

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145513.12 + phase: 0
(304 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
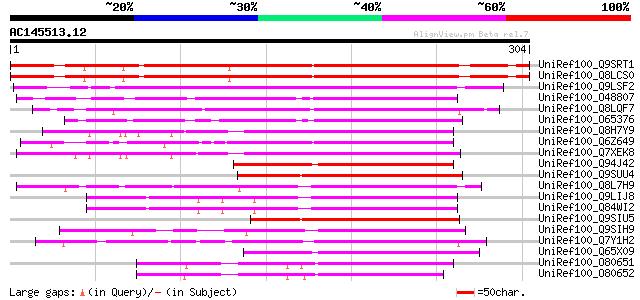
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SRT1 F21O3.6 protein [Arabidopsis thaliana] 260 3e-68
UniRef100_Q8LCS0 Hypothetical protein [Arabidopsis thaliana] 260 3e-68
UniRef100_Q9LSF2 Gb|AAF02145.1 [Arabidopsis thaliana] 210 4e-53
UniRef100_O48807 F2401.16 [Arabidopsis thaliana] 156 7e-37
UniRef100_Q8LQF7 Similar to Arabidopsis thaliana chromosome 3, A... 150 5e-35
UniRef100_O65376 F12F1.10 protein [Arabidopsis thaliana] 149 1e-34
UniRef100_Q8H7Y9 Hypothetical protein OJ1607A12.9 [Oryza sativa] 122 9e-27
UniRef100_Q6Z649 Hypothetical protein P0654B04.2 [Oryza sativa] 122 2e-26
UniRef100_Q7XEK8 Hypothetical protein OSJNBa0061K21.17 [Oryza sa... 118 2e-25
UniRef100_Q94J42 P0481E12.6 protein [Oryza sativa] 115 2e-24
UniRef100_Q9SUU4 Hypothetical protein F8B4.180 [Arabidopsis thal... 110 6e-23
UniRef100_Q8L7H9 Hypothetical protein At4g14620 [Arabidopsis tha... 109 8e-23
UniRef100_Q9LIJ8 Arabidopsis thaliana genomic DNA, chromosome 3,... 109 1e-22
UniRef100_Q84WI2 Hypothetical protein At3g22970 [Arabidopsis tha... 107 4e-22
UniRef100_Q9SIU5 Expressed protein [Arabidopsis thaliana] 105 1e-21
UniRef100_Q9SIH9 Expressed protein [Arabidopsis thaliana] 105 2e-21
UniRef100_Q7Y1H2 Hypothetical protein OSJNBa0094F01.21 [Oryza sa... 103 6e-21
UniRef100_Q65X09 Hypothetical protein P0599F04.9 [Oryza sativa] 102 1e-20
UniRef100_O80651 T14N5.3 protein [Arabidopsis thaliana] 102 1e-20
UniRef100_O80652 T14N5.4 protein [Arabidopsis thaliana] 102 1e-20
>UniRef100_Q9SRT1 F21O3.6 protein [Arabidopsis thaliana]
Length = 298
Score = 260 bits (665), Expect = 3e-68
Identities = 148/317 (46%), Positives = 196/317 (61%), Gaps = 32/317 (10%)
Query: 1 MNFARKKRVTDPFNDEAKARLVGADLRRLSDVSSGSEHSGTG----ECDSSPSLSELVHG 56
M F+R KRVTDP +E +ARLVG SSGSEH+G G E D SP LS+LV G
Sbjct: 1 MGFSRAKRVTDPLAEEVRARLVGCSF------SSGSEHTGDGIEDYEDDDSPCLSDLVQG 54
Query: 57 FLEEDDGNV------CDS-TGNDFDSERVDSVSDSMDSVEDLLRLSAENANADSYLNMLR 109
FLE++ V CD +G+D DS+ + + D +D+ +L + DSY +
Sbjct: 55 FLEDEVDTVDDESCWCDQDSGSDSDSD--SELGELPDFADDIAKLLRNSLREDSYGRTVL 112
Query: 110 LHVSEAAVKFDFLKKQSV--SVYNRNVMSFLREKGHNAAICKTRWDSSGGLTAGNHEFID 167
+HV+ A L Q +V+ R VMS LRE GHNAAICKT+W SSGGLTAGNHEFID
Sbjct: 113 VHVARAMEMLSSLGSQPEQRAVFQRKVMSLLRELGHNAAICKTKWKSSGGLTAGNHEFID 172
Query: 168 VVRMRSGSSTWQNRYFVELDFAVQFEIARPTSRYSEIMSYVPGIFVGNSEELKRTVLALC 227
VV S SS R+ V+LDF+ +F+IARPTS+Y+ ++ +P +FVG ++LKR + +C
Sbjct: 173 VVYTPSASSQ-SVRFIVDLDFSSRFQIARPTSQYARVLQSLPAVFVGKGDDLKRILRLVC 231
Query: 228 GAVKLCLRSRGLSIPPWRKNRYMQNKWFGPYRRTTNPVHGNPVPDVVSSFSGVKCRLVGF 287
A ++ LR+RGL++PPWRKNRYMQ +W GPY+RTTN + S V CR +GF
Sbjct: 232 DAARISLRNRGLTLPPWRKNRYMQTRWLGPYKRTTN------LTPSTSGVDTVMCRAIGF 285
Query: 288 DNAMSEIKHGVAFVRIK 304
DNA+ G FVR +
Sbjct: 286 DNAVG----GRLFVRTR 298
>UniRef100_Q8LCS0 Hypothetical protein [Arabidopsis thaliana]
Length = 298
Score = 260 bits (665), Expect = 3e-68
Identities = 148/317 (46%), Positives = 196/317 (61%), Gaps = 32/317 (10%)
Query: 1 MNFARKKRVTDPFNDEAKARLVGADLRRLSDVSSGSEHSGTG----ECDSSPSLSELVHG 56
M F+R KRVTDP +E +ARLVG SSGSEH+G G E D SP LS+LV G
Sbjct: 1 MGFSRAKRVTDPLAEEVRARLVGCSF------SSGSEHTGDGIEDYEDDDSPCLSDLVQG 54
Query: 57 FLEEDDGNV------CDS-TGNDFDSERVDSVSDSMDSVEDLLRLSAENANADSYLNMLR 109
FLE++ V CD +G+D DS+ + + D +D+ +L + DSY +
Sbjct: 55 FLEDEVDTVDDESCWCDQDSGSDSDSD--SELGELPDFADDIAKLLRNSLREDSYGRTVL 112
Query: 110 LHVSEAAVKFDFLKKQSV--SVYNRNVMSFLREKGHNAAICKTRWDSSGGLTAGNHEFID 167
+HV+ A L Q +V+ R VMS LRE GHNAAICKT+W SSGGLTAGNHEFID
Sbjct: 113 VHVARAMZXLSSLGSQPEQRAVFQRKVMSLLRELGHNAAICKTKWKSSGGLTAGNHEFID 172
Query: 168 VVRMRSGSSTWQNRYFVELDFAVQFEIARPTSRYSEIMSYVPGIFVGNSEELKRTVLALC 227
VV S SS R+ V+LDF+ +F+IARPTS+Y+ ++ +P +FVG ++LKR + +C
Sbjct: 173 VVYTPSASSQ-SVRFIVDLDFSSRFQIARPTSQYARVLQSLPAVFVGKGDDLKRILRLVC 231
Query: 228 GAVKLCLRSRGLSIPPWRKNRYMQNKWFGPYRRTTNPVHGNPVPDVVSSFSGVKCRLVGF 287
A ++ LR+RGL++PPWRKNRYMQ +W GPY+RTTN + S V CR +GF
Sbjct: 232 DAARISLRNRGLTLPPWRKNRYMQTRWLGPYKRTTN------LTPSTSGVDTVMCRAIGF 285
Query: 288 DNAMSEIKHGVAFVRIK 304
DNA+ G FVR +
Sbjct: 286 DNAVG----GRLFVRTR 298
>UniRef100_Q9LSF2 Gb|AAF02145.1 [Arabidopsis thaliana]
Length = 281
Score = 210 bits (534), Expect = 4e-53
Identities = 117/287 (40%), Positives = 168/287 (57%), Gaps = 20/287 (6%)
Query: 3 FARKKRVTDPFNDEAKARLVGADLRRLSDVSSGSEHSGTGECDSSPSLSELVHGFLEEDD 62
F+R KRVTDP +D+AKAR++ S HS + DS P L ELVHGFLE D
Sbjct: 4 FSRTKRVTDPLDDDAKARIL-------------SSHSYILDQDS-PRLCELVHGFLE--D 47
Query: 63 GNVCDSTGNDFDSERVDSVSDSMDSVEDLLRLSAENANADSYLNMLRLHVSEAAVKFDFL 122
G +D D S S ++ +++R++ +++D Y N+L HV A +
Sbjct: 48 GPEESFYDSDPDLSETSSAEHSGEASVEIVRMAVSFSDSDPYRNLLLAHVLRAVEVYSGF 107
Query: 123 KKQSVSVYNRNVMSFLREKGHNAAICKTRWDSSGGLTAGNHEFIDVVRMRSGSSTWQNRY 182
+ ++ +V+ V SFLRE GH+AA+C ++W SS L AG++ FIDVV S + RY
Sbjct: 108 RSRNKTVFRDKVASFLRELGHDAAVCVSKWTSSSKLIAGSYHFIDVVHKPSDNDQKAVRY 167
Query: 183 FVELDFAVQFEIARPTSRYSEIMSYVPGIFVGNSEELKRTVLALCGAVKLCLRSRGLSIP 242
V+LDFA +FEIARPT Y+ + +P +FVGN E L+ V C A K ++SRGLS+P
Sbjct: 168 LVDLDFASEFEIARPTREYTRGLQLLPNVFVGNEENLRTIVRESCDAAKRSMKSRGLSLP 227
Query: 243 PWRKNRYMQNKWFGPYRRTTNPVHGNPVPDVVSSFSGVKCRLVGFDN 289
PWR++ Y+Q+KWFGPY+R G+ + + V CR +GFD+
Sbjct: 228 PWRRSSYLQHKWFGPYKRKV----GSSLGVKPLNSDAVSCRSLGFDD 270
>UniRef100_O48807 F2401.16 [Arabidopsis thaliana]
Length = 283
Score = 156 bits (394), Expect = 7e-37
Identities = 95/258 (36%), Positives = 140/258 (53%), Gaps = 28/258 (10%)
Query: 5 RKKRVTDPFNDEAKARLVGADLRRLSDVSSGSEHSGTGECDSSPSLSELVHGFLEEDDGN 64
R KR+ FN + A + D SSGS+HS P LS+LV F+E++
Sbjct: 7 RFKRIESAFN------VAAARVHPPCDNSSGSDHS--------PDLSDLVASFIEKEGQI 52
Query: 65 VCDSTGNDFDSERVDSVSDSMDSVEDLLRLSAENANADSYLNMLRLHVSEAAVKFDFLKK 124
V + E S ++++ V + LR E + R+ + A ++
Sbjct: 53 VLR------EEEETSSDDNNLEDVNERLRKLLEGLSCGEE----RMRILSATMEVAGTFV 102
Query: 125 QSVSVYNRNVMSFLREKGHNAAICKTRWDSSGGLTAGNHEFIDVVRMRSGSSTWQNRYFV 184
+S R++M+FLR KG +A +CK+ W+ G T G +E++DV R G + NRYFV
Sbjct: 103 GDISSSKRHLMAFLRNKGFDAGLCKSSWERFGKNTGGKYEYVDV---RCGGD-YNNRYFV 158
Query: 185 ELDFAVQFEIARPTSRYSEIMSYVPGIFVGNSEELKRTVLALCGAVKLCLRSRGLSIPPW 244
E + A +FEIARPT RY I+S VP +FVG SEELK V +C ++ ++ G+ +PPW
Sbjct: 159 ETNLAGEFEIARPTKRYLSILSQVPRVFVGTSEELKLLVRIMCHEMRRSMKHVGIHVPPW 218
Query: 245 RKNRYMQNKWFGPYRRTT 262
R+N YMQ KWFG Y+RT+
Sbjct: 219 RRNGYMQAKWFGFYKRTS 236
>UniRef100_Q8LQF7 Similar to Arabidopsis thaliana chromosome 3, At3g07350 [Oryza
sativa]
Length = 293
Score = 150 bits (378), Expect = 5e-35
Identities = 99/286 (34%), Positives = 146/286 (50%), Gaps = 25/286 (8%)
Query: 14 NDEAKARLVGADLRRLSDVSSGSEHSGTGECDSSPSLSELVHGFLE-------EDDGNVC 66
+D +ARL G S SSGSEH ++S LS LV FLE ED
Sbjct: 8 DDAVRARLRGD---AGSSASSGSEH------EASACLSGLVQAFLETEGAAAGEDGAGPA 58
Query: 67 DSTGNDFDSERVDSVSDSMDSVEDLLRLSAENANADSYLNMLRLHVSEAAVKFDFLKKQS 126
G +DS+ D + + E + L D + L V+ AA++ + ++
Sbjct: 59 SKGGEGYDSDDGDGPERAAAAAESVRELLDPPVEEDPFRVRLAAAVA-AAMEAEPALRRY 117
Query: 127 VSVYNRNVMSFLREKGHNAAICKTRWDSSGGLTAGNHEFIDVVRMRSGSSTWQNRYFVEL 186
+ + R V LR G++A +CK+RW++SGG+TAG +E++DVV + ++RY V+
Sbjct: 118 GAAFRRAVARRLRAAGYDAGVCKSRWEASGGITAGTYEYVDVVAPAARGQ--KSRYIVDA 175
Query: 187 DFAVQFEIARPTSRYSEIMSYVPG-IFVGNSEELKRTVLALCGAVKLCLRSRGLSIPPWR 245
DF E+AR T+ Y+ +++ VP + V E + R V A + LRS GL +PPWR
Sbjct: 176 DFRAGLEVARATAEYAVVVAAVPASVVVAREEAVGRAVRVAADAARRSLRSHGLHVPPWR 235
Query: 246 KNRYMQNKWFGPYRRTT----NPVHGNPVPDVVSSFSGVKCRLVGF 287
K RYM KW GPY+R+T + P+P + VKCR VGF
Sbjct: 236 KTRYMLAKWLGPYKRSTATSPSAAGAMPMPAAAAGMD-VKCRAVGF 280
>UniRef100_O65376 F12F1.10 protein [Arabidopsis thaliana]
Length = 295
Score = 149 bits (375), Expect = 1e-34
Identities = 84/233 (36%), Positives = 132/233 (56%), Gaps = 22/233 (9%)
Query: 33 SSGSEHSGTGECDSSPSLSELVHGFLEEDDGNVCDSTGNDFDSERVDSVSDSMDSVEDLL 92
SSGS+HS D + L +LV F++ + + + F E D + + V++ L
Sbjct: 29 SSGSDHSP----DDTEDLWDLVESFIDREVETLPEDA---FQEEEDDKSDEDYEDVKERL 81
Query: 93 RLSAENANADSYLNMLRLHVSEAAVKFDFLKKQSVSVYNRNVMSFLREKGHNAAICKTRW 152
R EN + R + + AV + + R+ M++LR KG +A +CK+RW
Sbjct: 82 REILENHGGEE-----RQRIMDEAVN----ASRVFAGEKRHFMAYLRNKGFDAGLCKSRW 132
Query: 153 DSSGGLTAGNHEFIDVVRMRSGSSTWQNRYFVELDFAVQFEIARPTSRYSEIMSYVPGIF 212
+ G TAG +E++DV ++G +NRY VE + A +FEIARPT+RY +++ VP +F
Sbjct: 133 EKFGKNTAGKYEYVDV---KAGD---KNRYIVETNLAGEFEIARPTTRYLSVLAQVPRVF 186
Query: 213 VGNSEELKRTVLALCGAVKLCLRSRGLSIPPWRKNRYMQNKWFGPYRRTTNPV 265
VG EELK+ V +C ++ ++ + +PPWR+N YMQ KWFG Y+RT+N V
Sbjct: 187 VGTPEELKQLVRIMCFEIRRSMKRADIFVPPWRRNGYMQAKWFGHYKRTSNEV 239
>UniRef100_Q8H7Y9 Hypothetical protein OJ1607A12.9 [Oryza sativa]
Length = 307
Score = 122 bits (307), Expect = 9e-27
Identities = 83/266 (31%), Positives = 126/266 (47%), Gaps = 35/266 (13%)
Query: 20 RLVGADLRRLSDVSSGSEHSGTGECD-----SSPSLSELVHGFLEEDDG--------NVC 66
RL L R+S +G GE D SS L +V FLE+ G C
Sbjct: 25 RLFERQLLRVSPAERLPSVAGVGEKDESSEPSSVCLDGMVRSFLEDGVGVERPAGAARCC 84
Query: 67 ------DSTGNDFD---SERVDSVSDSMDSVEDLLR---LSAENANADSYLNMLRLHVSE 114
+++ +D D + + SD+ ++++ L+ L N AD ++ H +
Sbjct: 85 NCFHGGEASDDDDDGPAAAEAAATSDAAETIKGLVHCASLRERNLLAD-VSTLVERHRAA 143
Query: 115 AAVKFDFLKKQSVSVYNRNVMSFLREKGHNAAICKTRWDSSGGLTAGNHEFIDVVRMRSG 174
A K D L+ + S LR GH+AA+C +RWD S G H ++DV+ +
Sbjct: 144 GARKRDLLRLLADS---------LRAAGHDAAVCISRWDKSSSHPKGEHAYLDVLLPPAS 194
Query: 175 SSTWQNRYFVELDFAVQFEIARPTSRYSEIMSYVPGIFVGNSEELKRTVLALCGAVKLCL 234
+ R V++DF +FE+ARPT Y ++ +P +FVG + L+ V A A + L
Sbjct: 195 DRAERERILVDVDFRSEFEVARPTKAYRAVLQRLPSVFVGKEDRLRLLVAAAADAARASL 254
Query: 235 RSRGLSIPPWRKNRYMQNKWFGPYRR 260
+ RGL +PPWRK YM+ KW PY R
Sbjct: 255 KKRGLHLPPWRKPEYMRAKWLSPYER 280
>UniRef100_Q6Z649 Hypothetical protein P0654B04.2 [Oryza sativa]
Length = 287
Score = 122 bits (305), Expect = 2e-26
Identities = 84/262 (32%), Positives = 132/262 (50%), Gaps = 30/262 (11%)
Query: 7 KRVTDPFNDEAKARLVG------ADLRRLSDVSSGSEHSGTGECDSSPSLSELVHGFLEE 60
K+ T ++ A+ARL G A LRR D D L +LVH F
Sbjct: 4 KKATAVLDEAARARLRGPFASGAASLRRDQDD------------DDDDLLVDLVHEFY-- 49
Query: 61 DDGNVCDSTGNDFDSERVDSVSDSMDSVE--DLLRLSAENANADSYLNMLRLHVSEAAVK 118
DDG G D + S S + E D LR + +A +D+ +R +E AV+
Sbjct: 50 DDGE----RGADATARGGVSSSPEPEPTEWKDALREALADATSDAAAARIRAE-AERAVR 104
Query: 119 FDFLKKQSVSVYNRNVMSFLREKGHNAAICKTRWDSSGGLTAGNHEFIDVVRMRSGSSTW 178
D ++ V + V+ LR +G +A +C++ W+ +G + AG+HE++DV S +
Sbjct: 105 -DAVRNGG-DVIRKRVVERLRARGFDAGVCRSSWERTGSVPAGSHEYVDVTAAASATGR- 161
Query: 179 QNRYFVELDFAVQFEIARPTSRYSEIMSYVPGIFVGNSEELKRTVLALCGAVKLCLRSRG 238
+ RY VE++ A +FEIARP++ Y +++ +P + V E + A+C A +R G
Sbjct: 162 RARYIVEVNVAGEFEIARPSAEYQDLLLSLPPVLVATPEAFRGVAAAMCAAAAESIRGAG 221
Query: 239 LSIPPWRKNRYMQNKWFGPYRR 260
+ +PPWR+ RY+Q KW PY R
Sbjct: 222 MHLPPWRRARYVQAKWSAPYER 243
>UniRef100_Q7XEK8 Hypothetical protein OSJNBa0061K21.17 [Oryza sativa]
Length = 301
Score = 118 bits (295), Expect = 2e-25
Identities = 90/288 (31%), Positives = 131/288 (45%), Gaps = 38/288 (13%)
Query: 5 RKKRVTDPFNDEAKARLVGADLRRLSDVSSGSE--HSGTGECD----SSPSLSELVHGFL 58
R DP A++ L R+L VS G GE D SS L +V FL
Sbjct: 2 RAAAAPDPSPAPARSMLKRLFDRQLLRVSPAERIVAVGGGEKDEVEPSSVCLDGMVRSFL 61
Query: 59 EEDDG-------------NVCD------STGNDFDSERVDSVSDSMDSVEDLLR---LSA 96
E+ G C+ S+ +D D + + SD ++++ L+ L
Sbjct: 62 EDGSGVGAAVERAGGHGARRCNCFHGGGSSDDDDDEDDAAASSDVAETIKGLVHCATLRE 121
Query: 97 ENANADSYLNMLRLHVSEAAVKFDFLKKQSVSVYNRNVMSFLREKGHNAAICKTRWDSSG 156
N AD ++ R H + A + + L + S LR GH+AA+C +RWD S
Sbjct: 122 RNLLADVCGHVER-HRAGGARRRELLGLVAAS---------LRAAGHDAAVCVSRWDKSP 171
Query: 157 GLTAGNHEFIDVVRMRSGSSTWQNRYFVELDFAVQFEIARPTSRYSEIMSYVPGIFVGNS 216
AG H ++DV+ + + R V++DF FE+ARPT Y ++ +P +FVG
Sbjct: 172 THPAGEHAYVDVLLPPASDRGARERVLVDVDFRSAFEVARPTKAYRALLQRLPAVFVGKD 231
Query: 217 EELKRTVLALCGAVKLCLRSRGLSIPPWRKNRYMQNKWFGPYRRTTNP 264
+ L+ V A A + LR RGL +PPWRK YM+ KW PY R P
Sbjct: 232 DRLRLLVAASADAARASLRKRGLHLPPWRKPEYMRAKWLSPYDREPAP 279
>UniRef100_Q94J42 P0481E12.6 protein [Oryza sativa]
Length = 267
Score = 115 bits (287), Expect = 2e-24
Identities = 52/130 (40%), Positives = 85/130 (65%), Gaps = 4/130 (3%)
Query: 132 RNVMSFLREKGHNAAICKTRWDSSGGLTAGNHEFIDVV-RMRSGSSTWQNRYFVELDFAV 190
R+V LR+ G+N+AICK++W S + +G H ++DVV + RSG + R VEL+F
Sbjct: 104 RHVDERLRDAGYNSAICKSKWTRSPDIPSGEHSYVDVVVQTRSGKAV---RVVVELNFRA 160
Query: 191 QFEIARPTSRYSEIMSYVPGIFVGNSEELKRTVLALCGAVKLCLRSRGLSIPPWRKNRYM 250
+FE+AR ++ Y +++ +P +FVG ++ L+ V A+C A K C++ + + PWRK++YM
Sbjct: 161 EFEVARASAEYRALVTALPEVFVGRADRLRAVVKAMCAAAKQCMKENNMHMGPWRKHKYM 220
Query: 251 QNKWFGPYRR 260
Q+KW G R
Sbjct: 221 QSKWLGTPER 230
>UniRef100_Q9SUU4 Hypothetical protein F8B4.180 [Arabidopsis thaliana]
Length = 287
Score = 110 bits (274), Expect = 6e-23
Identities = 53/133 (39%), Positives = 84/133 (62%), Gaps = 2/133 (1%)
Query: 134 VMSFLREKGHNAAICKTRWDSSGGLTAGNHEFIDVVRMRSGSSTWQNRYFVELDFAVQFE 193
V S LRE G++ I K++W SS + AG HE+++VV +S S + R +EL F +FE
Sbjct: 120 VSSLLREAGYDCVISKSKWRSSHEIPAGEHEYLEVVD-KSVSKKGEIRVVIELCFRAEFE 178
Query: 194 IARPTSRYSEIMSYVPGIFVGNSEELKRTVLALCGAVKLCLRSRGLSIPPWRKNRYMQNK 253
+AR + Y ++ +P ++VG +E LK + LC A K C++ + + + PWRK++YMQ K
Sbjct: 179 MARGSEEYKRLIGMLPEVYVGKTERLKSLIKILCTAAKKCMKDKKMHMGPWRKHKYMQAK 238
Query: 254 WFGP-YRRTTNPV 265
WFG R++ +PV
Sbjct: 239 WFGTCERKSVSPV 251
>UniRef100_Q8L7H9 Hypothetical protein At4g14620 [Arabidopsis thaliana]
Length = 341
Score = 109 bits (273), Expect = 8e-23
Identities = 79/282 (28%), Positives = 138/282 (48%), Gaps = 28/282 (9%)
Query: 5 RKKRVTDPFNDEAKARLVGADLRRLSD-----VSSGSEHSGTGECDSSPSLSELVHGFLE 59
R + T P RL+ R+S+ +S +GT + PSL+++V ++E
Sbjct: 15 RVESSTKPVLKSRLKRLLDRPFTRISNSEKLLISGDGVVAGT---EFEPSLAKMVQNYME 71
Query: 60 EDDGNVCDSTGNDFDSERVDSVSDSMDSVEDLLRLSAENANADSYLNMLRLHVSEAAVKF 119
E++ T N ++ R + + + D +D L + N S + V+
Sbjct: 72 ENNDK---QTKNGRNTHRCNCFNGNNDISDDELDFF-DYDNFKSLIQCGSFVEKSLLVEA 127
Query: 120 DFLKKQSVSVYNRN-----VMSFLREKGHNAAICKTRWDSSGGLTAGNHEFIDVVRMRSG 174
+ +++ SV ++ V+ L G++++ICK++WD + + AG +E+IDV+
Sbjct: 128 TKIIEKNKSVKRKDELRKIVVDELSSLGYDSSICKSKWDKTRSIPAGEYEYIDVI----- 182
Query: 175 SSTWQNRYFVELDFAVQFEIARPTSRYSEIMSYVPGIFVGNSEELKRTVLALCGAVKLCL 234
R +++DF +FEIAR TS Y E++ +P IFVG S+ +++ V + A K L
Sbjct: 183 --VNGERLIIDIDFRSEFEIARQTSGYKELLQSLPLIFVGKSDRIRQIVSIVSEASKQSL 240
Query: 235 RSRGLSIPPWRKNRYMQNKWFGPYRRTTNPVHGNPVPDVVSS 276
+ +G+ PPWRK YM+ KW Y R + G P V S+
Sbjct: 241 KKKGMHFPPWRKADYMRAKWLSSYTRNS----GEKKPTVTSA 278
>UniRef100_Q9LIJ8 Arabidopsis thaliana genomic DNA, chromosome 3, BAC clone:F5N5
[Arabidopsis thaliana]
Length = 370
Score = 109 bits (272), Expect = 1e-22
Identities = 70/226 (30%), Positives = 115/226 (49%), Gaps = 17/226 (7%)
Query: 46 SSPSLSELVHGFLEEDDGNVCDSTGNDFDSERVDSVSDSMDSVEDLLRLSAENANADSYL 105
SS L+++V F+EE++ N + ++ S D DL S + +A +L
Sbjct: 76 SSVCLAKMVQNFIEENNEKQAKCGRNRCNCFNGNN-DGSSDDESDLFGGSIDGCDASDHL 134
Query: 106 NMLR--LHVSEAAVKFDFLK--KQSVSVYNRNVMSFLREKG-----HNAAICKTRWDSSG 156
L V E + D K ++ SV ++ M + +G +N++ICK++WD S
Sbjct: 135 KSLIPCTTVGERNLLADAAKIVDKNKSVKRKDDMKKIVNEGLLSLNYNSSICKSKWDKSP 194
Query: 157 GLTAGNHEFIDVVRMRSGSSTWQNRYFVELDFAVQFEIARPTSRYSEIMSYVPGIFVGNS 216
AG +E+IDV+ + R +++DF +F+IAR TS Y ++ +P IFVG S
Sbjct: 195 SFPAGEYEYIDVI-------IGEERLIIDVDFRSEFDIARQTSGYKVLLQSLPFIFVGKS 247
Query: 217 EELKRTVLALCGAVKLCLRSRGLSIPPWRKNRYMQNKWFGPYRRTT 262
+ L + V + A K L+ +G+ PPWRK YM++KW Y R +
Sbjct: 248 DRLSQIVFLISEAAKQSLKKKGMPFPPWRKAEYMRSKWLSSYTRAS 293
>UniRef100_Q84WI2 Hypothetical protein At3g22970 [Arabidopsis thaliana]
Length = 370
Score = 107 bits (267), Expect = 4e-22
Identities = 69/226 (30%), Positives = 115/226 (50%), Gaps = 17/226 (7%)
Query: 46 SSPSLSELVHGFLEEDDGNVCDSTGNDFDSERVDSVSDSMDSVEDLLRLSAENANADSYL 105
SS L+++V F+EE++ N + ++ S D DL S + +A +L
Sbjct: 76 SSVCLAKMVQNFIEENNEKQAKCGRNRCNCFNGNN-DGSSDDESDLFGGSIDGCDASDHL 134
Query: 106 NMLR--LHVSEAAVKFDFLK--KQSVSVYNRNVMSFLREKG-----HNAAICKTRWDSSG 156
L V E + D K ++ SV ++ M + +G +N++ICK++WD S
Sbjct: 135 KSLIPCTTVGERNLLADAAKIVDKNKSVKRKDDMKKIVNEGLLSLNYNSSICKSKWDKSP 194
Query: 157 GLTAGNHEFIDVVRMRSGSSTWQNRYFVELDFAVQFEIARPTSRYSEIMSYVPGIFVGNS 216
AG +E+IDV+ + R ++++F +F+IAR TS Y ++ +P IFVG S
Sbjct: 195 SFPAGEYEYIDVI-------IGEERLIIDVNFRSEFDIARQTSGYKVLLQSLPFIFVGKS 247
Query: 217 EELKRTVLALCGAVKLCLRSRGLSIPPWRKNRYMQNKWFGPYRRTT 262
+ L + V + A K L+ +G+ PPWRK YM++KW Y R +
Sbjct: 248 DRLSQIVFLISEAAKQSLKKKGMPFPPWRKAEYMRSKWLSSYTRAS 293
>UniRef100_Q9SIU5 Expressed protein [Arabidopsis thaliana]
Length = 294
Score = 105 bits (262), Expect = 1e-21
Identities = 49/122 (40%), Positives = 77/122 (62%), Gaps = 1/122 (0%)
Query: 142 GHNAAICKTRWDSSGGLTAGNHEFIDVVRMRSGSSTWQNRYFVELDFAVQFEIARPTSRY 201
G++ I K++W S + AG HEFI++V RSGS + R +EL F +FEIA+ + Y
Sbjct: 130 GYDCVISKSKWRSCQDIPAGEHEFIEIVD-RSGSKKSEMRVVIELSFRAEFEIAKGSEEY 188
Query: 202 SEIMSYVPGIFVGNSEELKRTVLALCGAVKLCLRSRGLSIPPWRKNRYMQNKWFGPYRRT 261
++S +P ++VG +E L+ + LC A K CLR + + + PWRK++YMQ KW G R+
Sbjct: 189 KRLISRLPEVYVGKTERLRSLIKILCIAGKKCLRDKKMHMAPWRKHKYMQAKWLGTCDRS 248
Query: 262 TN 263
++
Sbjct: 249 SS 250
>UniRef100_Q9SIH9 Expressed protein [Arabidopsis thaliana]
Length = 288
Score = 105 bits (261), Expect = 2e-21
Identities = 67/246 (27%), Positives = 118/246 (47%), Gaps = 39/246 (15%)
Query: 30 SDVSSGSEHSGTGECD-SSPSLSELVHGFLEEDDGNVCDSTG----NDFDSERVDSVSDS 84
SDV + +G+ + SS L+++V F+E+++G G N F +S D
Sbjct: 53 SDVEAPLSRGNSGDFEPSSVCLAKMVLNFMEDNNGGEKQRCGRSRCNCFSGSGTESSDDE 112
Query: 85 MDSVEDLLRLSAENANADSYLNMLRLHVSEAAVKFDFLKKQSVSVYNRNVMSF---LREK 141
+ + A L L L S+ RN+++ + E
Sbjct: 113 TE---------CSSGEACEILKSLVL---------------CKSIRVRNLLTDVTKIAET 148
Query: 142 GHNAAICKTRWDSSGGLTAGNHEFIDVVRMRSGSSTWQNRYFVELDFAVQFEIARPTSRY 201
++AA+CK+RW+ S AG +E++DV+ R +++DF +FEIAR T Y
Sbjct: 149 SYDAALCKSRWEKSPSCPAGEYEYVDVIMKGE-------RLLIDIDFKSKFEIARATKTY 201
Query: 202 SEIMSYVPGIFVGNSEELKRTVLALCGAVKLCLRSRGLSIPPWRKNRYMQNKWFGPYRRT 261
++ +P IFVG ++ L++ ++ +C A K L+ +GL +PPWR+ Y+++KW + R
Sbjct: 202 KSMLQTLPYIFVGKADRLQKIIVLICKAAKQSLKKKGLHVPPWRRAEYVKSKWLSSHVRV 261
Query: 262 TNPVHG 267
+G
Sbjct: 262 DQNSNG 267
>UniRef100_Q7Y1H2 Hypothetical protein OSJNBa0094F01.21 [Oryza sativa]
Length = 402
Score = 103 bits (257), Expect = 6e-21
Identities = 79/277 (28%), Positives = 126/277 (44%), Gaps = 31/277 (11%)
Query: 16 EAKARLVGADLRRLS--DVSSGSEHSGTGECDSSPSLSELVHGFLEEDDGNVCDSTGNDF 73
E +A VG D LS + S+ E G C+ + + E DD +
Sbjct: 75 EVEAGSVGLDRMVLSFMEDSAAVERPQRGRCNCFNGSN-----YEESDDEEGFFLPSDHS 129
Query: 74 DSERVDSVSDSMDSVEDLLRLSAENANADSYLNMLRLHVSEAAVKFDFLKKQSVSVYNRN 133
+ + D+++S++ L++ SA A + + R ++E K K + R
Sbjct: 130 SASAPAAAGDALESLKGLVQ-SASVAERNLLADASR--IAERCCKGSKGKAEC----RRA 182
Query: 134 VMSFLREKGHNAAICKTRWDSSGGLTAGNHEFIDVVRMRSGSSTWQNRYFVELDFAVQFE 193
V LR G++AA+C++RW+ + AG HE+ID V + R VE+DF +FE
Sbjct: 183 VADGLRALGYDAAVCRSRWEKTSSYPAGEHEYIDAVVGE------EVRLIVEVDFRSEFE 236
Query: 194 IARPTSRYSEIMSYVPGIFVGNSEELKRTVLALCGAVKLCLRSRGLSIPPWRKNRYMQNK 253
+AR T Y + +P +FVG + L + V + A + L+ +GL PPWRK YM+ K
Sbjct: 237 VARSTKAYRAALQALPPLFVGTPDRLGQIVAVVAEAARQSLKKKGLHFPPWRKPEYMRAK 296
Query: 254 WFGPYRRT-----------TNPVHGNPVPDVVSSFSG 279
W P+ R+ T+ P +SFSG
Sbjct: 297 WLSPHVRSGDKAAAAAKTVTSTTSATATPVSAASFSG 333
>UniRef100_Q65X09 Hypothetical protein P0599F04.9 [Oryza sativa]
Length = 306
Score = 102 bits (255), Expect = 1e-20
Identities = 51/139 (36%), Positives = 78/139 (55%), Gaps = 4/139 (2%)
Query: 138 LREKGHNAAICKTRWDSSGGLTAGNHEFIDVVR-MRSGSSTWQNRYFVELDFAVQFEIAR 196
LR+ G+N+AIC+++W S + +G H ++DVV RSG + R VE F +FE+AR
Sbjct: 132 LRDAGYNSAICRSKWPRSPEIPSGEHSYVDVVAPTRSGKAV---RVVVEPSFRGEFEMAR 188
Query: 197 PTSRYSEIMSYVPGIFVGNSEELKRTVLALCGAVKLCLRSRGLSIPPWRKNRYMQNKWFG 256
+ Y +++ +P FVG ++ L+ V +C A K C R G+ + PWRK RYM+ KW
Sbjct: 189 GGAGYRALVASLPEAFVGRADRLRGVVRVMCAAAKQCARESGMHMAPWRKQRYMEAKWLA 248
Query: 257 PYRRTTNPVHGNPVPDVVS 275
R P + D V+
Sbjct: 249 TPERVAPPGNAGGAGDAVA 267
>UniRef100_O80651 T14N5.3 protein [Arabidopsis thaliana]
Length = 260
Score = 102 bits (254), Expect = 1e-20
Identities = 65/203 (32%), Positives = 104/203 (51%), Gaps = 27/203 (13%)
Query: 75 SERVDSVSDSMDSVEDLLRLSAENANAD-----SYLNMLRLHVSEAAVKFDFLKKQSVSV 129
S+ ++S + ++ +++++LR E S++N RL K D + K
Sbjct: 24 SQHIESSTTALITLKEILRAKGEKEKEMEEKIWSFINGGRLSYEGDDEKRDVMNK----- 78
Query: 130 YNRNVMSFLREKGHNAAICKTRWDSSGGLTAG--------NHEFIDVV----RMRSGSST 177
++S LR +G++A++ KT WDSS G +E+IDV+ R G S
Sbjct: 79 ----IVSKLRSEGYDASLSKTSWDSSFDHREGCRVFRCSRKYEYIDVMVKGDSNRDGVSK 134
Query: 178 WQNRYFVELDFAVQFEIARPTSRYSEIMSYVPGIFVGNSEELKRTVLALCGAVKLCLRSR 237
+ R ++LDF QFE+AR T Y ++ +P +FV L+R V +CG +K ++
Sbjct: 135 LK-RVIIDLDFKTQFELARQTEAYKDMTEMLPLVFVATEGRLRRVVSLVCGEMKKSMKKE 193
Query: 238 GLSIPPWRKNRYMQNKWFGPYRR 260
G+S PPWR RYMQ+KW RR
Sbjct: 194 GMSRPPWRTTRYMQSKWLPENRR 216
>UniRef100_O80652 T14N5.4 protein [Arabidopsis thaliana]
Length = 258
Score = 102 bits (254), Expect = 1e-20
Identities = 63/197 (31%), Positives = 102/197 (50%), Gaps = 27/197 (13%)
Query: 75 SERVDSVSDSMDSVEDLLRLSAENANA-----DSYLNMLRLHVSEAAVKFDFLKKQSVSV 129
S+ +++ S ++ +++++LR SY+N RL K D + K
Sbjct: 24 SQHINTSSKALITLQEILRAKGVEEKEMEEKIRSYINRGRLSYEGDDEKRDVMNK----- 78
Query: 130 YNRNVMSFLREKGHNAAICKTRWDSSGGLTAG--------NHEFIDVVRM----RSGSST 177
++S LR +G+NA++ KT WDSS G +E+ID + + R G S
Sbjct: 79 ----IVSKLRSEGYNASLSKTSWDSSFDHREGCRVFTCSRKYEYIDAMVIGDSDRDGVSK 134
Query: 178 WQNRYFVELDFAVQFEIARPTSRYSEIMSYVPGIFVGNSEELKRTVLALCGAVKLCLRSR 237
+ R ++LDF QFE+AR T Y ++ +P +FV L+R V +CG +K ++
Sbjct: 135 LK-RVIIDLDFKTQFELARQTEAYKDMTEMLPTVFVATEGRLRRVVSLVCGEMKKSMKKE 193
Query: 238 GLSIPPWRKNRYMQNKW 254
G+S PPWR +RYMQ+KW
Sbjct: 194 GMSRPPWRTSRYMQSKW 210
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.318 0.133 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 508,771,812
Number of Sequences: 2790947
Number of extensions: 20957072
Number of successful extensions: 53761
Number of sequences better than 10.0: 83
Number of HSP's better than 10.0 without gapping: 40
Number of HSP's successfully gapped in prelim test: 44
Number of HSP's that attempted gapping in prelim test: 53641
Number of HSP's gapped (non-prelim): 135
length of query: 304
length of database: 848,049,833
effective HSP length: 127
effective length of query: 177
effective length of database: 493,599,564
effective search space: 87367122828
effective search space used: 87367122828
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 74 (33.1 bits)
Medicago: description of AC145513.12


