

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145330.19 - phase: 0 /pseudo
(469 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
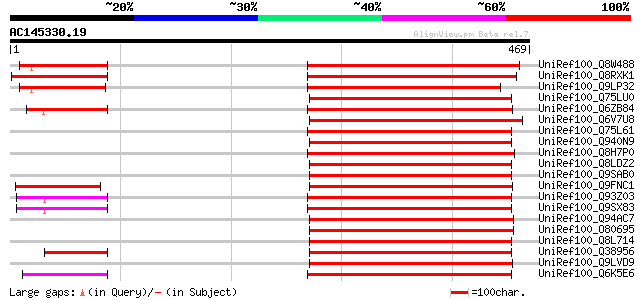
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8W488 Hypothetical protein T9L6.1 [Arabidopsis thaliana] 249 9e-65
UniRef100_Q8RXK1 Hypothetical protein At1g33080 [Arabidopsis tha... 240 7e-62
UniRef100_Q9LP32 T9L6.1 protein [Arabidopsis thaliana] 235 2e-60
UniRef100_Q75LU0 Putative MATE family protein [Oryza sativa] 211 5e-53
UniRef100_Q6ZB84 Putative ripening regulated protein [Oryza sativa] 210 6e-53
UniRef100_Q6V7U8 Putative anthocyanin permease [Lycopersicon esc... 209 1e-52
UniRef100_Q75L61 Putative MATE efflux family protein [Oryza sativa] 208 3e-52
UniRef100_Q940N9 AT4g21910/T8O5_120 [Arabidopsis thaliana] 206 9e-52
UniRef100_Q8H7P0 Putative ripening regulated protein [Oryza sativa] 206 9e-52
UniRef100_Q8LDZ2 Hypothetical protein [Arabidopsis thaliana] 206 9e-52
UniRef100_Q9SAB0 F25C20.18 protein [Arabidopsis thaliana] 206 2e-51
UniRef100_Q9FNC1 Emb|CAB89401.1 [Arabidopsis thaliana] 206 2e-51
UniRef100_Q93Z03 At1g47530/F16N3_20 [Arabidopsis thaliana] 204 6e-51
UniRef100_Q9SX83 F16N3.20 [Arabidopsis thaliana] 204 6e-51
UniRef100_Q94AC7 At1g61890/F8K4_9 [Arabidopsis thaliana] 204 6e-51
UniRef100_O80695 Hypothetical protein F8K4.9 [Arabidopsis thaliana] 203 8e-51
UniRef100_Q8L714 Hypothetical protein At1g11670 [Arabidopsis tha... 203 1e-50
UniRef100_Q38956 Hypothetical protein [Arabidopsis thaliana] 202 2e-50
UniRef100_Q9LVD9 Gb|AAC28507.1 [Arabidopsis thaliana] 202 2e-50
UniRef100_Q6K5E6 Putative ripening regulated protein DDTFR18 [Or... 202 2e-50
>UniRef100_Q8W488 Hypothetical protein T9L6.1 [Arabidopsis thaliana]
Length = 494
Score = 249 bits (637), Expect = 9e-65
Identities = 116/191 (60%), Positives = 154/191 (79%)
Query: 270 NVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFL 329
+ ++ICLNING EMMI+LGF+AAASVRVSNELG G+ K AKF+ + V TSL++G LF
Sbjct: 296 DALAICLNINGLEMMIALGFLAAASVRVSNELGSGNPKGAKFATLTAVFTSLSLGIVLFF 355
Query: 330 FFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVN 389
FLF R R++YIFT+++ VAA V +LSPLL+ SIL+NSVQPVLSGVA+GAGWQ V YVN
Sbjct: 356 VFLFLRGRVSYIFTTSEAVAAEVADLSPLLAFSILMNSVQPVLSGVAVGAGWQGYVTYVN 415
Query: 390 IGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKR 449
+ CYY++GIP+GI+LG ++ QVKG+W+GMLFG +QT VL ++T +T+WD+QV+ + +R
Sbjct: 416 LACYYLVGIPIGIILGYVVGLQVKGVWIGMLFGIFVQTCVLTVMTLRTDWDQQVSTSLRR 475
Query: 450 VNRWSKVESTD 460
+NRW ES D
Sbjct: 476 LNRWVVPESRD 486
Score = 105 bits (263), Expect = 2e-21
Identities = 50/81 (61%), Positives = 67/81 (81%), Gaps = 2/81 (2%)
Query: 10 LLKKPEEEN--EQEELSLGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSR 67
LLKK E E++EL L ++VW E+K +W+VAAPAIFTRFSTFG+ IISQ+F+GH+G
Sbjct: 12 LLKKTAENGGEEKDELGLKQKVWIESKKLWIVAAPAIFTRFSTFGVSIISQSFIGHLGPI 71
Query: 68 ELAAFALVFTVLIRFANGILM 88
ELAA+++ FTVL+RF+NGIL+
Sbjct: 72 ELAAYSITFTVLLRFSNGILL 92
>UniRef100_Q8RXK1 Hypothetical protein At1g33080 [Arabidopsis thaliana]
Length = 494
Score = 240 bits (612), Expect = 7e-62
Identities = 110/189 (58%), Positives = 149/189 (78%)
Query: 270 NVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFL 329
N ++IC+NIN EMM++ GFMAAASVRVSNE+G G++ AKF+ +V V TSL+IG F
Sbjct: 296 NALAICININALEMMVAFGFMAAASVRVSNEIGSGNSNGAKFATMVVVSTSLSIGIIFFF 355
Query: 330 FFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVN 389
FLF RER++YIFT+++ VA V +LSPLL+ SILLNS+QPVLSGVA+GAGWQ V VN
Sbjct: 356 IFLFLRERVSYIFTTSEAVATQVADLSPLLAFSILLNSIQPVLSGVAVGAGWQKYVTVVN 415
Query: 390 IGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKR 449
+ CYY++GIP G+ LG ++ QVKG+W+GM+FG +QT VL ++T +T+WD+QV+ + KR
Sbjct: 416 LACYYLVGIPSGLFLGYVVGLQVKGVWLGMIFGIFVQTCVLTVMTMRTDWDQQVSSSLKR 475
Query: 450 VNRWSKVES 458
+NRW + ES
Sbjct: 476 LNRWVEPES 484
Score = 100 bits (250), Expect = 7e-20
Identities = 47/88 (53%), Positives = 66/88 (74%), Gaps = 1/88 (1%)
Query: 2 ERDLKQNLLLKKPEEENEQEE-LSLGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAF 60
E ++ + LL K E E + L + ++VW E+K +WVVA PAIFTRFST G+ +ISQAF
Sbjct: 5 EGEVTETLLKKSTENRGEDRDGLGMKEKVWRESKKLWVVAGPAIFTRFSTSGLSLISQAF 64
Query: 61 VGHIGSRELAAFALVFTVLIRFANGILM 88
+GH+GS ELAA+++ TVL+RF+NGIL+
Sbjct: 65 IGHLGSTELAAYSITLTVLLRFSNGILL 92
>UniRef100_Q9LP32 T9L6.1 protein [Arabidopsis thaliana]
Length = 424
Score = 235 bits (599), Expect = 2e-60
Identities = 109/174 (62%), Positives = 143/174 (81%)
Query: 270 NVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFL 329
+ ++ICLNING EMMI+LGF+AAASVRVSNELG G+ K AKF+ + V TSL++G LF
Sbjct: 240 DALAICLNINGLEMMIALGFLAAASVRVSNELGSGNPKGAKFATLTAVFTSLSLGIVLFF 299
Query: 330 FFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVN 389
FLF R R++YIFT+++ VAA V +LSPLL+ SIL+NSVQPVLSGVA+GAGWQ V YVN
Sbjct: 300 VFLFLRGRVSYIFTTSEAVAAEVADLSPLLAFSILMNSVQPVLSGVAVGAGWQGYVTYVN 359
Query: 390 IGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQV 443
+ CYY++GIP+GI+LG ++ QVKG+W+GMLFG +QT VL ++T +T+WD+QV
Sbjct: 360 LACYYLVGIPIGIILGYVVGLQVKGVWIGMLFGIFVQTCVLTVMTLRTDWDQQV 413
Score = 100 bits (249), Expect = 9e-20
Identities = 47/79 (59%), Positives = 64/79 (80%), Gaps = 2/79 (2%)
Query: 10 LLKKPEEEN--EQEELSLGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSR 67
LLKK E E++EL L ++VW E+K +W+VAAPAIFTRFSTFG+ IISQ+F+GH+G
Sbjct: 12 LLKKTAENGGEEKDELGLKQKVWIESKKLWIVAAPAIFTRFSTFGVSIISQSFIGHLGPI 71
Query: 68 ELAAFALVFTVLIRFANGI 86
ELAA+++ FTVL+RF+N +
Sbjct: 72 ELAAYSITFTVLLRFSNAL 90
>UniRef100_Q75LU0 Putative MATE family protein [Oryza sativa]
Length = 401
Score = 211 bits (536), Expect = 5e-53
Identities = 95/182 (52%), Positives = 133/182 (72%)
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
+++C+ + GW MMIS+GF AAASVRV NELG G +AA FS+VV S I + + F
Sbjct: 206 LTVCMTLAGWVMMISIGFNAAASVRVGNELGAGHPRAAAFSVVVVTAVSFVITVVMAVVF 265
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
L FR+ ++YIFT + VA AV +L P L+ +++LN +QPVLSGVA+G GWQ VAY+N+G
Sbjct: 266 LMFRDYISYIFTEGETVARAVSDLCPFLAATLILNGIQPVLSGVAVGCGWQKIVAYINVG 325
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVN 451
CYY +GIP+G +LG H KGIW GML GT +QT++L IT++T+W+++V A+KR+N
Sbjct: 326 CYYFVGIPLGFLLGFKFHLGAKGIWTGMLGGTCMQTLILFWITFRTDWNKEVEEAKKRLN 385
Query: 452 RW 453
+W
Sbjct: 386 QW 387
>UniRef100_Q6ZB84 Putative ripening regulated protein [Oryza sativa]
Length = 489
Score = 210 bits (535), Expect = 6e-53
Identities = 97/185 (52%), Positives = 138/185 (74%)
Query: 270 NVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFL 329
+ +SICL INGWEMMI GF+AA VRV+NELG GS K A+F+IVV+V TS+AIG +
Sbjct: 302 DALSICLTINGWEMMIPFGFLAATGVRVANELGAGSGKGARFAIVVSVTTSVAIGLVFWC 361
Query: 330 FFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVN 389
+ + +++A +F+S+K V AV +LS LL+ ++LLNSVQPVLSGVAIG+GWQ+ VAYVN
Sbjct: 362 LIIAYNDKIALLFSSSKVVLDAVSDLSVLLAFTVLLNSVQPVLSGVAIGSGWQALVAYVN 421
Query: 390 IGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKR 449
+G YY++G+P+G +LG +H+ V GIW G++ GT +QT++L +T +WDE+ A R
Sbjct: 422 VGSYYLVGVPIGAILGWPLHFGVGGIWSGLIGGTAVQTLILAYLTISCDWDEEAKKASTR 481
Query: 450 VNRWS 454
+ W+
Sbjct: 482 MEVWA 486
Score = 66.6 bits (161), Expect = 1e-09
Identities = 34/77 (44%), Positives = 48/77 (62%), Gaps = 4/77 (5%)
Query: 16 EENEQEELSLGKRV----WNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAA 71
E +Q+ G RV W E+K +W V PAIF R + +GI ++SQAF+GH+G ELAA
Sbjct: 22 EAVKQQLWPAGARVAGEWWVESKKLWRVVGPAIFQRIALYGINVVSQAFIGHMGDLELAA 81
Query: 72 FALVFTVLIRFANGILM 88
F++ TV+ F G L+
Sbjct: 82 FSIASTVVAGFNFGFLL 98
>UniRef100_Q6V7U8 Putative anthocyanin permease [Lycopersicon esculentum]
Length = 506
Score = 209 bits (533), Expect = 1e-52
Identities = 94/192 (48%), Positives = 140/192 (71%)
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
+SIC+NINGWE M+ +G AA S+RVSNELG+G +A K+S+ +TV SL IG +
Sbjct: 302 ISICMNINGWESMLFIGINAAISIRVSNELGQGHPRATKYSVYITVFQSLLIGILCMVIV 361
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
L R+ LA IF+++KE+ AV +L+ LL I+++LNSVQPV+SGVA+G GWQ+ VAY+N+G
Sbjct: 362 LVARDHLAIIFSNSKEMQEAVADLAYLLGITMVLNSVQPVISGVAVGGGWQALVAYINLG 421
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVN 451
CYY+ G+P+G +LG + KG+W+GM+ G +QT++LLII YKTNW+++V +R+
Sbjct: 422 CYYVFGLPLGYLLGYVAKLGTKGLWLGMIAGAALQTLLLLIILYKTNWNKEVNDTTERMR 481
Query: 452 RWSKVESTDQET 463
+W + Q++
Sbjct: 482 KWGGQDFETQKS 493
Score = 45.4 bits (106), Expect = 0.003
Identities = 19/57 (33%), Positives = 33/57 (57%)
Query: 32 ETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVLIRFANGILM 88
ET +W + P F +G+ ++ FVGH+G+ EL+A ++ TV+ F+ G +M
Sbjct: 41 ETLKLWRIGGPIAFNIICQYGVNSLTNIFVGHLGNVELSAISIAQTVISTFSFGFMM 97
>UniRef100_Q75L61 Putative MATE efflux family protein [Oryza sativa]
Length = 500
Score = 208 bits (529), Expect = 3e-52
Identities = 96/184 (52%), Positives = 137/184 (74%)
Query: 270 NVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFL 329
+ +S+C+ I+GW MIS+GF AAASVRVSNELG G+ KAA FS+ V ++ I + L +
Sbjct: 301 DALSVCMTISGWVFMISVGFNAAASVRVSNELGAGNPKAAYFSVWVVTISCAIISAILAV 360
Query: 330 FFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVN 389
L R ++Y+FT + V+ AV +L PLL+I+++LN +QPVLSGVA+G GWQ VAYVN
Sbjct: 361 VILCLRNYISYLFTEGEVVSNAVADLCPLLAITLILNGIQPVLSGVAVGCGWQQFVAYVN 420
Query: 390 IGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKR 449
IGCYYI+G+P+G++LG + VKGIW GML GT +QT +L+ +T +T+W+ +V A+KR
Sbjct: 421 IGCYYIVGVPLGVLLGFVFKLGVKGIWGGMLGGTCMQTAILVWVTLRTDWNNEVEEAQKR 480
Query: 450 VNRW 453
+N+W
Sbjct: 481 LNKW 484
Score = 38.1 bits (87), Expect = 0.55
Identities = 22/70 (31%), Positives = 38/70 (53%), Gaps = 4/70 (5%)
Query: 23 LSLGKRVWNETK----LMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTV 78
+ L +R W T L+ +AAPA+ + + + +Q F GH+G+ ELAA +L
Sbjct: 27 MPLARRAWAATTIELGLLTRIAAPAVVMYMINYLMSMSTQIFSGHLGNLELAAASLGNNG 86
Query: 79 LIRFANGILM 88
+ FA G+++
Sbjct: 87 IQMFAYGLML 96
>UniRef100_Q940N9 AT4g21910/T8O5_120 [Arabidopsis thaliana]
Length = 507
Score = 206 bits (525), Expect = 9e-52
Identities = 96/182 (52%), Positives = 134/182 (72%)
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
+SIC++I+ M+S+GF AA SVR SNELG G+ K+A FS S I L
Sbjct: 317 LSICMSISALSFMVSVGFNAAVSVRTSNELGAGNPKSAWFSTWTATFVSFVISVTEALAV 376
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
++FR+ ++YIFT + +VA AV +L P L+I+I+LN +QPVLSGVA+G GWQ+ VAYVN+G
Sbjct: 377 IWFRDYVSYIFTEDADVAKAVSDLCPFLAITIILNGIQPVLSGVAVGCGWQTYVAYVNVG 436
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVN 451
CYY++GIPVG +LG +Q KGIW GM+ GTL+QT++LL +TY+T+WD++V ARKR++
Sbjct: 437 CYYVVGIPVGCILGFTFDFQAKGIWTGMIGGTLMQTLILLYVTYRTDWDKEVEKARKRLD 496
Query: 452 RW 453
W
Sbjct: 497 LW 498
Score = 37.7 bits (86), Expect = 0.71
Identities = 24/78 (30%), Positives = 42/78 (53%), Gaps = 11/78 (14%)
Query: 20 QEELSLGKRVWN----ETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAA---- 71
+ LS +RV+ E K+++ +A PAI G+ I ++ F GH+GS+ELAA
Sbjct: 39 ESSLSYRRRVYLGACIELKVLFRLALPAILIYLVNSGMGISARVFAGHVGSQELAAASIG 98
Query: 72 ---FALVFTVLIRFANGI 86
F LV+ +++ + +
Sbjct: 99 NSCFNLVYGLMLGMGSAV 116
>UniRef100_Q8H7P0 Putative ripening regulated protein [Oryza sativa]
Length = 369
Score = 206 bits (525), Expect = 9e-52
Identities = 98/187 (52%), Positives = 137/187 (72%)
Query: 270 NVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFL 329
+ +SIC+ IN WE+MI L F A VRV+NELG G+ K A+F+ +V+ +TSL IG F ++
Sbjct: 182 DALSICMTINAWELMIPLAFFAGTGVRVANELGAGNGKGARFATIVSSVTSLVIGLFFWV 241
Query: 330 FFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVN 389
+ ++ A IFTS+ V AV LS LL+ +ILLNS+QPVLSGVA+G+GWQS VAYVN
Sbjct: 242 LIVGLHDKFALIFTSSDVVLDAVDNLSVLLAFTILLNSIQPVLSGVAVGSGWQSMVAYVN 301
Query: 390 IGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKR 449
IG YY+IGIP+GI+LG + V GIW GM+ GT +QT++L IIT + +WD++ +A R
Sbjct: 302 IGTYYLIGIPMGILLGWLFKLGVLGIWAGMIGGTAVQTLILAIITIRCDWDKEAMIASTR 361
Query: 450 VNRWSKV 456
+++WS+V
Sbjct: 362 MDKWSQV 368
>UniRef100_Q8LDZ2 Hypothetical protein [Arabidopsis thaliana]
Length = 507
Score = 206 bits (525), Expect = 9e-52
Identities = 96/182 (52%), Positives = 134/182 (72%)
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
+SIC++I+ M+S+GF AA SVR SNELG G+ K+A FS S I L
Sbjct: 317 LSICMSISALSFMVSVGFNAAVSVRTSNELGAGNPKSAWFSTWTATFVSFVISVTEALAV 376
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
++FR+ ++YIFT + +VA AV +L P L+I+I+LN +QPVLSGVA+G GWQ+ VAYVN+G
Sbjct: 377 IWFRDYVSYIFTEDADVAKAVSDLCPFLAITIILNGIQPVLSGVAVGCGWQTYVAYVNVG 436
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVN 451
CYY++GIPVG +LG +Q KGIW GM+ GTL+QT++LL +TY+T+WD++V ARKR++
Sbjct: 437 CYYVVGIPVGCILGFTFDFQAKGIWTGMIGGTLMQTLILLYVTYRTDWDKEVEKARKRLD 496
Query: 452 RW 453
W
Sbjct: 497 LW 498
Score = 37.7 bits (86), Expect = 0.71
Identities = 24/78 (30%), Positives = 42/78 (53%), Gaps = 11/78 (14%)
Query: 20 QEELSLGKRVWN----ETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAA---- 71
+ LS +RV+ E K+++ +A PAI G+ I ++ F GH+GS+ELAA
Sbjct: 39 ESSLSYRRRVYLGAGIELKVLFRLALPAILIYLVNSGMGISARVFAGHVGSQELAAASIG 98
Query: 72 ---FALVFTVLIRFANGI 86
F LV+ +++ + +
Sbjct: 99 NSCFNLVYGLMLGMGSAV 116
>UniRef100_Q9SAB0 F25C20.18 protein [Arabidopsis thaliana]
Length = 503
Score = 206 bits (523), Expect = 2e-51
Identities = 98/182 (53%), Positives = 136/182 (73%)
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
++IC++I+ M+S+GF AAASVRVSNELG G+ ++A FS VT S + F +
Sbjct: 313 LAICMSISAMSFMVSVGFNAAASVRVSNELGAGNPRSAAFSTAVTTGVSFLLSLFEAIVI 372
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
L +R ++YIFT + VA AV ELSP L+I+I+LN VQPVLSGVA+G GWQ+ VAYVNIG
Sbjct: 373 LSWRHVISYIFTDSPAVAEAVAELSPFLAITIVLNGVQPVLSGVAVGCGWQAYVAYVNIG 432
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVN 451
CYYI+GIP+G VLG +GIW GM+ GTL+QTI+L+I+T++T+WD++V A +R++
Sbjct: 433 CYYIVGIPIGYVLGFTYDMGARGIWTGMIGGTLMQTIILVIVTFRTDWDKEVEKASRRLD 492
Query: 452 RW 453
+W
Sbjct: 493 QW 494
Score = 37.0 bits (84), Expect = 1.2
Identities = 19/57 (33%), Positives = 34/57 (59%)
Query: 32 ETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVLIRFANGILM 88
E K ++ +AAPAIF G+ ++++ F G +GS +LAA +L + F G+++
Sbjct: 50 EMKYLFHLAAPAIFVYVINNGMSMLTRIFAGRLGSMQLAAASLGNSGFNMFTLGLML 106
>UniRef100_Q9FNC1 Emb|CAB89401.1 [Arabidopsis thaliana]
Length = 491
Score = 206 bits (523), Expect = 2e-51
Identities = 95/184 (51%), Positives = 141/184 (76%), Gaps = 1/184 (0%)
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
+SIC++ING EMM+ L F A SVRV+NELG G+ K A+F+++++V SL IG + +
Sbjct: 302 MSICMSINGLEMMVPLAFFAGTSVRVANELGAGNGKRARFAMIISVTQSLIIGIIISVLI 361
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
F +++ ++F+S++ V AV LS LLS +ILLNSVQPVLSGVA+G+GWQS VA++N+G
Sbjct: 362 YFLLDQIGWMFSSSETVLKAVNNLSILLSFAILLNSVQPVLSGVAVGSGWQSLVAFINLG 421
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLF-GTLIQTIVLLIITYKTNWDEQVTVARKRV 450
CYY IG+P+GIV+G + + VKGIW GM+F GT++QT++L+ IT + +W+++ A+ RV
Sbjct: 422 CYYFIGLPLGIVMGWMFKFGVKGIWAGMIFGGTMVQTLILIFITMRCDWEKEAQNAKVRV 481
Query: 451 NRWS 454
N+WS
Sbjct: 482 NKWS 485
Score = 68.2 bits (165), Expect = 5e-10
Identities = 34/77 (44%), Positives = 48/77 (62%)
Query: 6 KQNLLLKKPEEENEQEELSLGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIG 65
K + L K + E+E + K +W ETK +W + PAIFTR +T I +I+QAF GH+G
Sbjct: 14 KAKIPLLKDQNVAEEENGEIKKEIWLETKKLWRIVGPAIFTRVTTNLIFVITQAFAGHLG 73
Query: 66 SRELAAFALVFTVLIRF 82
ELAA ++V V+I F
Sbjct: 74 ELELAAISIVNNVIIGF 90
>UniRef100_Q93Z03 At1g47530/F16N3_20 [Arabidopsis thaliana]
Length = 484
Score = 204 bits (518), Expect = 6e-51
Identities = 98/184 (53%), Positives = 131/184 (70%)
Query: 270 NVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFL 329
+ +SIC+NI GW MIS+GF AA SVRVSNELG G+A AKFS++V +TS IG +
Sbjct: 295 DAISICMNIEGWTAMISIGFNAAISVRVSNELGAGNAALAKFSVIVVSITSTLIGIVCMI 354
Query: 330 FFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVN 389
L ++ Y+FTS++ VAA ++ LL ++LLNS+QPVLSGVA+GAGWQ+ VAYVN
Sbjct: 355 VVLATKDSFPYLFTSSEAVAAETTRIAVLLGFTVLLNSLQPVLSGVAVGAGWQALVAYVN 414
Query: 390 IGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKR 449
I CYYIIG+P G+VLG + V+GIW GM+ G +QT++L+ I Y TNW+++ A R
Sbjct: 415 IACYYIIGLPAGLVLGFTLDLGVQGIWGGMVAGICLQTLILIGIIYFTNWNKEAEQAESR 474
Query: 450 VNRW 453
V RW
Sbjct: 475 VQRW 478
Score = 48.9 bits (115), Expect = 3e-04
Identities = 27/87 (31%), Positives = 44/87 (50%), Gaps = 5/87 (5%)
Query: 7 QNLLLKKPEEENEQEELSLGKRVW-----NETKLMWVVAAPAIFTRFSTFGIQIISQAFV 61
+ L L P E E +VW E+K +W +A PAIFT + + ++Q F
Sbjct: 5 KTLPLLDPREPPELTGTKSASKVWAKEFGEESKRLWELAGPAIFTAIGQYSLGALTQTFS 64
Query: 62 GHIGSRELAAFALVFTVLIRFANGILM 88
G +G ELAA ++ +V+ A G+++
Sbjct: 65 GRLGELELAAVSVENSVISGLAFGVML 91
>UniRef100_Q9SX83 F16N3.20 [Arabidopsis thaliana]
Length = 484
Score = 204 bits (518), Expect = 6e-51
Identities = 98/184 (53%), Positives = 131/184 (70%)
Query: 270 NVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFL 329
+ +SIC+NI GW MIS+GF AA SVRVSNELG G+A AKFS++V +TS IG +
Sbjct: 295 DAISICMNIEGWTAMISIGFNAAISVRVSNELGAGNAALAKFSVIVVSITSTLIGIVCMI 354
Query: 330 FFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVN 389
L ++ Y+FTS++ VAA ++ LL ++LLNS+QPVLSGVA+GAGWQ+ VAYVN
Sbjct: 355 VVLATKDSFPYLFTSSEAVAAETTRIAVLLGFTVLLNSLQPVLSGVAVGAGWQALVAYVN 414
Query: 390 IGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKR 449
I CYYIIG+P G+VLG + V+GIW GM+ G +QT++L+ I Y TNW+++ A R
Sbjct: 415 IACYYIIGLPAGLVLGFTLDLGVQGIWGGMVAGICLQTLILIGIIYFTNWNKEAEQAESR 474
Query: 450 VNRW 453
V RW
Sbjct: 475 VQRW 478
Score = 50.4 bits (119), Expect = 1e-04
Identities = 28/87 (32%), Positives = 45/87 (51%), Gaps = 5/87 (5%)
Query: 7 QNLLLKKPEEENEQEELSLGKRVW-----NETKLMWVVAAPAIFTRFSTFGIQIISQAFV 61
+ L L P E E +VW E+K +W +A PAIFT S + + ++Q F
Sbjct: 5 KTLPLLDPREPPELTGTKSASKVWAKEFGEESKRLWELAGPAIFTAISQYSLGALTQTFS 64
Query: 62 GHIGSRELAAFALVFTVLIRFANGILM 88
G +G ELAA ++ +V+ A G+++
Sbjct: 65 GRLGELELAAVSVENSVISGLAFGVML 91
>UniRef100_Q94AC7 At1g61890/F8K4_9 [Arabidopsis thaliana]
Length = 501
Score = 204 bits (518), Expect = 6e-51
Identities = 99/184 (53%), Positives = 135/184 (72%)
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
++IC++I+ M+S+GF AAASVRVSNELG G+ +AA FS VVT S + F +
Sbjct: 310 LAICMSISAISFMVSVGFNAAASVRVSNELGAGNPRAAAFSTVVTTGVSFLLSVFEAIVV 369
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
L +R ++Y FT + VA AV +LSP L+I+I+LN +QPVLSGVA+G GWQ+ VAYVNIG
Sbjct: 370 LSWRHVISYAFTDSPAVAEAVADLSPFLAITIVLNGIQPVLSGVAVGCGWQAFVAYVNIG 429
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVN 451
CYY++GIPVG VLG KGIW GM+ GTL+QTIVL+I+T +T+WD++V A R++
Sbjct: 430 CYYVVGIPVGFVLGFTYDMGAKGIWTGMIGGTLMQTIVLVIVTLRTDWDKEVEKASSRLD 489
Query: 452 RWSK 455
+W +
Sbjct: 490 QWEE 493
Score = 43.5 bits (101), Expect = 0.013
Identities = 23/57 (40%), Positives = 35/57 (61%)
Query: 32 ETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVLIRFANGILM 88
E K ++ +AAPAIF G+ I+++ F GH+GS ELAA +L + F G+L+
Sbjct: 47 EMKFLFHLAAPAIFVYVINNGMSILTRIFAGHVGSFELAAASLGNSGFNMFTYGLLL 103
>UniRef100_O80695 Hypothetical protein F8K4.9 [Arabidopsis thaliana]
Length = 501
Score = 203 bits (517), Expect = 8e-51
Identities = 98/184 (53%), Positives = 135/184 (73%)
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
++IC++I+ M+S+GF AAASVRVSNELG G+ +AA FS VVT S + F +
Sbjct: 310 LAICMSISAISFMVSVGFNAAASVRVSNELGAGNPRAAAFSTVVTTGVSFLLSVFEAIVV 369
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
L +R ++Y FT + VA AV +LSP L+I+I+LN +QPVLSGVA+G GWQ+ VAYVNIG
Sbjct: 370 LSWRHVISYAFTDSPAVAEAVADLSPFLAITIVLNGIQPVLSGVAVGCGWQAFVAYVNIG 429
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVN 451
CYY++GIPVG VLG KGIW GM+ GTL+QTI+L+I+T +T+WD++V A R++
Sbjct: 430 CYYVVGIPVGFVLGFTYDMGAKGIWTGMIGGTLMQTIILVIVTLRTDWDKEVEKASSRLD 489
Query: 452 RWSK 455
+W +
Sbjct: 490 QWEE 493
Score = 43.5 bits (101), Expect = 0.013
Identities = 23/57 (40%), Positives = 35/57 (61%)
Query: 32 ETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVLIRFANGILM 88
E K ++ +AAPAIF G+ I+++ F GH+GS ELAA +L + F G+L+
Sbjct: 47 EMKFLFHLAAPAIFVYVINNGMSILTRIFAGHVGSFELAAASLGNSGFNMFTYGLLL 103
>UniRef100_Q8L714 Hypothetical protein At1g11670 [Arabidopsis thaliana]
Length = 503
Score = 203 bits (516), Expect = 1e-50
Identities = 97/182 (53%), Positives = 135/182 (73%)
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
++IC++I+ M+S+GF AAASVRVSNELG G+ ++A FS VT S + F +
Sbjct: 313 LAICMSISAMSFMVSVGFNAAASVRVSNELGAGNPRSAAFSTAVTTGVSFLLSLFEAIVI 372
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
L +R ++YIFT + VA AV ELSP L+I+I+LN VQPVLSGVA+G GWQ+ VAYVNIG
Sbjct: 373 LSWRHVISYIFTDSPAVAEAVAELSPFLAITIVLNGVQPVLSGVAVGCGWQAYVAYVNIG 432
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVN 451
CYYI+GIP+G VLG +GIW GM+ TL+QTI+L+I+T++T+WD++V A +R++
Sbjct: 433 CYYIVGIPIGYVLGFTYDMGARGIWTGMIGDTLMQTIILVIVTFRTDWDKEVEKASRRLD 492
Query: 452 RW 453
+W
Sbjct: 493 QW 494
Score = 37.0 bits (84), Expect = 1.2
Identities = 19/57 (33%), Positives = 34/57 (59%)
Query: 32 ETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVLIRFANGILM 88
E K ++ +AAPAIF G+ ++++ F G +GS +LAA +L + F G+++
Sbjct: 50 EMKYLFHLAAPAIFVYVINNGMSMLTRIFAGRLGSMQLAAASLGNSGFNMFTLGLML 106
>UniRef100_Q38956 Hypothetical protein [Arabidopsis thaliana]
Length = 500
Score = 202 bits (514), Expect = 2e-50
Identities = 94/182 (51%), Positives = 132/182 (71%)
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
+SIC+NI GW MI++G A SVRVSNELG + AKFS++V V+TS IG + +
Sbjct: 307 LSICMNILGWTAMIAIGMNTAVSVRVSNELGANHPRTAKFSLLVAVITSTLIGFIVSMIL 366
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
L FR++ +F +++V V EL+P+L++SI++N+VQPVLSGVA+GAGWQ+ VAYVNI
Sbjct: 367 LIFRDQYPSLFVKDEKVIILVKELTPILALSIVINNVQPVLSGVAVGAGWQAVVAYVNIA 426
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVN 451
CYY+ GIP G++LG +++ V GIW GML GT++QTIVL + KTNWD + ++A R+
Sbjct: 427 CYYVFGIPFGLLLGYKLNYGVMGIWCGMLTGTVVQTIVLTWMICKTNWDTEASMAEDRIR 486
Query: 452 RW 453
W
Sbjct: 487 EW 488
Score = 49.3 bits (116), Expect = 2e-04
Identities = 23/57 (40%), Positives = 37/57 (64%)
Query: 32 ETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVLIRFANGILM 88
ETK +W +A PAIFT + + + I+Q F GHI + LAA ++ +V+ F+ GI++
Sbjct: 45 ETKKLWYLAGPAIFTSVNQYSLGAITQVFAGHISTIALAAVSVENSVVAGFSFGIML 101
>UniRef100_Q9LVD9 Gb|AAC28507.1 [Arabidopsis thaliana]
Length = 506
Score = 202 bits (514), Expect = 2e-50
Identities = 95/183 (51%), Positives = 135/183 (72%)
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
+SIC+ I+GW MIS+GF AA SVRVSNELG G+ K+A FS+++ + SL L +
Sbjct: 315 LSICMTISGWVFMISVGFNAAISVRVSNELGAGNPKSAAFSVIIVNIYSLITCVILAIVI 374
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
L R+ L+Y FT KEV+ AV +L PLL+++++LN +QPVLSGVA+G GWQ+ VA VN+G
Sbjct: 375 LACRDVLSYAFTEGKEVSDAVSDLCPLLAVTLVLNGIQPVLSGVAVGCGWQTFVAKVNVG 434
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVN 451
CYYIIGIP+G + G ++ KGIW GM+ GT+IQT +L +T++T+W ++V A KR++
Sbjct: 435 CYYIIGIPLGALFGFYFNFGAKGIWTGMIGGTVIQTFILAWVTFRTDWTKEVEEASKRLD 494
Query: 452 RWS 454
+WS
Sbjct: 495 KWS 497
Score = 42.4 bits (98), Expect = 0.029
Identities = 24/66 (36%), Positives = 40/66 (60%)
Query: 23 LSLGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVLIRF 82
L L K E+KL++ +AAPA+ + + + +Q F GH+G+ ELAA +L T + F
Sbjct: 43 LRLRKATIIESKLLFNLAAPAVIVYMINYLMSMSTQIFSGHLGNLELAAASLGNTGIQVF 102
Query: 83 ANGILM 88
A G+++
Sbjct: 103 AYGLML 108
>UniRef100_Q6K5E6 Putative ripening regulated protein DDTFR18 [Oryza sativa]
Length = 482
Score = 202 bits (514), Expect = 2e-50
Identities = 96/184 (52%), Positives = 132/184 (71%)
Query: 270 NVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFL 329
N +SIC++ WEMMI +GF+A VRV+NELG G+ K AKF+ +V+ TS IG F
Sbjct: 295 NALSICMSFQSWEMMIPVGFLAGTGVRVANELGAGNGKGAKFATIVSTTTSFLIGLFFSA 354
Query: 330 FFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVN 389
L F +++A +F+S+ V AV +S LL+++ILLN VQPVLSGVAIG+GWQ+ VAYVN
Sbjct: 355 LALAFHDKIALVFSSSNAVIDAVDNISFLLAVTILLNGVQPVLSGVAIGSGWQAAVAYVN 414
Query: 390 IGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKR 449
IGCYY IG+P+G++LG + V GIW GM+ GT IQTI+L +T + +W+++V A +R
Sbjct: 415 IGCYYFIGVPIGVLLGWSFNLGVFGIWAGMIAGTAIQTIILAHMTIQCDWNKEVLQASER 474
Query: 450 VNRW 453
V RW
Sbjct: 475 VQRW 478
Score = 62.8 bits (151), Expect = 2e-08
Identities = 31/78 (39%), Positives = 44/78 (55%), Gaps = 1/78 (1%)
Query: 12 KKPEEENEQEEL-SLGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELA 70
K+ EE + L +L + W E+K +W + PA+F R + IISQAF GHIG ELA
Sbjct: 14 KEKEEGRVRRRLPALAREAWEESKKLWEIVGPAVFLRLVLYSFNIISQAFAGHIGDLELA 73
Query: 71 AFALVFTVLIRFANGILM 88
AF++ V+ G L+
Sbjct: 74 AFSIANNVITGLNFGFLL 91
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.329 0.141 0.463
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 773,851,073
Number of Sequences: 2790947
Number of extensions: 32111034
Number of successful extensions: 106969
Number of sequences better than 10.0: 493
Number of HSP's better than 10.0 without gapping: 309
Number of HSP's successfully gapped in prelim test: 184
Number of HSP's that attempted gapping in prelim test: 106231
Number of HSP's gapped (non-prelim): 792
length of query: 469
length of database: 848,049,833
effective HSP length: 131
effective length of query: 338
effective length of database: 482,435,776
effective search space: 163063292288
effective search space used: 163063292288
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 77 (34.3 bits)
Medicago: description of AC145330.19


