

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145512.11 - phase: 0
(127 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
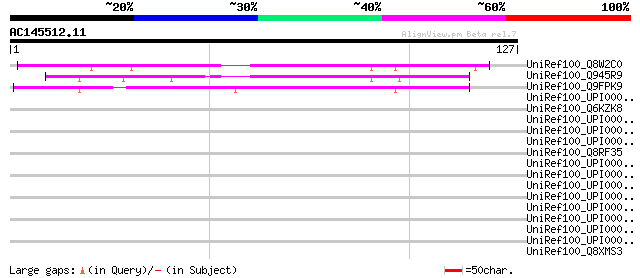
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8W2C0 Functional candidate resistance protein KR1 [Gl... 79 2e-14
UniRef100_Q945R9 NR1 [Glycine max] 65 3e-10
UniRef100_Q9FPK9 Putative resistance protein [Glycine max] 61 6e-09
UniRef100_UPI000029FE96 UPI000029FE96 UniRef100 entry 33 1.1
UniRef100_Q6KZK8 Beta-galactosidase [Picrophilus torridus] 33 1.1
UniRef100_UPI00002E7110 UPI00002E7110 UniRef100 entry 33 1.9
UniRef100_UPI00002969B1 UPI00002969B1 UniRef100 entry 33 1.9
UniRef100_UPI000030FBA0 UPI000030FBA0 UniRef100 entry 32 3.2
UniRef100_Q8RF35 Hemin-binding periplasmic protein hmuT [Fusobac... 32 4.2
UniRef100_UPI000032B958 UPI000032B958 UniRef100 entry 31 5.5
UniRef100_UPI000031ED9A UPI000031ED9A UniRef100 entry 31 5.5
UniRef100_UPI0000317D9A UPI0000317D9A UniRef100 entry 31 5.5
UniRef100_UPI000031324A UPI000031324A UniRef100 entry 31 5.5
UniRef100_UPI000028D5C3 UPI000028D5C3 UniRef100 entry 31 5.5
UniRef100_UPI00002B3DC8 UPI00002B3DC8 UniRef100 entry 31 7.2
UniRef100_UPI000027A12A UPI000027A12A UniRef100 entry 31 7.2
UniRef100_UPI0000288BC2 UPI0000288BC2 UniRef100 entry 30 9.4
UniRef100_Q8XMS3 Probable rhamnosyl transferase [Clostridium per... 30 9.4
>UniRef100_Q8W2C0 Functional candidate resistance protein KR1 [Glycine max]
Length = 1124
Score = 79.0 bits (193), Expect = 2e-14
Identities = 54/140 (38%), Positives = 77/140 (54%), Gaps = 29/140 (20%)
Query: 3 VSFWFCNKFPSIALGVV--------SAYTWGYLEH-PARVIINDNTFFYTHGRKIDRCSR 53
+SFWF NKFP+IA+ + S+ W + + +VIIN N + C+
Sbjct: 936 ISFWFRNKFPAIAICHIIKRVAEFSSSRGWTFRPNIRTKVIINGNANLFNSVVLGSDCTC 995
Query: 54 PDTYHLHLFHMQVEYFNGNMDKALLENKWNHAEVDF-GFPFMF------SGIHVLKEKSN 106
LF ++ E N+D+ALLEN+WNHAEV GF F F +G+HVLK++SN
Sbjct: 996 -------LFDLRGERVTDNLDEALLENEWNHAEVTCPGFTFTFAPTFIKTGLHVLKQESN 1048
Query: 107 MKDIRFTNP------ENDAN 120
M+DIRF++P +ND N
Sbjct: 1049 MEDIRFSDPCRKTKLDNDFN 1068
>UniRef100_Q945R9 NR1 [Glycine max]
Length = 175
Score = 65.5 bits (158), Expect = 3e-10
Identities = 50/123 (40%), Positives = 68/123 (54%), Gaps = 25/123 (20%)
Query: 10 KFPSIAL--------GVVSAYTWGYL-EHPARVIINDNT-FFYTHGRKIDRCSRPDTYHL 59
KFP+IA+ S+ W Y + +VIIN N FY+ D CS
Sbjct: 1 KFPAIAICHSIKRVPEFSSSRGWTYRPDIRTKVIINGNANLFYSMDLGSD-CSV------ 53
Query: 60 HLFHMQVEYFNGNMDKALLENKWNHAEVDF-GFPFMFS------GIHVLKEKSNMKDIRF 112
LF + + N+D+ALLEN+WNHAEV GF F F+ G+HVLK++SNM+DIRF
Sbjct: 54 -LFDPRDDKVTDNLDEALLENEWNHAEVTCPGFTFTFAPTFIKTGLHVLKQESNMEDIRF 112
Query: 113 TNP 115
++P
Sbjct: 113 SDP 115
>UniRef100_Q9FPK9 Putative resistance protein [Glycine max]
Length = 1093
Score = 60.8 bits (146), Expect = 6e-09
Identities = 43/127 (33%), Positives = 64/127 (49%), Gaps = 16/127 (12%)
Query: 2 SVSFWFCNKFPSIAL---GVVSAYTWGYLEHPARVIINDNTFFYTHGRKIDRCSRPD-TY 57
S+SFWF NKFP I+L G++ + +G V IN N R+ P T
Sbjct: 964 SISFWFRNKFPVISLCLAGLMHKHPFGL---KPIVSINGNKMKTEFQRRWFYFEFPVLTD 1020
Query: 58 HLHLFHMQVEYFNGNMDKALLENKWNHAEVDFGFPFMF---------SGIHVLKEKSNMK 108
H+ +F + F N+D+ + EN WNH V F + +G+HV+K KS+++
Sbjct: 1021 HILIFGERQIKFEDNVDEVVSENDWNHVVVSVDVDFKWNPTEPLVVRTGLHVIKPKSSVE 1080
Query: 109 DIRFTNP 115
DIRF +P
Sbjct: 1081 DIRFIDP 1087
>UniRef100_UPI000029FE96 UPI000029FE96 UniRef100 entry
Length = 214
Score = 33.5 bits (75), Expect = 1.1
Identities = 24/120 (20%), Positives = 52/120 (43%), Gaps = 33/120 (27%)
Query: 1 MSVSFWFCNKFPSIALGVVSAYTWGYLEHPARVIINDNTFFYT----------------- 43
+++ WF + F + + +++ + + + P RV INDN+F +
Sbjct: 22 ITIIIWFFSNFLATVMLLITIWVYYFFRDPDRVSINDNSFLVSPADGLITDISERNGPEE 81
Query: 44 ---HGRKIDRCS-RPDTYHLH------------LFHMQVEYFNGNMDKALLENKWNHAEV 87
+K R S + ++ H +F+ ++FN ++DKA EN+ N+ ++
Sbjct: 82 LGLENKKYTRISIFMNIFNCHVNRSPVEGKIEEIFYKPGKFFNASLDKASKENERNYYKI 141
>UniRef100_Q6KZK8 Beta-galactosidase [Picrophilus torridus]
Length = 453
Score = 33.5 bits (75), Expect = 1.1
Identities = 20/80 (25%), Positives = 41/80 (51%), Gaps = 4/80 (5%)
Query: 17 GVVSAYTWGYLEHPARVIINDNTFFYTHGRKIDRCSR-PDTYHLHLFHMQVEYFNGNMDK 75
G +S + GYL++ + N + + +TH + + + D Y ++EY N N+ +
Sbjct: 8 GFMSGFHLGYLQNNKNLPENSDWYQWTHDKNVRMMNYIRDGYPED---NKIEYLNDNVIR 64
Query: 76 ALLENKWNHAEVDFGFPFMF 95
L +N +N ++D +P +F
Sbjct: 65 DLHDNNFNLIKIDMDWPSLF 84
>UniRef100_UPI00002E7110 UPI00002E7110 UniRef100 entry
Length = 171
Score = 32.7 bits (73), Expect = 1.9
Identities = 16/57 (28%), Positives = 30/57 (52%), Gaps = 2/57 (3%)
Query: 30 PARVIINDNTFFYTHGRKIDRCSRPDTYHLHLFHMQ--VEYFNGNMDKALLENKWNH 84
P V N TF+YT+ +K + ++ ++ + H + YFNG+ K+ +E N+
Sbjct: 4 PDEVPSNFTTFYYTYLKKDEEVNKDIKFNKDILHQSKIINYFNGDYAKSKIEKDLNN 60
>UniRef100_UPI00002969B1 UPI00002969B1 UniRef100 entry
Length = 264
Score = 32.7 bits (73), Expect = 1.9
Identities = 21/67 (31%), Positives = 33/67 (48%), Gaps = 4/67 (5%)
Query: 30 PARVIINDNTFFYTHGRKIDRCSRPDTYHLHLFHMQVEYFNGNMDKALLENKWNHAEVDF 89
PA+ + N +F+THG D +R T H + F N D L+EN + H + F
Sbjct: 190 PAKKLSNTQYYFFTHG---DSDTRILTRHFNYFKDYTNQNKINADFWLVENSY-HVDAMF 245
Query: 90 GFPFMFS 96
+P ++S
Sbjct: 246 KYPEVYS 252
>UniRef100_UPI000030FBA0 UPI000030FBA0 UniRef100 entry
Length = 217
Score = 32.0 bits (71), Expect = 3.2
Identities = 19/72 (26%), Positives = 34/72 (46%), Gaps = 9/72 (12%)
Query: 36 NDNTFFYTHGRKIDRCSRPDTYHLHLFHMQVEYFNGNMDKALLENKWNHAEVDFGFPFMF 95
N+N FY Y+LH + ++ FN N + +L NK+ + ++ GF F
Sbjct: 131 NNNLSFYV--------GTSQEYYLHDYKLEGSLFNENAELTMLSNKYRNT-IEVGFQKRF 181
Query: 96 SGIHVLKEKSNM 107
+ + VL ++M
Sbjct: 182 NKLQVLTTFNSM 193
>UniRef100_Q8RF35 Hemin-binding periplasmic protein hmuT [Fusobacterium nucleatum]
Length = 286
Score = 31.6 bits (70), Expect = 4.2
Identities = 17/42 (40%), Positives = 25/42 (59%)
Query: 70 NGNMDKALLENKWNHAEVDFGFPFMFSGIHVLKEKSNMKDIR 111
+GN +K L+EN +D+G S I LKEKS +KD++
Sbjct: 207 DGNWEKVLVENPDIIVIIDYGDQSAESKIKFLKEKSPIKDLK 248
>UniRef100_UPI000032B958 UPI000032B958 UniRef100 entry
Length = 111
Score = 31.2 bits (69), Expect = 5.5
Identities = 20/67 (29%), Positives = 33/67 (48%), Gaps = 4/67 (5%)
Query: 30 PARVIINDNTFFYTHGRKIDRCSRPDTYHLHLFHMQVEYFNGNMDKALLENKWNHAEVDF 89
PA+ + N +F+THG D +R T H + F N + L+EN + H + F
Sbjct: 37 PAKKLSNKQYYFFTHG---DSDTRILTRHFNYFKDYTSQNKINAEFWLVENSY-HVDAMF 92
Query: 90 GFPFMFS 96
+P ++S
Sbjct: 93 KYPEVYS 99
>UniRef100_UPI000031ED9A UPI000031ED9A UniRef100 entry
Length = 79
Score = 31.2 bits (69), Expect = 5.5
Identities = 20/67 (29%), Positives = 33/67 (48%), Gaps = 4/67 (5%)
Query: 30 PARVIINDNTFFYTHGRKIDRCSRPDTYHLHLFHMQVEYFNGNMDKALLENKWNHAEVDF 89
PA+ + N +F+THG D +R T H + F N + L+EN + H + F
Sbjct: 5 PAKKLSNKQYYFFTHG---DSDTRILTRHFNYFKDYTNQNKINAEFWLVENSY-HVDAMF 60
Query: 90 GFPFMFS 96
+P ++S
Sbjct: 61 KYPEVYS 67
>UniRef100_UPI0000317D9A UPI0000317D9A UniRef100 entry
Length = 147
Score = 31.2 bits (69), Expect = 5.5
Identities = 20/67 (29%), Positives = 33/67 (48%), Gaps = 4/67 (5%)
Query: 30 PARVIINDNTFFYTHGRKIDRCSRPDTYHLHLFHMQVEYFNGNMDKALLENKWNHAEVDF 89
PA+ + N +F+THG D +R T H + F N + L+EN + H + F
Sbjct: 73 PAKKLSNKQHYFFTHG---DSDTRILTRHFNYFKDYTNQNKINAEFWLVENSY-HVDAMF 128
Query: 90 GFPFMFS 96
+P ++S
Sbjct: 129 KYPEVYS 135
>UniRef100_UPI000031324A UPI000031324A UniRef100 entry
Length = 329
Score = 31.2 bits (69), Expect = 5.5
Identities = 19/68 (27%), Positives = 33/68 (47%), Gaps = 4/68 (5%)
Query: 29 HPARVIINDNTFFYTHGRKIDRCSRPDTYHLHLFHMQVEYFNGNMDKALLENKWNHAEVD 88
+PA+ + +D +F+THG D+ R H + F E N L EN + H +
Sbjct: 254 NPAKKLTSDQNYFFTHG---DKDQRMLVRHYYYFKKYTEENNIKSKFWLAENSY-HVDGM 309
Query: 89 FGFPFMFS 96
F +P +++
Sbjct: 310 FKYPDLYA 317
>UniRef100_UPI000028D5C3 UPI000028D5C3 UniRef100 entry
Length = 120
Score = 31.2 bits (69), Expect = 5.5
Identities = 19/68 (27%), Positives = 33/68 (47%), Gaps = 4/68 (5%)
Query: 29 HPARVIINDNTFFYTHGRKIDRCSRPDTYHLHLFHMQVEYFNGNMDKALLENKWNHAEVD 88
+PA+ + +D +F+THG D+ R H + F E N L EN + H +
Sbjct: 45 NPAKKLTSDQNYFFTHG---DKDERMLVRHYYYFKKYTEENNIKSKFWLAENSY-HVDGM 100
Query: 89 FGFPFMFS 96
F +P +++
Sbjct: 101 FKYPDLYA 108
>UniRef100_UPI00002B3DC8 UPI00002B3DC8 UniRef100 entry
Length = 158
Score = 30.8 bits (68), Expect = 7.2
Identities = 32/110 (29%), Positives = 49/110 (44%), Gaps = 15/110 (13%)
Query: 12 PSIALG---VVSAYTWGYLEH--PARVIINDNTFFYTHGRKIDRCSRPDTYHLHLFHMQV 66
P++ +G VS YT+GY +H P I N FY S+PD L +F
Sbjct: 39 PTLLVGDRIFVSKYTYGYSKHSFPFSPPIIKNRIFY---------SKPDYGDLVVFKTPS 89
Query: 67 EYFNGNMDKALLENKWNHAEVDFGFPFMFSGIHVLKEKSNMKDIRFTNPE 116
+ + K L+ + ++ G F+ + I + K+K N DIR N E
Sbjct: 90 DN-RTDFIKRLIGLPGDEIQLKDGDLFVNNEIVLKKKKQNRNDIRCANNE 138
>UniRef100_UPI000027A12A UPI000027A12A UniRef100 entry
Length = 328
Score = 30.8 bits (68), Expect = 7.2
Identities = 23/70 (32%), Positives = 36/70 (50%), Gaps = 10/70 (14%)
Query: 30 PARVIINDNTFFYTHGRKIDRCSRPDTYHLHLFHMQVEYFNGNMDKA---LLENKWNHAE 86
PA+ + + +F+THG D +R T H + F EY N N A L+EN + H +
Sbjct: 254 PAKKLSKNQYYFFTHG---DSDTRILTRHFNYFK---EYTNQNKINAEFWLVENSY-HVD 306
Query: 87 VDFGFPFMFS 96
F +P ++S
Sbjct: 307 AMFKYPEVYS 316
>UniRef100_UPI0000288BC2 UPI0000288BC2 UniRef100 entry
Length = 540
Score = 30.4 bits (67), Expect = 9.4
Identities = 17/55 (30%), Positives = 32/55 (57%), Gaps = 1/55 (1%)
Query: 72 NMDKALLENKWNHAEVDFGFPFMFSGIHVLKEKSNMKDIRFTNPENDANIVLHSG 126
N+++AL+E ++ V FG FMFS + K+N K +F+N + + + ++G
Sbjct: 471 NVEEALIEPFFHQMLVSFGVAFMFSYL-FSDSKTNQKVFQFSNEDFRTSKLFNAG 524
>UniRef100_Q8XMS3 Probable rhamnosyl transferase [Clostridium perfringens]
Length = 304
Score = 30.4 bits (67), Expect = 9.4
Identities = 25/93 (26%), Positives = 41/93 (43%), Gaps = 18/93 (19%)
Query: 32 RVIINDNTFFYTH----GRKIDRCSRPDTYHLHLFHMQ---VEYFNGNM----DKALLEN 80
R++IN++ +F G +I CS + YH H+F ++ YF+ + + L
Sbjct: 178 RLLINEDMYFAEKLIQAGYRIKYCSDSEVYHSHVFTLKSLYERYFDTGVFFRQNSQFLNY 237
Query: 81 KWNHAEVDFGFPFMFSGIHVLKEKSNMKDIRFT 113
K N + D IHVLK K+ + T
Sbjct: 238 KGNKSGKDL-------AIHVLKRSLEDKNFKVT 263
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.326 0.140 0.463
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 239,379,777
Number of Sequences: 2790947
Number of extensions: 9942202
Number of successful extensions: 22842
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 22828
Number of HSP's gapped (non-prelim): 19
length of query: 127
length of database: 848,049,833
effective HSP length: 103
effective length of query: 24
effective length of database: 560,582,292
effective search space: 13453975008
effective search space used: 13453975008
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 67 (30.4 bits)
Medicago: description of AC145512.11


