

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145331.12 - phase: 0
(419 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
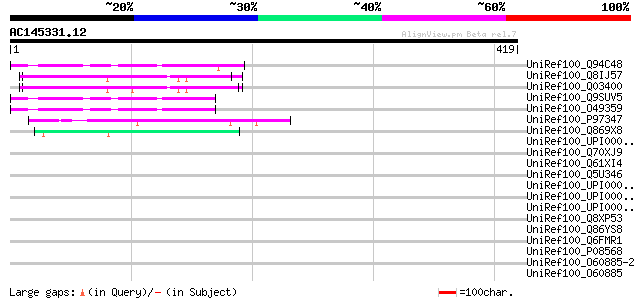
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q94C48 Hypothetical protein At4g32330 [Arabidopsis tha... 57 1e-06
UniRef100_Q8IJ57 S-antigen [Plasmodium falciparum] 54 6e-06
UniRef100_Q03400 S-antigen protein precursor [Plasmodium falcipa... 54 6e-06
UniRef100_Q9SUV5 Hypothetical protein F8B4.30 [Arabidopsis thali... 54 1e-05
UniRef100_O49359 Hypothetical protein F10M6.40 [Arabidopsis thal... 54 1e-05
UniRef100_P97347 Repetin [Mus musculus] 51 7e-05
UniRef100_Q869X8 Similar to Homo sapiens (Human). dentin sialoph... 47 8e-04
UniRef100_UPI000042E5EA UPI000042E5EA UniRef100 entry 47 0.001
UniRef100_Q70XJ9 Levansucrase [Lactobacillus sanfranciscensis] 47 0.001
UniRef100_Q61XI4 Hypothetical protein CBG03972 [Caenorhabditis b... 47 0.001
UniRef100_Q5U346 Hypothetical protein [Rattus norvegicus] 46 0.002
UniRef100_UPI00001AFFBD UPI00001AFFBD UniRef100 entry 45 0.003
UniRef100_UPI0000170992 UPI0000170992 UniRef100 entry 45 0.003
UniRef100_UPI000046BA7C UPI000046BA7C UniRef100 entry 45 0.003
UniRef100_Q8XP53 Hypothetical protein CPE0112 [Clostridium perfr... 45 0.003
UniRef100_Q86YS8 BRD4-NUT fusion oncoprotein [Homo sapiens] 45 0.003
UniRef100_Q6FMR1 Candida glabrata strain CBS138 chromosome K com... 45 0.003
UniRef100_P08568 Submandibular gland secretory Glx-rich protein ... 45 0.003
UniRef100_O60885-2 Splice isoform 2 of O60885 [Homo sapiens] 45 0.003
UniRef100_O60885 Bromodomain-containing protein 4 [Homo sapiens] 45 0.003
>UniRef100_Q94C48 Hypothetical protein At4g32330 [Arabidopsis thaliana]
Length = 437
Score = 57.0 bits (136), Expect = 1e-06
Identities = 55/210 (26%), Positives = 92/210 (43%), Gaps = 37/210 (17%)
Query: 1 MDPVNSLPADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLE 60
MDP + + ADG D P+N G + + + V+ ET S+N N
Sbjct: 1 MDPESIMAADGTDSA--------PANGGLAMENVCVKENGAVSVETVDTTSESQNENSAN 52
Query: 61 STATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSG 120
S S + IE + +G+ + + V + +AQ+ P K + ++S
Sbjct: 53 S-----STLDTIEHVKEAAEGTQV-----EHVDDSKCMKGEKAQRKPRHEKLSGGKNNSS 102
Query: 121 VNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKK--------- 171
V+ +K SK GK A A SNG+ A + + P+K++S N R++Q++K+
Sbjct: 103 VH---IKKSKEGKSADAKVAASNGSVAPNVQTTNPLKSKSFNGREAQVTKQGKHDSAPAE 159
Query: 172 -------KPKSLKKGPLDKVQGEGESSLYP 194
KPKS KK + + + +SS P
Sbjct: 160 SADGEKVKPKSQKKQAHETSEDDTQSSNSP 189
>UniRef100_Q8IJ57 S-antigen [Plasmodium falciparum]
Length = 585
Score = 54.3 bits (129), Expect = 6e-06
Identities = 49/189 (25%), Positives = 81/189 (41%), Gaps = 10/189 (5%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSA 68
++G +D NG D+ SN GED VSN + V+ E V +G + ++ N
Sbjct: 298 SNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGREDKVSNGGEDEVSNGREDKVSNGG 357
Query: 69 MKEI-EGSNDNV-DGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLV 126
E+ G D V +G VS +E ++ E + G + N + S V
Sbjct: 358 EDEVSNGREDKVSNGREDKVSNGREDEVSNGREDKVSNGGEDEVSNGREDKVSNGREDKV 417
Query: 127 KNSKIGKDKQAS---PAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDK 183
N G+D+ ++ VSNG S R+ ++ SN R+ ++S + + G D+
Sbjct: 418 SNG--GEDEVSNGREDKVSNGGEDEVSNGRE---DKVSNGREDKVSNGREDKVSNGGEDE 472
Query: 184 VQGEGESSL 192
V GE +
Sbjct: 473 VSNGGEDEV 481
Score = 54.3 bits (129), Expect = 6e-06
Identities = 49/186 (26%), Positives = 75/186 (39%), Gaps = 12/186 (6%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSA 68
++G +D NG D+ SN GED VSN + V+ E V +G + ++ N
Sbjct: 322 SNGREDKVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGREDKVSNGR 381
Query: 69 MKEI-EGSNDNV-DGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLV 126
E+ G D V +G VS +E K+ E + G + N + S V
Sbjct: 382 EDEVSNGREDKVSNGGEDEVSNGREDKVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEV 441
Query: 127 KNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQG 186
N + K VSNG S R+ ++ SN + ++S + G DKV
Sbjct: 442 SNGREDK-------VSNGREDKVSNGRE---DKVSNGGEDEVSNGGEDEVSNGREDKVSN 491
Query: 187 EGESSL 192
GE +
Sbjct: 492 GGEDEV 497
Score = 52.4 bits (124), Expect = 2e-05
Identities = 51/202 (25%), Positives = 82/202 (40%), Gaps = 20/202 (9%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENI-NQLESTATGNS 67
++G +D NG DE SN GED VSN + V+ E V +G + + N E +
Sbjct: 106 SNGREDKVSNGGEDEVSNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGR 165
Query: 68 AMKEIEGSNDNV---------DGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSS 118
K G D V +G VS +E K+ E + G + N +
Sbjct: 166 EDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGREDKVSNGGEDEVSNGREDKV 225
Query: 119 SGVNASLVKNSKIGKDKQAS---PAVSNG-----TSALDSRPRQPIKNRSSNDRQSQLSK 170
S V N G+D+ ++ VSNG ++ + + ++ SN R+ ++S
Sbjct: 226 SNGREDKVSNG--GEDEVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSN 283
Query: 171 KKPKSLKKGPLDKVQGEGESSL 192
+ + G DKV GE +
Sbjct: 284 GREDEVSNGREDKVSNGGEDEV 305
Score = 51.6 bits (122), Expect = 4e-05
Identities = 50/186 (26%), Positives = 75/186 (39%), Gaps = 20/186 (10%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSA 68
++G +D NG D+ SN GED VSN + V+ E V +G + ++ N
Sbjct: 194 SNGREDKVSNGREDKVSNGGEDEVSNGREDKVSNGREDKVSNGGEDEVSNGREDKVSNGG 253
Query: 69 MKEI-EGSNDNV-DGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLV 126
E+ G D V +G VS +E K+ E + K N G N
Sbjct: 254 EDEVSNGREDKVSNGGEDEVSNGREDKVSNGREDEVSNGREDKVSNG--GEDEVSNGRED 311
Query: 127 KNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQG 186
K S G+D+ VSNG +++ SN R+ ++S + G DKV
Sbjct: 312 KVSNGGEDE-----VSNGR-----------EDKVSNGREDKVSNGGEDEVSNGREDKVSN 355
Query: 187 EGESSL 192
GE +
Sbjct: 356 GGEDEV 361
Score = 51.2 bits (121), Expect = 5e-05
Identities = 48/184 (26%), Positives = 72/184 (39%), Gaps = 12/184 (6%)
Query: 11 GLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMK 70
G +D NG D+ SN GED VSN + V+ E V +G + ++ N
Sbjct: 100 GSEDEVSNGREDKVSNGGEDEVSNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGGED 159
Query: 71 EI-EGSNDNV-DGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKN 128
E+ G D V +G VS +E K+ E + K N + S V N
Sbjct: 160 EVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGREDKVSNGGEDEVSN 219
Query: 129 SKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEG 188
+ K VSNG + + ++ SN R+ ++S + G DKV G
Sbjct: 220 GREDK-------VSNGR---EDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGG 269
Query: 189 ESSL 192
E +
Sbjct: 270 EDEV 273
Score = 51.2 bits (121), Expect = 5e-05
Identities = 42/177 (23%), Positives = 74/177 (41%), Gaps = 4/177 (2%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSA 68
++G +D NG D+ SN GED VSN + V+ E V +G + ++ N
Sbjct: 378 SNGREDEVSNGREDKVSNGGEDEVSNGREDKVSNGREDKVSNGGEDEVSNGREDKVSNGG 437
Query: 69 MKEI-EGSNDNV-DGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLV 126
E+ G D V +G VS +E K+ E + G + N + S V
Sbjct: 438 EDEVSNGREDKVSNGREDKVSNGREDKVSNGGEDEVSNGGEDEVSNGREDKVSNGGEDEV 497
Query: 127 KNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDK 183
N + +DK ++ ++ + + ++ SN R+ ++S + + G DK
Sbjct: 498 SNGR--EDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGREDEVSNGREDK 552
Score = 50.8 bits (120), Expect = 7e-05
Identities = 46/186 (24%), Positives = 75/186 (39%), Gaps = 4/186 (2%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENI-NQLESTATGNS 67
++G +D NG DE SN ED VSN + V+ E V +G + + N E +
Sbjct: 346 SNGREDKVSNGGEDEVSNGREDKVSNGREDKVSNGREDEVSNGREDKVSNGGEDEVSNGR 405
Query: 68 AMKEIEGSNDNV-DGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLV 126
K G D V +G VS +E K+ E + K N + S V
Sbjct: 406 EDKVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGREDKVSNGREDKV 465
Query: 127 KNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQG 186
N G+D+ ++ ++ + + ++ SN R+ ++S + G DKV
Sbjct: 466 SNG--GEDEVSNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSN 523
Query: 187 EGESSL 192
GE +
Sbjct: 524 GGEDEV 529
Score = 50.1 bits (118), Expect = 1e-04
Identities = 47/184 (25%), Positives = 75/184 (40%), Gaps = 16/184 (8%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSA 68
++G +D NG D+ SN GED VSN + V+ E V +G + ++ N
Sbjct: 154 SNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGREDKVSNGG 213
Query: 69 MKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKN 128
E+ ++ ++ +E +V E S ++ V N G N K
Sbjct: 214 EDEVSNGRED----KVSNGREDKVSNGGEDEVSNGREDKVSNG----GEDEVSNGREDKV 265
Query: 129 SKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEG 188
S G+D+ VSNG S R+ + SN R+ ++S + G DKV G
Sbjct: 266 SNGGEDE-----VSNGREDKVSNGRE---DEVSNGREDKVSNGGEDEVSNGREDKVSNGG 317
Query: 189 ESSL 192
E +
Sbjct: 318 EDEV 321
Score = 37.7 bits (86), Expect = 0.61
Identities = 25/93 (26%), Positives = 40/93 (42%), Gaps = 4/93 (4%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSA 68
++G +D NG DE SN ED VSN + V+ E V +G + ++ N
Sbjct: 482 SNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGR 541
Query: 69 MKEIEGSNDNVDGS----NLTVSKEKEVKIKVS 97
E+ ++ G+ L+ + E K K S
Sbjct: 542 EDEVSNGREDKGGAGTDGELSHNSESHTKNKKS 574
>UniRef100_Q03400 S-antigen protein precursor [Plasmodium falciparum]
Length = 593
Score = 54.3 bits (129), Expect = 6e-06
Identities = 48/186 (25%), Positives = 78/186 (41%), Gaps = 12/186 (6%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSA 68
++G +D NG D+ SN GED VSN + V+ E V +G + ++ N
Sbjct: 298 SNGGEDEVSNGREDKVSNGGEDEVSNGREDEVSNGREDKVSNGGEDEVSNGREDKVSNGG 357
Query: 69 MKEI-EGSNDNV-DGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLV 126
E+ G D V +G VS +E ++ E + G + N + S V
Sbjct: 358 EDEVSNGREDEVSNGREDEVSNGREDEVSNGREDKVSNGGEDEVSNGREDEVSNGREDKV 417
Query: 127 KNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQG 186
N G+D+ VSNG + + ++ SN R+ ++S + + G DKV
Sbjct: 418 SNG--GEDE-----VSNGR---EDKVSNGGEDEVSNGREDEVSNGREDEVSNGREDKVSN 467
Query: 187 EGESSL 192
GE +
Sbjct: 468 GGEDEV 473
Score = 52.4 bits (124), Expect = 2e-05
Identities = 43/183 (23%), Positives = 76/183 (41%), Gaps = 4/183 (2%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSA 68
++G +D NG D+ SN GED VSN + V+ E V +G + ++ N
Sbjct: 378 SNGREDEVSNGREDKVSNGGEDEVSNGREDEVSNGREDKVSNGGEDEVSNGREDKVSNGG 437
Query: 69 MKEI-EGSNDNV-DGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLV 126
E+ G D V +G VS +E K+ E + G + N + S V
Sbjct: 438 EDEVSNGREDEVSNGREDEVSNGREDKVSNGGEDEVSNGGEDEVSNGREDKVSNGGEDEV 497
Query: 127 KNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQG 186
N + +DK ++ ++ + + ++ SN R+ ++S + + G D+V
Sbjct: 498 SNGR--EDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDEVSNGREDEVSNGREDEVSN 555
Query: 187 EGE 189
E
Sbjct: 556 GRE 558
Score = 51.2 bits (121), Expect = 5e-05
Identities = 50/185 (27%), Positives = 73/185 (39%), Gaps = 18/185 (9%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSA 68
++G +D NG D+ SN GED VSN + V+ E V +G + ++ N
Sbjct: 194 SNGREDEVSNGREDKVSNGGEDEVSNGREDEVSNGREDKVSNGGEDEVSNGREDKVSNGG 253
Query: 69 MKEI-EGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVK 127
E+ G D V E EV E S ++ V N S+ G + V
Sbjct: 254 EDEVSNGREDKVSNGG-----EDEVSNGREDEVSNGREDEVSNGREDKVSNGGEDE--VS 306
Query: 128 NSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGE 187
N + K VSNG S R+ + SN R+ ++S + G DKV
Sbjct: 307 NGREDK-------VSNGGEDEVSNGRE---DEVSNGREDKVSNGGEDEVSNGREDKVSNG 356
Query: 188 GESSL 192
GE +
Sbjct: 357 GEDEV 361
Score = 51.2 bits (121), Expect = 5e-05
Identities = 50/202 (24%), Positives = 82/202 (39%), Gaps = 20/202 (9%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENI-NQLESTATGNS 67
++G +D NG DE SN GED VSN + V+ E V +G + + N E +
Sbjct: 106 SNGREDKVSNGGEDEVSNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGR 165
Query: 68 AMKEIEGSNDNV---------DGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSS 118
K G D V +G VS +E ++ E + G + N +
Sbjct: 166 EDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDEVSNGREDKVSNGGEDEVSNGREDEV 225
Query: 119 SGVNASLVKNSKIGKDKQAS---PAVSNG-----TSALDSRPRQPIKNRSSNDRQSQLSK 170
S V N G+D+ ++ VSNG ++ + + ++ SN R+ ++S
Sbjct: 226 SNGREDKVSNG--GEDEVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDEVSN 283
Query: 171 KKPKSLKKGPLDKVQGEGESSL 192
+ + G DKV GE +
Sbjct: 284 GREDEVSNGREDKVSNGGEDEV 305
Score = 50.8 bits (120), Expect = 7e-05
Identities = 47/192 (24%), Positives = 76/192 (39%), Gaps = 8/192 (4%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSA 68
++G +D NG D+ SN GED VSN + V+ E V +G + ++ N
Sbjct: 338 SNGGEDEVSNGREDKVSNGGEDEVSNGREDEVSNGREDEVSNGREDEVSNGREDKVSNGG 397
Query: 69 MKEI-EGSNDNV-DGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLV 126
E+ G D V +G VS E ++ E + G + N + S V
Sbjct: 398 EDEVSNGREDEVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDEVSNGREDEV 457
Query: 127 KNSKIGK-DKQASPAVSNG-----TSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGP 180
N + K VSNG ++ + + ++ SN R+ ++S + G
Sbjct: 458 SNGREDKVSNGGEDEVSNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGR 517
Query: 181 LDKVQGEGESSL 192
DKV GE +
Sbjct: 518 EDKVSNGGEDEV 529
Score = 49.7 bits (117), Expect = 2e-04
Identities = 48/187 (25%), Positives = 76/187 (39%), Gaps = 18/187 (9%)
Query: 11 GLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMK 70
G +D NG D+ SN GED VSN + V+ E V +G + ++ N
Sbjct: 100 GTEDEVSNGREDKVSNGGEDEVSNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGGED 159
Query: 71 EI-EGSNDNV-DGSNLTVSKEKEVKIKVSTEQ--SRAQKGPVKN-KNAKVGSSSGVNASL 125
E+ G D V +G VS +E K+ E S ++ V N + KV + S
Sbjct: 160 EVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDEVSNGREDKVSNGGEDEVSN 219
Query: 126 VKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQ 185
+ ++ ++ VSNG ++ SN R+ ++S + G DKV
Sbjct: 220 GREDEVSNGRE--DKVSNGG-----------EDEVSNGREDKVSNGGEDEVSNGREDKVS 266
Query: 186 GEGESSL 192
GE +
Sbjct: 267 NGGEDEV 273
Score = 49.7 bits (117), Expect = 2e-04
Identities = 47/184 (25%), Positives = 75/184 (40%), Gaps = 16/184 (8%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSA 68
++G +D NG D+ SN GED VSN + V+ E V +G + ++ N
Sbjct: 154 SNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDEVSNGREDKVSNGG 213
Query: 69 MKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKN 128
E+ ++ ++ +E +V E S ++ V N G N K
Sbjct: 214 EDEVSNGRED----EVSNGREDKVSNGGEDEVSNGREDKVSNG----GEDEVSNGREDKV 265
Query: 129 SKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEG 188
S G+D+ VSNG S R+ + SN R+ ++S + G DKV G
Sbjct: 266 SNGGEDE-----VSNGREDEVSNGRE---DEVSNGREDKVSNGGEDEVSNGREDKVSNGG 317
Query: 189 ESSL 192
E +
Sbjct: 318 EDEV 321
Score = 38.5 bits (88), Expect = 0.36
Identities = 25/93 (26%), Positives = 41/93 (43%), Gaps = 4/93 (4%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSA 68
++G +D NG D+ SN GED VSN + V+ E V +G + ++ N
Sbjct: 490 SNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDEVSNGREDEVSNGR 549
Query: 69 MKEIEGSNDNVDGS----NLTVSKEKEVKIKVS 97
E+ ++ G+ L+ + E K K S
Sbjct: 550 EDEVSNGREDKGGAGTDGELSHNSESHTKNKKS 582
>UniRef100_Q9SUV5 Hypothetical protein F8B4.30 [Arabidopsis thaliana]
Length = 423
Score = 53.5 bits (127), Expect = 1e-05
Identities = 46/170 (27%), Positives = 78/170 (45%), Gaps = 21/170 (12%)
Query: 1 MDPVNSLPADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLE 60
MDP + + ADG D P+N G + + + V+ ET S+N N
Sbjct: 24 MDPESIMAADGTDSA--------PANGGLAMENVCVKENGAVSVETVDTTSESQNENSAN 75
Query: 61 STATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSG 120
S S + IE + +G+ + + V + +AQ+ P K + ++S
Sbjct: 76 S-----STLDTIEHVKEAAEGTQV-----EHVDDSKCMKGEKAQRKPRHEKLSGGKNNSS 125
Query: 121 VNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSK 170
V+ +K SK GK A A SNG+ A + + P+K++S N R++Q++K
Sbjct: 126 VH---IKKSKEGKSADAKVAASNGSVAPNVQTTNPLKSKSFNGREAQVTK 172
>UniRef100_O49359 Hypothetical protein F10M6.40 [Arabidopsis thaliana]
Length = 423
Score = 53.5 bits (127), Expect = 1e-05
Identities = 46/170 (27%), Positives = 78/170 (45%), Gaps = 21/170 (12%)
Query: 1 MDPVNSLPADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLE 60
MDP + + ADG D P+N G + + + V+ ET S+N N
Sbjct: 24 MDPESIMAADGTDSA--------PANGGLAMENVCVKENGAVSVETVDTTSESQNENSAN 75
Query: 61 STATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSG 120
S S + IE + +G+ + + V + +AQ+ P K + ++S
Sbjct: 76 S-----STLDTIEHVKEAAEGTQV-----EHVDDSKCMKGEKAQRKPRHEKLSGGKNNSS 125
Query: 121 VNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSK 170
V+ +K SK GK A A SNG+ A + + P+K++S N R++Q++K
Sbjct: 126 VH---IKKSKEGKSADAKVAASNGSVAPNVQTTNPLKSKSFNGREAQVTK 172
>UniRef100_P97347 Repetin [Mus musculus]
Length = 1130
Score = 50.8 bits (120), Expect = 7e-05
Identities = 53/231 (22%), Positives = 98/231 (41%), Gaps = 26/231 (11%)
Query: 16 HQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEIEGS 75
HQN HD+ +D+ +D DPH +ETF D + G+S ++ + S
Sbjct: 142 HQNVQHDQSQRQDKDSERHDTDPHCG-QSETFHGDSH-----------YGHSERQDTDYS 189
Query: 76 NDNVDGSNLTVSKEKEVKIKVSTEQSRAQ---------KGPVKNKNAKVGSSSGVNASLV 126
+D + N + S + + K S EQ + Q K P + + +SG +S
Sbjct: 190 SDQSESDNESSSSSQRLGYKSSHEQPKGQGYVFALSQSKNPEQAFHYGQSKTSGQQSSHG 249
Query: 127 KNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPL---DK 183
++ + KD +S + + + Q K+ +S + + + ++ KKG ++
Sbjct: 250 QSGRFRKDSYSSQTSQQESDSYEQYGSQHQKSGNSQTERQGQNSQYGQTNKKGHSSYHEQ 309
Query: 184 VQGEGESSLYPFSLRKGES--SSLRLRKRFRQRRRRRVACKQNPRRVKKPR 232
+G+G+S Y RK +S + RK ++ Q+P R +K R
Sbjct: 310 TEGQGQSFHYGQKGRKDQSFQQGQKGRKDQSPHLGQKGRQDQSPHRGQKGR 360
>UniRef100_Q869X8 Similar to Homo sapiens (Human). dentin sialophosphoprotein
[Dictyostelium discoideum]
Length = 1206
Score = 47.4 bits (111), Expect = 8e-04
Identities = 40/178 (22%), Positives = 72/178 (39%), Gaps = 8/178 (4%)
Query: 21 HDEPSN--SGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEIEGSNDN 78
H+EPSN S ++ +N+ + + N E+ N+E+ N S N+ E +N++
Sbjct: 922 HNEPSNQSSNNESSNNESSNNESSNNESSSKGSNNESSNNESSNQGSNNESSNNESNNES 981
Query: 79 VD------GSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIG 132
+ GSN S + + S++ S ++ N + S + S ++S G
Sbjct: 982 SNNESSSQGSNNESSNNESSNNESSSQSSNNGSSNNESSNQGSNNESSSHGSNNESSSKG 1041
Query: 133 KDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGES 190
+ ++S SN S+ + N SSN + S S K + E S
Sbjct: 1042 SNNESSSHGSNNESSSQGSNNESSNNESSNQNSNNESSNNESSSKGSNNESSNNESSS 1099
Score = 40.0 bits (92), Expect = 0.12
Identities = 44/183 (24%), Positives = 67/183 (36%), Gaps = 17/183 (9%)
Query: 3 PVNSLPADGLDDVHQNGVHDEPSNSGEDAVSNDL----------DPHVTVNTETFVPDGN 52
P N G N ++ S+SG+D+ S L D H + N E+ N
Sbjct: 516 PTNDSSNQGSGSHSNNHSSNQGSSSGQDSSSETLKIQNSNNESSDDHSSTNNESSSQGSN 575
Query: 53 SENINQLESTATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKN 112
+E+ N S N+ E SN GSN S E + S +S + + N
Sbjct: 576 NESSNNESSNQGSNNESSHNETSN---QGSN-NESSHNESSSQGSNNESAHNESSNQGSN 631
Query: 113 AKVGSSSGVNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKK 172
SS +S ++ + ++S SN S+ + Q N SSN+ S
Sbjct: 632 ---NESSHNESSNQGSNNESSNNESSNQGSNNESSHNESSNQGSNNESSNNESSNQGSNN 688
Query: 173 PKS 175
S
Sbjct: 689 ESS 691
Score = 36.6 bits (83), Expect = 1.4
Identities = 37/186 (19%), Positives = 73/186 (38%), Gaps = 4/186 (2%)
Query: 19 GVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEIEGSNDN 78
G ++E SN+ ++ +N+ + N + N + N+ S + N + + + +
Sbjct: 990 GSNNESSNN--ESSNNESSSQSSNNGSSNNESSNQGSNNESSSHGSNNESSSKGSNNESS 1047
Query: 79 VDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQAS 138
GSN S + + E S + N S +S ++S G + ++S
Sbjct: 1048 SHGSNNESSSQGSNNESSNNESSNQNSNNESSNNESSSKGSNNESSNNESSSQGSNNESS 1107
Query: 139 PAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGESSLYPFSLR 198
SN ++ + + + SN QS ++ K + G +++ ESS
Sbjct: 1108 NNESNNQNSNNESSHK--EESESNSHQSSETESKDQVSNSGSMNQSSESSESSSLSEQSS 1165
Query: 199 KGESSS 204
ESSS
Sbjct: 1166 AFESSS 1171
Score = 36.2 bits (82), Expect = 1.8
Identities = 34/160 (21%), Positives = 59/160 (36%), Gaps = 13/160 (8%)
Query: 13 DDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEI 72
D +QN ++ +N + SN+L N+E+ N S N+
Sbjct: 750 DSNNQNSNNEPSNNESSNQSSNNLS-------------SNNESSNNESSNQGSNNESSHS 796
Query: 73 EGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIG 132
E SN+ +L E ++E S N + SSS + S N++
Sbjct: 797 ESSNNESSNQSLNSEYNNEPSQGSNSESSNESSNQSLNSESNNDSSSHGSNSESSNNESS 856
Query: 133 KDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKK 172
+ + +N +S+ DS S+N+ SQ S +
Sbjct: 857 SHGSNNESSNNESSSQDSSNYSSNNGSSNNESSSQGSNNE 896
Score = 36.2 bits (82), Expect = 1.8
Identities = 35/164 (21%), Positives = 70/164 (42%), Gaps = 5/164 (3%)
Query: 16 HQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEIEGS 75
H ++E SN ++ N+ +P N+E+ N ++ + ++ + + E +
Sbjct: 795 HSESSNNESSNQSLNSEYNN-EPSQGSNSESSNESSNQSLNSESNNDSSSHGSNSESSNN 853
Query: 76 NDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDK 135
+ GSN S E + S+ S + G N+++ GS++ +S ++ +
Sbjct: 854 ESSSHGSN-NESSNNESSSQDSSNYS-SNNGSSNNESSSQGSNN--ESSNQGSNNESSNN 909
Query: 136 QASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKG 179
++S SN S+ + Q N SSN+ S +S KG
Sbjct: 910 ESSSQGSNNESSHNEPSNQSSNNESSNNESSNNESSNNESSSKG 953
Score = 35.4 bits (80), Expect = 3.0
Identities = 34/161 (21%), Positives = 67/161 (41%), Gaps = 10/161 (6%)
Query: 16 HQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEIEGS 75
+++ ++ S + SN+ + N E+ + N+E+ N S+ N+ E S
Sbjct: 942 NESSNNESSSKGSNNESSNNESSNQGSNNESSNNESNNESSNNESSSQGSNNESSNNESS 1001
Query: 76 -NDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKD 134
N++ S+ S E + S +S + N+++ GS++ ++S G +
Sbjct: 1002 NNESSSQSSNNGSSNNESSNQGSNNESSSHGS--NNESSSKGSNN-------ESSSHGSN 1052
Query: 135 KQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKS 175
++S SN S+ + Q N SSN+ S S
Sbjct: 1053 NESSSQGSNNESSNNESSNQNSNNESSNNESSSKGSNNESS 1093
Score = 35.0 bits (79), Expect = 4.0
Identities = 35/157 (22%), Positives = 66/157 (41%), Gaps = 13/157 (8%)
Query: 19 GVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEIEGSNDN 78
G ++E SN+ SN + + N E+ N+E+ N S+ N+ E +N N
Sbjct: 1059 GSNNESSNNES---SNQNSNNESSNNESSSKGSNNESSNNESSSQGSNNESSNNESNNQN 1115
Query: 79 VDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQAS 138
E K +E + Q ++K+ +V +S +N S + +Q+S
Sbjct: 1116 ---------SNNESSHKEESESNSHQSSETESKD-QVSNSGSMNQSSESSESSSLSEQSS 1165
Query: 139 PAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKS 175
S+ +S+ +S+ + + SS+ S+ S S
Sbjct: 1166 AFESSSSSSGNSQSKSSENDDSSSSSGSKNSNSAQSS 1202
Score = 35.0 bits (79), Expect = 4.0
Identities = 33/151 (21%), Positives = 66/151 (42%), Gaps = 5/151 (3%)
Query: 21 HDEPSNSGEDAVSNDLDPH-VTVNTETFVPDGNSENINQLESTATGNSAMKEIEGSNDNV 79
H+E S+ G + S++ D + N E + ++++ N L S ++ +GSN+
Sbjct: 734 HNESSSQGSNNESSNNDSNNQNSNNEPSNNESSNQSSNNLSSNNESSNNESSNQGSNNES 793
Query: 80 DGSNLT--VSKEKEVKIKVSTEQSRAQKGPVKNK--NAKVGSSSGVNASLVKNSKIGKDK 135
S + S + + + + E S+ N+ N + S S ++S ++ +
Sbjct: 794 SHSESSNNESSNQSLNSEYNNEPSQGSNSESSNESSNQSLNSESNNDSSSHGSNSESSNN 853
Query: 136 QASPAVSNGTSALDSRPRQPIKNRSSNDRQS 166
++S SN S+ + Q N SSN+ S
Sbjct: 854 ESSSHGSNNESSNNESSSQDSSNYSSNNGSS 884
Score = 33.9 bits (76), Expect = 8.8
Identities = 34/172 (19%), Positives = 64/172 (36%), Gaps = 8/172 (4%)
Query: 16 HQNGVHDEPSNSGED------AVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAM 69
+ H+E SN G + SN + + N E+ N+E+ + S N+
Sbjct: 617 NNESAHNESSNQGSNNESSHNESSNQGSNNESSNNESSNQGSNNESSHNESSNQGSNNES 676
Query: 70 KEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVN--ASLVK 127
E SN + + + S+ + +G S+ G N +S +
Sbjct: 677 SNNESSNQGSNNESSHNETSNQGSNNESSHNESSSQGSNNESAHNESSNQGSNNESSHNE 736
Query: 128 NSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKG 179
+S G + ++S SN ++ + N+SSN+ S +S +G
Sbjct: 737 SSSQGSNNESSNNDSNNQNSNNEPSNNESSNQSSNNLSSNNESSNNESSNQG 788
>UniRef100_UPI000042E5EA UPI000042E5EA UniRef100 entry
Length = 2010
Score = 47.0 bits (110), Expect = 0.001
Identities = 40/182 (21%), Positives = 78/182 (41%), Gaps = 6/182 (3%)
Query: 52 NSENINQLESTATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVK-- 109
+S ++S+A +S+ S N + SKE +I+VS +SR+++ K
Sbjct: 688 SSVKSKSVQSSAPSSSSSSSSSSSKKNSKNVSSIPSKENLSEIEVSRGRSRSRRNSSKSN 747
Query: 110 ----NKNAKVGSSSGVNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQ 165
N+N+ +GSS S + + KDK S + + +S +S ++ SS+
Sbjct: 748 FSESNRNSIIGSSDSEKDSKIASKSKSKDKLKSESKNKLSSGSNSENSIESESSSSSSSS 807
Query: 166 SQLSKKKPKSLKKGPLDKVQGEGESSLYPFSLRKGESSSLRLRKRFRQRRRRRVACKQNP 225
S+ + + + + E +S L K SSS + K Q + + +++P
Sbjct: 808 SESGSRSASRSESNSSSRSESESSNSENEMKLNKKLSSSSNINKLDSQTQEDLNSFEKDP 867
Query: 226 RR 227
+
Sbjct: 868 TK 869
>UniRef100_Q70XJ9 Levansucrase [Lactobacillus sanfranciscensis]
Length = 879
Score = 47.0 bits (110), Expect = 0.001
Identities = 39/194 (20%), Positives = 77/194 (39%), Gaps = 5/194 (2%)
Query: 2 DPVNSLPADGLDDVH---QNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQ 58
D V + DG +V+ Q V + ++ ++A S ++ + + +
Sbjct: 36 DAVENNKYDGTANVNIDCQANVDGKIISTDDNATSGSTKQESSIANDNATSGSTKQESSI 95
Query: 59 LESTATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSS 118
AT S +E +NDN + +E V +T S Q+ V N NA GS+
Sbjct: 96 ANDNATSGSTKQESSIANDNATSGS--TKQESSVANDNATSGSTKQESSVANDNATSGST 153
Query: 119 SGVNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKK 178
++ N+ G KQ S ++ +A+ + + N + Q + ++P + +
Sbjct: 154 KQESSVANDNATSGSTKQESSVANDTKTAVVDESKNTSNTENDNSQLKQTNNEQPSAATQ 213
Query: 179 GPLDKVQGEGESSL 192
L K+ E ++
Sbjct: 214 ANLKKLNHEAAKAV 227
>UniRef100_Q61XI4 Hypothetical protein CBG03972 [Caenorhabditis briggsae]
Length = 720
Score = 47.0 bits (110), Expect = 0.001
Identities = 41/130 (31%), Positives = 64/130 (48%), Gaps = 8/130 (6%)
Query: 51 GNSENINQLESTATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSR-AQKGPVK 109
G+ EN S + + K + S + D + S+ K+ KV +E+S+ +QK K
Sbjct: 35 GHRENFRNRGSKSKRSKRPKRSKSSRNRSDKGKKSSSRRKKKLRKVKSEKSKKSQKRSKK 94
Query: 110 NKNAKVGSSSGVNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPI-KNRSSNDRQSQL 168
+K K GS S N+S K S+ K+AS + SR ++PI K+R +DR S
Sbjct: 95 HKKKKRGSKSKSNSSKSKKSR----KEASE--KRNSKRGSSRGQKPIDKDRRDDDRGSSR 148
Query: 169 SKKKPKSLKK 178
SK+ K +K
Sbjct: 149 SKRDSKRSQK 158
>UniRef100_Q5U346 Hypothetical protein [Rattus norvegicus]
Length = 752
Score = 45.8 bits (107), Expect = 0.002
Identities = 41/170 (24%), Positives = 71/170 (41%), Gaps = 24/170 (14%)
Query: 64 TGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNA 123
+G ++EIE N D ++ ++ + + +S+ +K K + SSS
Sbjct: 146 SGQEVVREIE--NQKTDAASKPFAEVRILSCGELVPKSKVKKEEKKRHKSSSSSSSS--- 200
Query: 124 SLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGP--- 180
+S D Q+S S+ SA + + R+ K N R+ + KKK K KK P
Sbjct: 201 ----DSDSSSDSQSSSDSSDSESASEEKSRKRKKKHRKNSRKHKKEKKKRKKSKKSPSSE 256
Query: 181 --LDKVQGEGESSLYP----------FSLRKGESSSLRLRKRFRQRRRRR 218
D V + +S++ P F +RK + ++ R+R R R
Sbjct: 257 SEADNVDAQPQSTVRPEEIPPIPENRFLMRKSPPKADDKERKNRERERER 306
>UniRef100_UPI00001AFFBD UPI00001AFFBD UniRef100 entry
Length = 794
Score = 45.4 bits (106), Expect = 0.003
Identities = 52/214 (24%), Positives = 81/214 (37%), Gaps = 26/214 (12%)
Query: 22 DEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEIEGSNDNVDG 81
DEP E+ V P V T+ P +S++ + S + + S D+ +
Sbjct: 459 DEP----EEPVVAVSSPAVPPPTKVVAPPSSSDSSSDSSSDS---------DSSTDDSEE 505
Query: 82 SNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQASPAV 141
E + ++K EQ A P +NK K K K K K+
Sbjct: 506 ERAQRLAELQEQLKAVHEQLAALSQPQQNKPKKKEKDK-------KEKKKEKHKRKEEVE 558
Query: 142 SNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGESSLYPFSLRKGE 201
N S P P K + +N S +SKK+P +K P + E E P S +
Sbjct: 559 ENKKSKAKEPP--PKKTKKNNSSNSNVSKKEPAPMKSKPPPTYESEEEDKCKPMSYEEKR 616
Query: 202 SSSLRLRKRFRQRRRRRVACKQNPRRVKKPRLKS 235
SL + K ++ R V Q+ ++P LK+
Sbjct: 617 QLSLDINKLPGEKLGRVVHIIQS----REPSLKN 646
>UniRef100_UPI0000170992 UPI0000170992 UniRef100 entry
Length = 255
Score = 45.4 bits (106), Expect = 0.003
Identities = 34/167 (20%), Positives = 66/167 (39%), Gaps = 9/167 (5%)
Query: 9 ADGLDDVHQNG-VHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNS 67
A G D+ N D P++S + V + DP D ++EN+ + ES A N
Sbjct: 16 AQGTDEEVNNAETSDVPADSEQQPVDSGSDPPSA--------DADAENVQEGESAAPANE 67
Query: 68 AMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVK 127
GS + T ++ +E +E+ + Q+ P + +N + ++SG +
Sbjct: 68 EPPATSGSEEEQQQQEPTQAENQEPPATSGSEEEQQQQEPTQAENQEPPATSGSEEEQQQ 127
Query: 128 NSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPK 174
+ Q PA S + +N+ +D + + +P+
Sbjct: 128 QEPTQAEDQQPPATSGSEEEQQQQESTQAENQEPSDSAGEGQETQPE 174
>UniRef100_UPI000046BA7C UPI000046BA7C UniRef100 entry
Length = 1370
Score = 45.4 bits (106), Expect = 0.003
Identities = 43/186 (23%), Positives = 80/186 (42%), Gaps = 9/186 (4%)
Query: 53 SENI-NQLESTATGNSAMKEIEGSND--NVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVK 109
SENI N++ S K +N+ NV+ + EK+ K+ E +K K
Sbjct: 266 SENISNEINCNDVEKSGEKGTNNNNNCGNVNSEHNKQKSEKQTNNKLPKEGKNREK---K 322
Query: 110 NKNAKVGSSSGVNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLS 169
N ++ SS + KN+K + + L + Q I+N+ S+D +++
Sbjct: 323 NNSSSSSSSENSSYEYNKNNKESDEDKKKKLKKKLKKKLKKKYMQKIRNKYSSD--DKIN 380
Query: 170 KKKPKSLKKGPLDKVQGEGESSLYPFSLRKGESSSLR-LRKRFRQRRRRRVACKQNPRRV 228
K K+LKK ++ + +S Y S + + R L ++R + ++ K ++
Sbjct: 381 KYNEKNLKKENINDIYSSSYNSNYESSYSENSDTKERELDDEKKKRHKNKILEKLRKKKE 440
Query: 229 KKPRLK 234
KK + K
Sbjct: 441 KKKKKK 446
Score = 35.8 bits (81), Expect = 2.3
Identities = 39/173 (22%), Positives = 76/173 (43%), Gaps = 19/173 (10%)
Query: 27 SGEDAVSNDLDPH-VTVNTETFVPD-------GNSENINQLESTATGNSAMKEIEGSNDN 78
SGE+ + ++ P+ VN F + +S N+ + + KE E ++
Sbjct: 213 SGENFFTANIRPNDANVNINEFRANKEKDANIASSPNVGNKQKDMDKDETEKESENISNE 272
Query: 79 VDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQAS 138
++ +++ S EK + G V +++ K S N L K GK+++
Sbjct: 273 INCNDVEKSGEKGTN-------NNNNCGNVNSEHNKQKSEKQTNNKLPKE---GKNREKK 322
Query: 139 PAVSNGTSALDSRPRQPIKNRSSN-DRQSQLSKKKPKSLKKGPLDKVQGEGES 190
S+ +S+ +S N+ S+ D++ +L KK K LKK + K++ + S
Sbjct: 323 NNSSSSSSSENSSYEYNKNNKESDEDKKKKLKKKLKKKLKKKYMQKIRNKYSS 375
>UniRef100_Q8XP53 Hypothetical protein CPE0112 [Clostridium perfringens]
Length = 521
Score = 45.4 bits (106), Expect = 0.003
Identities = 38/143 (26%), Positives = 63/143 (43%), Gaps = 11/143 (7%)
Query: 37 DPHVTVNTETFVPDGNSENINQLESTATG----NSAMKEIEGSNDNVDGSNLTVSKEKEV 92
D VT+N T+ DGN N +QL + T ++ E N+ D S + SKEKE
Sbjct: 305 DGEVTINGVTYDKDGNMINKDQLNNIGTSISDEDNDTSNNESKNNIEDKSQKSDSKEKET 364
Query: 93 -KIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQASPAVSNGTSALDSR 151
K + ++ +G +++KN +G + K++K K+K A A + S +
Sbjct: 365 DKANSNKQEDSNSQGELESKN------NGSQNNDTKSNKESKNKDAQGAKESKNSKPNKN 418
Query: 152 PRQPIKNRSSNDRQSQLSKKKPK 174
KN S+ + + K K
Sbjct: 419 SVSSSKNNSNKNNSGSSNNSKDK 441
>UniRef100_Q86YS8 BRD4-NUT fusion oncoprotein [Homo sapiens]
Length = 1846
Score = 45.4 bits (106), Expect = 0.003
Identities = 52/214 (24%), Positives = 81/214 (37%), Gaps = 26/214 (12%)
Query: 22 DEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEIEGSNDNVDG 81
DEP E+ V P V T+ P +S++ + S + + S D+ +
Sbjct: 459 DEP----EEPVVAVSSPAVPPPTKVVAPPSSSDSSSDSSSDS---------DSSTDDSEE 505
Query: 82 SNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQASPAV 141
E + ++K EQ A P +NK K K K K K+
Sbjct: 506 ERAQRLAELQEQLKAVHEQLAALSQPQQNKPKKKEKDK-------KEKKKEKHKRKEEVE 558
Query: 142 SNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGESSLYPFSLRKGE 201
N S P P K + +N S +SKK+P +K P + E E P S +
Sbjct: 559 ENKKSKAKEPP--PKKTKKNNSSNSNVSKKEPAPMKSKPPPTYESEEEDKCKPMSYEEKR 616
Query: 202 SSSLRLRKRFRQRRRRRVACKQNPRRVKKPRLKS 235
SL + K ++ R V Q+ ++P LK+
Sbjct: 617 QLSLDINKLPGEKLGRVVHIIQS----REPSLKN 646
>UniRef100_Q6FMR1 Candida glabrata strain CBS138 chromosome K complete sequence
[Candida glabrata]
Length = 794
Score = 45.4 bits (106), Expect = 0.003
Identities = 43/173 (24%), Positives = 77/173 (43%), Gaps = 12/173 (6%)
Query: 21 HDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEIEGSNDNVD 80
H + ++SG D +N LD + +N T +P + IN T T + +E SND +
Sbjct: 176 HRKKNHSGNDDTNNKLDTFLPIN--TVIP--YARIINAKIFTTTPQNQNQEPRTSNDFTN 231
Query: 81 GSNLT---VSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQA 137
S L+ VS + I V ++ R + K NA++ S + ++ N +
Sbjct: 232 QSILSRDDVSSQASKFISVPDDRPRRKSFKRKPSNARITSMTSIDGDTDNNPFSTQPSST 291
Query: 138 SPAVSNGTSALDSRPRQPIK-----NRSSNDRQSQLSKKKPKSLKKGPLDKVQ 185
S + S T+ D+ +K N N S++S K+ + L +G D+++
Sbjct: 292 SASASRTTTNNDNNDNVDLKNPFNTNDDDNVNNSKVSVKESEILYEGSQDEME 344
>UniRef100_P08568 Submandibular gland secretory Glx-rich protein CA precursor [Rattus
norvegicus]
Length = 246
Score = 45.4 bits (106), Expect = 0.003
Identities = 34/167 (20%), Positives = 66/167 (39%), Gaps = 9/167 (5%)
Query: 9 ADGLDDVHQNG-VHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNS 67
A G D+ N D P++S + V + DP D ++EN+ + ES A N
Sbjct: 16 AQGTDEEVNNAETSDVPADSEQQPVDSGSDPPSA--------DADAENVQEGESAAPANE 67
Query: 68 AMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVK 127
GS + T ++ +E +E+ + Q+ P + +N + ++SG +
Sbjct: 68 EPPATSGSEEEQQQQEPTQAENQEPPATSGSEEEQQQQEPTQAENQEPPATSGSEEEQQQ 127
Query: 128 NSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPK 174
+ Q PA S + +N+ +D + + +P+
Sbjct: 128 QEPTQAEDQQPPATSGSEEEQQQQESTQAENQEPSDSAGEGQETQPE 174
>UniRef100_O60885-2 Splice isoform 2 of O60885 [Homo sapiens]
Length = 722
Score = 45.4 bits (106), Expect = 0.003
Identities = 52/214 (24%), Positives = 81/214 (37%), Gaps = 26/214 (12%)
Query: 22 DEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEIEGSNDNVDG 81
DEP E+ V P V T+ P +S++ + S + + S D+ +
Sbjct: 459 DEP----EEPVVAVSSPAVPPPTKVVAPPSSSDSSSDSSSDS---------DSSTDDSEE 505
Query: 82 SNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQASPAV 141
E + ++K EQ A P +NK K K K K K+
Sbjct: 506 ERAQRLAELQEQLKAVHEQLAALSQPQQNKPKKKEKDK-------KEKKKEKHKRKEEVE 558
Query: 142 SNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGESSLYPFSLRKGE 201
N S P P K + +N S +SKK+P +K P + E E P S +
Sbjct: 559 ENKKSKAKEPP--PKKTKKNNSSNSNVSKKEPAPMKSKPPPTYESEEEDKCKPMSYEEKR 616
Query: 202 SSSLRLRKRFRQRRRRRVACKQNPRRVKKPRLKS 235
SL + K ++ R V Q+ ++P LK+
Sbjct: 617 QLSLDINKLPGEKLGRVVHIIQS----REPSLKN 646
>UniRef100_O60885 Bromodomain-containing protein 4 [Homo sapiens]
Length = 1362
Score = 45.4 bits (106), Expect = 0.003
Identities = 52/214 (24%), Positives = 81/214 (37%), Gaps = 26/214 (12%)
Query: 22 DEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEIEGSNDNVDG 81
DEP E+ V P V T+ P +S++ + S + + S D+ +
Sbjct: 459 DEP----EEPVVAVSSPAVPPPTKVVAPPSSSDSSSDSSSDS---------DSSTDDSEE 505
Query: 82 SNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQASPAV 141
E + ++K EQ A P +NK K K K K K+
Sbjct: 506 ERAQRLAELQEQLKAVHEQLAALSQPQQNKPKKKEKDK-------KEKKKEKHKRKEEVE 558
Query: 142 SNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGESSLYPFSLRKGE 201
N S P P K + +N S +SKK+P +K P + E E P S +
Sbjct: 559 ENKKSKAKEPP--PKKTKKNNSSNSNVSKKEPAPMKSKPPPTYESEEEDKCKPMSYEEKR 616
Query: 202 SSSLRLRKRFRQRRRRRVACKQNPRRVKKPRLKS 235
SL + K ++ R V Q+ ++P LK+
Sbjct: 617 QLSLDINKLPGEKLGRVVHIIQS----REPSLKN 646
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.324 0.137 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 628,401,498
Number of Sequences: 2790947
Number of extensions: 25037236
Number of successful extensions: 143961
Number of sequences better than 10.0: 1024
Number of HSP's better than 10.0 without gapping: 67
Number of HSP's successfully gapped in prelim test: 1022
Number of HSP's that attempted gapping in prelim test: 140756
Number of HSP's gapped (non-prelim): 3024
length of query: 419
length of database: 848,049,833
effective HSP length: 130
effective length of query: 289
effective length of database: 485,226,723
effective search space: 140230522947
effective search space used: 140230522947
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 76 (33.9 bits)
Medicago: description of AC145331.12


