

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146329.2 + phase: 0
(404 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
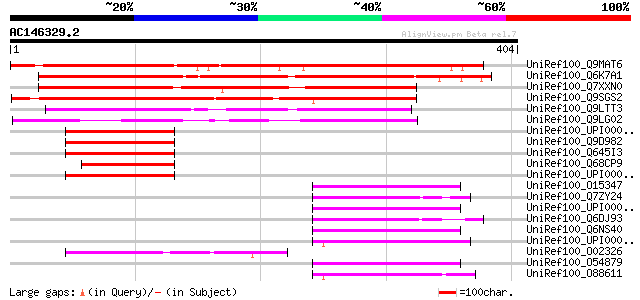
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9MAT6 F13M7.13 protein [Arabidopsis thaliana] 410 e-113
UniRef100_Q6K7A1 Glutathione S-transferase GST16-like protein [O... 293 6e-78
UniRef100_Q7XXN0 Glutathione S-transferase GST 16-like protein [... 269 9e-71
UniRef100_Q9SGS2 T23E18.4 [Arabidopsis thaliana] 268 3e-70
UniRef100_Q9LTT3 High mobility group protein-like [Arabidopsis t... 248 3e-64
UniRef100_Q9LG02 F20N2.8 [Arabidopsis thaliana] 129 1e-28
UniRef100_UPI00003A9660 UPI00003A9660 UniRef100 entry 84 7e-15
UniRef100_Q9D982 Mus musculus adult male testis cDNA, RIKEN full... 82 4e-14
UniRef100_Q645I3 ARID2 [Homo sapiens] 82 4e-14
UniRef100_Q68CP9 Hypothetical protein DKFZp779P0222 [Homo sapiens] 79 2e-13
UniRef100_UPI0000360A2D UPI0000360A2D UniRef100 entry 78 4e-13
UniRef100_O15347 High mobility group protein 4 [Homo sapiens] 67 9e-10
UniRef100_Q7ZY24 Hmgb3-prov protein [Xenopus laevis] 67 1e-09
UniRef100_UPI00001D1880 UPI00001D1880 UniRef100 entry 66 2e-09
UniRef100_Q6DJ93 MGC88931 protein [Xenopus tropicalis] 66 2e-09
UniRef100_Q6NS40 Hypothetical protein [Homo sapiens] 66 2e-09
UniRef100_UPI0000428F48 UPI0000428F48 UniRef100 entry 65 4e-09
UniRef100_O02326 Hypothetical protein T23D8.8 [Caenorhabditis el... 64 6e-09
UniRef100_O54879 High mobility group protein 4 [Mus musculus] 64 6e-09
UniRef100_O88611 High mobility group protein [Spalax leucodon eh... 64 8e-09
>UniRef100_Q9MAT6 F13M7.13 protein [Arabidopsis thaliana]
Length = 448
Score = 410 bits (1055), Expect = e-113
Identities = 228/402 (56%), Positives = 283/402 (69%), Gaps = 34/402 (8%)
Query: 1 MASTSCFSKSPLPMKETALSHGEYPPPMATYEEVVDNPKLFILCLEKLHTLMGTKFMIPV 60
MAS+SC + +PM ++ P ATYE VV +P+LF+ LE+LH+L+GTKFM+P+
Sbjct: 1 MASSSCLKQGSVPMNNVCVT------PEATYEAVVADPRLFMTSLERLHSLLGTKFMVPI 54
Query: 61 IGGRELDLHRLFVEVTSRGGFEKIIKDRKWKEVTLVFNFPSTATNASFVLRKYYTSLLYH 120
IGGR+LDLH+LFVEVTSRGG KI+ +R+WKEVT F FP TATNAS+VLRKYY SLL +
Sbjct: 55 IGGRDLDLHKLFVEVTSRGGINKILNERRWKEVTATFVFPPTATNASYVLRKYYFSLLNN 114
Query: 121 YEQIYYFKARDWTNTTSDVLQSQSSIPA-------PAPKMQFSH--PSPQVQPAVFPMGS 171
YEQIY+F++ D +QS S+ P P+ ++Q P P++ A F +G
Sbjct: 115 YEQIYFFRSNG--QIPPDSMQSPSARPCFIQGAIRPSQELQALTFTPQPKINTAEF-LGG 171
Query: 172 SSAGSQVVGVIDGKFDSGYLVTVTIGSEKLKGVLYQA-PQNPV---LPASHHSVPANNNN 227
S AGS VVGVIDGKF+SGYLVTVTIGSE+LKGVLYQ PQN V P H V N N
Sbjct: 172 SLAGSNVVGVIDGKFESGYLVTVTIGSEQLKGVLYQLLPQNTVSYQTPQQSHGVLPNTLN 231
Query: 228 VTASV-----GV-HRRRRRKKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKDRE 281
++A+ GV RRRRRKKSE+K+RDP HPKPNRSGYNFFFAEQH RLKPLH GKDR+
Sbjct: 232 ISANPQGVAGGVTKRRRRRKKSEIKRRDPDHPKPNRSGYNFFFAEQHARLKPLHPGKDRD 291
Query: 282 ISRTIGELWNKLPESEKAVYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPE 341
ISR IGELWNKL E EK +YQ KA++DKERY TEME YREK KN ++IS+AVPL+QRLPE
Sbjct: 292 ISRMIGELWNKLNEDEKLIYQGKAMEDKERYRTEMEDYREKKKNGQLISNAVPLQQRLPE 351
Query: 342 PDTDMLNAE--ADSLQTPEQ----SSLDGSDDYEDDKAKEKD 377
+ DM A+ D ++ ++ S G + DD++ E D
Sbjct: 352 QNVDMAEADLPIDEVEEDDEEGDSSGSSGESEPHDDQSIETD 393
>UniRef100_Q6K7A1 Glutathione S-transferase GST16-like protein [Oryza sativa]
Length = 467
Score = 293 bits (750), Expect = 6e-78
Identities = 161/372 (43%), Positives = 234/372 (62%), Gaps = 20/372 (5%)
Query: 24 YPPPMATYEEVVDNPKLFILCLEKLHTLMGTKFMIPVIGGRELDLHRLFVEVTSRGGFEK 83
YP +A Y++VV + +F LE LH MGTK +P+IGG++LDLH+LF EVTSRGG +K
Sbjct: 64 YPARVAGYKDVVADAAVFRRALEGLHAQMGTKLKVPIIGGKDLDLHQLFKEVTSRGGIDK 123
Query: 84 IIKDRKWKEVTLVFNFPSTATNASFVLRKYYTSLLYHYEQIYYFKARDWTNTTSDVLQSQ 143
+ D +W+EVT F FP+TATNASF+L+KYY SLLYH+E++Y F+A+ W T +S
Sbjct: 124 VKSDNRWREVTASFIFPATATNASFMLKKYYMSLLYHFERLYLFEAQGWYQETDS--RSI 181
Query: 144 SSIPAPAPKMQFSHPSPQVQPAVFPMGSSSAGSQVVGVIDGKFDSGYLVTVTIGSEKLKG 203
S I A + Q S + + ++S + V +IDGKF+ GY+VTV +GS+ K
Sbjct: 182 SCIEMKA-EGQASRKRKRGSNSCSSDLAASLDNDVQVIIDGKFEHGYIVTVIMGSKSTKA 240
Query: 204 VLYQAPQNPVLPASHHSVPANNNNVTASVGVHRRRRRKKSEMKKRDPAHPKPNRSGYNFF 263
VLY + P +P + V + ++ G+ RRRR++ ++ DP HPKPNRSGYNFF
Sbjct: 241 VLYNCTEEPAVPTAVPHVA-----IDSAEGIRPRRRRRRKKLSTTDPNHPKPNRSGYNFF 295
Query: 264 FAEQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVYQDKAVKDKERYITEMEYYREKL 323
F +QH +LKP + G+DR IS+ IGE WN L +KAVYQ+K V+DK RY ++ YRE+
Sbjct: 296 FQDQHRKLKPEYPGQDRLISKMIGERWNNLGPEDKAVYQEKGVEDKARYQRQLALYREQ- 354
Query: 324 KNDEVISDAVPLRQRLPE-----PDTDMLNAEADSLQTPE--QSSLDGSDDYEDDKAK-- 374
+ + IS+AVP++QRLP+ + D +E D L + + SS SD+ D K
Sbjct: 355 RTGQPISNAVPIQQRLPQKEVTIDEVDSKVSEGDILLSNQGYSSSTSSSDETADSGEKNV 414
Query: 375 --EKDFSVDSLP 384
+++F+ ++ P
Sbjct: 415 EDDEEFNTETSP 426
>UniRef100_Q7XXN0 Glutathione S-transferase GST 16-like protein [Oryza sativa]
Length = 306
Score = 269 bits (688), Expect = 9e-71
Identities = 140/310 (45%), Positives = 191/310 (61%), Gaps = 27/310 (8%)
Query: 24 YPPPMATYEEVVDNPKLFILCLEKLHTLMGTKFMIPVIGGRELDLHRLFVEVTSRGGFEK 83
YPPP+ ++EEV ++ F+ L + H+LMGTKFMIPVIGG+E+DLH L+VEVTSRGG K
Sbjct: 7 YPPPLLSHEEVANDRAAFMDTLRRFHSLMGTKFMIPVIGGKEMDLHALYVEVTSRGGLAK 66
Query: 84 IIKDRKWKEVTLVFNFPSTATNASFVLRKYYTSLLYHYEQIYYFKARDWTNTTSDVLQSQ 143
++++RKW+EV F+FP+T T+AS+VLR+YY SLL+HYEQ+Y+F+A +L+
Sbjct: 67 VMEERKWREVMARFSFPATTTSASYVLRRYYLSLLHHYEQVYFFRAH------GALLRPA 120
Query: 144 SSIPAPAPKMQFSHPSPQVQPAVFP---------MGSSSAGSQVVGVIDGKFDSGYLVTV 194
+S P+ + S Q A +G V G IDGKF+ GYLVTV
Sbjct: 121 ASALTKTPRRKMRGTSDQSPAAAEAGKRMALPERLGGEPCSFSVTGSIDGKFEHGYLVTV 180
Query: 195 TIGSEKLKGVLYQAPQNPVLPASHHSVPANNNNVTASVGVHRRRRRKKSEMKKRDPAHPK 254
I +E L+GVLY+ P PA+ P R R++ ++RDPA P+
Sbjct: 181 KIAAETLRGVLYRVAPPPPPPAAPPPPPP------------PARGRRRRGRRQRDPAQPR 228
Query: 255 PNRSGYNFFFAEQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVYQDKAVKDKERYIT 314
PNRS YNFFF E+HP LK H ++RE SR IG+ WN+L +K VY + +DKERY
Sbjct: 229 PNRSAYNFFFKEKHPELKATHPHREREYSRMIGDAWNRLAADDKMVYYRHSAEDKERYKR 288
Query: 315 EMEYYREKLK 324
EM+ Y E+LK
Sbjct: 289 EMQEYNERLK 298
>UniRef100_Q9SGS2 T23E18.4 [Arabidopsis thaliana]
Length = 338
Score = 268 bits (684), Expect = 3e-70
Identities = 148/327 (45%), Positives = 200/327 (60%), Gaps = 16/327 (4%)
Query: 2 ASTSCFSKSPLPMKETALSHGEYPPPMATYEEVVDNPKLFILCLEKLHTLMGTKFMIPVI 61
A+ + SP +KE YP P+A +E VV + +F L + H++M TKFMIPVI
Sbjct: 12 ATVEMMATSPAKIKE-------YPEPLALHEVVVKDSSVFWDTLRRFHSIMSTKFMIPVI 64
Query: 62 GGRELDLHRLFVEVTSRGGFEKIIKDRKWKEVTLVFNFPSTATNASFVLRKYYTSLLYHY 121
GG+ELDLH L+VEVT RGG+EK++ ++KW+EV VF F +T T+ASFVLRK+Y +LL+HY
Sbjct: 65 GGKELDLHVLYVEVTRRGGYEKVVVEKKWREVGGVFRFSATTTSASFVLRKHYLNLLFHY 124
Query: 122 EQIYYFKARDWTNTTSDVLQSQSSIPAPAPKMQFSHPSPQVQPAVFPMGSSSAGSQVVGV 181
EQ++ F AR + S ++++ PS + P S+ +G
Sbjct: 125 EQVHLFTARGPLLHPIATFHANPSTSKEMALVEYTPPSIRYH-NTHPPSQGSSSFTAIGT 183
Query: 182 IDGKFDSGYLVTVTIGSEKLKGVLYQAPQNPVLPASHHSVPA-NNNNVTASVGVHRRRRR 240
I+GKFD GYLV V +GSE L GVLY + Q P S A NN V V RRRRR
Sbjct: 184 IEGKFDCGYLVKVKLGSEILNGVLYHSAQ----PGPSSSPTAVLNNAVVPYVETGRRRRR 239
Query: 241 ---KKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKDREISRTIGELWNKLPESE 297
++ ++ DP +PKPNRSGYNFFFAE+H +LK L+ K+RE ++ IGE W+ L E
Sbjct: 240 LGKRRRSRRREDPNYPKPNRSGYNFFFAEKHCKLKSLYPNKEREFTKLIGESWSNLSTEE 299
Query: 298 KAVYQDKAVKDKERYITEMEYYREKLK 324
+ VYQD +KDKERY E+ YRE L+
Sbjct: 300 RMVYQDIGLKDKERYQRELNEYRETLR 326
>UniRef100_Q9LTT3 High mobility group protein-like [Arabidopsis thaliana]
Length = 319
Score = 248 bits (632), Expect = 3e-64
Identities = 137/293 (46%), Positives = 177/293 (59%), Gaps = 22/293 (7%)
Query: 29 ATYEEVVDNPKLFILCLEKLHTLMGTKFMIPVIGGRELDLHRLFVEVTSRGGFEKIIKDR 88
A Y+++V N LF L L +P +GG LDLHRLF+EVTSRGG E+++KDR
Sbjct: 34 AKYDDLVRNSALFWEKLRAFLGLTSKTLKVPTVGGNTLDLHRLFIEVTSRGGIERVVKDR 93
Query: 89 KWKEVTLVFNFPSTATNASFVLRKYYTSLLYHYEQIYYF-KARDWTNTTSDVLQSQSSIP 147
KWKEV F+FP+T T+ASFVLRKYY L+ E +YY K +T + L+S ++
Sbjct: 94 KWKEVIGAFSFPTTITSASFVLRKYYLKFLFQLEHVYYLEKPVSSLQSTDEALKSLAN-E 152
Query: 148 APAPKMQFSHPSPQVQPAVFPMGSSSAGSQVVGVIDGKFDSGYLVTVTIGSEKLKGVLYQ 207
+P P+ P G +V G IDGKFDSGYLVT+ +GS++LKGVLY
Sbjct: 153 SPNPEEGIDEP--------------QVGYEVQGFIDGKFDSGYLVTMKLGSQELKGVLYH 198
Query: 208 APQNPVLPASHHSVPANNNNVTASVGVHRRRRRKKSEMKKRDPAHPKPNRSGYNFFFAEQ 267
PQ P P +A V +RR RKKS++ D PK +RSGYNFFFAEQ
Sbjct: 199 IPQTPSQSQQTMETP------SAIVQSSQRRHRKKSKLAVVDTQKPKCHRSGYNFFFAEQ 252
Query: 268 HPRLKPLHRGKDREISRTIGELWNKLPESEKAVYQDKAVKDKERYITEMEYYR 320
+ RLKP + G++R I++ IG +W+ L ESEK VYQDK VKD ERY EM Y+
Sbjct: 253 YARLKPEYHGQERSITKKIGHMWSNLTESEKQVYQDKGVKDVERYRIEMLEYK 305
>UniRef100_Q9LG02 F20N2.8 [Arabidopsis thaliana]
Length = 315
Score = 129 bits (325), Expect = 1e-28
Identities = 93/324 (28%), Positives = 141/324 (42%), Gaps = 87/324 (26%)
Query: 3 STSCFSKSPLPMKETALSHGEYPPPMATYEEVVDNPKLFILCLEKLHTLMGTKFMIPVIG 62
ST +P +AL + Y+++V NP+LF L H KF
Sbjct: 28 STDSSQFEVVPANASALDNDVSSHMSMLYQDIVRNPELFWEMLRDFHESSDKKF------ 81
Query: 63 GRELDLHRLFVEVTSRGGFEKIIKDRKWKEVTLVFNFPSTATNASFVLRKYYTSLLYHYE 122
K KEV FNF +T TN++FVLRK Y +L+ +E
Sbjct: 82 --------------------------KCKEVIDAFNFKTTITNSAFVLRKSYLKMLFEFE 115
Query: 123 QIYYFKARDWTNTTSDVLQSQSSIPAPAPKMQFSHPSPQVQPAVFPMGSSSAGSQVVGVI 182
+YYF+A T D S +++P G+ + G+I
Sbjct: 116 HLYYFQAPLSTFWEKD--------------------SQELKP----------GTVITGII 145
Query: 183 DGKFDSGYLVTVTIGSEKLKGVLYQAPQNPVLPASHHSVPANNNNVTASVGVHRRRRRKK 242
DGKF+SGYL++ +GSEKLKG+LY + +R +KK
Sbjct: 146 DGKFESGYLISTKVGSEKLKGMLYH------------------------ISPETKRGKKK 181
Query: 243 SEMKKRDPAHP-KPNRSGYNFFFAEQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVY 301
++ + D P K R+GYNFF AEQ R+K + G+ + G +W L ES++ VY
Sbjct: 182 AKSSQGDSHKPPKRQRTGYNFFVAEQSVRIKAENAGQKVSSPKNFGNMWTNLSESDRKVY 241
Query: 302 QDKAVKDKERYITEMEYYREKLKN 325
+K+ +D +RY E+ YR +++
Sbjct: 242 YEKSREDGKRYKMEILQYRSLMES 265
>UniRef100_UPI00003A9660 UPI00003A9660 UniRef100 entry
Length = 141
Score = 84.0 bits (206), Expect = 7e-15
Identities = 38/88 (43%), Positives = 57/88 (64%), Gaps = 1/88 (1%)
Query: 45 LEKLHTLMGTKFM-IPVIGGRELDLHRLFVEVTSRGGFEKIIKDRKWKEVTLVFNFPSTA 103
L + H G+ F IPV+GG+ELDLH L+ VT+ GGF K+ + +W E+ FNFP +
Sbjct: 23 LRQFHHSRGSPFKKIPVVGGKELDLHALYTRVTTLGGFGKVSEKNQWGEIVEEFNFPRSC 82
Query: 104 TNASFVLRKYYTSLLYHYEQIYYFKARD 131
+NA+F L++YY L YE++++F D
Sbjct: 83 SNAAFALKQYYLRYLEKYEKVHHFGEED 110
>UniRef100_Q9D982 Mus musculus adult male testis cDNA, RIKEN full-length enriched
library, clone:1700124K17 product:hypothetical AT-rich
interaction domain (ARID) containing protein, full
insert sequence [Mus musculus]
Length = 145
Score = 81.6 bits (200), Expect = 4e-14
Identities = 37/88 (42%), Positives = 56/88 (63%), Gaps = 1/88 (1%)
Query: 45 LEKLHTLMGTKFM-IPVIGGRELDLHRLFVEVTSRGGFEKIIKDRKWKEVTLVFNFPSTA 103
L + H G+ F IP +GG+ELDLH L+ VT+ GGF K+ + +W E+ FNFP +
Sbjct: 23 LRQFHHSRGSPFKKIPAVGGKELDLHGLYTRVTTLGGFAKVSEKNQWGEIVEEFNFPRSC 82
Query: 104 TNASFVLRKYYTSLLYHYEQIYYFKARD 131
+NA+F L++YY L YE++++F D
Sbjct: 83 SNAAFALKQYYLRYLEKYEKVHHFGEDD 110
>UniRef100_Q645I3 ARID2 [Homo sapiens]
Length = 209
Score = 81.6 bits (200), Expect = 4e-14
Identities = 37/88 (42%), Positives = 56/88 (63%), Gaps = 1/88 (1%)
Query: 45 LEKLHTLMGTKFM-IPVIGGRELDLHRLFVEVTSRGGFEKIIKDRKWKEVTLVFNFPSTA 103
L + H G+ F IP +GG+ELDLH L+ VT+ GGF K+ + +W E+ FNFP +
Sbjct: 23 LRQFHHSRGSPFKKIPAVGGKELDLHGLYTRVTTLGGFAKVSEKNQWGEIVEEFNFPRSC 82
Query: 104 TNASFVLRKYYTSLLYHYEQIYYFKARD 131
+NA+F L++YY L YE++++F D
Sbjct: 83 SNAAFALKQYYLRYLEKYEKVHHFGEDD 110
>UniRef100_Q68CP9 Hypothetical protein DKFZp779P0222 [Homo sapiens]
Length = 1756
Score = 79.0 bits (193), Expect = 2e-13
Identities = 33/74 (44%), Positives = 50/74 (66%)
Query: 58 IPVIGGRELDLHRLFVEVTSRGGFEKIIKDRKWKEVTLVFNFPSTATNASFVLRKYYTSL 117
IP +GG+ELDLH L+ VT+ GGF K+ + +W E+ FNFP + +NA+F L++YY
Sbjct: 10 IPAVGGKELDLHGLYTRVTTLGGFAKVSEKNQWGEIVEEFNFPRSCSNAAFALKQYYLRY 69
Query: 118 LYHYEQIYYFKARD 131
L YE++++F D
Sbjct: 70 LEKYEKVHHFGEDD 83
>UniRef100_UPI0000360A2D UPI0000360A2D UniRef100 entry
Length = 140
Score = 78.2 bits (191), Expect = 4e-13
Identities = 36/88 (40%), Positives = 55/88 (61%), Gaps = 1/88 (1%)
Query: 45 LEKLHTLMGTKFM-IPVIGGRELDLHRLFVEVTSRGGFEKIIKDRKWKEVTLVFNFPSTA 103
L + H G+ F IP +GG+ELDL+ L++ V S GGF K+ + +W E+ FNFP
Sbjct: 23 LRQFHQSRGSPFRKIPFVGGKELDLNALYIRVVSLGGFAKVSEKNQWMELGEEFNFPRNC 82
Query: 104 TNASFVLRKYYTSLLYHYEQIYYFKARD 131
+NA+F L++YY L YE++++F D
Sbjct: 83 SNAAFALKQYYLRYLEKYEKVHHFGEDD 110
>UniRef100_O15347 High mobility group protein 4 [Homo sapiens]
Length = 199
Score = 67.0 bits (162), Expect = 9e-10
Identities = 38/119 (31%), Positives = 63/119 (52%), Gaps = 1/119 (0%)
Query: 242 KSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEKAV 300
K KK+DP PK SG+ F +E P++K + G ++++ +GE+WN L +SEK
Sbjct: 81 KGGKKKKDPNAPKRPPSGFFLFCSEFRPKIKSTNPGISIGDVAKKLGEMWNNLNDSEKQP 140
Query: 301 YQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPEQ 359
Y KA K KE+Y ++ Y+ K K D A R+++ E D + E + + ++
Sbjct: 141 YITKAAKLKEKYEKDVADYKSKGKFDGAKGPAKVARKKVEEEDEEQEEEEEEEEEEEDE 199
Score = 43.1 bits (100), Expect = 0.014
Identities = 26/76 (34%), Positives = 34/76 (44%), Gaps = 3/76 (3%)
Query: 247 KRDPAHPKPNRSGYNFFFA---EQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVYQD 303
K DP PK S Y FF E+H + P E S+ E W + EK+ + +
Sbjct: 2 KGDPKKPKGKTSAYAFFVQTCREEHKKKNPEVPVNFAEFSKKCSERWKTVSGKEKSKFDE 61
Query: 304 KAVKDKERYITEMEYY 319
A DK RY EM+ Y
Sbjct: 62 MAKADKVRYDREMKDY 77
>UniRef100_Q7ZY24 Hmgb3-prov protein [Xenopus laevis]
Length = 202
Score = 66.6 bits (161), Expect = 1e-09
Identities = 43/127 (33%), Positives = 64/127 (49%), Gaps = 8/127 (6%)
Query: 242 KSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEKAV 300
K KK+DP PK SG+ F +E P++K + G ++++ +GE+WN L +SEK
Sbjct: 82 KKGKKKKDPNAPKRPPSGFFLFCSEFRPKIKSTNPGITIGDVAKKLGEMWNNLSDSEKQP 141
Query: 301 YQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPEQS 360
Y +K K KE+Y ++ Y+ K K D + A P R E D D D + E+
Sbjct: 142 YNNKGAKLKEKYEKDVADYKSKGKFDG--AKAAPKLARKKEEDDD-----DDDEEDEEED 194
Query: 361 SLDGSDD 367
D DD
Sbjct: 195 EEDEDDD 201
Score = 43.5 bits (101), Expect = 0.011
Identities = 26/76 (34%), Positives = 34/76 (44%), Gaps = 3/76 (3%)
Query: 245 MKKRDPAHPKPNRSGYNFFFA---EQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVY 301
M K DP PK S Y +F E+H + P E S+ E W + EK+ +
Sbjct: 1 MAKGDPKKPKGKMSAYAYFVQTCREEHKKKNPEIPVNFSEFSKKCSERWRGMSGKEKSKF 60
Query: 302 QDKAVKDKERYITEME 317
D A DK RY EM+
Sbjct: 61 DDLAKADKVRYDREMQ 76
>UniRef100_UPI00001D1880 UPI00001D1880 UniRef100 entry
Length = 200
Score = 66.2 bits (160), Expect = 2e-09
Identities = 37/119 (31%), Positives = 63/119 (52%), Gaps = 1/119 (0%)
Query: 242 KSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEKAV 300
K KK+DP PK SG+ F +E P++K + G ++++ +GE+WN L +SEK
Sbjct: 82 KGGKKKKDPNAPKRPPSGFFLFCSEFRPKIKSANPGISIGDVAKKLGEMWNNLSDSEKQP 141
Query: 301 YQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPEQ 359
Y KA K KE+Y ++ Y+ K K D A R+++ E + + E + + ++
Sbjct: 142 YMTKAAKLKEKYEKDVADYKSKGKFDGAKGPAKVARKKVEEEEEEEEEEEEEEEEEEDE 200
Score = 45.8 bits (107), Expect = 0.002
Identities = 27/78 (34%), Positives = 35/78 (44%), Gaps = 3/78 (3%)
Query: 245 MKKRDPAHPKPNRSGYNFFFA---EQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVY 301
M K DP PK S Y FF E+H + P E S+ E W + EK+ +
Sbjct: 1 MAKGDPKKPKGKMSAYAFFVQTCREEHKKKNPEVPVNFAEFSKKCSERWKTMSSKEKSKF 60
Query: 302 QDKAVKDKERYITEMEYY 319
+ A DK RY EM+ Y
Sbjct: 61 DEMAKADKVRYDREMKDY 78
>UniRef100_Q6DJ93 MGC88931 protein [Xenopus tropicalis]
Length = 202
Score = 66.2 bits (160), Expect = 2e-09
Identities = 43/137 (31%), Positives = 66/137 (47%), Gaps = 20/137 (14%)
Query: 242 KSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEKAV 300
K KK+DP PK SG+ F +E P++K + G ++++ +GE+WN L + EK
Sbjct: 82 KKGKKKKDPNAPKRPPSGFFLFCSEFRPKIKSTNPGISIGDVAKKLGEMWNNLNDGEKQP 141
Query: 301 YQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPEQS 360
Y +KA K KE+Y ++ Y+ K K D + P R E D D
Sbjct: 142 YNNKAAKLKEKYEKDVADYKSKGKFD--CAKGAPKLARKKEEDED--------------- 184
Query: 361 SLDGSDDYEDDKAKEKD 377
D +D E+D+ E+D
Sbjct: 185 --DDDEDEEEDEEDEED 199
Score = 43.9 bits (102), Expect = 0.008
Identities = 26/76 (34%), Positives = 34/76 (44%), Gaps = 3/76 (3%)
Query: 245 MKKRDPAHPKPNRSGYNFFFA---EQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVY 301
M K DP PK S Y +F E+H + P E S+ E W + EK+ +
Sbjct: 1 MAKGDPKKPKGKMSAYAYFVQTCREEHKKKNPEIPVNFAEFSKKCSERWKTMSAKEKSKF 60
Query: 302 QDKAVKDKERYITEME 317
D A DK RY EM+
Sbjct: 61 DDLAKADKVRYDREMK 76
>UniRef100_Q6NS40 Hypothetical protein [Homo sapiens]
Length = 200
Score = 66.2 bits (160), Expect = 2e-09
Identities = 38/119 (31%), Positives = 63/119 (52%), Gaps = 1/119 (0%)
Query: 242 KSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEKAV 300
K KK+DP PK SG+ F +E P++K + G ++++ +GE+WN L +SEK
Sbjct: 82 KGGKKKKDPNAPKRPPSGFFLFCSEFRPKIKSTNPGISIGDVAKKLGEMWNNLNDSEKQP 141
Query: 301 YQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPEQ 359
Y KA K KE+Y ++ Y+ K K D A R+++ E D + E + + ++
Sbjct: 142 YITKAAKLKEKYEKDVADYKSKGKFDGAKGPAKVARKKVEEEDEEEEEEEEEEEEEEDE 200
Score = 45.1 bits (105), Expect = 0.004
Identities = 27/78 (34%), Positives = 35/78 (44%), Gaps = 3/78 (3%)
Query: 245 MKKRDPAHPKPNRSGYNFFFA---EQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVY 301
M K DP PK S Y FF E+H + P E S+ E W + EK+ +
Sbjct: 1 MAKGDPKKPKGKMSAYAFFVQTCREEHKKKNPEVPVNFAEFSKKCSERWKTMSGKEKSKF 60
Query: 302 QDKAVKDKERYITEMEYY 319
+ A DK RY EM+ Y
Sbjct: 61 DEMAKADKVRYDREMKDY 78
>UniRef100_UPI0000428F48 UPI0000428F48 UniRef100 entry
Length = 201
Score = 64.7 bits (156), Expect = 4e-09
Identities = 40/129 (31%), Positives = 62/129 (48%), Gaps = 3/129 (2%)
Query: 242 KSEMKKR--DPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEK 298
K E KK+ DP PK S + F +E HP++K H G ++++ +GE+WN +K
Sbjct: 72 KGETKKKFKDPNPPKRPPSAFFLFCSEYHPKIKGEHPGLSIGDVAKKLGEMWNNTAADDK 131
Query: 299 AVYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPE 358
Y+ KA K KE+Y ++ YR K K D + + + + D E D + E
Sbjct: 132 QPYEKKAAKLKEKYEKDIAAYRAKGKPDAGKKVVKAEKSKKKKEEEDDEEDEEDEEEEDE 191
Query: 359 QSSLDGSDD 367
+ D DD
Sbjct: 192 EEDEDEDDD 200
>UniRef100_O02326 Hypothetical protein T23D8.8 [Caenorhabditis elegans]
Length = 467
Score = 64.3 bits (155), Expect = 6e-09
Identities = 49/181 (27%), Positives = 84/181 (46%), Gaps = 11/181 (6%)
Query: 45 LEKLHTLMGTKFMIPVIGGRELDLHRLFVEVTSRGGFEKIIKDRKWKEVTLVFNFPSTAT 104
L +H + IP++ + LDL+ L+ V GG +II + W+E+T N PS+ T
Sbjct: 192 LNFMHRIGKPVTRIPIMAKQVLDLYELYRLVVQHGGLVEIINKKLWREITKGLNLPSSIT 251
Query: 105 NASFVLRKYYTSLLYHYEQIYYFKARDWTNTTSDVLQSQSSIPAPAPKMQFSHPSPQVQP 164
+A+F LR Y LY YE ++ + SD+ Q+ AP + + P P
Sbjct: 252 SAAFTLRTQYQKYLYDYE-----CEKEKLSNQSDLQQAIDGNRREAPGRRTAPSFP--LP 304
Query: 165 AVFPMGSSSAGSQVVGVIDGKFDSGYLV----TVTIGSEKLKGVLYQAPQNPVLPASHHS 220
P +S+A + + ++G L+ T+++ + L G Y A Q +L A +
Sbjct: 305 FQLPHAASAAATMLNNQLNGLGMRNDLLDDENTLSLQASGLFGTSYGAEQMAILEAHQRN 364
Query: 221 V 221
+
Sbjct: 365 L 365
>UniRef100_O54879 High mobility group protein 4 [Mus musculus]
Length = 199
Score = 64.3 bits (155), Expect = 6e-09
Identities = 36/119 (30%), Positives = 63/119 (52%), Gaps = 1/119 (0%)
Query: 242 KSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEKAV 300
K KK+DP PK SG+ F +E P++K + G ++++ +GE+WN L ++EK
Sbjct: 81 KGGKKKKDPNAPKRPPSGFFLFCSEFRPKIKSTNPGISIGDVAKKLGEMWNNLSDNEKQP 140
Query: 301 YQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPEQ 359
Y KA K KE+Y ++ Y+ K K D A R+++ E + + E + + ++
Sbjct: 141 YVTKAAKLKEKYEKDVADYKSKGKFDGAKGPAKVARKKVEEEEEEEEEEEEEEEEEEDE 199
Score = 44.3 bits (103), Expect = 0.006
Identities = 26/76 (34%), Positives = 34/76 (44%), Gaps = 3/76 (3%)
Query: 247 KRDPAHPKPNRSGYNFFFA---EQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVYQD 303
K DP PK S Y FF E+H + P E S+ E W + EK+ + +
Sbjct: 2 KGDPKKPKGKMSAYAFFVQTCREEHKKKNPEVPVNFAEFSKKCSERWKTMSSKEKSKFDE 61
Query: 304 KAVKDKERYITEMEYY 319
A DK RY EM+ Y
Sbjct: 62 MAKADKVRYDREMKDY 77
>UniRef100_O88611 High mobility group protein [Spalax leucodon ehrenbergi]
Length = 215
Score = 63.9 bits (154), Expect = 8e-09
Identities = 40/133 (30%), Positives = 64/133 (48%), Gaps = 5/133 (3%)
Query: 242 KSEMKKR--DPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEK 298
K E KK+ DP PK S + F +E P++K H G ++++ +GE+WN +K
Sbjct: 82 KGETKKKFKDPNAPKRPPSAFFLFCSEYRPKIKGEHPGLSIGDVAKKLGEMWNNTAADDK 141
Query: 299 AVYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPE 358
Y+ KA K KE+Y ++ YR K K D V + + + + E D + E
Sbjct: 142 QPYEKKAAKLKEKYEKDIAAYRAKGKPDAAKKGVVKAEKSKKKKEEE--EDEEDEEEEEE 199
Query: 359 QSSLDGSDDYEDD 371
+ + D+ EDD
Sbjct: 200 EEDEEDEDEEEDD 212
Score = 45.1 bits (105), Expect = 0.004
Identities = 27/78 (34%), Positives = 34/78 (42%), Gaps = 3/78 (3%)
Query: 245 MKKRDPAHPKPNRSGYNFFFA---EQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVY 301
M K DP P+ S Y FF E+H + P E S+ E W EK +
Sbjct: 1 MGKGDPKKPRGKMSSYAFFVQTCREEHKKKHPDASVNFSEFSKKCSERWKTKSAKEKGKF 60
Query: 302 QDKAVKDKERYITEMEYY 319
+D A DK RY EM+ Y
Sbjct: 61 EDMAKADKARYEREMKTY 78
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.314 0.132 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 716,340,450
Number of Sequences: 2790947
Number of extensions: 32518339
Number of successful extensions: 85155
Number of sequences better than 10.0: 1045
Number of HSP's better than 10.0 without gapping: 612
Number of HSP's successfully gapped in prelim test: 437
Number of HSP's that attempted gapping in prelim test: 83544
Number of HSP's gapped (non-prelim): 1597
length of query: 404
length of database: 848,049,833
effective HSP length: 130
effective length of query: 274
effective length of database: 485,226,723
effective search space: 132952122102
effective search space used: 132952122102
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 76 (33.9 bits)
Medicago: description of AC146329.2


