

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146329.15 - phase: 0
(204 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
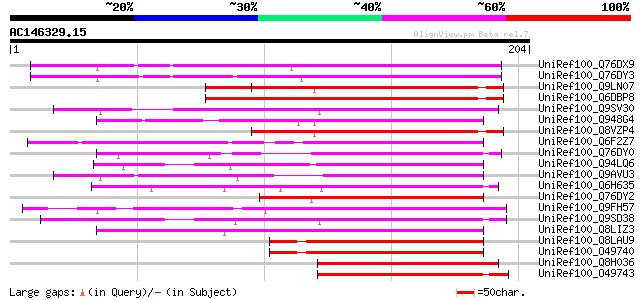
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q76DX9 AG-motif binding protein-5 [Nicotiana tabacum] 153 3e-36
UniRef100_Q76DY3 AG-motif binding protein-1 [Nicotiana tabacum] 152 4e-36
UniRef100_Q9LN07 T6D22.9 [Arabidopsis thaliana] 130 2e-29
UniRef100_Q6DBP8 At1g08010 [Arabidopsis thaliana] 130 2e-29
UniRef100_Q9SV30 Hypothetical protein F28P10.210 [Arabidopsis th... 129 6e-29
UniRef100_Q948G4 Putative GATA-1 zinc finger protein [Oryza sativa] 129 6e-29
UniRef100_Q8VZP4 Putative GATA transcription factor 3 [Arabidops... 125 7e-28
UniRef100_Q6F2Z7 Hypothetical protein P0483D07.7 [Oryza sativa] 125 9e-28
UniRef100_Q76DY0 AG-motif binding protein-4 [Nicotiana tabacum] 124 1e-27
UniRef100_Q94LQ6 Putative transcription factor [Oryza sativa] 123 3e-27
UniRef100_Q9AVU3 GATA-1 zinc finger protein [Nicotiana tabacum] 123 3e-27
UniRef100_Q6H635 Putative AG-motif binding protein-4 [Oryza sativa] 122 4e-27
UniRef100_Q76DY2 AG-motif binding protein-2 [Nicotiana tabacum] 122 6e-27
UniRef100_Q9FH57 GATA-binding transcription factor-like protein ... 121 1e-26
UniRef100_Q9SD38 Putative transcription factor [Arabidopsis thal... 121 1e-26
UniRef100_Q8LIZ3 OSJNBa0014K08.18 protein [Oryza sativa] 119 6e-26
UniRef100_Q8LAU9 GATA transcription factor 1 [Arabidopsis thaliana] 118 8e-26
UniRef100_O49740 Protein homologous to GATA-binding transcriptio... 118 8e-26
UniRef100_Q8H036 Hypothetical protein OJ1172F09.7 [Oryza sativa] 117 1e-25
UniRef100_O49743 Homologous to GATA-binding transcription factor... 116 4e-25
>UniRef100_Q76DX9 AG-motif binding protein-5 [Nicotiana tabacum]
Length = 342
Score = 153 bits (386), Expect = 3e-36
Identities = 93/196 (47%), Positives = 114/196 (57%), Gaps = 13/196 (6%)
Query: 9 SSSVNKEDFVLPKSNSSPTCEKTTV---------RRTRSKRPRLATFSSHHSTMQLISST 59
SS V+ + S+SS + EKT +R RSKRPR ATF+ +QLIS T
Sbjct: 118 SSPVSVLESSSSSSSSSCSVEKTVPLSSPCHRGPQRARSKRPRPATFNPA-PVIQLISPT 176
Query: 60 SSFVGENMQDSVISNKGASTEKFPDSQIAAKKQKLSSGESKKNKKTKAPL--LAALDHNA 117
SSF E Q V + +E F +S + + + + KK K P + A +
Sbjct: 177 SSFT-EIPQPFVARGIASESENFAESPMKKILKPAVAEQKKKKLKLSFPSARVEANQNPV 235
Query: 118 LGLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSN 177
+R+C HCE TKTPQWR GP GPKTLCNACGVRYKSGRL PEYRPAAS TF P +HSN
Sbjct: 236 AQTIRKCQHCEMTKTPQWRAGPMGPKTLCNACGVRYKSGRLFPEYRPAASPTFVPSIHSN 295
Query: 178 SHKKILEMRVMRRKDN 193
SHKK++EMR DN
Sbjct: 296 SHKKVIEMRTKFVPDN 311
>UniRef100_Q76DY3 AG-motif binding protein-1 [Nicotiana tabacum]
Length = 343
Score = 152 bits (385), Expect = 4e-36
Identities = 96/198 (48%), Positives = 117/198 (58%), Gaps = 16/198 (8%)
Query: 9 SSSVNKEDFVLPKSNSSPTCEKTTV---------RRTRSKRPRLATFSSHHSTMQLISST 59
SS V+ + S+SS + EKT +R RSKRPR ATF+ + +QLIS T
Sbjct: 118 SSPVSVLESSSSSSSSSCSVEKTVPLSSPCHRGPQRARSKRPRPATFNPAPA-IQLISPT 176
Query: 60 SSFVGENMQDSVISNKGASTEKFPDSQIAAKKQKLSSGESKKNKKTKAPLLAAL----DH 115
SSF E Q V + +E F +S + K K + E K KK K ++L +
Sbjct: 177 SSFT-EIPQPFVAPKITSESENFAESPMK-KILKPAVAEQKTKKKLKLSFPSSLVKTNQN 234
Query: 116 NALGLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLH 175
+R+C HCE TKTPQWR GP GPKTLCNACGVRYKSGRL PEYRPAAS TF P +H
Sbjct: 235 PVAQTIRKCQHCEITKTPQWRAGPMGPKTLCNACGVRYKSGRLFPEYRPAASPTFVPAIH 294
Query: 176 SNSHKKILEMRVMRRKDN 193
SNSHKK++EMR DN
Sbjct: 295 SNSHKKVIEMRTKVVPDN 312
>UniRef100_Q9LN07 T6D22.9 [Arabidopsis thaliana]
Length = 821
Score = 130 bits (327), Expect = 2e-29
Identities = 63/117 (53%), Positives = 82/117 (69%), Gaps = 3/117 (2%)
Query: 78 STEKFPDSQIAAKKQKLSSGESKKNKKTKAPLLAALDHNALGLVRQCTHCEATKTPQWRT 137
ST P+S+ ++ + + K + T+ N+ G+VR+CTHCE TKTPQWR
Sbjct: 251 STPGKPESECYFSSEQHAKKKRKIHLTTRTVSSTLEASNSDGIVRKCTHCETTKTPQWRE 310
Query: 138 GPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKKILEMRVMRRKDNK 194
GP GPKTLCNACGVR++SGRL PEYRPA+S TF P +HSNSH+KI+E MRRKD++
Sbjct: 311 GPSGPKTLCNACGVRFRSGRLVPEYRPASSPTFIPAVHSNSHRKIIE---MRRKDDE 364
Score = 125 bits (314), Expect = 7e-28
Identities = 61/105 (58%), Positives = 73/105 (69%), Gaps = 9/105 (8%)
Query: 96 SGESKKNKKTKAPLLAALDHNAL------GLVRQCTHCEATKTPQWRTGPEGPKTLCNAC 149
S E KK K L+ + + L G+VR CTHCE TPQWR GP GPKTLCNAC
Sbjct: 699 SSEQHAKKKRKIHLITHTESSTLESSKSDGIVRICTHCETITTPQWRQGPSGPKTLCNAC 758
Query: 150 GVRYKSGRLCPEYRPAASSTFSPDLHSNSHKKILEMRVMRRKDNK 194
GVR+KSGRL PEYRPA+S TF P +HSNSH+KI+E MR+KD++
Sbjct: 759 GVRFKSGRLVPEYRPASSPTFIPSVHSNSHRKIIE---MRKKDDE 800
>UniRef100_Q6DBP8 At1g08010 [Arabidopsis thaliana]
Length = 303
Score = 130 bits (327), Expect = 2e-29
Identities = 63/117 (53%), Positives = 82/117 (69%), Gaps = 3/117 (2%)
Query: 78 STEKFPDSQIAAKKQKLSSGESKKNKKTKAPLLAALDHNALGLVRQCTHCEATKTPQWRT 137
ST P+S+ ++ + + K + T+ N+ G+VR+CTHCE TKTPQWR
Sbjct: 176 STPGKPESECYFSSEQHAKKKRKIHLTTRTVSSTLEASNSDGIVRKCTHCETTKTPQWRE 235
Query: 138 GPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKKILEMRVMRRKDNK 194
GP GPKTLCNACGVR++SGRL PEYRPA+S TF P +HSNSH+KI+E MRRKD++
Sbjct: 236 GPSGPKTLCNACGVRFRSGRLVPEYRPASSPTFIPAVHSNSHRKIIE---MRRKDDE 289
>UniRef100_Q9SV30 Hypothetical protein F28P10.210 [Arabidopsis thaliana]
Length = 322
Score = 129 bits (323), Expect = 6e-29
Identities = 78/193 (40%), Positives = 98/193 (50%), Gaps = 34/193 (17%)
Query: 18 VLPKSNSSPTCEKTTVR-------RTRSKRPRLATFSSHHSTMQLISSTSSFVGENMQDS 70
VL S+SS TT R R+KRPR + ++D+
Sbjct: 123 VLESSSSSSQTTNTTSLVLPGKHGRPRTKRPRPPVQDK----------------DRVKDN 166
Query: 71 VISNKGASTEKFPDSQIAAKKQKLSSGESKKNKKTKAPLLAALDHNALGL---------- 120
V + P ++ + ++ + KK K T + + +D G
Sbjct: 167 VCGGDSRLIIRIPKQFLSDHNKMINKKKKKKAKITSSSSSSGIDLEVNGNNVDSYSSEQY 226
Query: 121 -VRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSH 179
+R+C HCE TKTPQWR GP GPKTLCNACGVRYKSGRL PEYRPAAS TF+P LHSNSH
Sbjct: 227 PLRKCMHCEVTKTPQWRLGPMGPKTLCNACGVRYKSGRLFPEYRPAASPTFTPALHSNSH 286
Query: 180 KKILEMRVMRRKD 192
KK+ EMR R D
Sbjct: 287 KKVAEMRNKRCSD 299
>UniRef100_Q948G4 Putative GATA-1 zinc finger protein [Oryza sativa]
Length = 418
Score = 129 bits (323), Expect = 6e-29
Identities = 77/169 (45%), Positives = 92/169 (53%), Gaps = 24/169 (14%)
Query: 35 RTRSKRPRLATFSSHHSTMQLISSTSSFVGENMQDSVISNKGASTEKFPDSQIAAKKQKL 94
R RSKR R + F + + T+ V M S S+ P+S +
Sbjct: 236 RARSKRSRPSAFPAVRGA-PAATETTILVPTPMYSSTSSHSD------PESIAESNPHPP 288
Query: 95 SSGESKKNKKTKAPLLAA---LDHNAL--------------GLVRQCTHCEATKTPQWRT 137
+ KK KK AP A+ D +A G VR+CTHC+ KTPQWR
Sbjct: 289 PMKKKKKAKKPAAPAAASDAEADADAADADYEEGGALALPPGTVRRCTHCQIEKTPQWRA 348
Query: 138 GPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKKILEMR 186
GP GPKTLCNACGVRYKSGRL PEYRPAAS TF P +HSNSHKK++EMR
Sbjct: 349 GPLGPKTLCNACGVRYKSGRLFPEYRPAASPTFMPSIHSNSHKKVVEMR 397
>UniRef100_Q8VZP4 Putative GATA transcription factor 3 [Arabidopsis thaliana]
Length = 308
Score = 125 bits (314), Expect = 7e-28
Identities = 61/105 (58%), Positives = 73/105 (69%), Gaps = 9/105 (8%)
Query: 96 SGESKKNKKTKAPLLAALDHNAL------GLVRQCTHCEATKTPQWRTGPEGPKTLCNAC 149
S E KK K L+ + + L G+VR CTHCE TPQWR GP GPKTLCNAC
Sbjct: 186 SSEQHAKKKRKIHLITHTESSTLESSKSDGIVRICTHCETITTPQWRQGPSGPKTLCNAC 245
Query: 150 GVRYKSGRLCPEYRPAASSTFSPDLHSNSHKKILEMRVMRRKDNK 194
GVR+KSGRL PEYRPA+S TF P +HSNSH+KI+E MR+KD++
Sbjct: 246 GVRFKSGRLVPEYRPASSPTFIPSVHSNSHRKIIE---MRKKDDE 287
>UniRef100_Q6F2Z7 Hypothetical protein P0483D07.7 [Oryza sativa]
Length = 386
Score = 125 bits (313), Expect = 9e-28
Identities = 80/179 (44%), Positives = 95/179 (52%), Gaps = 9/179 (5%)
Query: 8 PSSSVNKEDFVLPKSNSSPTCEKTTVRRTRSKRPRLATFSSHHSTMQLISSTSSFVGENM 67
P+ +V+ + + PT E + RSKR R+A S M L +S
Sbjct: 148 PALAVSAVQLPAGGAGALPT-EAPVPGKARSKRSRVAPCSWSSRLMVLPPPPASPPSPAS 206
Query: 68 QDSVISNKGASTEKFPDSQIAAKKQKLSSGESKKNKKTKAPLLAALDHNALGLVRQCTHC 127
S G + FP + AAK K G S AP AA A R+C HC
Sbjct: 207 AVISPSESGTAAPAFPAKK-AAKSAKKKDGPSP----APAPNAAA---QAAAEGRRCLHC 258
Query: 128 EATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKKILEMR 186
E KTPQWRTGP GPKTLCNACGVRYKSGRL PEYRPAAS TF HSNSH+K++E+R
Sbjct: 259 ETDKTPQWRTGPMGPKTLCNACGVRYKSGRLVPEYRPAASPTFVVSKHSNSHRKVVELR 317
>UniRef100_Q76DY0 AG-motif binding protein-4 [Nicotiana tabacum]
Length = 326
Score = 124 bits (312), Expect = 1e-27
Identities = 78/168 (46%), Positives = 95/168 (56%), Gaps = 34/168 (20%)
Query: 35 RTRSKRP--RLATFSSHHSTMQLISSTSSFVGENMQDSVISNKGA-------STEKFPDS 85
RT RP R +FSS + S TSS G + SV+ + S EK P
Sbjct: 169 RTYRSRPAGRKWSFSSPTVSADSCSPTSSSYGSSPFPSVLFSNPVLDGDLFCSVEKPP-- 226
Query: 86 QIAAKKQKLSSGESKKNKKTKAPLLAALDHNALGLVRQCTHCEATKTPQWRTGPEGPKTL 145
K +KLS+ E+ G R+CTHC+ KTPQWR GP GPKTL
Sbjct: 227 --LKKPKKLSTAET-------------------GSGRRCTHCQVQKTPQWRAGPLGPKTL 265
Query: 146 CNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKKILEMRVMRRKDN 193
CNACGVRYKSGRL PEYRPA S TFS ++HSNSH+K+LEMR R+K++
Sbjct: 266 CNACGVRYKSGRLFPEYRPACSPTFSQEVHSNSHRKVLEMR--RKKES 311
>UniRef100_Q94LQ6 Putative transcription factor [Oryza sativa]
Length = 387
Score = 123 bits (309), Expect = 3e-27
Identities = 73/156 (46%), Positives = 91/156 (57%), Gaps = 16/156 (10%)
Query: 34 RRTRSKRPRL--ATFSSHHSTMQLISSTSSFVGENMQDSVISNKGASTEKFPDS-QIAAK 90
R RSKR R A ++ HS + S SS + S+ FP S + +
Sbjct: 198 RGARSKRSRATAAAAAAWHSLVPRPPSQSS-----------PSSSCSSSDFPSSNKPSGT 246
Query: 91 KQKLSSGESKKNKKTKAPLLAALDHNALGLVRQCTHCEATKTPQWRTGPEGPKTLCNACG 150
+ SG + KK+ P A + A VR+CTHC + KTPQWRTGP GPKTLCNACG
Sbjct: 247 ARPNGSGGGSRGKKSPGPAGAEVGMEAG--VRRCTHCASEKTPQWRTGPLGPKTLCNACG 304
Query: 151 VRYKSGRLCPEYRPAASSTFSPDLHSNSHKKILEMR 186
VR+KSGRL PEYRPAAS TF HSNSH+K++E+R
Sbjct: 305 VRFKSGRLMPEYRPAASPTFVLTQHSNSHRKVMELR 340
>UniRef100_Q9AVU3 GATA-1 zinc finger protein [Nicotiana tabacum]
Length = 305
Score = 123 bits (308), Expect = 3e-27
Identities = 78/177 (44%), Positives = 93/177 (52%), Gaps = 28/177 (15%)
Query: 18 VLPKSNSSPTCEKTTVR-------RTRSKRPRLATFSSHHSTMQLISSTSSFVGENMQDS 70
VL SNS + +++ R RSKRPR + + M ISST +
Sbjct: 108 VLESSNSCSGGKSISIKHDIAIPVRPRSKRPRSSALNPW-ILMPPISSTRFASKKTCDAR 166
Query: 71 VISNKGASTEKFPDSQIA-AKKQKLSSGESKKNKKTKAPLLAALDHNALGLVRQCTHCEA 129
K QIA K+K +SG+ KK CTHC+
Sbjct: 167 KGKEKKRKMSLLSVPQIADVTKKKTTSGQQFSFKK-------------------CTHCQV 207
Query: 130 TKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKKILEMR 186
TKTPQWR GP GPKTLCNACGVRY+SGRL PEYRPAAS TF P LHSNSH+K++EMR
Sbjct: 208 TKTPQWREGPLGPKTLCNACGVRYRSGRLFPEYRPAASPTFVPTLHSNSHRKVVEMR 264
>UniRef100_Q6H635 Putative AG-motif binding protein-4 [Oryza sativa]
Length = 387
Score = 122 bits (307), Expect = 4e-27
Identities = 78/180 (43%), Positives = 96/180 (53%), Gaps = 22/180 (12%)
Query: 33 VRRTRSKRPRLATFSSHHSTMQ-LISSTSSFVGENMQDSVISNKGASTEKFP-------- 83
V+ RSKR R + +S + + SS+S+ + S + FP
Sbjct: 195 VKSRRSKRSRASVWSLSGAPLSDSTSSSSTATTSSCSSSASFSPFLQYVDFPALVASDLL 254
Query: 84 DSQIAAKKQKLSSGESKKNKKT-------KAPLLAALDHNALGLV----RQCTHCEATKT 132
D Q +KK K +K KK + P LAA L R+C+HC KT
Sbjct: 255 DEQPRSKKSKHGKNGKQKPKKRGRKPKHQQPPHLAAAAGGGAALPATGDRRCSHCGVQKT 314
Query: 133 PQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKKILEMRVMRRKD 192
PQWR GPEG KTLCNACGVRYKSGRL PEYRPA S TF LHSNSH+K+LEMR R+K+
Sbjct: 315 PQWRAGPEGAKTLCNACGVRYKSGRLLPEYRPACSPTFVSSLHSNSHRKVLEMR--RKKE 372
>UniRef100_Q76DY2 AG-motif binding protein-2 [Nicotiana tabacum]
Length = 289
Score = 122 bits (306), Expect = 6e-27
Identities = 59/90 (65%), Positives = 69/90 (76%), Gaps = 2/90 (2%)
Query: 99 SKKNKKTKAPLLAALDHNA--LGLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSG 156
S K+ K + PLL+ ++A + R+C HC A KTPQWR GP GPKTLCNACGVRYKSG
Sbjct: 180 SFKSVKQREPLLSLPLNSAKSASIGRRCQHCGADKTPQWRAGPLGPKTLCNACGVRYKSG 239
Query: 157 RLCPEYRPAASSTFSPDLHSNSHKKILEMR 186
RL PEYRPA S TFSP +HSNSH+K+LEMR
Sbjct: 240 RLLPEYRPANSPTFSPTVHSNSHRKVLEMR 269
>UniRef100_Q9FH57 GATA-binding transcription factor-like protein [Arabidopsis
thaliana]
Length = 339
Score = 121 bits (304), Expect = 1e-26
Identities = 76/195 (38%), Positives = 108/195 (54%), Gaps = 25/195 (12%)
Query: 6 QHPSSSVNKEDFVLPKSNSSPTCEKTTV-RRTRSKRPRLATFSSHHSTMQLISSTSSFVG 64
+HP ++V +E TC K+ V + RSKR R + + + S+SS
Sbjct: 148 KHPVTAVTEE-----------TCFKSPVPAKARSKRNR------NGLKVWSLGSSSSSGP 190
Query: 65 ENMQDSVISNKGASTEKFPDSQIAAKKQKLSSGES----KKNKKTKAPLLAALDHNALGL 120
+ + S+ G S+ F +++ + + + E KK+KK A + + + L
Sbjct: 191 SSSGSTSSSSSGPSSPWFSGAELL---EPVVTSERPPFPKKHKKRSAESVFSGELQQLQP 247
Query: 121 VRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHK 180
R+C+HC KTPQWR GP G KTLCNACGVRYKSGRL PEYRPA S TFS +LHSN H+
Sbjct: 248 QRKCSHCGVQKTPQWRAGPMGAKTLCNACGVRYKSGRLLPEYRPACSPTFSSELHSNHHR 307
Query: 181 KILEMRVMRRKDNKN 195
K++EMR + + N
Sbjct: 308 KVIEMRRKKEPTSDN 322
>UniRef100_Q9SD38 Putative transcription factor [Arabidopsis thaliana]
Length = 312
Score = 121 bits (303), Expect = 1e-26
Identities = 76/185 (41%), Positives = 92/185 (49%), Gaps = 15/185 (8%)
Query: 13 NKEDFVLPKSNSSPTCEKTTVRRTRSKRPRLATFSSHHSTMQLISSTSSFVGENMQDSVI 72
N+ V P + + +TR KR R H + L S+SS
Sbjct: 121 NRRHLVQPVKEETCFKSQHPAVKTRPKRARTGVRVWSHGSQSLTDSSSS----------- 169
Query: 73 SNKGASTEKFPDSQI-AAKKQKLSSGESKKNKKTKAPLLAALDHNALGL-VRQCTHCEAT 130
S +S+ P S + A Q L +K KK K A RQC HC
Sbjct: 170 STTSSSSSPRPSSPLWLASGQFLDEPMTKTQKKKKVWKNAGQTQTQTQTQTRQCGHCGVQ 229
Query: 131 KTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKKILEMRVMRR 190
KTPQWR GP G KTLCNACGVRYKSGRL PEYRPA S TFS +LHSN H K++EMR R+
Sbjct: 230 KTPQWRAGPLGAKTLCNACGVRYKSGRLLPEYRPACSPTFSSELHSNHHSKVIEMR--RK 287
Query: 191 KDNKN 195
K+ +
Sbjct: 288 KETSD 292
>UniRef100_Q8LIZ3 OSJNBa0014K08.18 protein [Oryza sativa]
Length = 387
Score = 119 bits (297), Expect = 6e-26
Identities = 69/155 (44%), Positives = 82/155 (52%), Gaps = 3/155 (1%)
Query: 35 RTRSKRPRLATFSSHHSTMQLISSTSSFVGENMQDSVISNKGASTEKFP---DSQIAAKK 91
+ RSKR R A + + L +S + G S FP S+ A KK
Sbjct: 172 KARSKRSRAAPGNWSSRLLVLPPPPASPPSPASMAISPAESGVSAHAFPIKKPSKPAKKK 231
Query: 92 QKLSSGESKKNKKTKAPLLAALDHNALGLVRQCTHCEATKTPQWRTGPEGPKTLCNACGV 151
+ + + A G R+C HCE KTPQWRTGP GPKTLCNACGV
Sbjct: 232 DAPAPPAQAQLSSVPVHSGGSAPAAAAGEGRRCLHCETDKTPQWRTGPMGPKTLCNACGV 291
Query: 152 RYKSGRLCPEYRPAASSTFSPDLHSNSHKKILEMR 186
RYKSGRL PEYRPAAS TF HSNSH+K+LE+R
Sbjct: 292 RYKSGRLVPEYRPAASPTFMVSKHSNSHRKVLELR 326
>UniRef100_Q8LAU9 GATA transcription factor 1 [Arabidopsis thaliana]
Length = 268
Score = 118 bits (296), Expect = 8e-26
Identities = 57/84 (67%), Positives = 65/84 (76%), Gaps = 3/84 (3%)
Query: 103 KKTKAPLLAALDHNALGLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEY 162
+K K +AA AL + R+C HC A KTPQWR GP GPKTLCNACGVRYKSGRL PEY
Sbjct: 172 QKKKTMTVAAA---ALIMGRKCQHCGAEKTPQWRAGPAGPKTLCNACGVRYKSGRLVPEY 228
Query: 163 RPAASSTFSPDLHSNSHKKILEMR 186
RPA S TF+ +LHSNSH+KI+EMR
Sbjct: 229 RPANSPTFTAELHSNSHRKIVEMR 252
>UniRef100_O49740 Protein homologous to GATA-binding transcription factors
[Arabidopsis thaliana]
Length = 274
Score = 118 bits (296), Expect = 8e-26
Identities = 57/84 (67%), Positives = 65/84 (76%), Gaps = 3/84 (3%)
Query: 103 KKTKAPLLAALDHNALGLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEY 162
+K K +AA AL + R+C HC A KTPQWR GP GPKTLCNACGVRYKSGRL PEY
Sbjct: 178 QKKKTMTVAAA---ALIMGRKCQHCGAEKTPQWRAGPAGPKTLCNACGVRYKSGRLVPEY 234
Query: 163 RPAASSTFSPDLHSNSHKKILEMR 186
RPA S TF+ +LHSNSH+KI+EMR
Sbjct: 235 RPANSPTFTAELHSNSHRKIVEMR 258
>UniRef100_Q8H036 Hypothetical protein OJ1172F09.7 [Oryza sativa]
Length = 219
Score = 117 bits (294), Expect = 1e-25
Identities = 49/71 (69%), Positives = 60/71 (84%)
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKK 181
R+CTHC +TPQWR GP+GP+TLCNACGVR+KSGRL PEYRPA S TFSP LHSNSH++
Sbjct: 122 RRCTHCAVDETPQWRLGPDGPRTLCNACGVRFKSGRLFPEYRPANSPTFSPLLHSNSHRR 181
Query: 182 ILEMRVMRRKD 192
++EMR+ +D
Sbjct: 182 VMEMRLQSEED 192
>UniRef100_O49743 Homologous to GATA-binding transcription factors [Arabidopsis
thaliana]
Length = 240
Score = 116 bits (290), Expect = 4e-25
Identities = 52/75 (69%), Positives = 62/75 (82%), Gaps = 2/75 (2%)
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKK 181
R+CTHC + KTPQWRTGP GPKTLCNACGVRYKSGRL PEYRPA+S TF HSNSH+K
Sbjct: 158 RRCTHCASEKTPQWRTGPLGPKTLCNACGVRYKSGRLVPEYRPASSPTFVLTQHSNSHRK 217
Query: 182 ILEMRVMRRKDNKNS 196
++E+R R+K+ + S
Sbjct: 218 VMELR--RQKEQQES 230
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.311 0.124 0.353
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 324,285,671
Number of Sequences: 2790947
Number of extensions: 12137218
Number of successful extensions: 31212
Number of sequences better than 10.0: 462
Number of HSP's better than 10.0 without gapping: 161
Number of HSP's successfully gapped in prelim test: 301
Number of HSP's that attempted gapping in prelim test: 30591
Number of HSP's gapped (non-prelim): 688
length of query: 204
length of database: 848,049,833
effective HSP length: 121
effective length of query: 83
effective length of database: 510,345,246
effective search space: 42358655418
effective search space used: 42358655418
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 72 (32.3 bits)
Medicago: description of AC146329.15


