

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146329.10 - phase: 0
(68 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
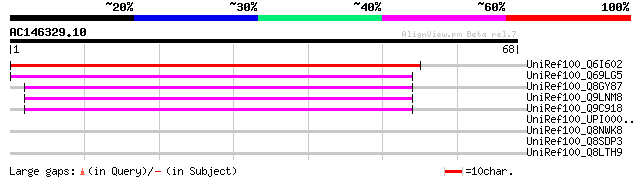
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6I602 Putative ubiquitin-conjugating enzyme [Oryza sa... 56 2e-07
UniRef100_Q69LG5 Ubiquitin-conjugating enzyme-like protein [Oryz... 54 1e-06
UniRef100_Q8GY87 Putative ubiquitin-conjugating enzyme [Arabidop... 45 4e-04
UniRef100_Q9LNM8 F8L10.11 protein [Arabidopsis thaliana] 45 4e-04
UniRef100_Q9C918 Putative ubiquitin-conjugating enzyme; 121361-1... 45 6e-04
UniRef100_UPI000032F31D UPI000032F31D UniRef100 entry 32 4.2
UniRef100_Q8NWK8 Hypothetical protein MW1390 [Staphylococcus aur... 31 7.1
UniRef100_Q8SDP3 Tail fiber protein [Staphylococcus aureus phage... 31 7.1
UniRef100_Q8LTH9 Phi12 tail fiber protein-like protein [Staphylo... 31 7.1
>UniRef100_Q6I602 Putative ubiquitin-conjugating enzyme [Oryza sativa]
Length = 509
Score = 56.2 bits (134), Expect = 2e-07
Identities = 27/55 (49%), Positives = 37/55 (67%)
Query: 1 MVAGNAYIEGAQVGCLVKGGVQDLDDGDKSSSRQFNNSLARHINVAAPNFTGIGS 55
++A AY++GAQVGCLV GVQD+D+GDKS S +F ++L R FT G+
Sbjct: 430 LIACRAYLDGAQVGCLVGNGVQDVDEGDKSCSARFKSALKRLFEELLMEFTVKGA 484
>UniRef100_Q69LG5 Ubiquitin-conjugating enzyme-like protein [Oryza sativa]
Length = 433
Score = 53.9 bits (128), Expect = 1e-06
Identities = 27/54 (50%), Positives = 32/54 (59%)
Query: 1 MVAGNAYIEGAQVGCLVKGGVQDLDDGDKSSSRQFNNSLARHINVAAPNFTGIG 54
+V AY+ GAQVGCL GVQD+D+GDKS S F SL + FT IG
Sbjct: 354 LVGCKAYMNGAQVGCLAGNGVQDVDEGDKSCSANFKGSLKTLFDELIKEFTRIG 407
>UniRef100_Q8GY87 Putative ubiquitin-conjugating enzyme [Arabidopsis thaliana]
Length = 543
Score = 45.4 bits (106), Expect = 4e-04
Identities = 21/52 (40%), Positives = 31/52 (59%)
Query: 3 AGNAYIEGAQVGCLVKGGVQDLDDGDKSSSRQFNNSLARHINVAAPNFTGIG 54
A NAY GA +G +VKGGVQDL++ +S S++F +A + F +G
Sbjct: 459 ACNAYKAGAPLGSMVKGGVQDLEEARQSGSKKFKTDVASFMQTVVDEFVKLG 510
>UniRef100_Q9LNM8 F8L10.11 protein [Arabidopsis thaliana]
Length = 543
Score = 45.4 bits (106), Expect = 4e-04
Identities = 21/52 (40%), Positives = 31/52 (59%)
Query: 3 AGNAYIEGAQVGCLVKGGVQDLDDGDKSSSRQFNNSLARHINVAAPNFTGIG 54
A NAY GA +G +VKGGVQDL++ +S S++F +A + F +G
Sbjct: 459 ACNAYKAGAPLGSMVKGGVQDLEEARQSGSKKFKTDVASFMQTVVDEFVKLG 510
>UniRef100_Q9C918 Putative ubiquitin-conjugating enzyme; 121361-120434 [Arabidopsis
thaliana]
Length = 244
Score = 44.7 bits (104), Expect = 6e-04
Identities = 21/52 (40%), Positives = 30/52 (57%)
Query: 3 AGNAYIEGAQVGCLVKGGVQDLDDGDKSSSRQFNNSLARHINVAAPNFTGIG 54
A NAY GA +G +VKGGVQDL+ +S S++F +A + F +G
Sbjct: 160 ACNAYKAGAPLGSMVKGGVQDLEQARQSGSKKFKTDVASFMQTVVDEFVKLG 211
>UniRef100_UPI000032F31D UPI000032F31D UniRef100 entry
Length = 384
Score = 32.0 bits (71), Expect = 4.2
Identities = 16/44 (36%), Positives = 25/44 (56%), Gaps = 3/44 (6%)
Query: 4 GNAYIEGAQVGCLVKGGVQDLDDG---DKSSSRQFNNSLARHIN 44
GN I+ + G L+KGG + G K+S +FNN L +H++
Sbjct: 248 GNIIIQAKEKGILIKGGGHKMAAGLKIKKTSLNEFNNFLVQHLD 291
>UniRef100_Q8NWK8 Hypothetical protein MW1390 [Staphylococcus aureus]
Length = 2066
Score = 31.2 bits (69), Expect = 7.1
Identities = 17/61 (27%), Positives = 32/61 (51%), Gaps = 1/61 (1%)
Query: 2 VAGNAYIEGAQVGCLVKGGVQDLDDGDKSSSRQFNNSLARHINVAAPNFTGIGSMLVQAP 61
V N+ +EG+Q G ++ L + KS++++ L H++ A F G+G ++ Q
Sbjct: 482 VLSNSGLEGSQAGTALRASFIRLANPSKSTAKEM-KKLGIHLSDAKGEFVGMGELIRQFQ 540
Query: 62 D 62
D
Sbjct: 541 D 541
>UniRef100_Q8SDP3 Tail fiber protein [Staphylococcus aureus phage phi 12]
Length = 2066
Score = 31.2 bits (69), Expect = 7.1
Identities = 17/61 (27%), Positives = 32/61 (51%), Gaps = 1/61 (1%)
Query: 2 VAGNAYIEGAQVGCLVKGGVQDLDDGDKSSSRQFNNSLARHINVAAPNFTGIGSMLVQAP 61
V N+ +EG+Q G ++ L + KS++++ L H++ A F G+G ++ Q
Sbjct: 482 VLSNSGLEGSQAGTALRASFIRLANPSKSTAKEM-KKLGIHLSDAKGEFVGMGELIRQFQ 540
Query: 62 D 62
D
Sbjct: 541 D 541
>UniRef100_Q8LTH9 Phi12 tail fiber protein-like protein [Staphylococcus aureus
bacteriophage phi 3A]
Length = 2066
Score = 31.2 bits (69), Expect = 7.1
Identities = 17/61 (27%), Positives = 32/61 (51%), Gaps = 1/61 (1%)
Query: 2 VAGNAYIEGAQVGCLVKGGVQDLDDGDKSSSRQFNNSLARHINVAAPNFTGIGSMLVQAP 61
V N+ +EG+Q G ++ L + KS++++ L H++ A F G+G ++ Q
Sbjct: 482 VLSNSGLEGSQAGTALRASFIRLANPSKSTAKEM-KKLGIHLSDAKGQFVGMGELIRQFQ 540
Query: 62 D 62
D
Sbjct: 541 D 541
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.135 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 110,461,813
Number of Sequences: 2790947
Number of extensions: 3401684
Number of successful extensions: 6817
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 6811
Number of HSP's gapped (non-prelim): 9
length of query: 68
length of database: 848,049,833
effective HSP length: 44
effective length of query: 24
effective length of database: 725,248,165
effective search space: 17405955960
effective search space used: 17405955960
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC146329.10


