

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145470.6 + phase: 2 /partial
(753 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
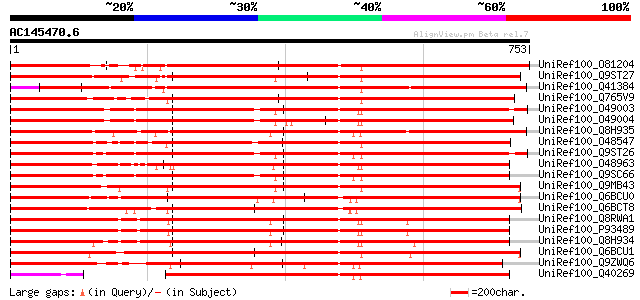
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O81204 Non phototropic hypocotyl 1-like [Arabidopsis t... 1037 0.0
UniRef100_Q9ST27 Nonphototrophic hypocotyl 1b [Oryza sativa] 982 0.0
UniRef100_Q41384 Protein kinase [Spinacia oleracea] 969 0.0
UniRef100_Q765V9 Phototropin 2 [Adiantum capillus-veneris] 903 0.0
UniRef100_O49003 NPH1-1 [Avena sativa] 853 0.0
UniRef100_O49004 NPH1-2 [Avena sativa] 852 0.0
UniRef100_Q8H935 Phototropin [Vicia faba] 847 0.0
UniRef100_O48547 Nonphototropic hypocotyl 1 [Zea mays] 842 0.0
UniRef100_Q9ST26 Nonphototrophic hypocotyl 1a [Oryza sativa] 837 0.0
UniRef100_O48963 Nonphototropic hypocotyl protein 1 [Arabidopsis... 835 0.0
UniRef100_Q9SC66 Non-phototropic hypocotyl NPH1 [Oryza sativa] 834 0.0
UniRef100_Q9MB43 Phototropin [Adiantum capillus-veneris] 826 0.0
UniRef100_Q6BCU0 Phototropin [Physcomitrella patens] 826 0.0
UniRef100_Q6BCT8 Phototropin [Physcomitrella patens] 826 0.0
UniRef100_Q8RWA1 Phototropin 1 [Pisum sativum] 825 0.0
UniRef100_P93489 Phototropin-like protein PsPK4 [Pisum sativum] 825 0.0
UniRef100_Q8H934 Phototropin [Vicia faba] 823 0.0
UniRef100_Q6BCU1 Phototropin [Physcomitrella patens] 811 0.0
UniRef100_Q9ZWQ6 PHY3 [Adiantum capillus-veneris] 702 0.0
UniRef100_Q40269 Protein kinase [Mesembryanthemum crystallinum] 656 0.0
>UniRef100_O81204 Non phototropic hypocotyl 1-like [Arabidopsis thaliana]
Length = 915
Score = 1037 bits (2682), Expect = 0.0
Identities = 546/779 (70%), Positives = 611/779 (78%), Gaps = 63/779 (8%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKFI 60
RFLQGP+TD+NEVAKIRD KNGKSYCGRLLNYKK+GTPFWNLLTVTPIKDD+GNTIKFI
Sbjct: 171 RFLQGPDTDKNEVAKIRDCVKNGKSYCGRLLNYKKDGTPFWNLLTVTPIKDDQGNTIKFI 230
Query: 61 GMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSITEVVQTVRNPKSIIRSKN 120
GMQVEVSKYTEGVN+KALRPNGL KSLIRYDARQKE+A+ SITEVVQT+R+ KS ++
Sbjct: 231 GMQVEVSKYTEGVNDKALRPNGLSKSLIRYDARQKEKALDSITEVVQTIRHRKSQVQ--- 287
Query: 121 DDTATIMHEEPENLNHDFVLPKSVEPVNDTT-TPGRQTPLKFHGDNNNMSRFSSYEERNN 179
E++++D + V+P + TT TPGRQT + +
Sbjct: 288 -----------ESVSNDTM----VKPDSSTTPTPGRQT---------------RQSDEAS 317
Query: 180 KSSRKSGITSLKGVKGKSMSSVGRDKDKTIVEPEVLMT------KEIEWSKYELRERDIR 233
KS R G S K SS R +D +EPE LM + W + RERDIR
Sbjct: 318 KSFRTPGRVSTP-TGSKLKSSNNRHEDLLRMEPEELMLSTEVIGQRDSWDLSD-RERDIR 375
Query: 234 QGIDLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETD 293
QGIDLATTLERIEKNFVISDPRLPD PIIFASDSFLELTEY+REEILGRNCRFLQGPETD
Sbjct: 376 QGIDLATTLERIEKNFVISDPRLPDNPIIFASDSFLELTEYSREEILGRNCRFLQGPETD 435
Query: 294 QATVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDH 353
QATV +IRDAI+DQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDH
Sbjct: 436 QATVQKIRDAIRDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDH 495
Query: 354 LEPLRNRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAVSPRPHKRD 413
+EPL+NRLSE +E+QS+KLVKATA NVD AVRELPDAN RPEDLWA HS+ V P PH ++
Sbjct: 496 VEPLQNRLSERTEMQSSKLVKATATNVDEAVRELPDANTRPEDLWAAHSKPVYPLPHNKE 555
Query: 414 NPSWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLN 473
+ SW AI+KI A GE +GLHHF PI+PLG GDTGSVHLVEL+GTGELYAMKAMEK++MLN
Sbjct: 556 STSWKAIKKIQASGETVGLHHFKPIKPLGSGDTGSVHLVELKGTGELYAMKAMEKTMMLN 615
Query: 474 RNKSVFYYRFIERALKERLYHYWIILFFQHY--TPHF-----------------KQPMKI 514
RNK+ + IER + L H ++ + + + H +QPMKI
Sbjct: 616 RNKA--HRACIEREIISLLDHPFLPTLYASFQTSTHVCLITDFCPGGELFALLDRQPMKI 673
Query: 515 LKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQV 574
L EDSARFYAAEVVIGLEYLHCLGI+YRDLKPEN+LL+KDGHIVL DFDLSF+T+C PQ+
Sbjct: 674 LTEDSARFYAAEVVIGLEYLHCLGIVYRDLKPENILLKKDGHIVLADFDLSFMTTCTPQL 733
Query: 575 VKQSLPGNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGIL 634
+ + P RRRS+SQP P FV+EP TQSNSFVGTEEYIAPEIITGA HTSAIDWW LGIL
Sbjct: 734 IIPAAPSKRRRSKSQPLPTFVAEPSTQSNSFVGTEEYIAPEIITGAGHTSAIDWWALGIL 793
Query: 635 LYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSAT 694
LYEMLYGRTPFRGKNRQKTF+NILHKDLTFPSSIP SL RQLIN LL RDP+SRLGS
Sbjct: 794 LYEMLYGRTPFRGKNRQKTFANILHKDLTFPSSIPVSLVGRQLINTLLNRDPSSRLGSKG 853
Query: 695 GSNEIKQHPFFRGINWPLIRNMSPPPLDVPLQFIGKDPTAKDKKWEDDGVLNTSIDMDI 753
G+NEIKQH FFRGINWPLIR MSPPPLD PL I KDP AKD KWEDDGVL S D+DI
Sbjct: 854 GANEIKQHAFFRGINWPLIRGMSPPPLDAPLSIIEKDPNAKDIKWEDDGVLVNSTDLDI 912
Score = 124 bits (310), Expect = 1e-26
Identities = 78/260 (30%), Positives = 134/260 (51%), Gaps = 24/260 (9%)
Query: 141 PKSVEPVNDTTTPGRQTPLKFHGDNNNMSRFSSYEERNNKS-----SRKSGITSLK---- 191
P S + + +T+ + PL D NN S + E + + + + G++++K
Sbjct: 28 PSSGKETHGSTSSSSKPPL----DGNNKGSSSKWMEFQDSAKITERTAEWGLSAVKPDSG 83
Query: 192 --GVKGKSMSSVGRDKDKTIVEPEVLMTKEIEWSKYELRERDIRQGIDLATTLERIEKNF 249
G+ K S V R K+ + + E S R +L T L +++ F
Sbjct: 84 DDGISFKLSSEVERSKN--------MSRRSSEESTSSESGAFPRVSQELKTALSTLQQTF 135
Query: 250 VISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQATVNRIRDAIKDQRE 309
V+SD P CPI++AS F +T Y+ +EI+GRNCRFLQGP+TD+ V +IRD +K+ +
Sbjct: 136 VVSDATQPHCPIVYASSGFFTMTGYSSKEIVGRNCRFLQGPDTDKNEVAKIRDCVKNGKS 195
Query: 310 ITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLEPLRNRLSEGSEIQS 369
+L+NY K G FWNL + P++D +G FIG+Q++ S + E + ++ + + S
Sbjct: 196 YCGRLLNYKKDGTPFWNLLTVTPIKDDQGNTIKFIGMQVEVSKYTEGVNDKALRPNGL-S 254
Query: 370 AKLVKATAENVDGAVRELPD 389
L++ A + A+ + +
Sbjct: 255 KSLIRYDARQKEKALDSITE 274
>UniRef100_Q9ST27 Nonphototrophic hypocotyl 1b [Oryza sativa]
Length = 907
Score = 982 bits (2538), Expect = 0.0
Identities = 517/774 (66%), Positives = 594/774 (75%), Gaps = 47/774 (6%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKFI 60
RFLQGP+TD EVAKIRDA K+G+S+CGRLLNY+K+G PFWNLLTVTPI+DD G IKFI
Sbjct: 140 RFLQGPDTDAAEVAKIRDAVKHGRSFCGRLLNYRKDGAPFWNLLTVTPIRDDNGKVIKFI 199
Query: 61 GMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSITEVVQTVRNPKSIIRSKN 120
GMQVEVSKYTEG+++K +RPN LP SLIRYD RQK++AM S+TEVVQTV+ P+
Sbjct: 200 GMQVEVSKYTEGLSDKRMRPNELPVSLIRYDERQKDKAMSSMTEVVQTVKQPRGARAPA- 258
Query: 121 DDTATIMHEEPENLNHDFVLPKSVEPVNDTTTPGRQTPLK-----FHGDNNNMSRFSSYE 175
D A + + + + + V P + G K + + +SR +S
Sbjct: 259 -DAALLTPPKMSDADKMAAMSPVVAPGTPSGGGGGAGSFKSPLWDLKKEESRLSRLAS-- 315
Query: 176 ERNNKSSRKSGITSLKGVKGKSMSSVGRDKDKTIVE---------PEVLMTKEIEWSKYE 226
RKSG +SL G K SSVG + +VE PEV+ + W + E
Sbjct: 316 ------GRKSGRSSLMGFKIGKRSSVGSREAPAVVEEPAPAPPPAPEVVERTD-SWERAE 368
Query: 227 LRERDIRQGIDLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRF 286
RE+DIRQGIDLATTLERIEKNFVI+DPR+PD PIIFASDSFLELTEYTREEILGRNCRF
Sbjct: 369 -REKDIRQGIDLATTLERIEKNFVITDPRIPDNPIIFASDSFLELTEYTREEILGRNCRF 427
Query: 287 LQGPETDQATVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGV 346
LQGPETDQ TV++IR+AI++Q+EITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGV
Sbjct: 428 LQGPETDQGTVDKIREAIREQKEITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGV 487
Query: 347 QLDGSDHLEPLRNRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAVS 406
QLDGSDH+EPLRNRLSE +EIQSAKLVKATAENVD AVRELPDANLRPEDLWAIHS VS
Sbjct: 488 QLDGSDHVEPLRNRLSENTEIQSAKLVKATAENVDDAVRELPDANLRPEDLWAIHSMRVS 547
Query: 407 PRPHKRDNPSWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAM 466
P+PHKR+NPSW+AI+K T GEKIGL HF P++PLGCGDTGSVHLVELQG+GEL+AMKAM
Sbjct: 548 PKPHKRNNPSWIAIEKATNLGEKIGLKHFKPVKPLGCGDTGSVHLVELQGSGELFAMKAM 607
Query: 467 EKSVMLNRNKSVFYYRFIERALKERLYHYWI-ILFFQHYTPHF----------------- 508
+KSVMLNRNK + IER + L H ++ L+ TP
Sbjct: 608 DKSVMLNRNK--VHRACIEREIYALLDHPFLPTLYTSFQTPTHVCLITDFCPGGELFAVL 665
Query: 509 -KQPMKILKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFI 567
+QPMKI +E+ ARFYAAEVVIGLEYLHCLGIIYRDLKPEN+LLQ DGHIVLTDFDLSF+
Sbjct: 666 DRQPMKIFREECARFYAAEVVIGLEYLHCLGIIYRDLKPENILLQADGHIVLTDFDLSFL 725
Query: 568 TSCKPQVVKQSLPGNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAID 627
T+ KP V+K S RRRS+ PP FVSEP T SNSFVGTEEYIAPE+ITGA HTSAID
Sbjct: 726 TTSKPHVIKNSTSLKRRRSQEFLPPTFVSEPSTPSNSFVGTEEYIAPEVITGAGHTSAID 785
Query: 628 WWTLGILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPA 687
WW LGILLYEMLYGRTPFRGKNR+KTF NILHKDLTFPSSIP SLAA+QLI+ LLQRDP+
Sbjct: 786 WWALGILLYEMLYGRTPFRGKNRKKTFYNILHKDLTFPSSIPVSLAAKQLIHGLLQRDPS 845
Query: 688 SRLGSATGSNEIKQHPFFRGINWPLIRNMSPPPLDVPLQFIGKDPTAKDKKWED 741
+R+GS G+N+IKQH FF+ INWPLIR MSPP LDVPL+ IGK+ K K ED
Sbjct: 846 NRIGSNAGANDIKQHSFFQDINWPLIRCMSPPELDVPLKLIGKETQPKAKPDED 899
Score = 131 bits (329), Expect = 8e-29
Identities = 72/198 (36%), Positives = 108/198 (54%), Gaps = 4/198 (2%)
Query: 237 DLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQAT 296
+L L +++ FV+SD PDCPII+AS+ F +T Y+ E++GRNCRFLQGP+TD A
Sbjct: 92 ELKDALSSLQQTFVVSDATRPDCPIIYASEGFFTMTGYSPREVVGRNCRFLQGPDTDAAE 151
Query: 297 VNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLEP 356
V +IRDA+K R +L+NY K G FWNL + P+RD G++ FIG+Q++ S + E
Sbjct: 152 VAKIRDAVKHGRSFCGRLLNYRKDGAPFWNLLTVTPIRDDNGKVIKFIGMQVEVSKYTEG 211
Query: 357 LRNRLSEGSEIQSAKLVKATAENVDGA---VRELPDANLRPEDLWAIHSQAVSPRPHKRD 413
L ++ +E+ L++ D A + E+ +P A A+ P D
Sbjct: 212 LSDKRMRPNEL-PVSLIRYDERQKDKAMSSMTEVVQTVKQPRGARAPADAALLTPPKMSD 270
Query: 414 NPSWVAIQKITARGEKIG 431
A+ + A G G
Sbjct: 271 ADKMAAMSPVVAPGTPSG 288
>UniRef100_Q41384 Protein kinase [Spinacia oleracea]
Length = 724
Score = 969 bits (2504), Expect = 0.0
Identities = 498/731 (68%), Positives = 581/731 (79%), Gaps = 33/731 (4%)
Query: 44 LTVTPIKDDRGNTIKFIGMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSIT 103
LTVTPI+DD+G IKFIGMQVEVSK+TEG+N+KALRPNGLPKSLIRYD RQKE A+GSI
Sbjct: 1 LTVTPIRDDKGCVIKFIGMQVEVSKFTEGINDKALRPNGLPKSLIRYDPRQKEAALGSII 60
Query: 104 EVVQTVRNPKSIIRSKNDDTATIMHEEPENLNHDFVLPKSVEPVNDTTTPGR-QTPLKFH 162
EVVQTV++P+S+ + +++ A E N D++LPK E +++ TPGR +P +
Sbjct: 61 EVVQTVKHPRSLSQPLSNNDAD-RREVAGKFNLDYMLPKLAE-IDNVGTPGRLSSPSRLS 118
Query: 163 GDNNNMSRFSSYEERNNKSSRKSGITSLKGVKGKSMSSVGRDKDKTIVEPEVLMTKEIE- 221
+ + ++KSSRKS SL G KG+S + R EPE L+ K+I+
Sbjct: 119 TPGRQTPKIDASSRDSDKSSRKSARISLLGFKGRSSAKHERPPSP---EPEFLIPKDIDR 175
Query: 222 ---WSKYELRERDIRQGIDLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREE 278
W + E RERD+RQGIDLATTLERIEKNFVISDPRLPD PIIFASDSFLELTEYTREE
Sbjct: 176 DDSWERAE-RERDVRQGIDLATTLERIEKNFVISDPRLPDNPIIFASDSFLELTEYTREE 234
Query: 279 ILGRNCRFLQGPETDQATVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKG 338
ILGRNCRFLQGPETDQ TV +IRDAIK+QR+ITVQLINYTKSG+KFWNLFHLQPMRDQKG
Sbjct: 235 ILGRNCRFLQGPETDQTTVQKIRDAIKEQRDITVQLINYTKSGRKFWNLFHLQPMRDQKG 294
Query: 339 ELQYFIGVQLDGSDHLEPLRNRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLW 398
ELQYFIGVQLDGSDH+EPLRNRLSE +EIQSAK+VKATAENVD AVRELPDAN RPEDLW
Sbjct: 295 ELQYFIGVQLDGSDHVEPLRNRLSERTEIQSAKVVKATAENVDEAVRELPDANSRPEDLW 354
Query: 399 AIHSQAVSPRPHKRDNPSWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTG 458
AIHS+ V PRPHKR + SW AIQKITA GEK+GL HF+PI+PLGCGDTGSVHLVEL+
Sbjct: 355 AIHSEPVYPRPHKRGSSSWAAIQKITAAGEKVGLEHFNPIKPLGCGDTGSVHLVELKVPE 414
Query: 459 ELYAMKAMEKSVMLNRNKSVFYYRFIERALKERLYHYWIILFFQHY--TPHF-------- 508
+AMKAM+KSVMLNRNK + +ER + L H ++ + + + H
Sbjct: 415 NWFAMKAMDKSVMLNRNK--VHRACVEREIISTLDHPFLPTLYASFQTSTHVCLITDFCP 472
Query: 509 ---------KQPMKILKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVL 559
KQP+KI KE+SARFYAAEVVIGLEYLHCLGIIYRDLKPEN+LLQKDGH+VL
Sbjct: 473 GGELFALLDKQPLKIFKEESARFYAAEVVIGLEYLHCLGIIYRDLKPENILLQKDGHLVL 532
Query: 560 TDFDLSFITSCKPQVVKQSLPGNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITG 619
TDFDLSF+TSC P ++ P +RRSRSQPPP F++EPVTQSNSFVGTEEYIAPE+ITG
Sbjct: 533 TDFDLSFLTSCNPHIINHPQP-KKRRSRSQPPPTFIAEPVTQSNSFVGTEEYIAPEVITG 591
Query: 620 ARHTSAIDWWTLGILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLIN 679
A HTSAIDWW LG+LLYEMLYGRTPFRGKNRQKTF+NI+HKDLTFPSSIP SL+ARQLI
Sbjct: 592 ASHTSAIDWWALGVLLYEMLYGRTPFRGKNRQKTFANIMHKDLTFPSSIPVSLSARQLIY 651
Query: 680 ALLQRDPASRLGSATGSNEIKQHPFFRGINWPLIRNMSPPPLDVPLQFIGKDPTAKDKKW 739
ALL RDPA+RLG+ G++EIK+HP+FRGINWPLIR M PP L+ P + IG+DP AK+ KW
Sbjct: 652 ALLNRDPATRLGTQGGASEIKEHPYFRGINWPLIRCMDPPTLEAPFKLIGRDPNAKEVKW 711
Query: 740 EDDGVLNTSID 750
+DDGVL + +D
Sbjct: 712 DDDGVLASPMD 722
Score = 76.3 bits (186), Expect = 3e-12
Identities = 40/105 (38%), Positives = 61/105 (58%), Gaps = 1/105 (0%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKFI 60
RFLQGPETDQ V KIRDA K + +L+NY K+G FWNL + P++D +G FI
Sbjct: 241 RFLQGPETDQTTVQKIRDAIKEQRDITVQLINYTKSGRKFWNLFHLQPMRDQKGELQYFI 300
Query: 61 GMQVEVSKYTEGV-NEKALRPNGLPKSLIRYDARQKEEAMGSITE 104
G+Q++ S + E + N + R +++ A +EA+ + +
Sbjct: 301 GVQLDGSDHVEPLRNRLSERTEIQSAKVVKATAENVDEAVRELPD 345
>UniRef100_Q765V9 Phototropin 2 [Adiantum capillus-veneris]
Length = 1019
Score = 903 bits (2334), Expect = 0.0
Identities = 467/759 (61%), Positives = 573/759 (74%), Gaps = 49/759 (6%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKFI 60
RFLQG +TD ++V +IR++ GK+YCGRLLNYKK+GT FWNLLT+ PIKD+ GN +KFI
Sbjct: 259 RFLQGADTDPDDVERIRESLAEGKNYCGRLLNYKKDGTAFWNLLTIAPIKDEEGNVLKFI 318
Query: 61 GMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSITEVVQTVRNPKSIIRSKN 120
GMQVEVSK+TEG KALRPNGLP+SLI+YDARQK+ A+ ++E++Q V++P +
Sbjct: 319 GMQVEVSKHTEGHKVKALRPNGLPESLIKYDARQKDRAVMDVSELIQAVKHP-------H 371
Query: 121 DDTATIMHEEPENLNHDFVLPKSVEPVNDTTTPGRQTPLKFHGDNNNMSRFSSYEERNNK 180
+ H P ++ SV V T R++ L G +++MS + K
Sbjct: 372 HNGHAPQHHPPSSVKSTIAEVPSVATVPPMTD--RRSSLG-PGKSDSMS------DGIPK 422
Query: 181 SSRKSGITSLKGVK--GKSMSSVGRDKDKTIVEPEVLMTKEIEWSKYEL-----RERDIR 233
R SG SL G+ GKS + + +EPE+LMT++ E + R ++IR
Sbjct: 423 RHRSSGFRSLIGLDKFGKS----AQQEPIEFIEPEILMTRDEETDSLDELDDKERLQEIR 478
Query: 234 QGIDLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETD 293
+GIDLATTLERIEKNFVI+DPRLPD PIIFASDSFLELTEYTREEI+GRNCRFLQG +TD
Sbjct: 479 RGIDLATTLERIEKNFVITDPRLPDNPIIFASDSFLELTEYTREEIIGRNCRFLQGQDTD 538
Query: 294 QATVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDH 353
Q TV +IRDAI++QREITVQL+NYTK+GK+FWNLFHLQPMRDQKGELQYFIGVQLDGS+
Sbjct: 539 QKTVQKIRDAIREQREITVQLLNYTKTGKRFWNLFHLQPMRDQKGELQYFIGVQLDGSEQ 598
Query: 354 LEPLRNRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAVSPRPHKRD 413
LEP++ RLSE +E + AK+V+ATA NV+ AV ELPDANL P+DLWA HS++VS +PHK
Sbjct: 599 LEPIQKRLSEKTEKEGAKIVRATALNVEEAVGELPDANLTPDDLWANHSKSVSAKPHKVH 658
Query: 414 NPSWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLN 473
+ W A+QKI RGEKIGL HF P++PLG GDTGSVHLVEL+G+GEL+A+KAMEKSVMLN
Sbjct: 659 SDLWKALQKIRERGEKIGLKHFRPVKPLGFGDTGSVHLVELRGSGELFAIKAMEKSVMLN 718
Query: 474 RNKSVFYYRFIERALKERLYHYWIILFFQHYTPHF-------------------KQPMKI 514
RNK + ER + L H ++ + + +QP K+
Sbjct: 719 RNK--VHRACAEREILAVLDHPFLPALYASFQTQTHVCLVTDFCPGGELFLLLDRQPRKV 776
Query: 515 LKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQV 574
E++ARFY AE++I LEYLHC GIIYRDLKPEN+LLQ+DGH+VLTDFDLSFITSC PQ+
Sbjct: 777 FSEETARFYLAEIIIALEYLHCQGIIYRDLKPENVLLQRDGHVVLTDFDLSFITSCNPQL 836
Query: 575 VKQ-SLPGNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGI 633
V+ S PG RR+ + PPP F++EPVT SNSFVGTEEYIAPE+ITGA H+SA+DWW +GI
Sbjct: 837 VRPPSPPGRRRKYKQMPPPFFMAEPVTTSNSFVGTEEYIAPEVITGAGHSSAVDWWAVGI 896
Query: 634 LLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSA 693
LLYEM+YGRTPFRGKNRQKTF+N+LHKDLTFPSSIPASLAARQLIN LL RDPA+RLGSA
Sbjct: 897 LLYEMIYGRTPFRGKNRQKTFANVLHKDLTFPSSIPASLAARQLINGLLHRDPANRLGSA 956
Query: 694 TGSNEIKQHPFFRGINWPLIRNMSPPPLDVPLQFIGKDP 732
TG+ EIK H FFRGINWPLIR+M PPPL+ PL+ +GKDP
Sbjct: 957 TGAYEIKNHAFFRGINWPLIRDMVPPPLEAPLELVGKDP 995
Score = 119 bits (299), Expect = 3e-25
Identities = 58/153 (37%), Positives = 95/153 (61%), Gaps = 1/153 (0%)
Query: 237 DLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQAT 296
DL LE ++ FV+SD PD PI++AS F ++T Y+ +E++GRNCRFLQG +TD
Sbjct: 211 DLKDALETFQQTFVVSDATRPDYPILYASAGFFKMTGYSSKEVIGRNCRFLQGADTDPDD 270
Query: 297 VNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLEP 356
V RIR+++ + + +L+NY K G FWNL + P++D++G + FIG+Q++ S H E
Sbjct: 271 VERIRESLAEGKNYCGRLLNYKKDGTAFWNLLTIAPIKDEEGNVLKFIGMQVEVSKHTEG 330
Query: 357 LRNRLSEGSEIQSAKLVKATAENVDGAVRELPD 389
+ + + + + L+K A D AV ++ +
Sbjct: 331 HKVKALRPNGLPES-LIKYDARQKDRAVMDVSE 362
>UniRef100_O49003 NPH1-1 [Avena sativa]
Length = 923
Score = 853 bits (2205), Expect = 0.0
Identities = 445/772 (57%), Positives = 548/772 (70%), Gaps = 44/772 (5%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKFI 60
RFLQG TD E+AKIR A NG +YCGR+LNYKK+GT FWNLLT+ PIKD+ G +KFI
Sbjct: 174 RFLQGSGTDPAEIAKIRQALANGSNYCGRVLNYKKDGTAFWNLLTIAPIKDEEGRVLKFI 233
Query: 61 GMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSITEVVQTVRNPKSIIRSKN 120
GMQVEVSKYTEG + +RPNGLP+SLI+YDARQK++A S++E++ ++NP+S+ S N
Sbjct: 234 GMQVEVSKYTEGNKDTVVRPNGLPESLIKYDARQKDQARSSVSELLLAIKNPRSLSESTN 293
Query: 121 DDTATIMHEEPENLNHDFVLPKSVEPVNDTTTPGRQTPLKFHGDNNNMSRFSSYEERNNK 180
E L D P ++ G + K G ++ + S ER +K
Sbjct: 294 STFKRKSQESVGALTGD-------RPGKRSSESGSRRNSK-SGARTSLQKISEVPERGSK 345
Query: 181 SSRKSGITSLKGVKGKSMSSVGRDKDKTIVEPEVLMTKEIEWSKY--ELRERDIRQGIDL 238
S RKSG+ SL + G ++ +D K E +L + + + ELR +++R+GIDL
Sbjct: 346 S-RKSGLYSLMSLLGMGPGNIEKDMLKPRDEDPLLDSDDERPESFDDELRRKEMRRGIDL 404
Query: 239 ATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQATVN 298
ATTLERIEKNFVI+DPRLPD PIIFASDSFL+LTEY+REEILGRNCRFLQGPETD+ATV
Sbjct: 405 ATTLERIEKNFVITDPRLPDNPIIFASDSFLQLTEYSREEILGRNCRFLQGPETDRATVR 464
Query: 299 RIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLEPLR 358
+IRDAI +Q E+TVQLINYTKSGKKFWNLFHLQPMRDQKG++QYFIGVQLDG++H+
Sbjct: 465 KIRDAIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRDQKGDVQYFIGVQLDGTEHVR--- 521
Query: 359 NRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAVSPRPHKRDNPSWV 418
+ +E + L+K TAEN+D A +ELPDANLRPEDLWA HS+ V P+PH +D+ SW
Sbjct: 522 ----DAAEREGVMLIKKTAENIDEAAKELPDANLRPEDLWANHSKVVLPKPHMKDSASWR 577
Query: 419 AIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRNKSV 478
AIQK+ GE I L HF P++PLG GDTGSVHLVEL TGE +AMKAM+K+VMLNRNK
Sbjct: 578 AIQKVLEGGENIDLKHFRPVKPLGSGDTGSVHLVELLNTGEYFAMKAMDKNVMLNRNK-- 635
Query: 479 FYYRFIERALKERLYHYWIILFFQHY---------TPHF----------KQPMKILKEDS 519
+ ER + + L H ++ + + T ++ +QP+K+L+ED+
Sbjct: 636 VHRANAEREILDMLDHPFLPTLYASFQTKTHICLITDYYPGGELFLLLDRQPLKVLREDA 695
Query: 520 ARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSL 579
RFYAAEVVI LEYLHC GIIYRDLKPEN+LL +DGHI LTDFDLS +TSC+PQV
Sbjct: 696 VRFYAAEVVIALEYLHCQGIIYRDLKPENILLHRDGHISLTDFDLSCLTSCRPQVFLPEE 755
Query: 580 PGNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEML 639
+ R +S+ PIF +EP+ SNSFVGTEEYIAPEIITGA HTSA+DWW LGILLYEML
Sbjct: 756 ANKKSRRKSRSSPIFFAEPMRASNSFVGTEEYIAPEIITGAGHTSAVDWWALGILLYEML 815
Query: 640 YGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSATGSNEI 699
YG TPFRGK RQ+TF+NILHKD+ FP+SI SL ARQLI LL RDP++RLGS GSNEI
Sbjct: 816 YGYTPFRGKTRQRTFANILHKDIRFPASISVSLPARQLIYRLLHRDPSNRLGSYEGSNEI 875
Query: 700 KQHPFFRGINWPLIRNMSPPPLDVPLQFIGKDPTAKDKKWEDDGVLNTSIDM 751
K+HPFFRGINW L+R +PP LD PL P DK D +T DM
Sbjct: 876 KEHPFFRGINWALVRGTAPPKLDAPL-----FPDDTDKGMGDAAAADTHTDM 922
Score = 106 bits (265), Expect = 2e-21
Identities = 59/164 (35%), Positives = 93/164 (55%), Gaps = 4/164 (2%)
Query: 237 DLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQAT 296
+L L ++ FV+SD P PI++AS F +T YT +E++GRNCRFLQG TD A
Sbjct: 126 ELRAALSAFQQTFVVSDASRPGHPIMYASAGFFNMTGYTSKEVVGRNCRFLQGSGTDPAE 185
Query: 297 VNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLEP 356
+ +IR A+ + +++NY K G FWNL + P++D++G + FIG+Q++ S + E
Sbjct: 186 IAKIRQALANGSNYCGRVLNYKKDGTAFWNLLTIAPIKDEEGRVLKFIGMQVEVSKYTEG 245
Query: 357 LRNRLSEGSEIQSAKLVKATAENVDGA---VRELPDANLRPEDL 397
++ + + + + L+K A D A V EL A P L
Sbjct: 246 NKDTVVRPNGLPES-LIKYDARQKDQARSSVSELLLAIKNPRSL 288
>UniRef100_O49004 NPH1-2 [Avena sativa]
Length = 927
Score = 852 bits (2201), Expect = 0.0
Identities = 440/752 (58%), Positives = 543/752 (71%), Gaps = 39/752 (5%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKFI 60
RFLQG TD E+AKIR A +G +YCGR+LNYKK+GT FWNLLT+ PIKD+ G +KFI
Sbjct: 177 RFLQGSGTDPAEIAKIRQALADGSNYCGRVLNYKKDGTAFWNLLTIAPIKDEEGRVLKFI 236
Query: 61 GMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSITEVVQTVRNPKSIIRSKN 120
GMQVEVSKYTEG + A+RPNGLP+SLI+YDARQK++A S++E++ ++NP+S+ S N
Sbjct: 237 GMQVEVSKYTEGNKDTAVRPNGLPESLIKYDARQKDQARSSVSELLLAIKNPRSLSESTN 296
Query: 121 DDTATIMHEEPENLNHDFVLPKSVEPVNDTTTPGRQTPLKFHGDNNNMSRFSSYEERNNK 180
E L D P ++ G + K G ++ + S ER NK
Sbjct: 297 STFKRKSQESVGPLTGD-------RPGKRSSESGSRRNSK-SGARTSLQKISEVPERGNK 348
Query: 181 SSRKSGITSLKGVKGKSMSSVGRDKDKTIVEPEVLMTKEIEWSKY--ELRERDIRQGIDL 238
S RKSG+ SL + G ++ +D K E +L + + + ELR +++R+GIDL
Sbjct: 349 S-RKSGLYSLMSLLGMGPGNIEKDMLKPRDEDPLLDSDDERPESFDDELRRKEMRRGIDL 407
Query: 239 ATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQATVN 298
ATTLERIEKNFVI+DPRLPD PIIFASDSFL+LTEY+REEILGRNCRFLQGPETD+ATV
Sbjct: 408 ATTLERIEKNFVITDPRLPDNPIIFASDSFLQLTEYSREEILGRNCRFLQGPETDRATVR 467
Query: 299 RIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLEPLR 358
+IRDAI +Q E+TVQLINYTKSGKKFWNLFHLQPMRDQKG++QYFIGVQLDG++H+
Sbjct: 468 KIRDAIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRDQKGDVQYFIGVQLDGTEHVR--- 524
Query: 359 NRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAVSPRPHKRDNPSWV 418
+ +E + L+K TAEN+D A +ELPDANLRPEDLWA HS+ V P+PH +D+ SW
Sbjct: 525 ----DAAEREGVMLIKKTAENIDEAAKELPDANLRPEDLWANHSKVVLPKPHMKDSASWR 580
Query: 419 AIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRNKSV 478
AIQK+ GE I L HF P++PLG GDTGSVHLVEL TGE +AMKAM+K+VMLNRNK
Sbjct: 581 AIQKVLEGGENIDLKHFRPVKPLGSGDTGSVHLVELLNTGEYFAMKAMDKNVMLNRNK-- 638
Query: 479 FYYRFIERALKERLYHYWIILFFQHY---------TPHF----------KQPMKILKEDS 519
+ ER + + L H ++ + + T ++ +QP+K+L+ED+
Sbjct: 639 VHRANAEREILDMLDHPFLPTLYASFQTKTHICLITDYYPGGELFLLLDRQPLKVLREDA 698
Query: 520 ARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSL 579
RFYAAEVVI LEYLHC GIIYRDLKPEN+LL +DGHI LTDFDLS +TSC+PQV
Sbjct: 699 VRFYAAEVVIALEYLHCQGIIYRDLKPENILLHRDGHISLTDFDLSCLTSCRPQVFLPEE 758
Query: 580 PGNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEML 639
+ R +S+ P+F +EP+ SNSFVGTEEYIAPEIITGA HTSA+DWW LGILLYEML
Sbjct: 759 ANKKSRRKSRSSPVFFAEPMRASNSFVGTEEYIAPEIITGAGHTSAVDWWALGILLYEML 818
Query: 640 YGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSATGSNEI 699
YG TPFRGK RQ+TF+NILHKD+ FP+SI SL ARQLI LL RDP++RLGS GSNEI
Sbjct: 819 YGYTPFRGKTRQRTFANILHKDIRFPASISVSLPARQLIYRLLHRDPSNRLGSYEGSNEI 878
Query: 700 KQHPFFRGINWPLIRNMSPPPLDVPLQFIGKD 731
K+HPFFRGINW L+R +PP LD PL G D
Sbjct: 879 KEHPFFRGINWALVRGTAPPKLDAPLFPDGTD 910
Score = 108 bits (269), Expect = 8e-22
Identities = 76/239 (31%), Positives = 118/239 (48%), Gaps = 20/239 (8%)
Query: 237 DLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQAT 296
+L L ++ FV+SD P PI++AS F +T YT +E++GRNCRFLQG TD A
Sbjct: 129 ELRAALSAFQQTFVVSDASRPGHPIMYASAGFFNMTGYTSKEVVGRNCRFLQGSGTDPAE 188
Query: 297 VNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLEP 356
+ +IR A+ D +++NY K G FWNL + P++D++G + FIG+Q++ S + E
Sbjct: 189 IAKIRQALADGSNYCGRVLNYKKDGTAFWNLLTIAPIKDEEGRVLKFIGMQVEVSKYTEG 248
Query: 357 LRNRLSEGSEIQSAKLVKATAENVDGA---VRELPDANLRPEDLWAI--------HSQAV 405
++ + + + L+K A D A V EL A P L ++V
Sbjct: 249 NKDTAVRPNGLPES-LIKYDARQKDQARSSVSELLLAIKNPRSLSESTNSTFKRKSQESV 307
Query: 406 SP----RPHKRDNPSWVAIQKITARGEKIGLHHFSPIRPLG--CGDTGSVHLVELQGTG 458
P RP KR + S ++ + G + L S + G +G L+ L G G
Sbjct: 308 GPLTGDRPGKRSSES--GSRRNSKSGARTSLQKISEVPERGNKSRKSGLYSLMSLLGMG 364
>UniRef100_Q8H935 Phototropin [Vicia faba]
Length = 963
Score = 847 bits (2188), Expect = 0.0
Identities = 449/778 (57%), Positives = 550/778 (69%), Gaps = 43/778 (5%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKFI 60
RF+QG +TD N+VAKIR+A G SYCGRLLNYKK+GT FWNLLT+ PIKD+ G +K I
Sbjct: 199 RFMQGADTDPNDVAKIREALAAGTSYCGRLLNYKKDGTTFWNLLTIAPIKDEHGKILKLI 258
Query: 61 GMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSITEVVQTV-RNPKSIIRSK 119
GMQVEVSK+TEG EK LRPNGLP+SLIRYDARQKE+A S+TE+V+ V + P+S+ S
Sbjct: 259 GMQVEVSKHTEGTKEKMLRPNGLPESLIRYDARQKEKANSSVTELVEAVSKRPRSLSESA 318
Query: 120 NDDTATIMHEEPENLNHDFVLPKSVEPVN----DTTTPGRQTPLKFHGDNNNMSRFSSYE 175
N +++P N ++D P + E + T R+ G++N+M +
Sbjct: 319 N---RLPFNKKPTNGSNDHATPPNSESSSRKSGSTLRSFRRKSHSGAGNSNSMHPITELP 375
Query: 176 ERNNKSSRKSGITSLKGVKGKSMSSVGRDKDKTIV-----EPEVLMTKEIEWSKYELRER 230
E NNKS R+S G KS+S+ R + ++ E E + E + + RE+
Sbjct: 376 ENNNKSRRRS----FMGFMRKSLSNNERFNHEQVIDRNSSEDEDRLDSFDEQNIAQKREK 431
Query: 231 DIRQGIDLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGP 290
R+G DLATTLERIEKNFVI+DPRLPD PIIFASDSFLELTEY+REEILGRNCRFLQGP
Sbjct: 432 --RKGFDLATTLERIEKNFVITDPRLPDNPIIFASDSFLELTEYSREEILGRNCRFLQGP 489
Query: 291 ETDQATVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDG 350
ETD ATV +IR AI +Q E+TVQLINYTK+GKKFWNLFHLQPMRDQKGE+QYFIGVQLDG
Sbjct: 490 ETDPATVKKIRYAIDNQTEVTVQLINYTKTGKKFWNLFHLQPMRDQKGEVQYFIGVQLDG 549
Query: 351 SDHLEPLRNRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAVSPRPH 410
S H+EPL N ++E + + LVK TAENVD A+RELPDAN++PEDLW HS+ V P+PH
Sbjct: 550 SQHVEPLHNGIAEDTAKEGENLVKKTAENVDDALRELPDANMKPEDLWMNHSKVVHPKPH 609
Query: 411 KRDNPSWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSV 470
+R++ +W AIQKI GE+IGL HF PI+PLG GDTGSVHLVEL GT +AMKAM+K V
Sbjct: 610 RREDSAWRAIQKIMESGEQIGLKHFKPIKPLGSGDTGSVHLVELCGTDHHFAMKAMDKGV 669
Query: 471 MLNRNKSVFYYRFIERALKERLYHYWI------------ILFFQHYTPH-------FKQP 511
M NRNK + ER + + L H ++ I Y P +QP
Sbjct: 670 MPNRNK--VHRACTEREILDMLDHPFLPALYASFQTKTHICLITDYCPGGELFMLLDRQP 727
Query: 512 MKILKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCK 571
K+LKED+ RFYA EVV+ LEYLHC GIIYRDLKPEN+LLQ GH+ LTDFDLS +TSCK
Sbjct: 728 AKVLKEDAVRFYATEVVVALEYLHCQGIIYRDLKPENVLLQSTGHVSLTDFDLSCLTSCK 787
Query: 572 PQVVKQSLPGNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTL 631
P+++ +P + + Q PIF++EP+ SNSFVGTEEYIAPEIITG+ HT A+DWW L
Sbjct: 788 PELI---VPSTNDKKKGQHGPIFMAEPMRASNSFVGTEEYIAPEIITGSGHTCAVDWWAL 844
Query: 632 GILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLG 691
GILLYEM YG TPFRGKNRQ+TF+NILHKDL P S SL+A+QLI LLQRDP SRLG
Sbjct: 845 GILLYEMFYGYTPFRGKNRQRTFANILHKDLKLPKSKQVSLSAKQLIYHLLQRDPTSRLG 904
Query: 692 SATGSNEIKQHPFFRGINWPLIRNMSPPPLDVPLQFIGKDPTAKDKKWEDDGVLNTSI 749
S G+N+IK H FF+GINW L+R PP LD PL K+ KD K+ D+G + S+
Sbjct: 905 SKGGANDIKHHSFFKGINWALVRCTKPPELDAPLFDTNKEEKEKDDKYVDNGQEDMSV 962
Score = 113 bits (283), Expect = 2e-23
Identities = 60/165 (36%), Positives = 97/165 (58%), Gaps = 5/165 (3%)
Query: 237 DLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQAT 296
DL L ++ FV+SD PD PI++AS F +T YT +E++GRNCRF+QG +TD
Sbjct: 151 DLRDALSAFQQTFVVSDATKPDYPIMYASAGFFSMTGYTSKEVIGRNCRFMQGADTDPND 210
Query: 297 VNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLEP 356
V +IR+A+ +L+NY K G FWNL + P++D+ G++ IG+Q++ S H E
Sbjct: 211 VAKIREALAAGTSYCGRLLNYKKDGTTFWNLLTIAPIKDEHGKILKLIGMQVEVSKHTEG 270
Query: 357 LRNRLSEGSEIQSAKLVKATA---ENVDGAVRELPDA-NLRPEDL 397
+ ++ + + + L++ A E + +V EL +A + RP L
Sbjct: 271 TKEKMLRPNGLPES-LIRYDARQKEKANSSVTELVEAVSKRPRSL 314
>UniRef100_O48547 Nonphototropic hypocotyl 1 [Zea mays]
Length = 911
Score = 842 bits (2174), Expect = 0.0
Identities = 443/751 (58%), Positives = 542/751 (71%), Gaps = 49/751 (6%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKFI 60
RFLQG TD E++KIR A G +YCGR+LNYKK+GTPFWNLLTV PIKD+ G +KFI
Sbjct: 165 RFLQGSGTDPVEISKIRQALAAGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFI 224
Query: 61 GMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSITEVVQTVRNPKSIIRSKN 120
GMQVEVSKYTEG + ALRPNGLP+SLI+YDARQK+ A S++E++ +++P+S+ S+N
Sbjct: 225 GMQVEVSKYTEGNKDTALRPNGLPESLIKYDARQKDHARSSVSELLLALKDPRSLSESRN 284
Query: 121 DDTATIMHEEPENLNHDFVLPKSVEPVNDTTTPGRQTPLKFHGDNNNMSRFSSYEERNNK 180
+ T+ + E+ V K + T R+ G N++ + S E NK
Sbjct: 285 N---TLKRKSQESAGSALVPGK-----RSSETGSRRN--SHSGMRNSLQKISEVPEGGNK 334
Query: 181 SSRKSGITSLKGVKGKSMSSVGRDKDKTIVEP--EVLMTKEIEWSKY---ELRERDIRQG 235
+ RKSG+ S G+ G +V +K I++P + L+ + E + R++++R+G
Sbjct: 335 T-RKSGLRSFMGLIGMGHGNV----EKNILKPREDPLLDSDDERPDSFDDDFRKKEMRKG 389
Query: 236 IDLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQA 295
IDLATTLERIEKNFVI+DPRLPD PIIFASDSFL LTEY REEILGRNCRFLQGPETD+
Sbjct: 390 IDLATTLERIEKNFVITDPRLPDNPIIFASDSFLRLTEYCREEILGRNCRFLQGPETDRG 449
Query: 296 TVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLE 355
TV +IRDAI +Q E+TVQLINYTKSGKKFWNLFHLQPMRDQKG++QYFIGVQLDG++
Sbjct: 450 TVKKIRDAIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRDQKGDVQYFIGVQLDGTE--- 506
Query: 356 PLRNRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAVSPRPHKRDNP 415
R+ E + A LVK TA+N+D A +ELPDANLRPEDLWA HS+ V P+PH +D
Sbjct: 507 ----RVREAAAKDGAILVKKTADNIDEAAKELPDANLRPEDLWANHSKPVLPKPHMKDTA 562
Query: 416 SWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRN 475
SW AIQK+ GE I L HF P+RPLG GDTGSVHLVEL GTGE +AMKAM+KSVMLNRN
Sbjct: 563 SWRAIQKVLENGESIDLKHFRPVRPLGSGDTGSVHLVELLGTGEYFAMKAMDKSVMLNRN 622
Query: 476 KSVFYYRFIERALKERLYHYWIILFFQHYTPHF-------------------KQPMKILK 516
K + ER + + L H ++ + + +QPMK+LK
Sbjct: 623 K--VHRATAERQILDMLDHPFLPTLYASFQTKTHICLIVDYCAGGELFMLLDRQPMKVLK 680
Query: 517 EDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQV-V 575
ED+ RFYAAEVV LEYLHC GIIYRDLKPEN+LL +DGHI LTDFDLS +TSC+PQV +
Sbjct: 681 EDAVRFYAAEVVTALEYLHCQGIIYRDLKPENILLHRDGHISLTDFDLSCLTSCRPQVFL 740
Query: 576 KQSLPGNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILL 635
+ ++R +S+ PIF +EP+ SNSFVGTEEYIAPEIITGA HTSA+DWW LGILL
Sbjct: 741 PHDIDKKKKRRKSRSNPIFFAEPMRASNSFVGTEEYIAPEIITGAGHTSAVDWWALGILL 800
Query: 636 YEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSATG 695
YEMLYG TPFRGK RQ+TF+NILHKD+ FP+SI SLAARQLI LL RDPA+RLGS G
Sbjct: 801 YEMLYGYTPFRGKTRQRTFANILHKDIRFPASIQVSLAARQLIYRLLHRDPANRLGSYEG 860
Query: 696 SNEIKQHPFFRGINWPLIRNMSPPPLDVPLQ 726
+ EIKQHPFFRGINW L+R +PP L+ PLQ
Sbjct: 861 AMEIKQHPFFRGINWALVRAATPPELEAPLQ 891
Score = 104 bits (260), Expect = 8e-21
Identities = 54/160 (33%), Positives = 89/160 (54%), Gaps = 1/160 (0%)
Query: 237 DLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQAT 296
+L L ++ FV+SD PD PI++AS F +T Y+ E++GRNCRFLQG TD
Sbjct: 117 ELRAALSAFQQTFVVSDATRPDHPILYASAGFFNMTGYSSNEVVGRNCRFLQGSGTDPVE 176
Query: 297 VNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLEP 356
+++IR A+ +++NY K G FWNL + P++D+ G + FIG+Q++ S + E
Sbjct: 177 ISKIRQALAAGSNYCGRILNYKKDGTPFWNLLTVAPIKDEDGRVLKFIGMQVEVSKYTEG 236
Query: 357 LRNRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPED 396
++ + + + L+K A D A + + L +D
Sbjct: 237 NKDTALRPNGLPES-LIKYDARQKDHARSSVSELLLALKD 275
>UniRef100_Q9ST26 Nonphototrophic hypocotyl 1a [Oryza sativa]
Length = 921
Score = 837 bits (2161), Expect = 0.0
Identities = 436/772 (56%), Positives = 550/772 (70%), Gaps = 46/772 (5%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKFI 60
RFLQG TD +E+ KIR + NG +YCGR+LNYKK+GTPFWNLLT+ PIKD+ G +KFI
Sbjct: 174 RFLQGSGTDPHEIDKIRQSLANGSNYCGRILNYKKDGTPFWNLLTIAPIKDEDGRLLKFI 233
Query: 61 GMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSITEVVQTVRNPKSIIRSKN 120
GMQVEVSKYTEG + +RPNGL +SLI+YDARQK+ A S++E++ ++NP+S+ S N
Sbjct: 234 GMQVEVSKYTEGKKDTVVRPNGLSESLIKYDARQKDHARSSVSELLLALKNPRSLSESSN 293
Query: 121 DDTATIMHEEPENLNHDFVLPKSVEPVNDTTTPGRQTPLKFHGDNNNMSRFSSYEERNNK 180
+ T+ + E+L+ + + P ++ G + + G +++ + + ++ N+
Sbjct: 294 N---TLKRKSQESLS----MSMTEVPSKRSSESGSRRNSR-SGTRSSLQKINEVPDQGNR 345
Query: 181 SSRKSGITSLKGVKGKSMSSVGRDKDKTIVEPEVLMTKEIEWSKYE--LRERDIRQGIDL 238
+ RKSG+ + G G SV ++ K E ++ + + +E R +++R+GIDL
Sbjct: 346 T-RKSGLRAFMGFLGMGHGSVEKNMLKPRDEDPLIDSDDERPESFEDEFRRKEMRRGIDL 404
Query: 239 ATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQATVN 298
ATTLERIEKNFVI+DPRLPD PIIFASDSFL+LTEY REEILGRNCRFLQGPETD+ATV
Sbjct: 405 ATTLERIEKNFVITDPRLPDNPIIFASDSFLQLTEYNREEILGRNCRFLQGPETDRATVR 464
Query: 299 RIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLEPLR 358
+IRDAI +Q E+TVQLINYTKSGKKFWNLFHLQPMRDQKG++QYFIGVQLDG++H++
Sbjct: 465 KIRDAIDNQAEVTVQLINYTKSGKKFWNLFHLQPMRDQKGDVQYFIGVQLDGTEHVQ--- 521
Query: 359 NRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAVSPRPHKRDNPSWV 418
+ + + LVK TA+N+D A +ELPDANLRPEDLWA HS+ V P PH +D SW
Sbjct: 522 ----DDAAKEGVVLVKKTADNIDEAAKELPDANLRPEDLWANHSKVVLPNPHMKDTASWR 577
Query: 419 AIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRNKSV 478
AIQK+ GE IGL HF P++PLG GDTGSVHLVEL TGE +AMKAM+KS+MLNRNK
Sbjct: 578 AIQKVLESGESIGLKHFRPVKPLGSGDTGSVHLVELLNTGEYFAMKAMDKSIMLNRNK-- 635
Query: 479 FYYRFIERALKERLYHYWI------------ILFFQHYTPHFK-------QPMKILKEDS 519
+ ER + + L H ++ I Y P + QP+K+L ED+
Sbjct: 636 VHRATAERQILDLLDHPFLPTLYASFQTKTHICLITDYCPGGELFVLLDNQPLKVLHEDA 695
Query: 520 ARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSL 579
RFYAAEVV+ LEYLHC GIIYRDLKPEN+LL +DGHI LTDFDLS +TSC+PQV
Sbjct: 696 VRFYAAEVVVALEYLHCQGIIYRDLKPENILLHRDGHISLTDFDLSCLTSCRPQVFLPED 755
Query: 580 PGNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEML 639
++ ++ PIF +EP+ SNSFVGTEEYIAPEIITGA HTSA+DWW LGILLYEML
Sbjct: 756 ADEKKGRKNGSYPIFFAEPMRASNSFVGTEEYIAPEIITGAGHTSAVDWWALGILLYEML 815
Query: 640 YGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSATGSNEI 699
YG TPFRGK RQ+TF+NILHKD+ FP+SI SLAARQL+ LL RDPA+RLGS G+NEI
Sbjct: 816 YGYTPFRGKTRQRTFANILHKDIRFPASISVSLAARQLMYRLLHRDPANRLGSYEGANEI 875
Query: 700 KQHPFFRGINWPLIRNMSPPPLDVPLQFIGKDPTAKDKKWEDDGVLNTSIDM 751
K HPFFRGINWPLIR +PP L++PL +KD + V NT DM
Sbjct: 876 KGHPFFRGINWPLIRATAPPKLEIPL-------FSKDDMEKKGLVTNTRTDM 920
Score = 106 bits (264), Expect = 3e-21
Identities = 59/164 (35%), Positives = 93/164 (55%), Gaps = 4/164 (2%)
Query: 237 DLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQAT 296
+L L ++ FV+SD P+ PI++AS F +T YT +E++GRNCRFLQG TD
Sbjct: 126 ELRAALSVFQQTFVVSDATHPNHPIMYASAGFFNMTGYTSKEVVGRNCRFLQGSGTDPHE 185
Query: 297 VNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLEP 356
+++IR ++ + +++NY K G FWNL + P++D+ G L FIG+Q++ S + E
Sbjct: 186 IDKIRQSLANGSNYCGRILNYKKDGTPFWNLLTIAPIKDEDGRLLKFIGMQVEVSKYTEG 245
Query: 357 LRNRLSEGSEIQSAKLVKATAENVDGA---VRELPDANLRPEDL 397
++ + + + S L+K A D A V EL A P L
Sbjct: 246 KKDTVVRPNGL-SESLIKYDARQKDHARSSVSELLLALKNPRSL 288
>UniRef100_O48963 Nonphototropic hypocotyl protein 1 [Arabidopsis thaliana]
Length = 996
Score = 835 bits (2158), Expect = 0.0
Identities = 438/753 (58%), Positives = 547/753 (72%), Gaps = 43/753 (5%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKFI 60
RFLQG TD +E+AKIR+ G +YCGR+LNYKK+GT FWNLLT+ PIKD+ G +KFI
Sbjct: 235 RFLQGSGTDADELAKIRETLAAGNNYCGRILNYKKDGTSFWNLLTIAPIKDESGKVLKFI 294
Query: 61 GMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSITEVVQTVRNPKSIIRSKN 120
GMQVEVSK+TEG EKALRPNGLP+SLIRYDARQK+ A S+TE+V+ V+ P+++ S N
Sbjct: 295 GMQVEVSKHTEGAKEKALRPNGLPESLIRYDARQKDMATNSVTELVEAVKRPRALSESTN 354
Query: 121 DDTATIMHEEPENLNHDFVLPKSVEPVNDTTTP-GRQTPLKFHGDNNNMSRFSSYEERNN 179
+H D + K +++ P GR+ G N+M R + E
Sbjct: 355 ------LHPFMTKSESDELPKKPARRMSENVVPSGRRN--SGGGRRNSMQRINEIPE--- 403
Query: 180 KSSRKSGITSLKGVKGKSMSSVGRDKDKTIVE-----PEVLMTKEIEWSKYE-LRERDIR 233
K SRKS + S G+K KS S+ D +E E+ E S + +R++++R
Sbjct: 404 KKSRKSSL-SFMGIKKKS-ESLDESIDDGFIEYGEEDDEISDRDERPESVDDKVRQKEMR 461
Query: 234 QGIDLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETD 293
+GIDLATTLERIEKNFVI+DPRLPD PIIFASDSFLELTEY+REEILGRNCRFLQGPETD
Sbjct: 462 KGIDLATTLERIEKNFVITDPRLPDNPIIFASDSFLELTEYSREEILGRNCRFLQGPETD 521
Query: 294 QATVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDH 353
TV +IR+AI +Q E+TVQLINYTKSGKKFWN+FHLQPMRDQKGE+QYFIGVQLDGS H
Sbjct: 522 LTTVKKIRNAIDNQTEVTVQLINYTKSGKKFWNIFHLQPMRDQKGEVQYFIGVQLDGSKH 581
Query: 354 LEPLRNRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAVSPRPHKRD 413
+EP+RN + E + + LVK TA N+D AVRELPDAN+ PEDLWA HS+ V +PH++D
Sbjct: 582 VEPVRNVIEETAVKEGEDLVKKTAVNIDEAVRELPDANMTPEDLWANHSKVVHCKPHRKD 641
Query: 414 NPSWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLN 473
+P W+AIQK+ GE IGL HF P++PLG GDTGSVHLVEL GT +L+AMKAM+K+VMLN
Sbjct: 642 SPPWIAIQKVLESGEPIGLKHFKPVKPLGSGDTGSVHLVELVGTDQLFAMKAMDKAVMLN 701
Query: 474 RNKSVFYYRFIERALKERLYHYWIILFFQHY---------TPHF----------KQPMKI 514
RNK + ER + + L H ++ + + T ++ +QP K+
Sbjct: 702 RNK--VHRARAEREILDLLDHPFLPALYASFQTKTHICLITDYYPGGELFMLLDRQPRKV 759
Query: 515 LKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQV 574
LKED+ RFYAA+VV+ LEYLHC GIIYRDLKPEN+L+Q +G I L+DFDLS +TSCKPQ+
Sbjct: 760 LKEDAVRFYAAQVVVALEYLHCQGIIYRDLKPENVLIQGNGDISLSDFDLSCLTSCKPQL 819
Query: 575 VKQSLPGNRRR--SRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLG 632
+ S+ +++ +SQ PIF++EP+ SNSFVGTEEYIAPEII+GA HTSA+DWW LG
Sbjct: 820 LIPSIDEKKKKKQQKSQQTPIFMAEPMRASNSFVGTEEYIAPEIISGAGHTSAVDWWALG 879
Query: 633 ILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGS 692
IL+YEMLYG TPFRGK RQKTF+N+L KDL FP+SIPASL +QLI LLQRDP RLG
Sbjct: 880 ILMYEMLYGYTPFRGKTRQKTFTNVLQKDLKFPASIPASLQVKQLIFRLLQRDPKKRLGC 939
Query: 693 ATGSNEIKQHPFFRGINWPLIRNMSPPPLDVPL 725
G+NE+KQH FF+GINW LIR +PP L+ P+
Sbjct: 940 FEGANEVKQHSFFKGINWALIRCTNPPELETPI 972
Score = 112 bits (281), Expect = 3e-23
Identities = 65/186 (34%), Positives = 102/186 (53%), Gaps = 12/186 (6%)
Query: 223 SKYELRERDI---RQGI-----DLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEY 274
S E+ + D+ R GI DL L ++ FV+SD PD PI++AS F +T Y
Sbjct: 165 SSGEMSDGDVPGGRSGIPRVSEDLKDALSTFQQTFVVSDATKPDYPIMYASAGFFNMTGY 224
Query: 275 TREEILGRNCRFLQGPETDQATVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMR 334
T +E++GRNCRFLQG TD + +IR+ + +++NY K G FWNL + P++
Sbjct: 225 TSKEVVGRNCRFLQGSGTDADELAKIRETLAAGNNYCGRILNYKKDGTSFWNLLTIAPIK 284
Query: 335 DQKGELQYFIGVQLDGSDHLEPLRNRLSEGSEIQSAKLVKATAENVD---GAVRELPDAN 391
D+ G++ FIG+Q++ S H E + + + + + L++ A D +V EL +A
Sbjct: 285 DESGKVLKFIGMQVEVSKHTEGAKEKALRPNGLPES-LIRYDARQKDMATNSVTELVEAV 343
Query: 392 LRPEDL 397
RP L
Sbjct: 344 KRPRAL 349
>UniRef100_Q9SC66 Non-phototropic hypocotyl NPH1 [Oryza sativa]
Length = 921
Score = 834 bits (2155), Expect = 0.0
Identities = 432/746 (57%), Positives = 540/746 (71%), Gaps = 39/746 (5%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKFI 60
RFLQG TD +E+ KIR A NG +YCGR+LNYKK+GTPFWNLLT+ PIKD+ G +KFI
Sbjct: 174 RFLQGSGTDPHEIDKIRQALANGSNYCGRILNYKKDGTPFWNLLTIAPIKDEDGRLLKFI 233
Query: 61 GMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSITEVVQTVRNPKSIIRSKN 120
GMQVEVSKYTEG E +RPNGL +SLI+YDARQK+ A S++E++ ++NP+S+ S N
Sbjct: 234 GMQVEVSKYTEGKKETVVRPNGLSESLIKYDARQKDHARSSVSELLLALKNPRSLSESSN 293
Query: 121 DDTATIMHEEPENLNHDFVLPKSVEPVNDTTTPGRQTPLKFHGDNNNMSRFSSYEERNNK 180
+ T+ + E+L+ + S P ++ G + + G +++ + + ++ N+
Sbjct: 294 N---TLKRKSQESLS----MSMSEVPSKRSSESGSRRNSR-SGTRSSLQKINEVPDQVNR 345
Query: 181 SSRKSGITSLKGVKGKSMSSVGRDKDKTIVEPEVLMTKEIEWSKYE--LRERDIRQGIDL 238
+ RKSG+ + G G SV ++ K E ++ + + +E R +++R+GIDL
Sbjct: 346 T-RKSGLRAFMGFLGMGHGSVEKNMLKPRDEDPLIDSDDERPESFEDEFRRKEMRRGIDL 404
Query: 239 ATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQATVN 298
ATTLERIEKNFVI+DPRLPD PIIFASDSFL+LTEY REEILGRNCRFLQGPETD+A V
Sbjct: 405 ATTLERIEKNFVITDPRLPDNPIIFASDSFLQLTEYNREEILGRNCRFLQGPETDRAIVR 464
Query: 299 RIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLEPLR 358
+IRDAI +Q E+TVQLINYTKSGKKFWNLFHLQPMRDQKG++QYFIGVQLDG++H++
Sbjct: 465 KIRDAIDNQAEVTVQLINYTKSGKKFWNLFHLQPMRDQKGDVQYFIGVQLDGTEHVQ--- 521
Query: 359 NRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAVSPRPHKRDNPSWV 418
+ + + LVK TA+N+D A +ELPDANLRPEDLWA HS+ V P PH +D SW
Sbjct: 522 ----DDAAEEGVVLVKKTADNIDEAAKELPDANLRPEDLWANHSKVVLPNPHMKDTASWR 577
Query: 419 AIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRNKSV 478
AIQK+ GE IGL HF PI+PLG GDTGSVHLVEL TGE +AMKAM+KS+MLNRNK
Sbjct: 578 AIQKVLESGESIGLKHFRPIKPLGSGDTGSVHLVELLNTGEYFAMKAMDKSIMLNRNK-- 635
Query: 479 FYYRFIERALKERLYHYWI------------ILFFQHYTPHFK-------QPMKILKEDS 519
+ ER + + L H ++ I Y P + QP+K+L ED+
Sbjct: 636 VHRATAERQILDLLDHPFLPTLYASFQTKTHICLITDYCPGGELFVLLDNQPLKVLHEDA 695
Query: 520 ARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSL 579
RFY AEVVI LEYLHC GIIYRDLKPEN+LL +DGHI LTDFDLS +TSC+PQV
Sbjct: 696 VRFYVAEVVIALEYLHCQGIIYRDLKPENILLHRDGHISLTDFDLSCLTSCRPQVFLPED 755
Query: 580 PGNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEML 639
++ +++ PIF +EP+ SNSFVGTEEYIAPEIITGA HTSA+DWW LGILLYEML
Sbjct: 756 ADEKKGRKNRSYPIFFAEPMRASNSFVGTEEYIAPEIITGAGHTSAVDWWALGILLYEML 815
Query: 640 YGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSATGSNEI 699
YG TPFRGK RQ+TF+NILHKD+ FP+SI SLAARQL+ LL RDPA+RLGS G+NEI
Sbjct: 816 YGYTPFRGKTRQRTFANILHKDIRFPASILVSLAARQLMYRLLHRDPANRLGSYEGANEI 875
Query: 700 KQHPFFRGINWPLIRNMSPPPLDVPL 725
K HPFFRGINWPLIR +PP L++PL
Sbjct: 876 KGHPFFRGINWPLIRATAPPKLEIPL 901
Score = 108 bits (269), Expect = 8e-22
Identities = 60/164 (36%), Positives = 92/164 (55%), Gaps = 4/164 (2%)
Query: 237 DLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQAT 296
+L L ++ FV+SD P+ PI++AS F +T YT +E++GRNCRFLQG TD
Sbjct: 126 ELRAALSAFQQTFVVSDATRPNHPIMYASAGFFNMTGYTSKEVVGRNCRFLQGSGTDPHE 185
Query: 297 VNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLEP 356
+++IR A+ + +++NY K G FWNL + P++D+ G L FIG+Q++ S + E
Sbjct: 186 IDKIRQALANGSNYCGRILNYKKDGTPFWNLLTIAPIKDEDGRLLKFIGMQVEVSKYTEG 245
Query: 357 LRNRLSEGSEIQSAKLVKATAENVDGA---VRELPDANLRPEDL 397
+ + + + S L+K A D A V EL A P L
Sbjct: 246 KKETVVRPNGL-SESLIKYDARQKDHARSSVSELLLALKNPRSL 288
>UniRef100_Q9MB43 Phototropin [Adiantum capillus-veneris]
Length = 1092
Score = 826 bits (2134), Expect = 0.0
Identities = 424/775 (54%), Positives = 548/775 (70%), Gaps = 43/775 (5%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKFI 60
RFLQG ETD E+ +IR+ G YCGRLLNYKK+G+ FWNLLT++PIKD G+ +K+I
Sbjct: 317 RFLQGKETDPEEIDRIRECISKGSGYCGRLLNYKKDGSAFWNLLTISPIKDVDGSVLKYI 376
Query: 61 GMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSITEVVQTVRNPKSIIRSKN 120
GMQVEVS++TEG E A+RPNGL +SLI+YDARQKE A +TE+V+ +++PK + K
Sbjct: 377 GMQVEVSQFTEGTKENAMRPNGLSESLIKYDARQKERASFQVTELVEAIKDPKQVGDDKK 436
Query: 121 DDTA--TIMHEEPENLNHDFVLPKSVEPVNDTTTPGRQTP---LKFHGDNNNMSRFSSYE 175
+ + P P+ + + T + TP +N++ S+
Sbjct: 437 TSISLGVVRPVAPSG-------PRKGPDLRELLTAQQYTPRSATSVQPKGSNLTEPGSHR 489
Query: 176 ERNNKSSRKSGITSLKGVKGKSMSSVGRDKDKT-IVEPEVLMTKEIEWSK--YEL---RE 229
E +K +R SG S G+ + G D +EPE+LMTK+ + S+ +EL R
Sbjct: 490 ESISKRNRSSGFFSFLGLDKLAGKGPGNQHDAAEFIEPEILMTKDEDSSEASFELDKARL 549
Query: 230 RDIRQGIDLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQG 289
++IR+GIDLATTLERIEKNFVI+DPRLPD PIIFASD+FLELTEY+REEILGRNCRFLQG
Sbjct: 550 KEIRRGIDLATTLERIEKNFVITDPRLPDNPIIFASDNFLELTEYSREEILGRNCRFLQG 609
Query: 290 PETDQATVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLD 349
P+T++ TV IRDAI +++E+TVQL+NYTK+G+ FWNLFHLQPMRD KGELQYF GVQLD
Sbjct: 610 PDTNRETVKLIRDAIDNEKEVTVQLLNYTKTGRTFWNLFHLQPMRDHKGELQYFTGVQLD 669
Query: 350 GSDHLEPLRNRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAVSPRP 409
G+++LEPL RLS+ + AK+++ TA NV+ A+RELPDANL+ EDLW IHS+ V P+P
Sbjct: 670 GTEYLEPLTKRLSQQIASEGAKIIRETAANVNEALRELPDANLKVEDLWRIHSRLVLPKP 729
Query: 410 HKRDNPSWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKS 469
HK ++ SW I+KI A GEK+ L HF P+RPLG GDTGSVHLVEL+GTG+L+AMKAMEK+
Sbjct: 730 HKLNHDSWGVIRKIHASGEKVKLKHFRPLRPLGYGDTGSVHLVELRGTGKLFAMKAMEKN 789
Query: 470 VMLNRNKSVFYYRFIERALKERLYHYWIILFFQHYTPHF-------------------KQ 510
VM+ RNK + ER + + H ++ + + +Q
Sbjct: 790 VMVKRNK--VHRVCAEREILGMMDHPFLPTLYASFETQTHVCLITDFCAGGELFLLLERQ 847
Query: 511 PMKILKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSC 570
P KI +E++ARFY +EVV+ LEYLHC G+IYRDLKPEN+LLQ+DGH++L+DFDLS+++S
Sbjct: 848 PTKIFREETARFYTSEVVVALEYLHCQGVIYRDLKPENILLQQDGHVMLSDFDLSYLSSS 907
Query: 571 KPQ-VVKQSLPGNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWW 629
P+ VV L R + ++ PPPIF +EP+ NSFVGTEEYIAPE+ITG+ H S++DWW
Sbjct: 908 NPRLVVPPRLHKKRSKRKNFPPPIFRAEPIGACNSFVGTEEYIAPEVITGSGHNSSVDWW 967
Query: 630 TLGILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASR 689
LGIL+YEMLYGRTPFRGK RQKTF NILHKDL FP IP SLAARQLIN LLQ+DP +R
Sbjct: 968 ALGILMYEMLYGRTPFRGKTRQKTFGNILHKDLVFPRRIPTSLAARQLINGLLQKDPENR 1027
Query: 690 LGSATGSNEIKQHPFFRGINWPLIRNMSPPPLDVPLQFIGKDPTAKDKK---WED 741
LGS G+NEIK HPFF+G+NW LIR M PP D P+ F G + + + W+D
Sbjct: 1028 LGSQGGANEIKGHPFFQGVNWTLIRCMRPPTFDAPVCFAGSELDVEGDEALDWDD 1082
Score = 111 bits (278), Expect = 7e-23
Identities = 61/164 (37%), Positives = 92/164 (55%), Gaps = 4/164 (2%)
Query: 237 DLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQAT 296
D+ LE ++ FVI+D PD PI++AS F ++T YT E++GRNCRFLQG ETD
Sbjct: 269 DVLQALEGFQQTFVIADGTKPDLPIMYASAGFFKMTGYTSSEVIGRNCRFLQGKETDPEE 328
Query: 297 VNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLEP 356
++RIR+ I +L+NY K G FWNL + P++D G + +IG+Q++ S E
Sbjct: 329 IDRIRECISKGSGYCGRLLNYKKDGSAFWNLLTISPIKDVDGSVLKYIGMQVEVSQFTEG 388
Query: 357 LRNRLSEGSEIQSAKLVKATAENVDGA---VRELPDANLRPEDL 397
+ + + S L+K A + A V EL +A P+ +
Sbjct: 389 TKENAMRPNGL-SESLIKYDARQKERASFQVTELVEAIKDPKQV 431
>UniRef100_Q6BCU0 Phototropin [Physcomitrella patens]
Length = 1095
Score = 826 bits (2133), Expect = 0.0
Identities = 442/776 (56%), Positives = 545/776 (69%), Gaps = 48/776 (6%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKFI 60
RFLQGP+TD EV KIR A + GK +CGRLLNY+K+GT FWNLLT+TPIKD+ IKFI
Sbjct: 322 RFLQGPDTDPMEVEKIRQAVRTGKPFCGRLLNYRKDGTQFWNLLTITPIKDENDKVIKFI 381
Query: 61 GMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSITEVVQTVRNPKSIIRSKN 120
GMQVEVSKYTEG A RPNGLP+SLIRYDARQK++A S+TE+V + P +
Sbjct: 382 GMQVEVSKYTEGAKAVARRPNGLPESLIRYDARQKDKATESVTELVGAFKKPPPVT-PPT 440
Query: 121 DDTATIMHEEPENLNHDFVLPKSVEPVNDTTTPGRQTPLKFHGD--NNNMSRFSSYEERN 178
A P H PK V + + G P H + +S S ++ +
Sbjct: 441 QAIAADSFVSPLPGIHSISSPK-FSNVKEHSHRGSSEP---HPSLRHQKLSGVSEHDLMS 496
Query: 179 N-KSSRKSGITSLKGVKGKSMSSVGRDKDKTIVEPEVLMTK-EIEWSKYEL-RERDIRQG 235
+S R SG SL G+ GKS + D ++ +PEV+M E S+ + R IR+G
Sbjct: 497 TTRSKRTSGFLSLLGI-GKSQRLMHEDIPESEFDPEVVMLGYERPESQDDFDRTLGIRRG 555
Query: 236 IDLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQA 295
IDLATTLERI+KNFVI+DPRLPD PIIFASD FLELTEYTREE+LGRNCRFLQG +TDQ
Sbjct: 556 IDLATTLERIDKNFVITDPRLPDNPIIFASDEFLELTEYTREEVLGRNCRFLQGQDTDQN 615
Query: 296 TVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLE 355
TV +IRDAIK+ R+ITVQL+NYTKSGK FWNLFHLQ MRDQ+GELQYFIGVQLDGS ++E
Sbjct: 616 TVQQIRDAIKENRDITVQLLNYTKSGKPFWNLFHLQAMRDQRGELQYFIGVQLDGSQYVE 675
Query: 356 PLRNRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAVSPRPHKRDNP 415
P+R+RLS+ +E SAKLV+ TA N+D AVRELPDAN PEDLWA HS+ V P+PH
Sbjct: 676 PVRHRLSDNTEKASAKLVRETARNIDVAVRELPDANTSPEDLWANHSEFVKPKPHMGGTA 735
Query: 416 SWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRN 475
+W A+ K+ + G+K+GL HF PI+PLGCGDTGSVHLV L+GTG ++AMKAM+K+VML+RN
Sbjct: 736 AWKALIKVRSSGQKLGLKHFKPIKPLGCGDTGSVHLVSLRGTGHVFAMKAMDKTVMLDRN 795
Query: 476 KSVFYYRFIERALKERLY--------------------HYWIILFF----QHYTPHFKQP 511
K + RA ERL H +I F + + +QP
Sbjct: 796 K-------VHRACAERLILDLVDHPFLPTLYASFQTMTHVCLITDFCPGGELFLVLERQP 848
Query: 512 MKILKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCK 571
K +EDSARFYAAEVV+ LEYLHC+G++YRDLKPEN+L+ GH+ LTDFDLSF++S +
Sbjct: 849 KKHFQEDSARFYAAEVVLALEYLHCIGVVYRDLKPENILVTASGHVQLTDFDLSFVSSPR 908
Query: 572 PQVVKQSLPG--NRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWW 629
++V P +R++S++ P P+ +EPVT SNSFVGTEEYIAPEII+G H+SA+DWW
Sbjct: 909 VEMVTPPKPKKKSRKKSKNVPRPVIFAEPVTSSNSFVGTEEYIAPEIISGLGHSSAVDWW 968
Query: 630 TLGILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASR 689
LGILLYEML+GRTPFRGKNRQ TFSNIL K+L FPSSIP SL A+ LI LL +DP R
Sbjct: 969 ALGILLYEMLFGRTPFRGKNRQNTFSNILEKELYFPSSIPVSLEAKLLIRDLLNKDPLKR 1028
Query: 690 LGSATGSNEIKQHPFFRGINWPLIRNMSPPPLDVPLQFIGKDPTAKDK----KWED 741
LGS G+N+IK HPFF+ INWPLIR M+PPPL+VPL P +DK +W+D
Sbjct: 1029 LGSYRGANDIKNHPFFKDINWPLIRIMTPPPLEVPLNLTSSLPDFEDKDAELEWDD 1084
Score = 113 bits (283), Expect = 2e-23
Identities = 69/210 (32%), Positives = 111/210 (52%), Gaps = 13/210 (6%)
Query: 229 ERDIRQGIDLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQ 288
E +R DL L ++ FV+SD PD PI++AS F +T Y+ +E++G NCRFLQ
Sbjct: 266 EVPVRVSEDLLDALSSFKQTFVVSDATKPDYPIMYASAGFFSMTGYSPKEVIGYNCRFLQ 325
Query: 289 GPETDQATVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQL 348
GP+TD V +IR A++ + +L+NY K G +FWNL + P++D+ ++ FIG+Q+
Sbjct: 326 GPDTDPMEVEKIRQAVRTGKPFCGRLLNYRKDGTQFWNLLTITPIKDENDKVIKFIGMQV 385
Query: 349 DGSDHLEPLR------NRLSEGSEIQSAKLVKATAENVD---GAVRELPDANLRPEDLWA 399
+ S + E + N L E A+ E+V GA ++ P + + A
Sbjct: 386 EVSKYTEGAKAVARRPNGLPESLIRYDARQKDKATESVTELVGAFKKPPPVTPPTQAIAA 445
Query: 400 IHSQAVSPRP--HKRDNPSWVAIQKITARG 427
VSP P H +P + +++ + RG
Sbjct: 446 --DSFVSPLPGIHSISSPKFSNVKEHSHRG 473
>UniRef100_Q6BCT8 Phototropin [Physcomitrella patens]
Length = 1133
Score = 826 bits (2133), Expect = 0.0
Identities = 439/785 (55%), Positives = 554/785 (69%), Gaps = 57/785 (7%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKFI 60
RFLQG TD +VA+IR+A K GKS+CGRLLNYKK+G+ FWNLLT+TPIKDD G +KFI
Sbjct: 352 RFLQGAGTDNADVARIREALKEGKSFCGRLLNYKKDGSAFWNLLTITPIKDDDGKVLKFI 411
Query: 61 GMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSITEVVQTVRNPKSIIRSKN 120
GMQVEVSK+TEG EK+LRPNGLP+SLIRYDAR +++A ++ ++V +NP + S
Sbjct: 412 GMQVEVSKHTEGKKEKSLRPNGLPESLIRYDARLQDKATAAVGDLVGAFKNP--VAPSTP 469
Query: 121 DDTATIMHEEPENLNHDFVLPKSVEPVNDTTTPGRQTPLKFHGDNNNM-----SRFSSYE 175
DT + P +L D L S + + N+ SR S+
Sbjct: 470 PDTLNTGPKSPPSLPPDLQLVDSATESKLASKSSLHDSTRRRSTGTNIMTRSDSRLSNLP 529
Query: 176 ERNN-----KSSRKSGITSLKGVKGKSMSSVGRDKDKTIVEPEVLMTKEIEWSK-YEL-- 227
E ++ R SG S+ G+ S + D P++ M ++ + + +E+
Sbjct: 530 ENGQPAYPRRNRRSSGFLSIFGLGKPEPKSPDPEMD-----PQLRMLEDEDRPESFEVDL 584
Query: 228 -RERDIRQGIDLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRF 286
R ++IR+GIDLATTLERI KNFVI+DPRLPD PIIFASD FLELTEYTREEILGRNCRF
Sbjct: 585 ERSKEIRRGIDLATTLERIAKNFVITDPRLPDNPIIFASDEFLELTEYTREEILGRNCRF 644
Query: 287 LQGPETDQATVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGV 346
LQGP+TD A V++IRDAI +R+ITVQL+NYTKSGK FWNLFHLQ MRD GELQYFIGV
Sbjct: 645 LQGPDTDLAVVDQIRDAIAARRDITVQLLNYTKSGKPFWNLFHLQAMRDHDGELQYFIGV 704
Query: 347 QLDGSDHLEPLRNRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAVS 406
QLDGS++LEP R RLSE +E + AK+V+ TA N+D AVRELPDANL+PEDLW+ HS V
Sbjct: 705 QLDGSEYLEPERRRLSEKTEKEGAKVVQETANNIDDAVRELPDANLKPEDLWSKHSLPVH 764
Query: 407 PRPHKRDNPSWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAM 466
P+PH + + +W AI K+ G+ +GL F PI+PLG GDTGSVHLVEL+ TG ++AMKAM
Sbjct: 765 PKPHNKVSRAWDAIHKMKINGQGLGLKDFRPIKPLGSGDTGSVHLVELRETGLVFAMKAM 824
Query: 467 EKSVMLNRNKSVFYYRFIERALKER-------------LY-------HYWIILFF----Q 502
+KSVM+ RNK + RA ER LY H ++ F +
Sbjct: 825 DKSVMMQRNK-------VHRARAERDILALMDHPFLPTLYSTFQTQTHICLVTDFCPGGE 877
Query: 503 HYTPHFKQPMKILKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDF 562
+ +QP K+ ED RF+AAEVVI LEYLHCLG++YRDLKPEN+LL+ DGHI LTDF
Sbjct: 878 LFLLLERQPRKVFTEDVVRFFAAEVVIALEYLHCLGVVYRDLKPENVLLRADGHIQLTDF 937
Query: 563 DLSFITSCKPQVVKQSLPGNRRRSRSQPP-PIFVSEPVTQSNSFVGTEEYIAPEIITGAR 621
DLSF+TS KP++V+Q LP RRR +PP PIFV+EPVT SNSFVGTEEYIAPEIITG
Sbjct: 938 DLSFLTSAKPRLVEQDLPPGRRRKPKRPPSPIFVAEPVTPSNSFVGTEEYIAPEIITGQG 997
Query: 622 HTSAIDWWTLGILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINAL 681
H+SA+DWW LGIL+YEMLYGRTPFRGKNRQ+TF+N+L +D+ FP+SIP S++ARQL+ L
Sbjct: 998 HSSAVDWWALGILIYEMLYGRTPFRGKNRQRTFTNVLQRDIIFPASIPVSISARQLMRDL 1057
Query: 682 LQRDPASRLGSATGSNEIKQHPFFRGINWPLIRNMSPPPLDVPLQFIGKD--PTAKDK-- 737
LQR+P RLGS G++++K HPFFRGINWPL+R+ +PPPL+ P I D PT D+
Sbjct: 1058 LQRNPLKRLGSHRGASDVKNHPFFRGINWPLLRHTTPPPLETPALLITPDVEPTKADEDL 1117
Query: 738 KWEDD 742
+W+++
Sbjct: 1118 EWDEE 1122
Score = 117 bits (292), Expect = 2e-24
Identities = 53/119 (44%), Positives = 78/119 (65%)
Query: 237 DLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQAT 296
++ L ++ FV+SD PD PI++AS F +T YT +E++GRNCRFLQG TD A
Sbjct: 304 EVKEALSTFQQTFVVSDATQPDFPILYASAGFFNMTGYTPKEVIGRNCRFLQGAGTDNAD 363
Query: 297 VNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLE 355
V RIR+A+K+ + +L+NY K G FWNL + P++D G++ FIG+Q++ S H E
Sbjct: 364 VARIREALKEGKSFCGRLLNYKKDGSAFWNLLTITPIKDDDGKVLKFIGMQVEVSKHTE 422
>UniRef100_Q8RWA1 Phototropin 1 [Pisum sativum]
Length = 976
Score = 825 bits (2130), Expect = 0.0
Identities = 429/757 (56%), Positives = 538/757 (70%), Gaps = 40/757 (5%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKFI 60
RFLQG +TD ++VA+IR+A + GKS+CGRLLNYKK+GTPFWNLLT++PIKDD GN +K I
Sbjct: 203 RFLQGADTDPDDVARIREALEGGKSFCGRLLNYKKDGTPFWNLLTISPIKDDDGNVLKLI 262
Query: 61 GMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSITEVVQTVRNPKSIIRSKN 120
GM VEV+K+TEG EK LRPNGLP+SLIRYDARQKE+A S++E+++ ++ P+++ S
Sbjct: 263 GMLVEVNKHTEGSKEKKLRPNGLPESLIRYDARQKEKATSSVSELLEAMKRPRALSESGQ 322
Query: 121 DDTATIMHEEPENLNHDFVLPKSVEPVNDTTTPGRQTPLKFHGD-NNNMSRFSSYEERNN 179
+ + + +D+ R P G ++M R S E
Sbjct: 323 RPFIRKSGGGGGSEEDEEAVENKSRRKSDSVASFRPKP---QGKIRHSMERISELPENKQ 379
Query: 180 KSSRKSGITSLKGVKGKSMSSVGRDKDKTIVEPEVLMTKEIEWSKYELRE----RDIRQG 235
K+SR+ S G KS S ++ IV+ + +E + R+ R+G
Sbjct: 380 KNSRRG---SFMGFMRKSDSIDESIDNEVIVDVSSGSEDDERDDSFEFDDKEKLREKRKG 436
Query: 236 IDLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQA 295
+DLATTLERIEKNFVI+DPRLPD PIIFASDSFLELTEY+REEILG+NCRFLQGPETD A
Sbjct: 437 LDLATTLERIEKNFVITDPRLPDNPIIFASDSFLELTEYSREEILGKNCRFLQGPETDPA 496
Query: 296 TVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLE 355
TV +IR+AI +Q E+TVQLINYTKSGKKFWNLFHLQPMRD KGE+QYFIGVQLDGS H+E
Sbjct: 497 TVRKIREAIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRDHKGEVQYFIGVQLDGSQHVE 556
Query: 356 PLRNRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAVSPRPHKRDNP 415
PL N ++E + + LVK TAENV AV+ELPDAN +P+DLW HS+ V P+PH++D+
Sbjct: 557 PLHNCIAEDTAKEGELLVKETAENVGEAVKELPDANQKPDDLWMNHSKVVRPKPHRKDDD 616
Query: 416 SWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRN 475
+W AIQK+ GE++GL HF PI+PLG GDTGSVHLVEL+GTG+ +AMKAM+K VMLNRN
Sbjct: 617 AWRAIQKVLENGEQVGLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRN 676
Query: 476 KSVFYYRFIERALKERLYHYWIILFFQHY---------TPHF----------KQPMKILK 516
K + ER + + L H ++ + + T ++ +QP K+LK
Sbjct: 677 K--VHRACTEREILDMLDHPFLPALYASFQTKTHVCLITDYYSGGELFLLLDQQPTKVLK 734
Query: 517 EDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVV- 575
EDS RFYAAEVVI LEYLHCLGIIYRDLKPEN+L+Q +GH+ LTDFDLS +TSCKPQ++
Sbjct: 735 EDSVRFYAAEVVIALEYLHCLGIIYRDLKPENVLIQSNGHVSLTDFDLSCLTSCKPQLIL 794
Query: 576 -------KQSLPGNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDW 628
K+ N+ + ++Q P+F++EP+ SNSFVGTEEYIAPEIITG+ HTSA+DW
Sbjct: 795 PAIEEKKKRKKKKNKGQQKNQQVPMFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDW 854
Query: 629 WTLGILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPAS 688
W LGILLYEMLYG TPFRGK RQKTF+NILHKDL FP S P S +QLI LL RDP +
Sbjct: 855 WALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPHGKQLIYWLLHRDPKN 914
Query: 689 RLGSATGSNEIKQHPFFRGINWPLIRNMSPPPLDVPL 725
RLGS G+NEIK HPFF+ INW L+R PP LD P+
Sbjct: 915 RLGSLEGANEIKNHPFFKNINWALVRCTKPPELDGPI 951
Score = 113 bits (283), Expect = 2e-23
Identities = 60/164 (36%), Positives = 94/164 (56%), Gaps = 4/164 (2%)
Query: 237 DLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQAT 296
DL L ++ FV+SD PD PI++AS F ++T YT +E++GRNCRFLQG +TD
Sbjct: 155 DLKDALSAFQQTFVVSDATKPDYPILYASAGFFKMTGYTSKEVIGRNCRFLQGADTDPDD 214
Query: 297 VNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLEP 356
V RIR+A++ + +L+NY K G FWNL + P++D G + IG+ ++ + H E
Sbjct: 215 VARIREALEGGKSFCGRLLNYKKDGTPFWNLLTISPIKDDDGNVLKLIGMLVEVNKHTEG 274
Query: 357 LRNRLSEGSEIQSAKLVKATA---ENVDGAVRELPDANLRPEDL 397
+ + + + + L++ A E +V EL +A RP L
Sbjct: 275 SKEKKLRPNGLPES-LIRYDARQKEKATSSVSELLEAMKRPRAL 317
>UniRef100_P93489 Phototropin-like protein PsPK4 [Pisum sativum]
Length = 976
Score = 825 bits (2130), Expect = 0.0
Identities = 429/757 (56%), Positives = 538/757 (70%), Gaps = 40/757 (5%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKFI 60
RFLQG +TD ++VA+IR+A + GKS+CGRLLNYKK+GTPFWNLLT++PIKDD GN +K I
Sbjct: 203 RFLQGADTDPDDVARIREALEGGKSFCGRLLNYKKDGTPFWNLLTISPIKDDDGNVLKLI 262
Query: 61 GMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSITEVVQTVRNPKSIIRSKN 120
GM VEV+K+TEG EK LRPNGLP+SLIRYDARQKE+A S++E+++ ++ P+++ S
Sbjct: 263 GMLVEVNKHTEGSKEKKLRPNGLPESLIRYDARQKEKATSSVSELLEAMKRPRALSESGQ 322
Query: 121 DDTATIMHEEPENLNHDFVLPKSVEPVNDTTTPGRQTPLKFHGD-NNNMSRFSSYEERNN 179
+ + + +D+ R P G ++M R S E
Sbjct: 323 RPFIRKSGGGGGSEEDEEAVENKSRRKSDSVASFRPKP---QGKIRHSMERISELPENKQ 379
Query: 180 KSSRKSGITSLKGVKGKSMSSVGRDKDKTIVEPEVLMTKEIEWSKYELRE----RDIRQG 235
K+SR+ S G KS S ++ IV+ + +E + R+ R+G
Sbjct: 380 KNSRRG---SFMGFMRKSDSIDESIDNEVIVDVSSGSEDDERDDSFEFDDKEKLREKRKG 436
Query: 236 IDLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQA 295
+DLATTLERIEKNFVI+DPRLPD PIIFASDSFLELTEY+REEILG+NCRFLQGPETD A
Sbjct: 437 LDLATTLERIEKNFVITDPRLPDNPIIFASDSFLELTEYSREEILGKNCRFLQGPETDPA 496
Query: 296 TVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLE 355
TV +IR+AI +Q E+TVQLINYTKSGKKFWNLFHLQPMRD KGE+QYFIGVQLDGS H+E
Sbjct: 497 TVRKIREAIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRDHKGEVQYFIGVQLDGSQHVE 556
Query: 356 PLRNRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAVSPRPHKRDNP 415
PL N ++E + + LVK TAENV AV+ELPDAN +P+DLW HS+ V P+PH++D+
Sbjct: 557 PLHNCIAEDTAKEGELLVKETAENVGEAVKELPDANQKPDDLWMNHSKVVRPKPHRKDDD 616
Query: 416 SWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRN 475
+W AIQK+ GE++GL HF PI+PLG GDTGSVHLVEL+GTG+ +AMKAM+K VMLNRN
Sbjct: 617 AWRAIQKVLENGEQVGLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRN 676
Query: 476 KSVFYYRFIERALKERLYHYWIILFFQHY---------TPHF----------KQPMKILK 516
K + ER + + L H ++ + + T ++ +QP K+LK
Sbjct: 677 K--VHRACTEREILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLK 734
Query: 517 EDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVV- 575
EDS RFYAAEVVI LEYLHCLGIIYRDLKPEN+L+Q +GH+ LTDFDLS +TSCKPQ++
Sbjct: 735 EDSVRFYAAEVVIALEYLHCLGIIYRDLKPENVLIQSNGHVSLTDFDLSCLTSCKPQLIL 794
Query: 576 -------KQSLPGNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDW 628
K+ N+ + ++Q P+F++EP+ SNSFVGTEEYIAPEIITG+ HTSA+DW
Sbjct: 795 PAIEEKKKRKKKKNKGQQKNQQVPMFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDW 854
Query: 629 WTLGILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPAS 688
W LGILLYEMLYG TPFRGK RQKTF+NILHKDL FP S P S +QLI LL RDP +
Sbjct: 855 WALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPHGKQLIYWLLHRDPKN 914
Query: 689 RLGSATGSNEIKQHPFFRGINWPLIRNMSPPPLDVPL 725
RLGS G+NEIK HPFF+ INW L+R PP LD P+
Sbjct: 915 RLGSLEGANEIKNHPFFKNINWALVRCTKPPELDGPI 951
Score = 113 bits (283), Expect = 2e-23
Identities = 60/164 (36%), Positives = 94/164 (56%), Gaps = 4/164 (2%)
Query: 237 DLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQAT 296
DL L ++ FV+SD PD PI++AS F ++T YT +E++GRNCRFLQG +TD
Sbjct: 155 DLKDALSAFQQTFVVSDATKPDYPILYASAGFFKMTGYTSKEVIGRNCRFLQGADTDPDD 214
Query: 297 VNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLEP 356
V RIR+A++ + +L+NY K G FWNL + P++D G + IG+ ++ + H E
Sbjct: 215 VARIREALEGGKSFCGRLLNYKKDGTPFWNLLTISPIKDDDGNVLKLIGMLVEVNKHTEG 274
Query: 357 LRNRLSEGSEIQSAKLVKATA---ENVDGAVRELPDANLRPEDL 397
+ + + + + L++ A E +V EL +A RP L
Sbjct: 275 SKEKKLRPNGLPES-LIRYDARQKEKATSSVSELLEAMKRPRAL 317
>UniRef100_Q8H934 Phototropin [Vicia faba]
Length = 970
Score = 823 bits (2126), Expect = 0.0
Identities = 431/762 (56%), Positives = 538/762 (70%), Gaps = 51/762 (6%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKFI 60
RFLQG +TD N+VA+IR+A + GKS+CGRLLNYKK+GTPFWNLLT++PIKDD GN +K I
Sbjct: 198 RFLQGADTDPNDVARIREALEGGKSFCGRLLNYKKDGTPFWNLLTISPIKDDDGNVLKLI 257
Query: 61 GMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSITEVVQTVRNPKSIIRSKN 120
GM VEV+K+TEG EK LRPNGLP+SLIRYDARQKE+A S++E+++ ++ P+++ S +
Sbjct: 258 GMLVEVNKHTEGSKEKKLRPNGLPESLIRYDARQKEKATSSVSELLEAMKRPRAMSESGH 317
Query: 121 -----DDTATIMHEEPENLNHDFVLPKSVEPVNDTTTPGRQTPLKFHGD-NNNMSRFSSY 174
EE E L + +D+ R P G ++M R S
Sbjct: 318 RPFIRKSGGGGSSEEDERLEN------KSRRKSDSVASFRPKP---QGKIRHSMERISEL 368
Query: 175 EERNNKSSRKSGITSLKGVKGKSMSSVGRDKDKTIVEPEVLMTKEIEWSKYELRERDI-- 232
E K+SR+ S G KS S ++ IV+ + +E +++
Sbjct: 369 PENKQKNSRRG---SFMGFMRKSHSIDESIDNEVIVDVSSGSEDDERDDSFEFDDKEKLK 425
Query: 233 --RQGIDLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGP 290
R+G+DLATTLERIEKNFVI+DPRLPD PIIFASDSFLELTEY+REEILG+NCRFLQG
Sbjct: 426 EKRKGLDLATTLERIEKNFVITDPRLPDNPIIFASDSFLELTEYSREEILGKNCRFLQGQ 485
Query: 291 ETDQATVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDG 350
ETD ATV +IR+AI +Q E+TVQLINYTKSGKKFWNLFHLQPMRD KGE+QYFIGVQLDG
Sbjct: 486 ETDPATVRKIREAIDNQTEVTVQLINYTKSGKKFWNLFHLQPMRDHKGEVQYFIGVQLDG 545
Query: 351 SDHLEPLRNRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAVSPRPH 410
S H+EPL N ++E S + LVK TAENV AV+ELPDAN +P+DLW HS+ V P+PH
Sbjct: 546 SQHVEPLHNCIAEESAKEGELLVKETAENVGEAVKELPDANQKPDDLWKNHSKVVRPKPH 605
Query: 411 KRDNPSWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSV 470
++D+ +W AIQ + GE++GL HF PI+PLG GDTGSVHLVEL+GTG +AMKAM+K V
Sbjct: 606 RKDDDAWRAIQNVVGNGEQVGLKHFRPIKPLGSGDTGSVHLVELEGTGHYFAMKAMDKGV 665
Query: 471 MLNRNKSVFYYRFIERALKERLYHYWIILFFQHY---------TPHF----------KQP 511
MLNRNK + ER + + L H ++ + + T ++ +QP
Sbjct: 666 MLNRNK--VHRACTEREILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQP 723
Query: 512 MKILKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCK 571
K+LKEDS RFYAAEVVI LEYLHCLGIIYRDLKPEN+L+Q++GH+ LTDFDLS +TSCK
Sbjct: 724 TKVLKEDSVRFYAAEVVIALEYLHCLGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCK 783
Query: 572 PQVVKQSLPGNRRRS--------RSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHT 623
PQ++ + ++R ++Q P+F++EP+ SNSFVGTEEYIAPEIITG+ HT
Sbjct: 784 PQLILPATEEKKKRKNKKKKGQPKNQEVPMFMAEPMRASNSFVGTEEYIAPEIITGSGHT 843
Query: 624 SAIDWWTLGILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQ 683
SA+DWW LGILLYEMLYG TPFRGK RQKTF NILHKDL FP S P S +QLI LL
Sbjct: 844 SAVDWWALGILLYEMLYGYTPFRGKTRQKTFGNILHKDLKFPKSKPVSPHGKQLIYWLLH 903
Query: 684 RDPASRLGSATGSNEIKQHPFFRGINWPLIRNMSPPPLDVPL 725
RDP +RLGS G+NEIK HPFF+ +NW L+R M PP LD P+
Sbjct: 904 RDPKNRLGSLEGANEIKNHPFFKNVNWALVRCMKPPELDAPI 945
Score = 112 bits (281), Expect = 3e-23
Identities = 59/161 (36%), Positives = 93/161 (57%), Gaps = 4/161 (2%)
Query: 237 DLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQAT 296
DL L ++ FV+SD PD PI++AS F ++T YT +E++GRNCRFLQG +TD
Sbjct: 150 DLKDALSAFQQTFVVSDATKPDYPILYASAGFFKMTGYTSKEVIGRNCRFLQGADTDPND 209
Query: 297 VNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLEP 356
V RIR+A++ + +L+NY K G FWNL + P++D G + IG+ ++ + H E
Sbjct: 210 VARIREALEGGKSFCGRLLNYKKDGTPFWNLLTISPIKDDDGNVLKLIGMLVEVNKHTEG 269
Query: 357 LRNRLSEGSEIQSAKLVKATA---ENVDGAVRELPDANLRP 394
+ + + + + L++ A E +V EL +A RP
Sbjct: 270 SKEKKLRPNGLPES-LIRYDARQKEKATSSVSELLEAMKRP 309
>UniRef100_Q6BCU1 Phototropin [Physcomitrella patens]
Length = 1070
Score = 811 bits (2094), Expect = 0.0
Identities = 427/784 (54%), Positives = 541/784 (68%), Gaps = 55/784 (7%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKFI 60
RFLQGP+TD +V KIR A KNGK++CGRLLNY+K+G+ FWNLLT+TPIKD+ +KFI
Sbjct: 288 RFLQGPDTDPADVEKIRHAVKNGKNFCGRLLNYRKDGSTFWNLLTITPIKDENDKVVKFI 347
Query: 61 GMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSITEVVQTVRNPKS----II 116
GMQVEVSKYTEG A RPNGLP+SLIRYDARQK++A S+TE+V + P+ +
Sbjct: 348 GMQVEVSKYTEGAKAVARRPNGLPESLIRYDARQKDKATESVTELVGAFKKPQPPPTPAL 407
Query: 117 RSKNDDTATIMHEEPENLNHDFVLPKSVEPVNDTTTPGRQTPLKFHGDNNNMSRFSSYEE 176
+S D +++ P + V+ +T P ++ + S E
Sbjct: 408 QSIPAD-GSLVSPLPSQTISTAIHSNKVQGHRRSTEPYT---------SSRAQKLSGVSE 457
Query: 177 RNN--KSSRKSGITSLKGVKGKSMSSVGRDKDKTIVEPEVLMTKEIEWSKYELRERDIRQ 234
+ +S R SG SL G+ KS + ++ +E ++L+ E R IR+
Sbjct: 458 LGDTGRSKRTSGFLSLLGIGQKSERIMEEGNLESDLEADLLVLDRPESRDDFDRTLGIRR 517
Query: 235 GIDLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQ 294
GIDLATTLERIEKNFVI+DPRLPD PIIFASD FLELTEYTREEILGRNCRFLQGP+TDQ
Sbjct: 518 GIDLATTLERIEKNFVITDPRLPDNPIIFASDEFLELTEYTREEILGRNCRFLQGPDTDQ 577
Query: 295 ATVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHL 354
TV +IRDAIK+ R+ITVQL+NYTKSGK FWNLFHLQ MRD KGELQYFIGVQ+DGS+++
Sbjct: 578 NTVQKIRDAIKENRDITVQLLNYTKSGKPFWNLFHLQAMRDHKGELQYFIGVQMDGSEYV 637
Query: 355 EPLRNRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAVSPRPHKRDN 414
EP R+RLS+ +E SA LV+ TA N+D AVRELPDAN PEDLWA HS++V P+PH
Sbjct: 638 EPTRHRLSDKTEKASAMLVQETARNIDTAVRELPDANTTPEDLWANHSKSVMPKPHMGGT 697
Query: 415 PSWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNR 474
P W +I K+ G+K+GL +F PI+PLGCGDTGSVHLVEL+GT ++AMKAM+K+VM++R
Sbjct: 698 PEWQSILKVRTAGKKLGLKNFKPIKPLGCGDTGSVHLVELRGTDHVFAMKAMDKTVMMDR 757
Query: 475 NKSVFYYRFIERALKERLYHYWIILFFQHY--TPHF-----------------KQPMKIL 515
NK + +ER + + + H ++ + + H +QP K
Sbjct: 758 NK--VHRACVERQILDLMDHPFLPTLYASFQTATHVCLITDFCPGGELFLVLERQPKKHF 815
Query: 516 KEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVV 575
+EDSARFYAAEVV+ LEYLHC G+IYRDLKPEN+L+ + GHI LTDFDLSFIT+ + Q++
Sbjct: 816 REDSARFYAAEVVLALEYLHCKGVIYRDLKPENILVTESGHIQLTDFDLSFITTPRVQLI 875
Query: 576 KQSLP--------------GNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGAR 621
++P + +++ P PIF + PVT SNSF+GTEEYIAPEII+G
Sbjct: 876 PPAIPKTSTWDRARGAKKKAQQPQTKDIPRPIFFAAPVTPSNSFIGTEEYIAPEIISGQG 935
Query: 622 HTSAIDWWTLGILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINAL 681
H+SA+DWW LGIL+YEML+GRTPFRGKNRQ TF+N+L K+L FP+ IP SL A+ LI L
Sbjct: 936 HSSAVDWWGLGILIYEMLFGRTPFRGKNRQTTFANVLEKELCFPAHIPVSLEAKTLIRDL 995
Query: 682 LQRDPASRLGSATGSNEIKQHPFFRGINWPLIRNMSPPPLDVPLQFIGKDPTAKDK---- 737
L RDP RLGS G+N+IK HPFFRGI WPLIRNM+PP L+VPL P DK
Sbjct: 996 LIRDPLKRLGSYRGANDIKNHPFFRGIKWPLIRNMTPPSLEVPLLITSSVPELDDKGAEL 1055
Query: 738 KWED 741
+W+D
Sbjct: 1056 EWDD 1059
Score = 116 bits (291), Expect = 2e-24
Identities = 51/119 (42%), Positives = 79/119 (65%)
Query: 237 DLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQAT 296
+L L ++ FV+SD PDCPI++AS F ++ Y+ +EI+G NCRFLQGP+TD A
Sbjct: 240 ELLDALSSFKQTFVVSDATKPDCPIVYASAGFFTMSGYSAKEIIGHNCRFLQGPDTDPAD 299
Query: 297 VNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLE 355
V +IR A+K+ + +L+NY K G FWNL + P++D+ ++ FIG+Q++ S + E
Sbjct: 300 VEKIRHAVKNGKNFCGRLLNYRKDGSTFWNLLTITPIKDENDKVVKFIGMQVEVSKYTE 358
>UniRef100_Q9ZWQ6 PHY3 [Adiantum capillus-veneris]
Length = 1465
Score = 702 bits (1813), Expect = 0.0
Identities = 382/751 (50%), Positives = 496/751 (65%), Gaps = 78/751 (10%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGK-SYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKF 59
RFLQGP+T+ +VA IR+A G ++CGRLLNY+K+G+ FWNLLT+ PIKDD G+ +K
Sbjct: 713 RFLQGPDTNPADVASIREALAQGTGTFCGRLLNYRKDGSSFWNLLTIAPIKDDLGSIVKL 772
Query: 60 IGMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSITEVVQTVRNPKSIIRSK 119
IG+Q+EVSKYTEG+ RPNG+P+SLIRYD R +++ I ++V + P + +
Sbjct: 773 IGVQLEVSKYTEGIRANNRRPNGMPQSLIRYDVRHQDKVSAFIAQLVAALTKPDKVETPR 832
Query: 120 NDDTATIMHEEPENLNHDFVLPKSVEPVNDTTTPGRQTPLKFHGDNNNMSRFSSYEERNN 179
+ +++E + T R+ G R SS+
Sbjct: 833 LSSAMRFS-----------LTGQTIESLPQPTAIPREG-----GGRTRRPRSSSF----- 871
Query: 180 KSSRKSGITSLKGVKGKSMSSVGRDKDKTIVEPEVLMTKEIEWSKYELRERD-------- 231
+S +G +K+K I E + L E+ + R
Sbjct: 872 ------------------LSLLGMEKEKDIPEEDELQELEVIMLEDASVGRPGSLDDPER 913
Query: 232 IRQGIDLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPE 291
R+GIDLATTLERI K+FVI+DPRLPD PIIFASD FLELTEYTREE+LG NCRFLQG
Sbjct: 914 TRRGIDLATTLERIGKSFVITDPRLPDNPIIFASDRFLELTEYTREEVLGNNCRFLQGRG 973
Query: 292 TDQATVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLD-- 349
TD+ V IRDA+K+QR++TVQ++NYTK G+ FWNLFHLQ MRD+ G++QYFIGVQ +
Sbjct: 974 TDRKAVQLIRDAVKEQRDVTVQVLNYTKGGRAFWNLFHLQVMRDENGDVQYFIGVQQEMV 1033
Query: 350 --GSDHLEP-LRNRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAVS 406
H P L + L + E + A++V+ATA+ VD A RELPDANL P+ L+A HS+ V+
Sbjct: 1034 APRPVHQPPELPDILPDRVEQEKAEVVRATAQRVDAAARELPDANLVPDHLFAPHSKVVT 1093
Query: 407 PRPHKRDNPS-WVAIQKITAR---GEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYA 462
P PH + N S W AI+++ R GE++GL HF PI+PLG GDTGSVHLVEL+GTG+++A
Sbjct: 1094 PLPHSKTNSSSWFAIRRVQRRLRRGERLGLKHFRPIKPLGSGDTGSVHLVELRGTGQVFA 1153
Query: 463 MKAMEKSVMLNRNKSVFYYRFIERALKERLYHYWI------------ILFFQHYTP---- 506
+KAM+KS+ML RNK + ER + + H ++ + Y P
Sbjct: 1154 LKAMDKSMMLQRNK--VHRARAEREILAIMDHPFLPTLYASFQTKTHVCLITDYCPGGDL 1211
Query: 507 ---HFKQPMKILKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFD 563
KQP + L E +A FYAAEVV+ LEYLHC+G+IYRDLKPEN+LLQK+GHI+LTDFD
Sbjct: 1212 FLLQDKQPTQTLSERTASFYAAEVVVALEYLHCMGVIYRDLKPENVLLQKNGHILLTDFD 1271
Query: 564 LSFITSCKPQVVKQSLPGNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHT 623
LSF+TSC+PQ++ Q G RRS+ + F +EP SNSFVGTEEYIAPEII+G H+
Sbjct: 1272 LSFLTSCRPQLILQGGKGRSRRSKRRRRVTFCAEPRVSSNSFVGTEEYIAPEIISGEPHS 1331
Query: 624 SAIDWWTLGILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQ 683
SA+DWW LGILLYEMLYGRTPF G+NRQKTF N+L+K+L FP+SIP SLA RQLI LLQ
Sbjct: 1332 SAVDWWALGILLYEMLYGRTPFVGRNRQKTFYNVLNKELIFPTSIPVSLAGRQLIAGLLQ 1391
Query: 684 RDPASRLGSATGSNEIKQHPFFRGINWPLIR 714
RDP RLG+ G++E+K+HPFFR INWPLIR
Sbjct: 1392 RDPTIRLGTLRGASELKKHPFFREINWPLIR 1422
Score = 103 bits (256), Expect = 2e-20
Identities = 57/151 (37%), Positives = 83/151 (54%), Gaps = 3/151 (1%)
Query: 248 NFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQATVNRIRDAI-KD 306
+F++ D PD PII+AS F LT YT E++G NCRFLQGP+T+ A V IR+A+ +
Sbjct: 676 SFIVVDALKPDFPIIYASTGFFNLTGYTSREVIGGNCRFLQGPDTNPADVASIREALAQG 735
Query: 307 QREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLEPLR--NRLSEG 364
+L+NY K G FWNL + P++D G + IGVQL+ S + E +R NR G
Sbjct: 736 TGTFCGRLLNYRKDGSSFWNLLTIAPIKDDLGSIVKLIGVQLEVSKYTEGIRANNRRPNG 795
Query: 365 SEIQSAKLVKATAENVDGAVRELPDANLRPE 395
+ + V + +L A +P+
Sbjct: 796 MPQSLIRYDVRHQDKVSAFIAQLVAALTKPD 826
>UniRef100_Q40269 Protein kinase [Mesembryanthemum crystallinum]
Length = 572
Score = 656 bits (1693), Expect = 0.0
Identities = 330/520 (63%), Positives = 400/520 (76%), Gaps = 23/520 (4%)
Query: 226 ELRERDIRQGIDLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCR 285
++R++++R+GIDLATTLERIEKNFVI+DPRLPD PIIFASDSFLELTE++R EIL RN R
Sbjct: 30 KVRKKEMRKGIDLATTLERIEKNFVITDPRLPDNPIIFASDSFLELTEFSRAEILARNRR 89
Query: 286 FLQGPETDQATVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIG 345
FLQGPETD ATV +IRDAI ++ ++TVQLINYTK+GKKFWN+FHLQPMRDQKGE+QYFIG
Sbjct: 90 FLQGPETDPATVAKIRDAIDNETDVTVQLINYTKTGKKFWNVFHLQPMRDQKGEVQYFIG 149
Query: 346 VQLDGSDHLEPLRNRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAV 405
VQLDGS+H+EP++N + S + S K VK TA NVD AVRELPDAN +PEDLWA HS+ V
Sbjct: 150 VQLDGSEHVEPVQNSIPVASVMDSEKQVKETATNVDVAVRELPDANKKPEDLWANHSKVV 209
Query: 406 SPRPHKRDNPSWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKA 465
P+PH+++ SW AI+KI GE+IGL HF P++PLG GDTGSVHLVEL GTGE +AMKA
Sbjct: 210 QPKPHRKECSSWKAIEKIKESGEQIGLKHFRPVKPLGAGDTGSVHLVELCGTGEYFAMKA 269
Query: 466 MEKSVMLNRNKSVFYYRFIERALKERLYHYWI------------ILFFQHYTPH------ 507
M+K+VMLNRNK + ER + + L H ++ I Y P
Sbjct: 270 MDKNVMLNRNK--VHRACAEREILDMLDHPFLPALYASFQTNTHICLITEYCPGGELFLL 327
Query: 508 -FKQPMKILKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSF 566
+QP K+LKED+ RFYAAEV+I LEYLHC GIIYRDLKPEN+LLQ +GH+ LTDFDLS
Sbjct: 328 LDRQPTKVLKEDAVRFYAAEVIIALEYLHCQGIIYRDLKPENILLQSNGHVSLTDFDLSC 387
Query: 567 ITSCKPQVVKQSLPGNRRRSRS--QPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTS 624
+TSCKPQ++ + +++ ++ Q PIF++EP+ SNSFVGTEEYIAPEII GA
Sbjct: 388 LTSCKPQLLIPEIRDKKKQQKAQHQQTPIFMAEPMRASNSFVGTEEYIAPEIIAGAGIQG 447
Query: 625 AIDWWTLGILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQR 684
A+DWW LGILLYEMLYG TPFRGK RQKTFSN+L KDL FP++ SL A QLI LLQ+
Sbjct: 448 AVDWWALGILLYEMLYGFTPFRGKTRQKTFSNVLRKDLKFPATKQVSLDASQLIYQLLQK 507
Query: 685 DPASRLGSATGSNEIKQHPFFRGINWPLIRNMSPPPLDVP 724
DP RLG+ G+NEIK+HPFFRG NW L+R M PP LD P
Sbjct: 508 DPKDRLGACEGANEIKRHPFFRGANWALVRCMKPPVLDAP 547
Score = 75.9 bits (185), Expect = 4e-12
Identities = 40/107 (37%), Positives = 60/107 (55%), Gaps = 6/107 (5%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKFI 60
RFLQGPETD VAKIRDA N +L+NY K G FWN+ + P++D +G FI
Sbjct: 89 RFLQGPETDPATVAKIRDAIDNETDVTVQLINYTKTGKKFWNVFHLQPMRDQKGEVQYFI 148
Query: 61 GMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSITEVVQ 107
G+Q++ S++ E V N +P + + +Q +E ++ V+
Sbjct: 149 GVQLDGSEHVEPVQ------NSIPVASVMDSEKQVKETATNVDVAVR 189
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.136 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,315,879,183
Number of Sequences: 2790947
Number of extensions: 58048961
Number of successful extensions: 179595
Number of sequences better than 10.0: 14741
Number of HSP's better than 10.0 without gapping: 10394
Number of HSP's successfully gapped in prelim test: 4379
Number of HSP's that attempted gapping in prelim test: 148017
Number of HSP's gapped (non-prelim): 23669
length of query: 753
length of database: 848,049,833
effective HSP length: 135
effective length of query: 618
effective length of database: 471,271,988
effective search space: 291246088584
effective search space used: 291246088584
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 79 (35.0 bits)
Medicago: description of AC145470.6


