

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145470.4 + phase: 0
(457 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
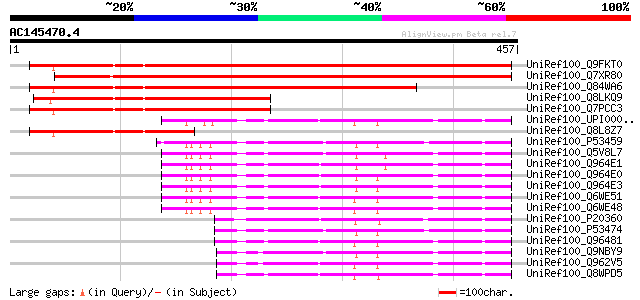
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FKT0 Similarity to actin [Arabidopsis thaliana] 572 e-161
UniRef100_Q7XR80 OSJNBa0043A12.5 protein [Oryza sativa] 531 e-149
UniRef100_Q84WA6 Putative F-box protein [Arabidopsis thaliana] 444 e-123
UniRef100_Q8LKQ9 Actin-related protein 8A [Arabidopsis thaliana] 242 2e-62
UniRef100_Q7PCC3 Actin-related protein 8A [Arabidopsis thaliana] 241 3e-62
UniRef100_UPI000032EAA0 UPI000032EAA0 UniRef100 entry 136 1e-30
UniRef100_Q8L8Z7 Hypothetical protein [Arabidopsis thaliana] 136 1e-30
UniRef100_P53459 Actin 6 [Diphyllobothrium dendriticum] 136 1e-30
UniRef100_Q5V8L7 Actin [Paxillus involutus] 134 4e-30
UniRef100_Q964E1 Actin, cytoplasmic [Biomphalaria obstructa] 134 5e-30
UniRef100_Q964E0 Actin, cytoplasmic [Biomphalaria tenagophila] 133 9e-30
UniRef100_Q964E3 Actin, cytoplasmic [Biomphalaria alexandrina] 133 9e-30
UniRef100_Q6WE51 Actin [Glaeseria mira] 133 1e-29
UniRef100_Q6WE48 Actin [Hartmannella cantabrigiensis] 133 1e-29
UniRef100_P20360 Actin, cytoplasmic [Euplotes crassus] 131 3e-29
UniRef100_P53474 Actin, cytoskeletal IIIA [Strongylocentrotus pu... 131 5e-29
UniRef100_Q96481 Actin 105 [Lycopersicon esculentum] 131 5e-29
UniRef100_Q9NBY9 Actin [Eledonella pygmaea] 130 1e-28
UniRef100_Q962V5 Alpha actin [Homarus americanus] 130 1e-28
UniRef100_Q8WPD5 Beta-actin [Gecarcinus lateralis] 129 1e-28
>UniRef100_Q9FKT0 Similarity to actin [Arabidopsis thaliana]
Length = 471
Score = 572 bits (1473), Expect = e-161
Identities = 290/441 (65%), Positives = 346/441 (77%), Gaps = 9/441 (2%)
Query: 19 NTTNSSSSSRSHQHHHHHAP------SPLGLFDSLPPDILLKITRLLGPKHAAKLCLVCK 72
N +NS +HQ P S LG FD LP DIL++I ++ PK A KL L CK
Sbjct: 12 NRSNSGKDLVNHQRAIDVPPLLLSSSSSLGAFDQLPMDILVQILMMMEPKDAVKLGLTCK 71
Query: 73 SWRSLVSDNELWAHFLQTHQPIHFHSILFSETNLTSGYPLPIFDTQTTPHVSFKHVFGHR 132
+W+ + S N LW +LQ Q + SI F+ET+L SGYPL + +Q+ +SF H++ R
Sbjct: 72 AWKCVASGNRLWIFYLQCSQE-PWDSIFFAETSLRSGYPLRMISSQSG-ELSFMHIYSQR 129
Query: 133 EQLPPAIIIDGGSGYCKFGWSKEDRPLGRVATFLEFGNVETPIYTRLRHFFATVYGRMKV 192
Q+P +IIIDGGSGYCKFGWSK P GR ATFLEFGN+E+PIY RL+ FFAT++ RM+V
Sbjct: 130 AQVPGSIIIDGGSGYCKFGWSKYASPSGRSATFLEFGNIESPIYARLQQFFATIFTRMQV 189
Query: 193 KPSSQVVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVPAVCALNQATLALYAANQ 252
KPS Q +VVSLPLCH+DDTESAKASR+QLK AI LFDMNVPAVCA+NQA LALYAA +
Sbjct: 190 KPSMQPIVVSLPLCHFDDTESAKASRRQLKTAIFNVLFDMNVPAVCAVNQAVLALYAARR 249
Query: 253 TSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNNLYFESLYTV 312
TSGI VNIGFQV +++PIL+GKVMR+VGVEV+G GALKLTGFL+EKMQ+NN+ F+SLYTV
Sbjct: 250 TSGIVVNIGFQVITILPILHGKVMRQVGVEVIGFGALKLTGFLKEKMQENNISFQSLYTV 309
Query: 313 RTLKEKLCYVAYDYEAELSKDTHASFE-AAEGKFTLSKERFQTGEILFQPRLAGVRAMSL 371
RTLKEKLCYVA DY+AELSKDT AS E + EG FTLSKERFQTGEILFQPRLAG+RAMSL
Sbjct: 310 RTLKEKLCYVALDYKAELSKDTQASVEVSGEGWFTLSKERFQTGEILFQPRLAGMRAMSL 369
Query: 372 HHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLEKELHSLLPPYMSNGIRV 431
H A++LCMDHC +A L G WFKTVVL+GGSACLPGL+ERLE+EL LP +SNGIRV
Sbjct: 370 HQAVSLCMDHCDAAGLTGDDSWFKTVVLTGGSACLPGLSERLERELQDHLPSSISNGIRV 429
Query: 432 IPPPHGADTAWFGAKLIGSVS 452
IPPP+G DT+W GAKLI ++S
Sbjct: 430 IPPPYGVDTSWHGAKLISNLS 450
>UniRef100_Q7XR80 OSJNBa0043A12.5 protein [Oryza sativa]
Length = 484
Score = 531 bits (1368), Expect = e-149
Identities = 261/414 (63%), Positives = 326/414 (78%), Gaps = 3/414 (0%)
Query: 41 LGLFDSLPPDILLKITRLLGPKHAAKLCLVCKSWRSLVSDNELWAHFLQTHQPIHFHSIL 100
LG D LP D+L +I RLLGP AA+ VC++WR L SDN LWA FL+ P + ++
Sbjct: 44 LGALDVLPIDVLAQILRLLGPADAARSTAVCRAWRLLASDNGLWAFFLRLG-PDPWELVV 102
Query: 101 FSETNLTSGYPL-PIFDTQTTPHVSFKHVFGHREQLPPAIIIDGGSGYCKFGWSKEDRPL 159
F+ET+L +G L P ++P +SFKHV+ R +P +II+DGGSGYCK+GWSK P
Sbjct: 103 FAETHLGAGPALHPGLYYDSSPQLSFKHVYTRRAVVPGSIIVDGGSGYCKYGWSKYAAPS 162
Query: 160 GRVATFLEFGNVETPIYTRLRHFFATVYGRMKVKPSSQVVVVSLPLCHYDDTESAKASRQ 219
GR ATFLEFGN+E+P+Y RLRHF +T+Y RM+VKPS+Q ++V LPLCH DDTESA+ASR+
Sbjct: 163 GRCATFLEFGNIESPMYARLRHFLSTIYTRMQVKPSTQPIIVVLPLCHSDDTESARASRK 222
Query: 220 QLKEAICAALFDMNVPAVCALNQATLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKV 279
Q ++ + + LFDMNVPAVC+++QA LALYAA +TSGI VNIGF TS+VPI G+VM ++
Sbjct: 223 QYRDTLYSVLFDMNVPAVCSVDQAVLALYAAKRTSGIVVNIGFNATSIVPIFQGRVMHEI 282
Query: 280 GVEVVGLGALKLTGFLREKMQQNNLYFESLYTVRTLKEKLCYVAYDYEAELSKDTHASFE 339
GVE VG GALKLTGFL+E MQQ N+ FESLYTVRT+KEKLCYVA DYEAE KDT AS E
Sbjct: 283 GVETVGQGALKLTGFLKELMQQRNITFESLYTVRTIKEKLCYVAADYEAEKRKDTQASCE 342
Query: 340 A-AEGKFTLSKERFQTGEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVV 398
EG FTLS+ERF+T EILFQP++ GVRAM LH A++LCMDHC+++E+ G +W+KTVV
Sbjct: 343 VDGEGWFTLSEERFKTAEILFQPQIGGVRAMGLHKAVSLCMDHCYNSEVFGDDNWYKTVV 402
Query: 399 LSGGSACLPGLAERLEKELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
LSGGS+CLPGL+ERLEKEL LLP ++S GIRVIPPP G D+AWFGAK+I +VS
Sbjct: 403 LSGGSSCLPGLSERLEKELRELLPAHISEGIRVIPPPFGTDSAWFGAKMISNVS 456
>UniRef100_Q84WA6 Putative F-box protein [Arabidopsis thaliana]
Length = 387
Score = 444 bits (1141), Expect = e-123
Identities = 229/355 (64%), Positives = 275/355 (76%), Gaps = 9/355 (2%)
Query: 19 NTTNSSSSSRSHQHHHHHAP------SPLGLFDSLPPDILLKITRLLGPKHAAKLCLVCK 72
N +NS +HQ P S LG FD LP DIL++I ++ PK A KL L CK
Sbjct: 12 NRSNSGKDLVNHQRAIDVPPLLLSSSSSLGAFDQLPMDILVQILMMMEPKDAVKLGLTCK 71
Query: 73 SWRSLVSDNELWAHFLQTHQPIHFHSILFSETNLTSGYPLPIFDTQTTPHVSFKHVFGHR 132
+W+ + S N LW +LQ Q + SI F+ET+L SGYPL + +Q+ +SF H++ R
Sbjct: 72 AWKCVASGNRLWIFYLQCSQE-PWDSIFFAETSLRSGYPLRMISSQSG-ELSFMHIYSQR 129
Query: 133 EQLPPAIIIDGGSGYCKFGWSKEDRPLGRVATFLEFGNVETPIYTRLRHFFATVYGRMKV 192
Q+P +IIIDGGSGYCKFGWSK P GR ATFLEFGN+E+PIY RL+ FFAT++ RM+V
Sbjct: 130 AQVPGSIIIDGGSGYCKFGWSKYASPSGRSATFLEFGNIESPIYARLQQFFATIFTRMQV 189
Query: 193 KPSSQVVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVPAVCALNQATLALYAANQ 252
KPS Q +VVSLPLCH+DDTESAKASR+QLK AI LFDMNVPAVCA+NQA LALYAA +
Sbjct: 190 KPSMQPIVVSLPLCHFDDTESAKASRRQLKTAIFNVLFDMNVPAVCAVNQAVLALYAARR 249
Query: 253 TSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNNLYFESLYTV 312
TSGI VNIGF+V +++PIL+GKVMR+VGVEV+G GALKLTGFL+EKMQ+NN+ F+SLYTV
Sbjct: 250 TSGIVVNIGFKVITILPILHGKVMRQVGVEVIGFGALKLTGFLKEKMQENNISFQSLYTV 309
Query: 313 RTLKEKLCYVAYDYEAELSKDTHASFE-AAEGKFTLSKERFQTGEILFQPRLAGV 366
RTLKEKLCYVA DY+AELSKDT AS E + EG FTLSKERFQTGEILFQPRLAG+
Sbjct: 310 RTLKEKLCYVALDYKAELSKDTQASVEVSGEGWFTLSKERFQTGEILFQPRLAGI 364
>UniRef100_Q8LKQ9 Actin-related protein 8A [Arabidopsis thaliana]
Length = 241
Score = 242 bits (617), Expect = 2e-62
Identities = 124/223 (55%), Positives = 156/223 (69%), Gaps = 11/223 (4%)
Query: 22 NSSSSSRSHQHHHH---------HAPSPLGLFDSLPPDILLKITRLLGPKHAAKLCLVCK 72
N S+S + +HH + S LG FD LP DIL++I ++ PK A KL L CK
Sbjct: 12 NRSNSGKDLVNHHRAIDVPPLLLSSSSSLGAFDQLPMDILVQILMMMEPKDAVKLGLTCK 71
Query: 73 SWRSLVSDNELWAHFLQTHQPIHFHSILFSETNLTSGYPLPIFDTQTTPHVSFKHVFGHR 132
+W+ + S N LW +LQ Q + SI F+ET+L SGYPL + +Q+ +SF H++ R
Sbjct: 72 AWKCVASGNRLWIFYLQCSQE-PWDSIFFAETSLRSGYPLRMISSQSG-ELSFMHIYSQR 129
Query: 133 EQLPPAIIIDGGSGYCKFGWSKEDRPLGRVATFLEFGNVETPIYTRLRHFFATVYGRMKV 192
Q+P +IIIDGGSGYCKFGWSK P GR ATFLEFGN+E+PIY RL+ FFAT++ RM+V
Sbjct: 130 AQVPGSIIIDGGSGYCKFGWSKYASPSGRSATFLEFGNIESPIYARLQQFFATIFTRMQV 189
Query: 193 KPSSQVVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVP 235
KPS Q +VVSLPLCH+DDTESAKASR+QLK AI LFDMNVP
Sbjct: 190 KPSMQPIVVSLPLCHFDDTESAKASRRQLKTAIFNVLFDMNVP 232
>UniRef100_Q7PCC3 Actin-related protein 8A [Arabidopsis thaliana]
Length = 241
Score = 241 bits (615), Expect = 3e-62
Identities = 125/223 (56%), Positives = 155/223 (69%), Gaps = 8/223 (3%)
Query: 19 NTTNSSSSSRSHQHHHHHAP------SPLGLFDSLPPDILLKITRLLGPKHAAKLCLVCK 72
N +NS +HQ P S LG FD LP DIL++I ++ PK A KL L CK
Sbjct: 12 NRSNSGKDLVNHQRAIDVPPLLLSSSSSLGAFDQLPMDILVQILMMMEPKDAVKLGLTCK 71
Query: 73 SWRSLVSDNELWAHFLQTHQPIHFHSILFSETNLTSGYPLPIFDTQTTPHVSFKHVFGHR 132
+W+ + S N LW +LQ Q + SI F+ET+L SGYPL + +Q+ +SF H++ R
Sbjct: 72 AWKCVASGNRLWIFYLQCSQE-PWDSIFFAETSLRSGYPLRMISSQSG-ELSFMHIYSQR 129
Query: 133 EQLPPAIIIDGGSGYCKFGWSKEDRPLGRVATFLEFGNVETPIYTRLRHFFATVYGRMKV 192
Q+P +IIIDGGSGYCKFGWSK P GR ATFLEFGN+E+PIY RL+ FFAT++ RM+V
Sbjct: 130 AQVPGSIIIDGGSGYCKFGWSKYASPSGRSATFLEFGNIESPIYARLQQFFATIFTRMQV 189
Query: 193 KPSSQVVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVP 235
KPS Q +VVSLPLCH+DDTESAKASR+QLK AI LFDMNVP
Sbjct: 190 KPSMQPIVVSLPLCHFDDTESAKASRRQLKTAIFNVLFDMNVP 232
>UniRef100_UPI000032EAA0 UPI000032EAA0 UniRef100 entry
Length = 451
Score = 136 bits (343), Expect = 1e-30
Identities = 103/340 (30%), Positives = 166/340 (48%), Gaps = 41/340 (12%)
Query: 138 AIIIDGGSGYCKFGWSKEDRP-------LGRVATFLEFGNVETP------IYTRLRH--F 182
A++ D GSG CK G++ +D P +GR F + P I R RH +
Sbjct: 13 ALVCDNGSGLCKAGFAGDDAPRAVFPSIVGRPRHQAGFAGDDAPRAVFPSIVGRPRHQIW 72
Query: 183 FATVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVPAVCALN 241
+ Y ++V P ++ PL + KA+R+++ + + NVPA+
Sbjct: 73 HHSFYNELRVAPEEHPTLLTEAPL-------NPKANREKMTQIMFETF---NVPAMYVAI 122
Query: 242 QATLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQ 301
QA L+LYA+ +T+GI ++ G VT VPI G + + + L LT +L + + +
Sbjct: 123 QAVLSLYASGRTTGIVLDSGDGVTHNVPIYEGYALPH-AIMRLDLAGRDLTDYLMKILTE 181
Query: 302 NNLYFESL---YTVRTLKEKLCYVAYDYEAEL-----SKDTHASFEAAEGK-FTLSKERF 352
F + VR +KEKLCYVA D+E E+ S S+E +G+ T+ ERF
Sbjct: 182 RGYSFVTTAEREIVRDIKEKLCYVALDFENEMGTAATSSSLEKSYELPDGQVITIGNERF 241
Query: 353 QTGEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAER 412
+ E LFQP G+ + +H + C ++ D + VLSGG+ PG+A+R
Sbjct: 242 RCPETLFQPSFIGMESAGIHETTYNSIMKC---DIDIRKDLYANNVLSGGTTMYPGIADR 298
Query: 413 LEKELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
++KE+ +L P M I++I PP + W G ++ S+S
Sbjct: 299 MQKEITALAPSTMK--IKIIAPPERKYSVWIGGSILASLS 336
>UniRef100_Q8L8Z7 Hypothetical protein [Arabidopsis thaliana]
Length = 165
Score = 136 bits (343), Expect = 1e-30
Identities = 74/154 (48%), Positives = 95/154 (61%), Gaps = 8/154 (5%)
Query: 19 NTTNSSSSSRSHQHHHHHAP------SPLGLFDSLPPDILLKITRLLGPKHAAKLCLVCK 72
N +NS +HQ P S LG FD LP DIL++I ++ PK A KL L CK
Sbjct: 12 NRSNSGKDLVNHQRAIDVPPLLLSSSSSLGAFDQLPMDILVQILMMMEPKDAVKLGLTCK 71
Query: 73 SWRSLVSDNELWAHFLQTHQPIHFHSILFSETNLTSGYPLPIFDTQTTPHVSFKHVFGHR 132
+W+ + S N LW +LQ Q + SI F+ET+L SGYPL + +Q+ +SF H++ R
Sbjct: 72 AWKCVASGNRLWIFYLQCSQE-PWDSIFFAETSLRSGYPLRMISSQSG-ELSFMHIYSQR 129
Query: 133 EQLPPAIIIDGGSGYCKFGWSKEDRPLGRVATFL 166
Q+P +IIIDGGSGYCKFGWSK P GR ATFL
Sbjct: 130 AQVPGSIIIDGGSGYCKFGWSKYASPSGRSATFL 163
>UniRef100_P53459 Actin 6 [Diphyllobothrium dendriticum]
Length = 373
Score = 136 bits (343), Expect = 1e-30
Identities = 108/365 (29%), Positives = 173/365 (46%), Gaps = 62/365 (16%)
Query: 133 EQLPPAIIIDGGSGYCKFGWSKEDRP-------LGRV---ATFLEFGN------------ 170
E++ P +++D GSG CK G++ +D P +GR + + GN
Sbjct: 1 EEVQP-LVVDNGSGMCKAGFAGDDSPRAVFPSIVGRPRQQSIMVGMGNKDSYVGDEAQSK 59
Query: 171 ---------VETPIYTRL----RHFFATVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKA 216
+E I T + + T Y ++V P V++ PL + KA
Sbjct: 60 RGILSLKYPIEHGIVTNWDDMEKIWHHTFYNELRVAPEEHPVLLTEAPL-------NPKA 112
Query: 217 SRQQLKEAICAALFDMNVPAVCALNQATLALYAANQTSGIAVNIGFQVTSVVPILNGKVM 276
+R+++ + N PA+ QA L+LYA+ +T+GI ++ G V+ VPI G +
Sbjct: 113 NREKMTSIMFETF---NCPAMYVAIQAVLSLYASGRTTGIVLDSGDGVSHTVPIYEGYAL 169
Query: 277 RKVGVEVVGLGALKLTGFLREKMQQNNLYFESLYT---VRTLKEKLCYVAYDYEAEL--- 330
+ + L LT +L + + + F + VR +KEKLCYVA D+E E+
Sbjct: 170 PHAILRL-DLAGRDLTDYLMKILTERGYSFTTTAEREIVRDIKEKLCYVALDFENEMATA 228
Query: 331 --SKDTHASFEAAEGK-FTLSKERFQTGEILFQPRLAGVRAMSLHHAIALCMDHCHSAEL 387
S S+E +G+ T+ ERF+ E LFQP G+ ++ +H C + +L
Sbjct: 229 ASSSSLEKSYELPDGQVITVGNERFRCPEALFQPSFLGLESVGIHET---CYNSIMKCDL 285
Query: 388 AGGSDWFKTVVLSGGSACLPGLAERLEKELHSLLPPYMSNGIRVIPPPHGADTAWFGAKL 447
D + +VLSGGS PG+A+R+ KEL SL P M I++I PP + W G +
Sbjct: 286 DIRKDLYSNIVLSGGSTMYPGIADRMNKELTSLAPSSMK--IKIIAPPERKYSVWIGGSI 343
Query: 448 IGSVS 452
+GS+S
Sbjct: 344 LGSLS 348
>UniRef100_Q5V8L7 Actin [Paxillus involutus]
Length = 375
Score = 134 bits (338), Expect = 4e-30
Identities = 107/360 (29%), Positives = 169/360 (46%), Gaps = 61/360 (16%)
Query: 138 AIIIDGGSGYCKFGWSKEDRP-------LGRV---ATFLEFGN----------------- 170
A++ID GSG CK G++ +D P +GR + G
Sbjct: 7 ALVIDNGSGMCKAGFAGDDAPRAVFPSIVGRPRHQGVMVGMGQKDSYVGDEAQSKRGILT 66
Query: 171 ----VETPIYTRL----RHFFATVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQL 221
+E I T + + T Y ++V P V++ PL + KA+R+++
Sbjct: 67 LKYPIEHGIVTNWDDMEKIWHHTFYNELRVAPEEHPVLLTEAPL-------NPKANREKM 119
Query: 222 KEAICAALFDMNVPAVCALNQATLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGV 281
+ + N PA QA L+LYA+ +T+GI ++ G VT VPI G + +
Sbjct: 120 TQIMFETF---NAPAFYVAIQAVLSLYASGRTTGIVMDSGDGVTHTVPIYEGFALPHAIL 176
Query: 282 EVVGLGALKLTGFLREKMQQNNLYFESLYT---VRTLKEKLCYVAYDYEAELSKDTHAS- 337
+ L LT +L + + + F + VR +KEKLCYVA D+E EL H+S
Sbjct: 177 RL-DLAGRDLTEYLIKNLMERGYAFTTTAEREIVRGIKEKLCYVALDFEQELQTAAHSSA 235
Query: 338 ----FEAAEGK-FTLSKERFQTGEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSD 392
+E +G+ T+ ERF++ E LFQP G+ A +H + C +L D
Sbjct: 236 LEKSYELPDGQVLTIGNERFRSPEALFQPAFLGLEAAGIHETTYNSIFKC---DLDIRRD 292
Query: 393 WFKTVVLSGGSACLPGLAERLEKELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
+ VVLSGG+ PG+A+R++KEL SL P M ++++ PP + W G ++ S+S
Sbjct: 293 LYGNVVLSGGTTMFPGIADRMQKELTSLSPSSMK--VKIVAPPERKYSVWIGGSILASLS 350
>UniRef100_Q964E1 Actin, cytoplasmic [Biomphalaria obstructa]
Length = 376
Score = 134 bits (337), Expect = 5e-30
Identities = 105/360 (29%), Positives = 169/360 (46%), Gaps = 61/360 (16%)
Query: 138 AIIIDGGSGYCKFGWSKEDRP-------LGRV---ATFLEFGN----------------- 170
A+++D GSG CK G++ +D P +GR + G
Sbjct: 8 ALVVDNGSGMCKAGFAGDDAPRAVFPSIVGRPRHQGVMVGMGQKDSYVGDEAQSKRGILT 67
Query: 171 ----VETPIYTRL----RHFFATVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQL 221
+E I T + + T Y ++V P V++ PL + KA+R+++
Sbjct: 68 LKYPIEHGIVTNWDDMEKIWHHTFYNELRVAPEEHPVLLTEAPL-------NPKANREKM 120
Query: 222 KEAICAALFDMNVPAVCALNQATLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGV 281
+ + N PA+ QA L+LYA+ +T+GI ++ G VT VPI G + +
Sbjct: 121 TQIMFETF---NTPAMYVAIQAVLSLYASGRTTGIVLDSGDGVTHTVPIYEGYALPHA-I 176
Query: 282 EVVGLGALKLTGFLREKMQQNNLYFESLYT---VRTLKEKLCYVAYDYEAELSKDTHAS- 337
+ L LT +L + + + F + VR +KEKLCYVA D+E E+ T +S
Sbjct: 177 MRLDLAGRDLTDYLMKILTERGYSFTTTAEREIVRDIKEKLCYVALDFEQEMQTATSSSS 236
Query: 338 ----FEAAEGK-FTLSKERFQTGEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSD 392
+E +G+ T+ ERF+ E LFQP G+ A +H + C ++ D
Sbjct: 237 LEKSYELPDGQVITIGNERFRCPEALFQPSFLGMEAAGIHETTYNSIMKC---DVDIRKD 293
Query: 393 WFKTVVLSGGSACLPGLAERLEKELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
+ VLSGGS PG+A+R++KE+ +L PP M I++I PP + W G ++ S+S
Sbjct: 294 LYANTVLSGGSTMFPGIADRMQKEITALAPPTMK--IKIIAPPERKYSVWIGGSILASLS 351
>UniRef100_Q964E0 Actin, cytoplasmic [Biomphalaria tenagophila]
Length = 376
Score = 133 bits (335), Expect = 9e-30
Identities = 104/360 (28%), Positives = 168/360 (45%), Gaps = 61/360 (16%)
Query: 138 AIIIDGGSGYCKFGWSKEDRP-------LGRV---ATFLEFGN----------------- 170
A+++D GSG CK G++ +D P +GR + G
Sbjct: 8 ALVVDNGSGMCKAGFAGDDAPRAVFPSIVGRPRHQGVMVGMGQKDSYVGDEAQSKRGILT 67
Query: 171 ----VETPIYTRL----RHFFATVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQL 221
+E I T + + T Y ++V P V++ PL + KA+R+++
Sbjct: 68 LKYPIEHGIVTNWDDMEKIWHHTFYNELRVAPEEHPVLLTEAPL-------NPKANREKM 120
Query: 222 KEAICAALFDMNVPAVCALNQATLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGV 281
+ + N PA+ QA L+LYA+ +T+GI ++ G VT VPI G + +
Sbjct: 121 TQIMFETF---NTPAMYVAIQAVLSLYASGRTTGIVLDSGDGVTHTVPIYEGYALPHA-I 176
Query: 282 EVVGLGALKLTGFLREKMQQNNLYFESLYT---VRTLKEKLCYVAYDYEAEL-----SKD 333
+ L LT +L + + + F + VR +KEKLCYVA D+E E+ S
Sbjct: 177 MRLDLAGRDLTDYLMKILTERGYSFTTTAEREIVRDIKEKLCYVALDFEQEMQTASTSSS 236
Query: 334 THASFEAAEGK-FTLSKERFQTGEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSD 392
S+E +G+ T+ ERF+ E +FQP G+ A +H + C ++ D
Sbjct: 237 LEKSYELPDGQVITIGNERFRCPEAMFQPSFLGMEAAGIHETTYNSIMKC---DVDIRKD 293
Query: 393 WFKTVVLSGGSACLPGLAERLEKELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
+ VLSGGS PG+A+R++KE+ +L PP M I++I PP + W G ++ S+S
Sbjct: 294 LYANTVLSGGSTMFPGIADRMQKEITALAPPTMK--IKIIAPPERKYSVWIGGSILASLS 351
>UniRef100_Q964E3 Actin, cytoplasmic [Biomphalaria alexandrina]
Length = 376
Score = 133 bits (335), Expect = 9e-30
Identities = 104/360 (28%), Positives = 168/360 (45%), Gaps = 61/360 (16%)
Query: 138 AIIIDGGSGYCKFGWSKEDRP-------LGRV---ATFLEFGN----------------- 170
A+++D GSG CK G++ +D P +GR + G
Sbjct: 8 ALVVDNGSGMCKAGFAGDDAPRAVFPSIVGRPRHQGVMVGMGQKDSYVGDEAQSKRGILT 67
Query: 171 ----VETPIYTRL----RHFFATVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQL 221
+E I T + + T Y ++V P V++ PL + KA+R+++
Sbjct: 68 LKYPIEHGIVTNWDDMEKIWHHTFYNELRVAPEEHPVLLTEAPL-------NPKANREKM 120
Query: 222 KEAICAALFDMNVPAVCALNQATLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGV 281
+ + N PA+ QA L+LYA+ +T+GI ++ G VT VPI G + +
Sbjct: 121 TQIMFETF---NTPAMYVAIQAVLSLYASGRTTGIVMDSGDGVTHTVPIYEGYALPHA-I 176
Query: 282 EVVGLGALKLTGFLREKMQQNNLYFESLYT---VRTLKEKLCYVAYDYEAEL-----SKD 333
+ L LT +L + + + F + VR +KEKLCYVA D+E E+ S
Sbjct: 177 MRLDLAGRDLTDYLMKILTERGYSFTTTAEREIVRDIKEKLCYVALDFEQEMQTASTSSS 236
Query: 334 THASFEAAEGK-FTLSKERFQTGEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSD 392
S+E +G+ T+ ERF+ E +FQP G+ A +H + C ++ D
Sbjct: 237 LEKSYELPDGQVITIGNERFRCPEAMFQPSFLGMEAAGIHETTYNSIMKC---DVDIRKD 293
Query: 393 WFKTVVLSGGSACLPGLAERLEKELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
+ VLSGGS PG+A+R++KE+ +L PP M I++I PP + W G ++ S+S
Sbjct: 294 LYANTVLSGGSTMFPGIADRMQKEITALAPPTMK--IKIIAPPERKYSVWIGGSILASLS 351
>UniRef100_Q6WE51 Actin [Glaeseria mira]
Length = 366
Score = 133 bits (334), Expect = 1e-29
Identities = 105/360 (29%), Positives = 170/360 (47%), Gaps = 61/360 (16%)
Query: 138 AIIIDGGSGYCKFGWSKEDRP-------LGRV---ATFLEFGN----------------- 170
A+++D GSG CK G++ +D P +GR + G
Sbjct: 6 ALVVDNGSGMCKAGFAGDDAPRAVFPSIVGRPRHKGVMVGMGQKDSYVGDEAQSKRGILT 65
Query: 171 ----VETPIYTRL----RHFFATVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQL 221
+E I T + + T Y ++V P V++ PL + KA+R+++
Sbjct: 66 LKYPIEHGIVTNWDDMEKIWHHTFYNELRVAPEEHPVLLTEAPL-------NPKANREKM 118
Query: 222 KEAICAALFDMNVPAVCALNQATLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGV 281
+ + NVPA+ QA L+LYA+ +T+GI ++IG V VPI G + +
Sbjct: 119 TQIMFETF---NVPAMYVAIQAVLSLYASGRTTGIVLDIGDGVCHTVPIYEGYALPH-AI 174
Query: 282 EVVGLGALKLTGFLREKMQQNNLYFESL---YTVRTLKEKLCYVAYDYEAEL-----SKD 333
+ L LT + + + + F + VR +KEKLCYVA D++AE+ S
Sbjct: 175 ARLDLAGRDLTDYCMKILTERGYAFTTTAEREIVRDIKEKLCYVALDFDAEMDTAASSST 234
Query: 334 THASFEAAEGK-FTLSKERFQTGEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSD 392
S+E +G+ T+ ERF+T E LFQP G + +H A + C ++ D
Sbjct: 235 IEKSYELPDGQVITVGNERFRTPEALFQPSFLGRESSGIHEATYNSIMKC---DIDIRKD 291
Query: 393 WFKTVVLSGGSACLPGLAERLEKELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
+ +VLSGG+ PG+A+RL+KEL +L P M +++I PP + W G ++ S+S
Sbjct: 292 LYGNIVLSGGTTMFPGIADRLQKELVNLAPSTMK--VKIIAPPERKYSVWIGGSILASLS 349
>UniRef100_Q6WE48 Actin [Hartmannella cantabrigiensis]
Length = 374
Score = 133 bits (334), Expect = 1e-29
Identities = 105/360 (29%), Positives = 170/360 (47%), Gaps = 61/360 (16%)
Query: 138 AIIIDGGSGYCKFGWSKEDRP-------LGRV---ATFLEFGN----------------- 170
A+++D GSG CK G++ +D P +GR + G
Sbjct: 6 ALVVDNGSGMCKAGFAGDDAPRAVFPSIVGRPRHKGVMVGMGQKDSYVGDEAQSKRGILT 65
Query: 171 ----VETPIYTRL----RHFFATVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQL 221
+E I T + + T Y ++V P V++ PL + KA+R+++
Sbjct: 66 LKYPIEHGIVTNWDDMEKIWHHTFYNELRVAPEEHPVLLTEAPL-------NPKANREKM 118
Query: 222 KEAICAALFDMNVPAVCALNQATLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGV 281
+ + NVPA+ QA L+LYA+ +T+GI ++IG V VPI G + +
Sbjct: 119 TQIMFETF---NVPAMYVAIQAVLSLYASGRTTGIVLDIGDGVCHTVPIYEGYALPH-AI 174
Query: 282 EVVGLGALKLTGFLREKMQQNNLYFESL---YTVRTLKEKLCYVAYDYEAEL-----SKD 333
+ L LT + + + + F + VR +KEKLCYVA D++AE+ S
Sbjct: 175 ARLDLAGRDLTDYCMKILTERGYAFTTTAEREIVRDIKEKLCYVALDFDAEMDTAASSST 234
Query: 334 THASFEAAEGK-FTLSKERFQTGEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSD 392
S+E +G+ T+ ERF+T E LFQP G + +H A + C ++ D
Sbjct: 235 IEKSYELPDGQVITVGNERFRTPEALFQPSFLGRESAGIHEATYNSIMKC---DIDIRKD 291
Query: 393 WFKTVVLSGGSACLPGLAERLEKELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
+ +VLSGG+ PG+A+RL+KEL +L P M +++I PP + W G ++ S+S
Sbjct: 292 LYGNIVLSGGTTMFPGIADRLQKELVNLAPSTMK--VKIIAPPERKYSVWIGGSILSSLS 349
>UniRef100_P20360 Actin, cytoplasmic [Euplotes crassus]
Length = 379
Score = 131 bits (330), Expect = 3e-29
Identities = 91/278 (32%), Positives = 143/278 (50%), Gaps = 26/278 (9%)
Query: 185 TVYGRMKVKPSSQVVVVSLPLCHYDDTESAKASRQQLKEAICAALFD-MNVPAVCALNQA 243
T Y ++++ + V+++ A + +Q +E +C +F+ + P++ QA
Sbjct: 93 TFYNELRIETENHPVLLT----------EAPLNPKQNRENMCRIMFEEYDFPSMYIQIQA 142
Query: 244 TLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNN 303
L+LY+A +T+GI V+ G VT VVPI G + +E + L LT ++ ++ ++
Sbjct: 143 VLSLYSAGRTTGIVVDSGDGVTHVVPIFEGYQIPHA-IEKILLAGRDLTDYMCRILKDDD 201
Query: 304 LYFESLY---TVRTLKEKLCYVAYDYEAELSK-----DTHASFEAAEGK-FTLSKERFQT 354
+FE+ TVR +KEKLCYVA DYEAEL K + S+ +G+ +S +RFQ
Sbjct: 202 YHFETTAEKETVRDIKEKLCYVADDYEAELKKAGEGGELEESYALPDGRPLKISTQRFQC 261
Query: 355 GEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLE 414
E LFQP L G S+H L D + +L D + ++LSGG+ PGL ERL
Sbjct: 262 PEFLFQPDLGGRECKSVHQ---LTYDSIMTCDLDVRKDLYANIILSGGTTMFPGLGERLY 318
Query: 415 KELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
KE+ L P M ++VI P W G + +S
Sbjct: 319 KEMKDLAPQTMK--VKVIASPDRKYAVWRGGSTLAKLS 354
>UniRef100_P53474 Actin, cytoskeletal IIIA [Strongylocentrotus purpuratus]
Length = 376
Score = 131 bits (329), Expect = 5e-29
Identities = 87/278 (31%), Positives = 140/278 (50%), Gaps = 26/278 (9%)
Query: 185 TVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVPAVCALNQA 243
T Y ++V P V++ PL + KA+R+++ + + N PA+ QA
Sbjct: 90 TFYNELRVAPEEHPVLLTEAPL-------NPKANREKMTQIMFETF---NSPAMYVAIQA 139
Query: 244 TLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNN 303
L+LYA+ +T+GI + G V+ VPI G + + + L LT +L + + +
Sbjct: 140 VLSLYASGRTTGIVFDSGDGVSHTVPIYEGYALPHAIIRL-DLAGRDLTDYLMKILTERG 198
Query: 304 LYFESLYT---VRTLKEKLCYVAYDYEAEL-----SKDTHASFEAAEGK-FTLSKERFQT 354
F + VR +KEKLCYVA D+E E+ S S+E +G+ T+ ERF+
Sbjct: 199 YSFTTTAEREIVRDIKEKLCYVALDFEQEMQTAASSSSLEKSYELPDGQVITIGNERFRA 258
Query: 355 GEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLE 414
E LFQP G+ + +H C + ++ D + VLSGGS PG+A+R++
Sbjct: 259 SETLFQPSFIGMESAGIHET---CYNRIMKCDVDIRKDLYVNTVLSGGSTMFPGIADRMQ 315
Query: 415 KELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
KE+ +L PP M I++I PP + W G ++ S+S
Sbjct: 316 KEISALAPPTMK--IKIIAPPERKYSVWIGGSILASLS 351
>UniRef100_Q96481 Actin 105 [Lycopersicon esculentum]
Length = 336
Score = 131 bits (329), Expect = 5e-29
Identities = 91/278 (32%), Positives = 141/278 (49%), Gaps = 26/278 (9%)
Query: 185 TVYGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVPAVCALNQA 243
T Y ++V P +++ PL + KA+R+++ + + NVPA+ QA
Sbjct: 71 TFYNELRVAPEDDSILLTEAPL-------NPKANREKMTQIMFETF---NVPAMYIAIQA 120
Query: 244 TLALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNN 303
L+LYA+ +T+GI ++ G VT VPI G + + + L LT +L + +
Sbjct: 121 VLSLYASGRTTGIVLDSGDGVTHTVPIYEGYALPH-AILRLDLAGRDLTDYLMMILTERG 179
Query: 304 LYFESL---YTVRTLKEKLCYVAYDYEAEL-----SKDTHASFEAAEGKF-TLSKERFQT 354
F + VR +KEKL YVA DYE EL KD S+E +G+ T+ ERF+
Sbjct: 180 YSFTTAAEREIVRDVKEKLAYVALDYEQELETTKNGKDAEKSYELPDGQIVTIGAERFRC 239
Query: 355 GEILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLE 414
E+LFQP L G+ A+ +H + C ++ D + VVLSGGS PG+ +R+
Sbjct: 240 PEVLFQPSLIGMEAVGIHETTYSSIMKC---DVDIRRDLYGNVVLSGGSTMFPGITDRMS 296
Query: 415 KELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
KE +L P M I+V+ PP + W G ++ S+S
Sbjct: 297 KEFTALAPSSMK--IKVVAPPESKYSVWIGGSILASLS 332
>UniRef100_Q9NBY9 Actin [Eledonella pygmaea]
Length = 261
Score = 130 bits (326), Expect = 1e-28
Identities = 88/277 (31%), Positives = 143/277 (50%), Gaps = 28/277 (10%)
Query: 187 YGRMKVKPSSQ-VVVVSLPLCHYDDTESAKASRQQLKEAICAALFD-MNVPAVCALNQAT 244
Y ++V P V++ PL + KA+R+++ + +FD N PA+ QA
Sbjct: 2 YNELRVAPEEHPVLLTEAPL-------NPKANREKMTQI----MFDTFNTPAMYVAIQAV 50
Query: 245 LALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNNL 304
L+LYA+ +T+GI ++ G VT VPI +G + + + L LT +L + + +
Sbjct: 51 LSLYASGRTTGIVMDSGDGVTHTVPIYHGYALPHAIIRL-DLAGRDLTDYLMKILTERGY 109
Query: 305 YFESLYT---VRTLKEKLCYVAYDYEAEL-----SKDTHASFEAAEGK-FTLSKERFQTG 355
F + VR +KEKLCYVA D+E E+ S S+E +G+ T+ ERF+
Sbjct: 110 SFTTTAEREIVRDIKEKLCYVALDFEQEMQTAASSSSLEKSYELPDGQVITIGNERFRAP 169
Query: 356 EILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLEK 415
E +FQP G+ + +H + C ++ D + VLSGGS PG+A+R++K
Sbjct: 170 EAMFQPSFIGMESAGIHETTYNSIMKC---DVDIRKDLYANTVLSGGSTMYPGIADRMQK 226
Query: 416 ELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
E+ +L PP M I++I PP + W G ++ S+S
Sbjct: 227 EISALAPPTMK--IKIIAPPERKYSVWIGGSILASLS 261
>UniRef100_Q962V5 Alpha actin [Homarus americanus]
Length = 377
Score = 130 bits (326), Expect = 1e-28
Identities = 88/277 (31%), Positives = 146/277 (51%), Gaps = 28/277 (10%)
Query: 187 YGRMKVKPS-SQVVVVSLPLCHYDDTESAKASRQQLKEAICAALFDM-NVPAVCALNQAT 244
Y +++ P S V++ PL + KA+R+++ + +F++ N PA+ QA
Sbjct: 93 YNELRIAPEESPVLLTEAPL-------NPKANREKMTQI----MFEIFNTPAMYVAIQAV 141
Query: 245 LALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNNL 304
L+LYA+ +T+GI ++ G VT VPI G + + + L LT +L + M +
Sbjct: 142 LSLYASGRTTGIVLDTGDGVTHTVPIYEGYCLPH-AILRLDLAGRDLTAYLTKIMTERGY 200
Query: 305 YFESL---YTVRTLKEKLCYVAYDYEAEL-----SKDTHASFEAAEGK-FTLSKERFQTG 355
F + VR +KEKLCYVA D+E E+ S S+E +G+ T+ ERF+
Sbjct: 201 SFTTTAEREIVRDIKEKLCYVALDFENEMNIAAASSSVEKSYELPDGQVITIGNERFRCP 260
Query: 356 EILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLEK 415
E LFQP G+ ++ +H + + C ++ D F VLSGG+ PG+A+R++K
Sbjct: 261 ESLFQPSFLGMESVGIHETVYNSIMRC---DIDIRKDLFANNVLSGGTTMYPGIADRMQK 317
Query: 416 ELHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
E+ +L PP + I++I PP + W G ++ S+S
Sbjct: 318 EITALAPPTIK--IKIIAPPERKYSVWIGGSILASLS 352
>UniRef100_Q8WPD5 Beta-actin [Gecarcinus lateralis]
Length = 376
Score = 129 bits (325), Expect = 1e-28
Identities = 87/276 (31%), Positives = 144/276 (51%), Gaps = 26/276 (9%)
Query: 187 YGRMKVKPS-SQVVVVSLPLCHYDDTESAKASRQQLKEAICAALFDMNVPAVCALNQATL 245
Y ++V P S V++ PL + KA+R+++ + + N PA+ QA L
Sbjct: 92 YNELRVAPEESPVLLTEAPL-------NPKANREKMTQIMFEVF---NTPAMYVAIQAVL 141
Query: 246 ALYAANQTSGIAVNIGFQVTSVVPILNGKVMRKVGVEVVGLGALKLTGFLREKMQQNNLY 305
+LYA+ +T+GI ++ G VT VPI G + + + L LT +L + M +
Sbjct: 142 SLYASGRTTGIVLDTGDGVTHTVPIYEGYCLPH-AILRLDLAGRDLTAYLTKIMTERGYS 200
Query: 306 FESL---YTVRTLKEKLCYVAYDYEAELS-----KDTHASFEAAEGKF-TLSKERFQTGE 356
F + VR +KEKLCYVA D+E+E++ S+E +G+F T+ ERF+ E
Sbjct: 201 FTTTAEREIVRDIKEKLCYVALDFESEMNVAAAPSSLEKSYELPDGRFITIGNERFRCPE 260
Query: 357 ILFQPRLAGVRAMSLHHAIALCMDHCHSAELAGGSDWFKTVVLSGGSACLPGLAERLEKE 416
LFQP G+ ++ +H + + C ++ D F VLSG + PG+A+R++KE
Sbjct: 261 SLFQPSFLGMESVGIHETVYNSIMRC---DIDIRKDLFANNVLSGRTTMYPGIADRMQKE 317
Query: 417 LHSLLPPYMSNGIRVIPPPHGADTAWFGAKLIGSVS 452
+ +L PP + I++I PP + W G ++ S+S
Sbjct: 318 ITALAPPTIK--IKIIAPPERKYSVWIGGSILASLS 351
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.135 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 768,721,442
Number of Sequences: 2790947
Number of extensions: 32066513
Number of successful extensions: 120872
Number of sequences better than 10.0: 1905
Number of HSP's better than 10.0 without gapping: 1556
Number of HSP's successfully gapped in prelim test: 349
Number of HSP's that attempted gapping in prelim test: 113048
Number of HSP's gapped (non-prelim): 2997
length of query: 457
length of database: 848,049,833
effective HSP length: 131
effective length of query: 326
effective length of database: 482,435,776
effective search space: 157274062976
effective search space used: 157274062976
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 76 (33.9 bits)
Medicago: description of AC145470.4


