++LG+G A +
Sbjct: 71 ATATATTGEISSRVAALAGLGHHYARCYWELSKAKLSMLVVATSGTGYILGTGNAAISFP 130
Query: 91 MLSYTCLGTMMVAASASTLNQVFEIKNDAIMRRTSQRPLPSGRITVPHAVGWASSVGLAG 150
L YTC GTMM+AASA++LNQ+FEI ND+ M+RT RPLPSGRI+VPHAV WA+ G +G
Sbjct: 131 GLCYTCAGTMMIAASANSLNQIFEISNDSKMKRTMLRPLPSGRISVPHAVAWATIAGASG 190
Query: 151 TALLATQTNMLAAGLAASNLVLYSFVYTPLKQIHPVNTWVGAVVGAIPPLLGWAAASGDI 210
LLA++TNMLAAGLA++NLVLY+FVYTPLKQ+HP+NTWVGAVVGAIPPLLGWAAASG I
Sbjct: 191 ACLLASKTNMLAAGLASANLVLYAFVYTPLKQLHPINTWVGAVVGAIPPLLGWAAASGQI 250
Query: 211 SLNSLILPAALYFWQIPHFMALAYMCREDYAAGGFKMYSLADPSGRKTAMVAMRNSIYLI 270
S NS+ILPAALYFWQIPHFMALA++CR DYAAGG+KM SL DPSG++ A VA+RN Y+I
Sbjct: 251 SYNSMILPAALYFWQIPHFMALAHLCRNDYAAGGYKMLSLFDPSGKRIAAVALRNCFYMI 310
Query: 271 PLGFLAYDWGVTSGWFCLESTALTLAISAAAFSFYRDRTKERARRMFHASLLYLPVFMAG 330
PLGF+AYDWG+TS WFCLEST LTLAI+A AFSFYRDRT +AR+MFHASLL+LPVFM+G
Sbjct: 311 PLGFIAYDWGLTSSWFCLESTLLTLAIAATAFSFYRDRTMHKARKMFHASLLFLPVFMSG 370
Query: 331 LLIHR-RTDNQHFLEVNAEGFMKSPSSVETSEMDDKNGNQKIKGRQGTRARPPVAYASIA 389
LL+HR DNQ L V G S S G K + R+ A+PPVAYAS A
Sbjct: 371 LLLHRVSNDNQQQL-VEEAGLTNSVS-----------GEVKTQRRKKRVAQPPVAYASAA 418
Query: 390 PFPFLPAPSY 399
PFPFLPAPS+
Sbjct: 419 PFPFLPAPSF 428
>UniRef100_Q5QLH9 Putative heme A:farnesyltransferase [Oryza sativa]
Length = 435
Score = 464 bits (1195), Expect = e-129
Identities = 240/401 (59%), Positives = 299/401 (73%), Gaps = 26/401 (6%)
Query: 14 RLVNSSSSSSLHNFTHIRSTFSSSPSVSASKSNDLISLA---RHYGNCYSQLSKARLSLL 70
R +SSSSSS + + + +A+ S S++ RHYG CY +LSKARLS L
Sbjct: 38 RSSSSSSSSSAPTIAAAVPAVAEATAAAAAVSRQAGSVSDALRHYGRCYFELSKARLSAL 97
Query: 71 VVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDAIMRRTSQRPLP 130
VVATSG G+VLGSG VD++ L TC GTMMVAASA+TLNQVFEIKNDA M+RT +RPLP
Sbjct: 98 VVATSGAGYVLGSGNMVDIAGLCCTCAGTMMVAASANTLNQVFEIKNDAKMKRTMRRPLP 157
Query: 131 SGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTPLKQIHPVNTWV 190
SGRI+ HA WA+SVG+AGTALLA + N LAAGLAASNL+LY+FVYTPLKQIHPVNTWV
Sbjct: 158 SGRISPAHAAMWATSVGVAGTALLAWKANGLAAGLAASNLILYAFVYTPLKQIHPVNTWV 217
Query: 191 GAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCREDYAAGGFKMYSL 250
GAVVGAIPPLLGWAAAS ++SLN++ILPAALY+WQIPHFMALAY+CR DY AGG++M+S
Sbjct: 218 GAVVGAIPPLLGWAAASSELSLNAMILPAALYYWQIPHFMALAYLCRNDYLAGGYRMFSF 277
Query: 251 ADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISAAAFSFYRDRTK 310
ADP+G++TA V++RN +Y++PLGF AY+WG+TS WF LE++ LTL ++ A SF + T
Sbjct: 278 ADPTGKRTAWVSLRNCLYMLPLGFFAYNWGLTSEWFSLEASLLTLGLTIGALSFVLEPTP 337
Query: 311 ERARRMFHASLLYLPVFMAGLLIHRRTDNQHFLEVNAEGFMKSPSSVETSEM-------- 362
+ ARRMF+ SLLYLP FMAGLL+HR + Q K + +TSE+
Sbjct: 338 KTARRMFYGSLLYLPAFMAGLLLHRLPNEQ-----------KEHNVTQTSEITGILYGAE 386
Query: 363 --DDKNGNQKIKGRQGTR--ARPPVAYASIAPFPFLPAPSY 399
D++ QK + R+ +R +RPPVAYAS+APFPFLP P Y
Sbjct: 387 QQDEERARQKREDRKPSRIHSRPPVAYASVAPFPFLPVPIY 427
>UniRef100_UPI0000259A1C UPI0000259A1C UniRef100 entry
Length = 454
Score = 274 bits (700), Expect = 4e-72
Identities = 158/351 (45%), Positives = 213/351 (60%), Gaps = 15/351 (4%)
Query: 58 CYSQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKN 117
CY LSK RLS VV+T+ GF LGS G +DL + Y C GTMM +ASA+TLNQV E
Sbjct: 86 CYG-LSKFRLSAFVVSTTAAGFALGSPGNIDLERMFYACAGTMMCSASANTLNQVIERVP 144
Query: 118 DAIMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVY 177
D +MRRT+ RP+PSGR + A+ +A + + G LA TN A L A N+ LY+ Y
Sbjct: 145 DGLMRRTAGRPMPSGRCGMGFALTFAGACAVGGVGTLAVTTNETTAALGAGNIALYAGAY 204
Query: 178 TPLKQIHPVNTWVGAVVGAIPPLLGWAAA--SGDISLNSLILPAALYFWQIPHFMALAYM 235
T LK++H +NTWVGA+VGAIPPL+GWAAA +G + + +L AALY WQ+PHFMALAYM
Sbjct: 205 TALKRVHFLNTWVGAIVGAIPPLMGWAAANETGALDPGAYVLGAALYLWQMPHFMALAYM 264
Query: 236 CREDYAAGGFKMYS--LADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTAL 293
++DY GG++M S + D +GR+ A VA+RNS+ L+PLG A GVT+ F E+ AL
Sbjct: 265 GKDDYFNGGYRMLSHPMYDKTGRRVAGVALRNSLALLPLGLAACAMGVTTTPFAYETLAL 324
Query: 294 TLAISAAAFSFYRDRTKERARRMFHASLLYLPVFMAGLLIHRRTDNQHFLEVNA-EGFMK 352
+ A +FY + ARRMF SLLYLP F A +HR ++ VNA E +
Sbjct: 325 GAPLVGTAAAFYASPSFPAARRMFFGSLLYLPAFQALACLHRIPRDE---SVNAYESLRR 381
Query: 353 SPSSVETSEMDDKNGN------QKIKGRQGTRARPPVAYASIAPFPFLPAP 397
+ + GN ++++ + P ++ S+APFP LP P
Sbjct: 382 VSVNFPWHCVPFAAGNDWTPERERLRQFRSRALGPALSETSVAPFPLLPLP 432
>UniRef100_UPI00003AB46C UPI00003AB46C UniRef100 entry
Length = 424
Score = 200 bits (508), Expect = 7e-50
Identities = 112/269 (41%), Positives = 162/269 (59%), Gaps = 3/269 (1%)
Query: 60 SQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDA 119
++LSK +L+ LVV+T+ GF + DL LGT + + +A+++NQ FE+ D+
Sbjct: 138 ARLSKIKLTALVVSTASAGFAMAPV-PFDLPCFLLASLGTGLASCAANSINQFFEVPFDS 196
Query: 120 IMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTP 179
M RT RPL G+I+ AV +A+S G+ G ALL N L L A N+ LY+ YTP
Sbjct: 197 NMNRTKNRPLVRGQISPLLAVCFAASCGVPGIALLTMGVNPLTGALGAFNIFLYTCCYTP 256
Query: 180 LKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCRED 239
+K++ NTWVGAVVGAIPP++GW AA+G + +L+L LY WQ PHF AL++ RED
Sbjct: 257 MKRMSIANTWVGAVVGAIPPVMGWTAATGSLDAGALLLGGILYSWQFPHFNALSWGLRED 316
Query: 240 YAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISA 299
Y+ GG+ M S+ P VA+R+ + LI L +A +T+ F + S L L IS
Sbjct: 317 YSRGGYCMMSVTHP--ELCRRVALRHCVALIGLCTVAPILDITTWTFPIISLPLNLYISY 374
Query: 300 AAFSFYRDRTKERARRMFHASLLYLPVFM 328
F FYRD + +R++F SL +LP+ +
Sbjct: 375 LGFRFYRDADRSSSRKLFFCSLWHLPLLL 403
>UniRef100_Q5ZJY8 Hypothetical protein [Gallus gallus]
Length = 448
Score = 200 bits (508), Expect = 7e-50
Identities = 112/269 (41%), Positives = 162/269 (59%), Gaps = 3/269 (1%)
Query: 60 SQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDA 119
++LSK +L+ LVV+T+ GF + DL LGT + + +A+++NQ FE+ D+
Sbjct: 150 ARLSKIKLTALVVSTASAGFAMAPV-PFDLPCFLLASLGTGLASCAANSINQFFEVPFDS 208
Query: 120 IMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTP 179
M RT RPL G+I+ AV +A+S G+ G ALL N L L A N+ LY+ YTP
Sbjct: 209 NMNRTKNRPLVRGQISPLLAVCFAASCGVPGIALLTMGVNPLTGALGAFNIFLYTCCYTP 268
Query: 180 LKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCRED 239
+K++ NTWVGAVVGAIPP++GW AA+G + +L+L LY WQ PHF AL++ RED
Sbjct: 269 MKRMSIANTWVGAVVGAIPPVMGWTAATGSLDAGALLLGGILYSWQFPHFNALSWGLRED 328
Query: 240 YAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISA 299
Y+ GG+ M S+ P VA+R+ + LI L +A +T+ F + S L L IS
Sbjct: 329 YSRGGYCMMSVTHP--ELCRRVALRHCVALIGLCTVAPILDITTWTFPIISLPLNLYISY 386
Query: 300 AAFSFYRDRTKERARRMFHASLLYLPVFM 328
F FYRD + +R++F SL +LP+ +
Sbjct: 387 LGFRFYRDADRSSSRKLFFCSLWHLPLLL 415
>UniRef100_Q8CFY5 COX10 homolog, cytochrome c oxidase assembly protein, heme A:
farnesyltransferase [Mus musculus]
Length = 443
Score = 200 bits (508), Expect = 7e-50
Identities = 113/281 (40%), Positives = 165/281 (58%), Gaps = 3/281 (1%)
Query: 60 SQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDA 119
++LSK +L+ LVV+T+ GF L G D S T LGT + + +A+++NQ FE+ D+
Sbjct: 157 ARLSKIKLTALVVSTTSAGFALAPG-PFDWSCFLLTSLGTGLASCAANSINQFFEVPFDS 215
Query: 120 IMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTP 179
M RT RPL G+I+ AV +A+ + G ALL N L L N+ LY+ YTP
Sbjct: 216 NMNRTKNRPLVRGQISPLLAVSFATCCAVPGVALLTWGVNPLTGALGVFNIFLYTCCYTP 275
Query: 180 LKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCRED 239
LK++ NTWVGAVVGAIPP++GW AA+G + +L+L LY WQ PHF AL++ RED
Sbjct: 276 LKRVSITNTWVGAVVGAIPPVMGWTAATGSLDAGALLLGGILYSWQFPHFNALSWGLRED 335
Query: 240 YAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISA 299
Y+ GG+ M S+ P+ VA+R+ + LI L A +T+ F + S + L IS
Sbjct: 336 YSRGGYCMMSVTHPA--LCRRVALRHCLALIALSTAAPVLDITTWVFPVISLPINLYISY 393
Query: 300 AAFSFYRDRTKERARRMFHASLLYLPVFMAGLLIHRRTDNQ 340
F FY D + +R++F SL +LP+ + +L ++ Q
Sbjct: 394 LGFRFYVDADRRSSRKLFFCSLWHLPLLLLLMLTCKQRPGQ 434
>UniRef100_UPI00003C25B8 UPI00003C25B8 UniRef100 entry
Length = 1527
Score = 197 bits (500), Expect = 6e-49
Identities = 117/281 (41%), Positives = 168/281 (59%), Gaps = 13/281 (4%)
Query: 59 YSQLSKARLSLLVVATSGTGFVLGSGG-----AVDLSMLSYTCLGTMMVAASASTLNQVF 113
Y QLSK+RL+ LVV T G+ L A ++ L G + +A+A+ LNQ+
Sbjct: 1216 YKQLSKSRLTFLVVLTGMAGYALCPASLTVAVASPVTTLLALTAGMTLCSAAANALNQLV 1275
Query: 114 EIKNDAIMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLY 173
E DA M+RT RPLPS +T HA +AS +G LL T N L A L A+N+VLY
Sbjct: 1276 ESPYDAQMQRTRARPLPSRSVTPLHAFTFASVSAASGVGLLLTTVNPLTALLGAANVVLY 1335
Query: 174 SFVYTPLKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLI----LPAALYFWQIPHF 229
SF YTP+K+ NTWVGAVVGA+PPL+GWAA +G + ++ + L A L+ WQ PHF
Sbjct: 1336 SFTYTPMKRSTIANTWVGAVVGALPPLMGWAACTGTLHTSTDVGGWSLAALLFAWQFPHF 1395
Query: 230 MALAYMCREDYAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWG--VTSGWFC 287
+LA+ R +YA GG++M ++ DP+ + V++R ++ L+P+ L V+ +
Sbjct: 1396 NSLAHTLRAEYARGGYRMMAVTDPALNR--RVSLRYAVALLPICTLMIPLSGIVSPVAYA 1453
Query: 288 LESTALTLAISAAAFSFYRDRTKERARRMFHASLLYLPVFM 328
+ ST L L ++ AA+ FYR+RT + AR F SL++LP M
Sbjct: 1454 VLSTPLNLLMAHAAWKFYRERTVKSARWCFWVSLIHLPAVM 1494
>UniRef100_Q9VKZ1 CG5037-PA [Drosophila melanogaster]
Length = 391
Score = 196 bits (498), Expect = 1e-48
Identities = 108/279 (38%), Positives = 167/279 (59%), Gaps = 5/279 (1%)
Query: 59 YSQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKND 118
Y +LSK RL+ LVV T+ G+ + A D + + LGT +V+A+A+ +NQ E+ D
Sbjct: 76 YKKLSKFRLTSLVVITTMGGYAMAPA-AFDPTTFAMCTLGTGLVSAAANAINQYHEVPFD 134
Query: 119 AIMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYT 178
+ M RT R L +G++T HAV +A+ AG ++L N L A L A NL LY+ +YT
Sbjct: 135 SQMSRTKNRVLVTGQMTPLHAVTFAAVSATAGLSMLYFGVNGLTAALGAGNLFLYTTIYT 194
Query: 179 PLKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCRE 238
P+K+I VNTWVG++VGAIPPL+GWA + + ++IL LY WQ PHF AL++ R
Sbjct: 195 PMKRISIVNTWVGSIVGAIPPLMGWAGCAATLDAGAMILAGVLYAWQFPHFNALSWNLRP 254
Query: 239 DYAAGGFKMYSLADPS-GRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAI 297
DY+ G++M ++ +P R+T A+R S+ ++ L +A VT+ WF LE+ L
Sbjct: 255 DYSRAGYRMMAVTNPGLCRRT---ALRYSVAIVGLSAMAPVLDVTNYWFALETLPLNAYF 311
Query: 298 SAAAFSFYRDRTKERARRMFHASLLYLPVFMAGLLIHRR 336
+ A+ F+ +R++F SL++LP M L +++
Sbjct: 312 AYLAYKFHEKSDSGSSRKLFRFSLIHLPALMLLFLTNKK 350
>UniRef100_Q12887 Protoheme IX farnesyltransferase, mitochondrial precursor [Homo
sapiens]
Length = 443
Score = 194 bits (494), Expect = 3e-48
Identities = 109/277 (39%), Positives = 160/277 (57%), Gaps = 3/277 (1%)
Query: 60 SQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDA 119
+QLSK +L+ LVV+T+ GF L G D T +GT + + +A+++NQ FE+ D+
Sbjct: 158 AQLSKIKLTALVVSTTAAGFALAPG-PFDWPCFLLTSVGTGLASCAANSINQFFEVPFDS 216
Query: 120 IMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTP 179
M RT RPL G+I+ AV +A+ + G A+L N L L N+ LY+ YTP
Sbjct: 217 NMNRTKNRPLVRGQISPLLAVSFATCCAVPGVAILTLGVNPLTGALGLFNIFLYTCCYTP 276
Query: 180 LKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCRED 239
LK+I NTWVGAVVGAIPP++GW AA+G + + +L LY WQ PHF AL++ RED
Sbjct: 277 LKRISIANTWVGAVVGAIPPVMGWTAATGSLDAGAFLLGGILYSWQFPHFNALSWGLRED 336
Query: 240 YAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISA 299
Y+ GG+ M S+ P VA+R+ + L+ L A +T+ F + + + IS
Sbjct: 337 YSRGGYCMMSVTHPG--LCRRVALRHCLALLVLSAAAPVLDITTWTFPIMALPINAYISY 394
Query: 300 AAFSFYRDRTKERARRMFHASLLYLPVFMAGLLIHRR 336
F FY D + +RR+F SL +LP+ + +L +R
Sbjct: 395 LGFRFYVDADRRSSRRLFFCSLWHLPLLLLLMLTCKR 431
>UniRef100_UPI000036B006 UPI000036B006 UniRef100 entry
Length = 443
Score = 193 bits (490), Expect = 8e-48
Identities = 108/277 (38%), Positives = 160/277 (56%), Gaps = 3/277 (1%)
Query: 60 SQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDA 119
++LSK +L+ LVV+T+ GF L G D T +GT + + +A+++NQ FE+ D+
Sbjct: 158 ARLSKIKLTALVVSTTAAGFALAPG-PFDWPCFLLTSVGTGLASCAANSINQFFEVPFDS 216
Query: 120 IMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTP 179
M RT RPL G+I+ AV +A+ + G A+L N L L N+ LY+ YTP
Sbjct: 217 NMNRTKNRPLVRGQISPLLAVSFATCCAVPGVAILTLGVNPLTGALGLFNIFLYTCCYTP 276
Query: 180 LKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCRED 239
LK+I NTWVGAVVGAIPP++GW AA+G + + +L LY WQ PHF AL++ RED
Sbjct: 277 LKRISIANTWVGAVVGAIPPVMGWTAATGSLDAGAFLLGGILYSWQFPHFNALSWGLRED 336
Query: 240 YAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISA 299
Y+ GG+ M S+ P VA+R+ + L+ L A +T+ F + + + IS
Sbjct: 337 YSRGGYCMMSVTHPG--LCRRVALRHCLALLVLSAAAPVLDITTWTFPIMALPINAYISY 394
Query: 300 AAFSFYRDRTKERARRMFHASLLYLPVFMAGLLIHRR 336
F FY D + +RR+F SL +LP+ + +L +R
Sbjct: 395 LGFRFYVDADRRSSRRLFFCSLWHLPLLLLLMLTCKR 431
>UniRef100_UPI00002CA06A UPI00002CA06A UniRef100 entry
Length = 296
Score = 192 bits (488), Expect = 1e-47
Identities = 105/277 (37%), Positives = 157/277 (55%), Gaps = 1/277 (0%)
Query: 59 YSQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKND 118
Y L+K RL +V+ ++ G++L +G ++ ++ GT + A A LNQ E D
Sbjct: 17 YFALTKPRLLFIVLMSTLIGYLLPAGSTINFTLFQLI-FGTALTGAGAHVLNQWQERLQD 75
Query: 119 AIMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYT 178
A M RT QRPLPSGR+ A+ + + + G + LA N++ A L A L Y F+YT
Sbjct: 76 ARMMRTKQRPLPSGRLEPEEALIFGIILSILGVSYLALTINLITAVLGALTLGSYIFIYT 135
Query: 179 PLKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCRE 238
PLK++ +NTW GAV G++PPL+GWAA +G + N + + LYFWQ+PHF A+A+M R+
Sbjct: 136 PLKRLTWMNTWFGAVTGSLPPLMGWAAQTGQLDWNCVPIFLLLYFWQLPHFFAIAWMYRD 195
Query: 239 DYAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAIS 298
DY GGF+M S D G +TA + N I L+ Y +F + S +
Sbjct: 196 DYRLGGFRMLSRDDQHGSRTAFHMLVNGILLLISSLTFYFVDQGGFFFLIVSIISGIGFL 255
Query: 299 AAAFSFYRDRTKERARRMFHASLLYLPVFMAGLLIHR 335
+ F ++R+ ERAR +F S++YLPV ++I R
Sbjct: 256 TSIVGFMKERSVERARMVFLVSIVYLPVLCTVMVIDR 292
>UniRef100_UPI000028260F UPI000028260F UniRef100 entry
Length = 301
Score = 192 bits (487), Expect = 2e-47
Identities = 108/275 (39%), Positives = 157/275 (56%), Gaps = 1/275 (0%)
Query: 59 YSQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKND 118
Y LSK RL VV ++ GFVL G + + L + GT ++ A A+TLNQ EI D
Sbjct: 22 YLSLSKPRLLATVVLSALLGFVLPMGLSNTIFPLLFLLFGTALMGAGANTLNQCMEIGPD 81
Query: 119 AIMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYT 178
+ M RT R LP G+++ A+ + + +G +L N L A L L+ Y VYT
Sbjct: 82 SRMYRTKNRALPMGKLSEKEAIIFGILISASGFFVLWFGLNFLTAVLGLLTLLSYIMVYT 141
Query: 179 PLKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCRE 238
PLKQ NTW+G + GA+PP++GW AA G + L + A LYFWQ+PHF A+A+M R+
Sbjct: 142 PLKQKTATNTWLGGITGALPPVMGWVAARGQLEWEVLPIFALLYFWQLPHFFAIAWMYRD 201
Query: 239 DYAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAIS 298
DY GGFKM S D SG++TAM + N L + S ++ ++AL L
Sbjct: 202 DYKRGGFKMLSHYDKSGKRTAMQMLINCGMLYAASLAVFVVNEGSIFYLAGASALILVFF 261
Query: 299 AAAFSFYRDRTKERARRMFHASLLYLPVFMAGLLI 333
F+++R+ E AR++F AS++YLP+ + G+L+
Sbjct: 262 GVIVLFHKERSVENARKVFLASIIYLPL-LIGILV 295
>UniRef100_Q5R460 Hypothetical protein DKFZp459I052 [Pongo pygmaeus]
Length = 443
Score = 190 bits (483), Expect = 5e-47
Identities = 107/277 (38%), Positives = 159/277 (56%), Gaps = 3/277 (1%)
Query: 60 SQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDA 119
++LSK +L+ LVV+T+ GF L G D T +GT + + +A+++NQ FE+ D+
Sbjct: 158 ARLSKIKLTALVVSTTAAGFALAPG-PFDWPCFLLTSVGTGLASCAANSINQFFEVPFDS 216
Query: 120 IMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTP 179
M RT RPL G+I+ AV +A+ + G A+L N L L N+ LY+ YTP
Sbjct: 217 NMNRTKNRPLVRGQISPLLAVSFATCCAVPGVAILTLGVNPLTGALGLFNIFLYTCCYTP 276
Query: 180 LKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCRED 239
LK+I NTWVGAVVGAIPP++GW AA+G + + +L LY WQ PHF AL++ RE
Sbjct: 277 LKRISIANTWVGAVVGAIPPVMGWTAATGSLDAGAFLLGGILYSWQFPHFNALSWGLREG 336
Query: 240 YAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISA 299
Y+ GG+ M S+ P VA+R+ + L+ L A +T+ F + + + IS
Sbjct: 337 YSRGGYCMMSVTHPG--LCRRVALRHCLALLVLSAAAPVLDITTWTFPIMALPINAYISY 394
Query: 300 AAFSFYRDRTKERARRMFHASLLYLPVFMAGLLIHRR 336
F FY D + +RR+F SL +LP+ + +L +R
Sbjct: 395 LGFRFYVDADRRSSRRLFFCSLWHLPLLLLLMLTCKR 431
>UniRef100_Q7PWQ5 ENSANGP00000013941 [Anopheles gambiae str. PEST]
Length = 271
Score = 187 bits (476), Expect = 3e-46
Identities = 104/268 (38%), Positives = 158/268 (58%), Gaps = 4/268 (1%)
Query: 63 SKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDAIMR 122
S A + LVV T+ G+ + G + +L GT +V+ +A+++NQV E DA M
Sbjct: 4 SLAPFAALVVMTAMAGYAMAPG-SFELGTFLLCSAGTTLVSGAANSINQVIETSFDAQMP 62
Query: 123 RTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTPLKQ 182
RT R L G ++ HAVG+A G ++L N + A L A+NL+LY+ +YTP+K+
Sbjct: 63 RTRNRVLVKGHLSRLHAVGFALGASTIGCSMLYYGVNEMTAALGAANLILYTSIYTPMKR 122
Query: 183 IHPVNTWVGAVVGAIPPLLGWAAAS-GDISLNSLILPAALYFWQIPHFMALAYMCREDYA 241
+NTWVG+VVGAIPPL+GWAA + GD+ + IL LY WQ PHF AL++ R +Y
Sbjct: 123 YSILNTWVGSVVGAIPPLMGWAACTGGDLGAGAWILAGLLYCWQFPHFNALSWNIRPEYL 182
Query: 242 AGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISAAA 301
G+KM + + P+ V++R++ ++ L LA VT+ WF LES L + A
Sbjct: 183 KAGYKMMANSHPA--LCTRVSLRHTGFIAGLSLLAPALDVTNVWFALESLPLNAYFAYLA 240
Query: 302 FSFYRDRTKERARRMFHASLLYLPVFMA 329
+ F++ + +R++F SLL+LP+ MA
Sbjct: 241 YDFHQKADSKSSRKLFRFSLLHLPLLMA 268
>UniRef100_Q9Y7Y4 SPBC365.02c protein [Schizosaccharomyces pombe]
Length = 387
Score = 187 bits (475), Expect = 5e-46
Identities = 102/273 (37%), Positives = 161/273 (58%), Gaps = 6/273 (2%)
Query: 57 NCYSQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIK 116
+ + +L K RL++LVV ++ + + L + + L++ +GT + + SA+ NQ E
Sbjct: 85 SAFLELGKPRLTVLVVLSTMSSYALAPYPGLSFNTLAWLTMGTALCSISANAFNQSMEPM 144
Query: 117 NDAIMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFV 176
D M RT RP+P G I +A +A+ G+AGT++ + N L N+VLY +
Sbjct: 145 LDCQMARTRSRPIPRGAIRPEYAWLFATLTGIAGTSM-SFLVNPTVGWLGLGNIVLYMGI 203
Query: 177 YTPLKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLN---SLILPAALYFWQIPHFMALA 233
YTPLK+I VNTWVG++VGAIPPL+GWAA SG L+ LI A L+ WQ PHF A +
Sbjct: 204 YTPLKRISIVNTWVGSLVGAIPPLMGWAACSGGDLLSHPGGLITAAMLFAWQFPHFNAFS 263
Query: 234 YMCREDYAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTAL 293
M ++DY G++M + +P+ A V++R ++ +PL + G+ W+ + +T
Sbjct: 264 TMVKDDYKKCGYQMMAWKNPA--LNARVSLRYALAFLPLSYAYISTGLVGPWYAVPATGT 321
Query: 294 TLAISAAAFSFYRDRTKERARRMFHASLLYLPV 326
+ + A A+ FYR+R + AR +F ASLL+LP+
Sbjct: 322 NMFLIARAWKFYRNRNYQNARSLFFASLLHLPL 354
>UniRef100_UPI0000433F63 UPI0000433F63 UniRef100 entry
Length = 279
Score = 186 bits (473), Expect = 8e-46
Identities = 113/277 (40%), Positives = 168/277 (59%), Gaps = 17/277 (6%)
Query: 62 LSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDAIM 121
LSK RL+ LVV T+ G+VL G D +GT +++A+A+ +NQ FE+ DA M
Sbjct: 9 LSKIRLTSLVVITTMAGYVLAPG-VFDAHTFIACSMGTGLLSATANAINQFFEVPFDAQM 67
Query: 122 RRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTPLK 181
RT R L G +T A+ +A+ G +G LL +Q N L A L ASNL+LY+ +YTP+K
Sbjct: 68 SRTKNRVLVRGHLTPGQAIIFATVSGFSGLLLLYSQVNELTAILGASNLILYTLIYTPMK 127
Query: 182 QIHPVNTWVGAVVGAIPPLLGWAAASGDI-SLNSLILPAALYFWQIPHFMALAYMCREDY 240
+I +NTW +VGAIPPL+GWA+ DI S + I+ LY WQ PHF AL++ R DY
Sbjct: 128 RISILNTW---IVGAIPPLMGWASCVNDIVSPGAWIMSGILYAWQFPHFNALSWNLRPDY 184
Query: 241 AAGGFKMYSLADPS-GRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISA 299
+ G++M ++ +P RKT A+R + L L +LA + VT+ WF L ST L +
Sbjct: 185 SRAGYRMMAVTNPKLCRKT---ALRYTAALTGLCYLAPVFNVTNWWFALTSTPLNV---- 237
Query: 300 AAFSFYRDRTKERARRMFHASLLYLPVFMAGLLIHRR 336
+ Y D + +R++FH SL++LPV + +L++++
Sbjct: 238 --YFLYLD--SKSSRKLFHFSLIHLPVLLILILVNKK 270
>UniRef100_UPI0000310A37 UPI0000310A37 UniRef100 entry
Length = 293
Score = 186 bits (473), Expect = 8e-46
Identities = 99/277 (35%), Positives = 159/277 (56%)
Query: 59 YSQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKND 118
Y +L+K ++ +++ ++ G+ LGS +DL +T LG+ +V++ A LN E++ D
Sbjct: 15 YFELTKPSITFMILISTALGYYLGSDDIIDLKRFFFTLLGSGLVSSGAGALNHFAEMETD 74
Query: 119 AIMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYT 178
+M RTSQRP+P+G IT HA + + L G LL + L A LA +LY F+YT
Sbjct: 75 RLMTRTSQRPIPTGLITPDHARVFGITFVLIGALLLYVFIDPLTALLALITALLYIFIYT 134
Query: 179 PLKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCRE 238
PLK++ +NT +GA+ G+IPPL GW AASG + + +L A L+FWQ PHF A+A M +E
Sbjct: 135 PLKKMTWLNTSIGAIPGSIPPLGGWVAASGTLEPEAWVLFAILFFWQHPHFYAIALMFKE 194
Query: 239 DYAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAIS 298
DY G KM ++ +P G++T + +S+ LIP+ + + + + + L L
Sbjct: 195 DYQKAGLKMLTVLEPDGKRTNRQIIWHSLLLIPVSLVPFYLNLLGIVYFYGALLLGLGYL 254
Query: 299 AAAFSFYRDRTKERARRMFHASLLYLPVFMAGLLIHR 335
+ F + + E AR + S++YLP +L+ R
Sbjct: 255 LSGFMLVKKYSVENARFLLKMSVIYLPALFGTILLDR 291
>UniRef100_UPI00002AFF7B UPI00002AFF7B UniRef100 entry
Length = 272
Score = 184 bits (467), Expect = 4e-45
Identities = 101/263 (38%), Positives = 152/263 (57%), Gaps = 1/263 (0%)
Query: 71 VVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDAIMRRTSQRPLP 130
VV ++ GFVL G + + L + GT ++ A A+TLNQ EI D+ M RT R LP
Sbjct: 5 VVLSALLGFVLPMGLSNTIPQLLFLLFGTALMGAGANTLNQCMEIGPDSRMYRTKNRALP 64
Query: 131 SGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTPLKQIHPVNTWV 190
G+++ A+ + + ++G +L N L A L L+ Y VYTPLKQ NTW+
Sbjct: 65 MGKLSEMEAIIFGILISVSGFLVLWFGLNFLTAVLGLLTLLSYIMVYTPLKQKTATNTWL 124
Query: 191 GAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCREDYAAGGFKMYSL 250
G + GA+PP++GW AA G + L + A LYFWQ+PHF A+A+M R+DY GGFKM S
Sbjct: 125 GGITGALPPVMGWVAARGQLEWEVLPIFALLYFWQLPHFFAIAWMYRDDYKRGGFKMLSH 184
Query: 251 ADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISAAAFSFYRDRTK 310
D +G++TAM + N L + S ++ ++AL L F+++R+
Sbjct: 185 YDKTGKRTAMQMLINCGMLYAASLAVFVVNEGSIFYLAGASALILVFFGVIVLFHKERSV 244
Query: 311 ERARRMFHASLLYLPVFMAGLLI 333
E AR++F AS++YLP+ + G+L+
Sbjct: 245 ENARKVFLASIIYLPL-LIGILV 266
>UniRef100_UPI00002362EA UPI00002362EA UniRef100 entry
Length = 506
Score = 182 bits (462), Expect = 1e-44
Identities = 122/335 (36%), Positives = 176/335 (52%), Gaps = 38/335 (11%)
Query: 38 PSVSASKSNDLISLA----RHYGNCYSQLSKARLSLLVV--ATSGTG-FVLGSGGAVD-- 88
P SA SN +L R + L+K RLS LV+ TS G + + S A+D
Sbjct: 128 PDASAQLSNFSSTLPKTSLRRKLAAFFALTKPRLSFLVLLSTTSAYGIYPVSSILALDPT 187
Query: 89 ---LSMLS-------YTCLGTMMVAASASTLNQVFEIKNDAIMRRTSQRPLPSGRITVPH 138
L LS Y GT + A SA+TLN +FE K DA M RT RPL G +T
Sbjct: 188 IAPLPTLSTSTLTFLYLTTGTFLSACSANTLNMIFEPKYDAQMSRTRNRPLVRGLVTRRA 247
Query: 139 AVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTPLKQIHPVNTWVGAVVGAIP 198
AV +A + G LL TN GL+A+N+ LY+FVYTPLK++H +NTW+GA+VG IP
Sbjct: 248 AVFFAIATAAVGLGLLYFGTNPTVTGLSAANIALYAFVYTPLKRMHVINTWIGAIVGGIP 307
Query: 199 PLLGWAAAS----------------GDISLNSLILPAALYFWQIPHFMALAYMCREDYAA 242
P++GW AA+ G+ SL +L L+ WQ PHF AL++ RE+Y
Sbjct: 308 PMMGWVAAAGQGATTGHDTWRDMLFGENSLGGWLLGGILFAWQFPHFNALSHTIREEYKG 367
Query: 243 GGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISAAAF 302
G+KM +P+ + A VA+R S+ + PL + G+ F + ST ++ A+
Sbjct: 368 AGYKMLCWTNPA--RNARVALRYSVLMFPLSIGLWWAGIVGHGFLVSSTVANGWLTKEAY 425
Query: 303 SFYRDR-TKERARRMFHASLLYLPVFMAGLLIHRR 336
F++ + AR +F AS+ LP+ + G L+ ++
Sbjct: 426 GFWKHQGANGTARGLFWASIWQLPILLVGALVTKK 460
>UniRef100_UPI0000274407 UPI0000274407 UniRef100 entry
Length = 293
Score = 181 bits (459), Expect = 3e-44
Identities = 96/279 (34%), Positives = 156/279 (55%)
Query: 57 NCYSQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIK 116
N Y L K + ++V+ T+ G+ LGS G + L +GT + A AS LNQ E +
Sbjct: 13 NSYIDLMKPNILIMVLITTILGYYLGSDGKILWDNLISLMIGTFLSAGGASVLNQYLERE 72
Query: 117 NDAIMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFV 176
D IM RT +RP+P G I+ +A+ + ++GT +LA N+L ++ +Y +
Sbjct: 73 QDKIMNRTCKRPIPMGIISPQNALIFGIISVISGTVILAIMINILTGFISLLTAFMYVLI 132
Query: 177 YTPLKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMC 236
YTP+K+I +NT +G++ GA+PP+ GW A++ I + IL A LY WQ PHF A+A+MC
Sbjct: 133 YTPMKRITWLNTSIGSIPGALPPVGGWVASTNSIDSGAWILFAILYLWQHPHFFAIAWMC 192
Query: 237 REDYAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLA 296
++DY GFKM + +P+GR+T + + L P+ L G+ + +TL
Sbjct: 193 KDDYEKAGFKMLPVIEPNGRRTVRQILWHLSLLFPITLLPVFIGMNGSIYMYGVLIITLY 252
Query: 297 ISAAAFSFYRDRTKERARRMFHASLLYLPVFMAGLLIHR 335
+AF +++ + A R+ AS++YLP M ++I +
Sbjct: 253 YFLSAFPMLYNKSHKNASRILKASVVYLPALMMIIIIDK 291
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.132 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 621,493,580
Number of Sequences: 2790947
Number of extensions: 24115690
Number of successful extensions: 88188
Number of sequences better than 10.0: 1049
Number of HSP's better than 10.0 without gapping: 768
Number of HSP's successfully gapped in prelim test: 281
Number of HSP's that attempted gapping in prelim test: 86001
Number of HSP's gapped (non-prelim): 1221
length of query: 402
length of database: 848,049,833
effective HSP length: 130
effective length of query: 272
effective length of database: 485,226,723
effective search space: 131981668656
effective search space used: 131981668656
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 76 (33.9 bits)
Medicago: description of AC146329.8


