

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145329.13 - phase: 2 /pseudo
(528 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
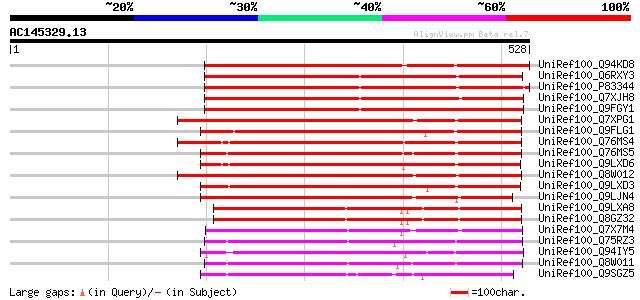
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q94KD8 At1g02640/T14P4_11 [Arabidopsis thaliana] 451 e-125
UniRef100_Q6RXY3 Beta xylosidase [Fragaria ananassa] 442 e-123
UniRef100_P83344 Putative beta-D-xylosidase [Prunus persica] 430 e-119
UniRef100_Q7XJH8 Auxin-induced beta-glucosidase [Chenopodium rub... 418 e-115
UniRef100_Q9FGY1 Xylosidase [Arabidopsis thaliana] 402 e-110
UniRef100_Q7XPG1 OSJNBb0003B01.22 protein [Oryza sativa] 329 1e-88
UniRef100_Q9FLG1 Beta-xylosidase [Arabidopsis thaliana] 319 1e-85
UniRef100_Q76MS4 LEXYL2 protein [Lycopersicon esculentum] 317 5e-85
UniRef100_Q76MS5 LEXYL1 protein [Lycopersicon esculentum] 311 2e-83
UniRef100_Q9LXD6 Beta-xylosidase [Arabidopsis thaliana] 307 6e-82
UniRef100_Q8W012 Alpha-L-arabinofuranosidase/beta-D-xylosidase i... 302 2e-80
UniRef100_Q9LXD3 Similarity to beta-glucosidase [Arabidopsis tha... 296 8e-79
UniRef100_Q9LJN4 Beta-1,4-xylosidase [Arabidopsis thaliana] 275 2e-72
UniRef100_Q9LXA8 Beta-xylosidase-like protein [Arabidopsis thali... 274 5e-72
UniRef100_Q8GZ32 Putative beta-xylosidase [Arabidopsis thaliana] 274 5e-72
UniRef100_Q7X7M4 OSJNBa0074L08.23 protein [Oryza sativa] 261 5e-68
UniRef100_Q75RZ3 Putative beta-xylosidase [Triticum aestivum] 245 2e-63
UniRef100_Q94IY5 Putative beta-xylosidase [Oryza sativa] 245 2e-63
UniRef100_Q8W011 Beta-D-xylosidase [Hordeum vulgare] 245 2e-63
UniRef100_Q9SGZ5 F28K19.27 [Arabidopsis thaliana] 239 1e-61
>UniRef100_Q94KD8 At1g02640/T14P4_11 [Arabidopsis thaliana]
Length = 768
Score = 451 bits (1159), Expect = e-125
Identities = 219/331 (66%), Positives = 264/331 (79%), Gaps = 7/331 (2%)
Query: 199 PLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETVDRTS 258
P+QGI YA+TIHQ+GC +V C DD+ F A++AAR ADAT+LV+GLDQSIEAE DR S
Sbjct: 443 PVQGITGYARTIHQKGCVDVHCMDDRLFDAAVEAARGADATVLVMGLDQSIEAEFKDRNS 502
Query: 259 LLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAI 318
LLLPG QQ+LVS+VA A+KGP ILVLMSGGP+DI+FA+ D K+ I+WAGYPGQ GG AI
Sbjct: 503 LLLPGKQQELVSRVAKAAKGPVILVLMSGGPIDISFAEKDRKIPAIVWAGYPGQEGGTAI 562
Query: 319 ADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRP-SKIGYPGRTYRFYKGPVVYPFGH 377
ADILFG+A+PGGKLP+TWYPQ+YL NL MT M+MRP PGRTYRFY GPVVYPFGH
Sbjct: 563 ADILFGSANPGGKLPMTWYPQDYLTNLPMTEMSMRPVHSKRIPGRTYRFYDGPVVYPFGH 622
Query: 378 GLTYTHFVHELSSAPTVVSVPVHGHRHGNNTNISNKAIRVTHARCGKLSIALHVDVKNVG 437
GL+YT F H ++ AP V+ + V G N +S K+IRVTHARC +LS+ +HV+V NVG
Sbjct: 623 GLSYTRFTHNIADAPKVIPIAV----RGRNGTVSGKSIRVTHARCDRLSLGVHVEVTNVG 678
Query: 438 SRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVVDKSG 497
SRDGTHT+LVFSAPP G W P+K LVAFE+VHV K+RV+VNIHVCK LSVVD++G
Sbjct: 679 SRDGTHTMLVFSAPP--GGEWAPKKQLVAFERVHVAVGEKKRVQVNIHVCKYLSVVDRAG 736
Query: 498 IRRIPMGEHSLHIGDVKHSVSLQAEALGIIK 528
RRIP+G+H +HIGD H+VSLQA LG+IK
Sbjct: 737 NRRIPIGDHGIHIGDESHTVSLQASTLGVIK 767
>UniRef100_Q6RXY3 Beta xylosidase [Fragaria ananassa]
Length = 491
Score = 442 bits (1138), Expect = e-123
Identities = 209/323 (64%), Positives = 259/323 (79%), Gaps = 3/323 (0%)
Query: 199 PLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETVDRTS 258
PLQGIGRY KTIHQQGC+NVAC ++ FG A AAR ADAT+LV+GLDQSIEAE DRT
Sbjct: 165 PLQGIGRYTKTIHQQGCTNVACTTNQLFGAAEAAARQADATVLVMGLDQSIEAEFRDRTD 224
Query: 259 LLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAI 318
L++PGHQQ+LVS+VA AS+GPT+LVLMSGGP+D++FAKNDPK+ I+W GYPGQAGG A+
Sbjct: 225 LVMPGHQQELVSRVARASRGPTVLVLMSGGPIDVSFAKNDPKIGAIIWVGYPGQAGGTAM 284
Query: 319 ADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPSKIGYPGRTYRFYKGPVVYPFGHG 378
AD+LFGT +P GKLP+TWYPQ+Y+ + MTNMAMR + GYPGRTYRFYKGPVV+PFG G
Sbjct: 285 ADVLFGTTNPSGKLPMTWYPQDYVSKVPMTNMAMRAGR-GYPGRTYRFYKGPVVFPFGLG 343
Query: 379 LTYTHFVHELSSAPTVVSVPVHGHRHGNNTNISNKAIRVTHARCGKLSIALHVDVKNVGS 438
L+YT F H L+ PT VSVP+ N+ + + A+RV+H C LS+ALHV VKN G+
Sbjct: 344 LSYTTFAHSLAQVPTSVSVPLTSLSATTNSTMLSSAVRVSHTNCNPLSLALHVVVKNTGA 403
Query: 439 RDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVVDKSGI 498
RDGTHTLLVFS+PP+G W K LV F KVH+ A + +RV+V++HVCK LSVVD+ GI
Sbjct: 404 RDGTHTLLVFSSPPSG--KWAANKQLVGFHKVHIVAGSHKRVKVDVHVCKHLSVVDQFGI 461
Query: 499 RRIPMGEHSLHIGDVKHSVSLQA 521
RRIP+GEH L IGD++H +S++A
Sbjct: 462 RRIPIGEHKLQIGDLEHHISVEA 484
>UniRef100_P83344 Putative beta-D-xylosidase [Prunus persica]
Length = 461
Score = 430 bits (1106), Expect = e-119
Identities = 211/333 (63%), Positives = 263/333 (78%), Gaps = 7/333 (2%)
Query: 199 PLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETVDRTS 258
PLQGIGRY +TIHQ GC++V C ++ FG A AAR ADAT+LV+GLDQSIEAE VDR
Sbjct: 132 PLQGIGRYTRTIHQAGCTDVHCNGNQLFGAAEAAARQADATVLVMGLDQSIEAEFVDRAG 191
Query: 259 LLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAI 318
LLLPGHQQ+LVS+VA AS+GPTILVLMSGGP+D+TFAKNDP+++ I+W GYPGQAGG AI
Sbjct: 192 LLLPGHQQELVSRVARASRGPTILVLMSGGPIDVTFAKNDPRISAIIWVGYPGQAGGTAI 251
Query: 319 ADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMR--PSKIGYPGRTYRFYKGPVVYPFG 376
A++LFGTA+PGGKLP+TWYPQ Y+ +L MT+MAMR P++ GYPGRTYRFY GPVV+PFG
Sbjct: 252 ANVLFGTANPGGKLPMTWYPQNYVTHLPMTDMAMRADPAR-GYPGRTYRFYIGPVVFPFG 310
Query: 377 HGLTYTHFVHELSSAPTVVSVPVHGHRHGNNTNISNKAIRVTHARCGKLS-IALHVDVKN 435
GL+YT F H L+ PT+VSVP+ + N+ + +K +RV+H C LS + +HVDVKN
Sbjct: 311 LGLSYTTFAHNLAHGPTLVSVPLTSLKATANSTMLSKTVRVSHPDCNALSPLDVHVDVKN 370
Query: 436 VGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVVDK 495
GS DGTHTLLVF++PP+G W K L+ F K+H+ +++RVR+ +HVCK LSVVD+
Sbjct: 371 TGSMDGTHTLLVFTSPPDG--KWASSKQLMGFHKIHIATGSEKRVRIAVHVCKHLSVVDR 428
Query: 496 SGIRRIPMGEHSLHIGDVKHSVSLQAEALGIIK 528
GIRRIP+GEH L IGD+ H VSLQ LG IK
Sbjct: 429 FGIRRIPLGEHKLQIGDLSHHVSLQTN-LGEIK 460
>UniRef100_Q7XJH8 Auxin-induced beta-glucosidase [Chenopodium rubrum]
Length = 767
Score = 418 bits (1074), Expect = e-115
Identities = 207/328 (63%), Positives = 252/328 (76%), Gaps = 7/328 (2%)
Query: 199 PLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETVDRTS 258
PLQGI RYAKT+HQ GC VAC ++QFG A AA HADAT+LV+GLDQSIEAE DR S
Sbjct: 440 PLQGIARYAKTVHQAGCIGVACTSNQQFGAATAAAAHADATVLVMGLDQSIEAEFRDRAS 499
Query: 259 LLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAI 318
+LLPGHQQ+LVSKVA AS+GPTILVLM GGPVD+TFAKNDPK++ ILW GYPGQAGG AI
Sbjct: 500 VLLPGHQQELVSKVALASRGPTILVLMCGGPVDVTFAKNDPKISAILWVGYPGQAGGTAI 559
Query: 319 ADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMR--PSKIGYPGRTYRFYKGPVVYPFG 376
AD+LFGT +PGGKLP TWYPQ Y+ + MT++AMR PS GYPGRTYRFYKGPVV+PFG
Sbjct: 560 ADVLFGTTNPGGKLPNTWYPQSYVAKVPMTDLAMRANPSN-GYPGRTYRFYKGPVVFPFG 618
Query: 377 HGLTYTHFVHELSSAPTVVSVPV-HGHRHGNNTNISNKAIRVTHARCGKLSIALHVDVKN 435
GL+YT F L+ APT V VP+ + + N T+ + A++V H C + ++LH+DVKN
Sbjct: 619 FGLSYTRFTQSLAHAPTKVMVPLANQFTNSNITSFNKDALKVLHTNCDNIPLSLHIDVKN 678
Query: 436 VGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVVDK 495
G DG+HT+LVFS PP G +K L+ F++VHV A +KQRVR+NIHVC LS D+
Sbjct: 679 KGKVDGSHTILVFSTPPKGTKS--SEKQLIGFKRVHVFAGSKQRVRMNIHVCNHLSRADE 736
Query: 496 SGIRRIPMGEHSLHIG-DVKHSVSLQAE 522
G+RRIP+GEH+LHIG D KH +SL +
Sbjct: 737 FGVRRIPIGEHTLHIGDDHKHKLSLHID 764
>UniRef100_Q9FGY1 Xylosidase [Arabidopsis thaliana]
Length = 774
Score = 402 bits (1032), Expect = e-110
Identities = 202/328 (61%), Positives = 249/328 (75%), Gaps = 5/328 (1%)
Query: 199 PLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETVDRTS 258
PLQGI RYA+T+HQ GC+ VAC+ ++ FG A AAR ADAT+LV+GLDQSIEAET DRT
Sbjct: 447 PLQGISRYARTLHQAGCAGVACKGNQGFGAAEAAAREADATVLVMGLDQSIEAETRDRTG 506
Query: 259 LLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAI 318
LLLPG+QQDLV++VA AS+GP ILVLMSGGP+D+TFAKNDP+VA I+WAGYPGQAGGAAI
Sbjct: 507 LLLPGYQQDLVTRVAQASRGPVILVLMSGGPIDVTFAKNDPRVAAIIWAGYPGQAGGAAI 566
Query: 319 ADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPSKIGYPGRTYRFYKGPVVYPFGHG 378
A+I+FG A+PGGKLP+TWYPQ+Y+ + MT MAMR S YPGRTYRFYKGPVV+PFG G
Sbjct: 567 ANIIFGAANPGGKLPMTWYPQDYVAKVPMTVMAMRASG-NYPGRTYRFYKGPVVFPFGFG 625
Query: 379 LTYTHFVHELSSAPTV-VSVPVHGHRHGNN-TNISNKAIRVTHARCGKL-SIALHVDVKN 435
L+YT F H L+ +P +SV + N N S+ +I+V+H C + LHV+V N
Sbjct: 626 LSYTTFTHSLAKSPLAQLSVSLSNLNSANTILNSSSHSIKVSHTNCNSFPKMPLHVEVSN 685
Query: 436 VGSRDGTHTLLVFSAPP-NGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVVD 494
G DGTHT+ VF+ PP NG K L+AFEKVHV A KQ V+V++ CK L VVD
Sbjct: 686 TGEFDGTHTVFVFAEPPINGIKGLGVNKQLIAFEKVHVMAGAKQTVQVDVDACKHLGVVD 745
Query: 495 KSGIRRIPMGEHSLHIGDVKHSVSLQAE 522
+ G RRIPMGEH LHIGD+KH++ +Q +
Sbjct: 746 EYGKRRIPMGEHKLHIGDLKHTILVQPQ 773
>UniRef100_Q7XPG1 OSJNBb0003B01.22 protein [Oryza sativa]
Length = 765
Score = 329 bits (844), Expect = 1e-88
Identities = 172/353 (48%), Positives = 224/353 (62%), Gaps = 8/353 (2%)
Query: 171 SLPTTPPNCGCHWAQFRCHSYHDWKLCRPLQGIGRYAKTIHQQGCSNVACRDDK-QFGPA 229
S+ PN + + K PLQG+G T++Q GC+NV C + Q A
Sbjct: 417 SMAVIGPNANASFTMIGNYEGTPCKYTTPLQGLGANVATVYQPGCTNVGCSGNSLQLSAA 476
Query: 230 LDAARHADATILVIGLDQSIEAETVDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGP 289
AA AD T+LV+G DQS+E E++DRTSLLLPG Q LVS VA AS+GP ILV+MSGGP
Sbjct: 477 TQAAASADVTVLVVGADQSVERESLDRTSLLLPGQQPQLVSAVANASRGPVILVVMSGGP 536
Query: 290 VDITFAKNDPKVAGILWAGYPGQAGGAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTN 349
DI+FAK+ K++ ILW GYPG+AGGAA+ADILFG +PGG+LPVTWYP + ++MT+
Sbjct: 537 FDISFAKSSDKISAILWVGYPGEAGGAALADILFGYHNPGGRLPVTWYPASFADKVSMTD 596
Query: 350 MAMRP-SKIGYPGRTYRFYKGPVVYPFGHGLTYTHFVHELSSAPTVVSVPV-HGHRHGNN 407
M MRP S GYPGRTYRFY G VY FG GL+YT F H L SAP V+V + GH
Sbjct: 597 MRMRPDSSTGYPGRTYRFYTGDTVYAFGDGLSYTKFAHSLVSAPEQVAVQLAEGHACHTE 656
Query: 408 TNISNKAIRVTHARCGKLSIALHVDVKNVGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAF 467
S + CG LS +H+ V+N G G HT+ +FS+PP+ H P K L+ F
Sbjct: 657 HCFS---VEAAGEHCGSLSFDVHLRVRNAGGMAGGHTVFLFSSPPS--VHSAPAKHLLGF 711
Query: 468 EKVHVPAKTKQRVRVNIHVCKLLSVVDKSGIRRIPMGEHSLHIGDVKHSVSLQ 520
EKV + V + VCK LSVVD+ G R++ +G H+LH+GD+KH+++L+
Sbjct: 712 EKVSLEPGQAGVVAFKVDVCKDLSVVDELGNRKVALGSHTLHVGDLKHTLNLR 764
>UniRef100_Q9FLG1 Beta-xylosidase [Arabidopsis thaliana]
Length = 784
Score = 319 bits (817), Expect = 1e-85
Identities = 167/331 (50%), Positives = 218/331 (65%), Gaps = 8/331 (2%)
Query: 195 KLCRPLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETV 254
K PLQG+ T + GCSNVAC G A A AD ++LVIG DQSIEAE+
Sbjct: 456 KYTTPLQGLAGTVSTTYLPGCSNVACAVADVAG-ATKLAATADVSVLVIGADQSIEAESR 514
Query: 255 DRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAG 314
DR L LPG QQ+LV +VA A+KGP +LV+MSGG DITFAKNDPK+AGILW GYPG+AG
Sbjct: 515 DRVDLHLPGQQQELVIQVAKAAKGPVLLVIMSGGGFDITFAKNDPKIAGILWVGYPGEAG 574
Query: 315 GAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPSKI-GYPGRTYRFYKGPVVY 373
G AIADI+FG +P GKLP+TWYPQ Y++ + MT M MRP K GYPGRTYRFY G VY
Sbjct: 575 GIAIADIIFGRYNPSGKLPMTWYPQSYVEKVPMTIMNMRPDKASGYPGRTYRFYTGETVY 634
Query: 374 PFGHGLTYTHFVHELSSAPTVVSVPVHGHRHGNNTNISNKAIRVTHAR----CGKLSIAL 429
FG GL+YT F H L AP++VS+ + + ++ + H G + +
Sbjct: 635 AFGDGLSYTKFSHTLVKAPSLVSLGLEENHVCRSSECQSLDAIGPHCENAVSGGGSAFEV 694
Query: 430 HVDVKNVGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKL 489
H+ V+N G R+G HT+ +F+ PP H P+K LV FEK+ + + + VR + +CK
Sbjct: 695 HIKVRNGGDREGIHTVFLFTTPP--AIHGSPRKHLVGFEKIRLGKREEAVVRFKVEICKD 752
Query: 490 LSVVDKSGIRRIPMGEHSLHIGDVKHSVSLQ 520
LSVVD+ G R+I +G+H LH+GD+KHS+S++
Sbjct: 753 LSVVDEIGKRKIGLGKHLLHVGDLKHSLSIR 783
>UniRef100_Q76MS4 LEXYL2 protein [Lycopersicon esculentum]
Length = 633
Score = 317 bits (812), Expect = 5e-85
Identities = 162/351 (46%), Positives = 228/351 (64%), Gaps = 6/351 (1%)
Query: 171 SLPTTPPNCGCHWAQFRCHSYHDWKLCRPLQGIGRYAKTIHQQGCSNVACRDDKQFGPAL 230
SL PN + + K PL G+G T++QQGC ++AC Q A
Sbjct: 287 SLAVIGPNANLAYTMVGSYEGSPCKYTTPLDGLGASVSTVYQQGC-DIACAT-AQVDNAK 344
Query: 231 DAARHADATILVIGLDQSIEAETVDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPV 290
A ADA +LV+G DQ+IE E+ DR ++ LPG Q LV++VA+ SKGP ILV+MSGG +
Sbjct: 345 KVAAAADAVVLVMGSDQTIERESKDRFNITLPGQQSLLVTEVASVSKGPVILVIMSGGGM 404
Query: 291 DITFAKNDPKVAGILWAGYPGQAGGAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNM 350
D+ FA ++PKV ILW G+PG+AGGAA+AD++FG +PGG+LP+TWYPQ Y+ + MTNM
Sbjct: 405 DVKFAVDNPKVTSILWVGFPGEAGGAALADVVFGYHNPGGRLPMTWYPQSYVDKVDMTNM 464
Query: 351 AMRPS-KIGYPGRTYRFYKGPVVYPFGHGLTYTHFVHELSSAPTVVSVPVHGHRHGNNTN 409
MR K G+PGR+YRFYKGP V+ FG GL+YT + H L AP VS+P+ H +
Sbjct: 465 NMRADPKTGFPGRSYRFYKGPTVFNFGDGLSYTQYKHHLVKAPKFVSIPLE-EGHACRST 523
Query: 410 ISNKAIRVTHARCGKLSIALHVDVKNVGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEK 469
V C L + +H+ V+NVG G+HT+L+F++PP+ H PQK L+ F+K
Sbjct: 524 KCKSIDAVNEQGCNNLGLDIHLKVQNVGKMRGSHTVLLFTSPPS--VHNAPQKHLLDFQK 581
Query: 470 VHVPAKTKQRVRVNIHVCKLLSVVDKSGIRRIPMGEHSLHIGDVKHSVSLQ 520
+H+ +++ V+ N+ VCK LSVVD+ G R++ +G H LHIGD+KHS++L+
Sbjct: 582 IHLTPQSEGVVKFNLDVCKHLSVVDEVGNRKVALGLHVLHIGDLKHSLTLR 632
>UniRef100_Q76MS5 LEXYL1 protein [Lycopersicon esculentum]
Length = 770
Score = 311 bits (798), Expect = 2e-83
Identities = 155/327 (47%), Positives = 221/327 (67%), Gaps = 6/327 (1%)
Query: 195 KLCRPLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETV 254
K PLQG+ TI++ GC++V+C + Q A A ADA +LV+G DQSIE E++
Sbjct: 448 KYTTPLQGLTASVATIYKPGCADVSC-NTAQIDDAKQIATTADAVVLVMGSDQSIEKESL 506
Query: 255 DRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAG 314
DRTS+ LPG Q LV++VA +KGP ILV+MSGG +D+ FA ++PK+ ILW G+PG+AG
Sbjct: 507 DRTSITLPGQQSILVAEVAKVAKGPVILVIMSGGGMDVQFAVDNPKITSILWVGFPGEAG 566
Query: 315 GAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPS-KIGYPGRTYRFYKGPVVY 373
GAA+AD++FG +P G+LP+TWYPQ Y + MT+M MRP+ YPGRTYRFY GP V+
Sbjct: 567 GAALADVIFGYYNPSGRLPMTWYPQSYADVVPMTDMNMRPNPATNYPGRTYRFYTGPTVF 626
Query: 374 PFGHGLTYTHFVHELSSAPTVVSVPVHGHRHGNNTNISNKAIRVTHARCGKLSIALHVDV 433
FGHGL+Y+ F H L AP VS+P+ G +H + K + C + +H+ V
Sbjct: 627 TFGHGLSYSQFKHHLDKAPQFVSLPL-GEKHTCRLS-KCKTVDAVGQSCSNMGFDIHLRV 684
Query: 434 KNVGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVV 493
KNVG G+H + +F++PP+ H P+K L+ FEKVH+ + + V+ N++VCK LSV
Sbjct: 685 KNVGKISGSHIIFLFTSPPS--VHNAPKKHLLGFEKVHLTPQGEGVVKFNVNVCKHLSVH 742
Query: 494 DKSGIRRIPMGEHSLHIGDVKHSVSLQ 520
D+ G R++ +G H LHIGD+KHS++++
Sbjct: 743 DELGNRKVALGPHVLHIGDLKHSLTVR 769
>UniRef100_Q9LXD6 Beta-xylosidase [Arabidopsis thaliana]
Length = 773
Score = 307 bits (786), Expect = 6e-82
Identities = 159/329 (48%), Positives = 213/329 (64%), Gaps = 8/329 (2%)
Query: 195 KLCRPLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETV 254
K PLQG+ + +Q GC NVAC D G A+D A ADA +LV+G DQSIE E
Sbjct: 447 KYTTPLQGLAETVSSTYQLGC-NVACVD-ADIGSAVDLAASADAVVLVVGADQSIEREGH 504
Query: 255 DRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAG 314
DR L LPG QQ+LV++VA A++GP +LV+MSGG DITFAKND K+ I+W GYPG+AG
Sbjct: 505 DRVDLYLPGKQQELVTRVAMAARGPVVLVIMSGGGFDITFAKNDKKITSIMWVGYPGEAG 564
Query: 315 GAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPSKI-GYPGRTYRFYKGPVVY 373
G AIAD++FG +P G LP+TWYPQ Y++ + M+NM MRP K GYPGR+YRFY G VY
Sbjct: 565 GLAIADVIFGRHNPSGNLPMTWYPQSYVEKVPMSNMNMRPDKSKGYPGRSYRFYTGETVY 624
Query: 374 PFGHGLTYTHFVHELSSAPTVVSVPV---HGHRHGNNTNISNKAIRVTHARCGKLSIALH 430
F LTYT F H+L AP +VS+ + H R ++ +A G +H
Sbjct: 625 AFADALTYTKFDHQLIKAPRLVSLSLDENHPCRSSECQSLDAIGPHCENAVEGGSDFEVH 684
Query: 431 VDVKNVGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLL 490
++VKN G R G+HT+ +F+ P H P K L+ FEK+ + + VR N++VCK L
Sbjct: 685 LNVKNTGDRAGSHTVFLFTTSPQ--VHGSPIKQLLGFEKIRLGKSEEAVVRFNVNVCKDL 742
Query: 491 SVVDKSGIRRIPMGEHSLHIGDVKHSVSL 519
SVVD++G R+I +G H LH+G +KHS+++
Sbjct: 743 SVVDETGKRKIALGHHLLHVGSLKHSLNI 771
>UniRef100_Q8W012 Alpha-L-arabinofuranosidase/beta-D-xylosidase isoenzyme ARA-I
[Hordeum vulgare]
Length = 777
Score = 302 bits (773), Expect = 2e-80
Identities = 163/353 (46%), Positives = 215/353 (60%), Gaps = 7/353 (1%)
Query: 171 SLPTTPPNCGCHWAQFRCHSYHDWKLCRPLQGIGRYAKTIHQQGCSNVACRDDK-QFGPA 229
S+ PN + + K PLQG+G T++Q GC+NV C + Q A
Sbjct: 428 SMAVIGPNANASFTMIGNYEGTPCKYTTPLQGLGAKVNTVYQPGCTNVGCSGNSLQLSTA 487
Query: 230 LDAARHADATILVIGLDQSIEAETVDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGP 289
+ AA AD T+LV+G DQSIE E++DRTSLLLPG Q LVS VA AS GP ILV+MSGGP
Sbjct: 488 VAAAASADVTVLVVGADQSIERESLDRTSLLLPGQQTQLVSAVANASSGPVILVVMSGGP 547
Query: 290 VDITFAKNDPKVAGILWAGYPGQAGGAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTN 349
DI+FAK K+A LW GYPG+AGGAA+ D LFG+ +P G+LPVTWYP Y + MT+
Sbjct: 548 FDISFAKASDKIAATLWVGYPGEAGGAALDDTLFGSHNPSGRLPVTWYPASYADTVTMTD 607
Query: 350 MAMRP-SKIGYPGRTYRFYKGPVVYPFGHGLTYTHFVHELSSA-PTVVSVPVHGHRHGNN 407
M MRP + GYPGRTYRFY G V+ FG GL+YT H L SA P+ VS+ +
Sbjct: 608 MRMRPDTSTGYPGRTYRFYTGDTVFAFGDGLSYTKMSHSLVSAPPSYVSMRLAEDHLCRA 667
Query: 408 TNISNKAIRVTHARCGKLSIALHVDVKNVGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAF 467
+ ++ C L++ + + V+N G G H++L+FS+PP H P K LV F
Sbjct: 668 EECA--SVEAAGDHCDDLALDVKLQVRNAGEVAGAHSVLLFSSPPPA--HNAPAKHLVGF 723
Query: 468 EKVHVPAKTKQRVRVNIHVCKLLSVVDKSGIRRIPMGEHSLHIGDVKHSVSLQ 520
EKV + V + VC+ LSVVD+ G R++ +G H+LH GD+KH+V L+
Sbjct: 724 EKVSLAPGEAGTVAFRVDVCRDLSVVDELGGRKVALGGHTLHDGDLKHTVELR 776
>UniRef100_Q9LXD3 Similarity to beta-glucosidase [Arabidopsis thaliana]
Length = 411
Score = 296 bits (759), Expect = 8e-79
Identities = 155/330 (46%), Positives = 210/330 (62%), Gaps = 8/330 (2%)
Query: 195 KLCRPLQGIGRYAKTI-HQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAET 253
K PLQG+ R T + +GC NV C + A A ADAT+LV+G DQ+IE ET
Sbjct: 83 KYTTPLQGLERTVLTTKYHRGCFNVTCTE-ADLDSAKTLAASADATVLVMGADQTIEKET 141
Query: 254 VDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQA 313
+DR L LPG QQ+LV++VA A++GP +LV+MSGG DITFAKND K+ I+W GYPG+A
Sbjct: 142 LDRIDLNLPGKQQELVTQVAKAARGPVVLVIMSGGGFDITFAKNDEKITSIMWVGYPGEA 201
Query: 314 GGAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPSKI-GYPGRTYRFYKGPVV 372
GG AIAD++FG +P GKLP+TWYPQ Y++ + MTNM MRP K GY GRTYRFY G V
Sbjct: 202 GGIAIADVIFGRHNPSGKLPMTWYPQSYVEKVPMTNMNMRPDKSNGYLGRTYRFYIGETV 261
Query: 373 YPFGHGLTYTHFVHELSSAPTVVSVPVHGHRHGNNTNISNKAIRVTHARCG---KLSIAL 429
Y FG GL+YT+F H+L AP VS+ + + + + H + +
Sbjct: 262 YAFGDGLSYTNFSHQLIKAPKFVSLNLDESQSCRSPECQSLDAIGPHCEKAVGERSDFEV 321
Query: 430 HVDVKNVGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKL 489
+ V+NVG R+GT T+ +F+ PP H P+K L+ FEK+ + K + VR + VCK
Sbjct: 322 QLKVRNVGDREGTETVFLFTTPPE--VHGSPRKQLLGFEKIRLGKKEETVVRFKVDVCKD 379
Query: 490 LSVVDKSGIRRIPMGEHSLHIGDVKHSVSL 519
L VVD+ G R++ +G H LH+G +KHS ++
Sbjct: 380 LGVVDEIGKRKLALGHHLLHVGSLKHSFNI 409
>UniRef100_Q9LJN4 Beta-1,4-xylosidase [Arabidopsis thaliana]
Length = 876
Score = 275 bits (704), Expect = 2e-72
Identities = 143/326 (43%), Positives = 202/326 (61%), Gaps = 15/326 (4%)
Query: 195 KLCRPLQGIGRYA--KTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAE 252
K P+QG+ +Y K +++ GC +V C D A+ A AD T+LV+GLDQ++EAE
Sbjct: 439 KYTSPIQGLQKYVPEKIVYEPGCKDVKCGDQTLISAAVKAVSEADVTVLVVGLDQTVEAE 498
Query: 253 TVDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQ 312
+DR +L LPG+Q+ LV VA A+K +LV+MS GP+DI+FAKN + +LW GYPG+
Sbjct: 499 GLDRVNLTLPGYQEKLVRDVANAAKKTVVLVIMSAGPIDISFAKNLSTIRAVLWVGYPGE 558
Query: 313 AGGAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRP-SKIGYPGRTYRFYKGPV 371
AGG AIA ++FG +P G+LP TWYPQE+ +AMT+M MRP S G+PGR+YRFY G
Sbjct: 559 AGGDAIAQVIFGDYNPSGRLPETWYPQEFADKVAMTDMNMRPNSTSGFPGRSYRFYTGKP 618
Query: 372 VYPFGHGLTYTHFVHELSSAPTVVSVPVHGHRHGNNTNISNKAIRVTHARCGKLSIALHV 431
+Y FG+GL+Y+ F + SAP+++ + + + N T ++ ++ C L I + +
Sbjct: 619 IYKFGYGLSYSSFSTFVLSAPSIIHIKTNPIMNLNKTT----SVDISTVNCHDLKIRIVI 674
Query: 432 DVKNVGSRDGTHTLLVFSAPPN------GGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIH 485
VKN G R G+H +LVF PP GG VP LV FE+V V ++ V+
Sbjct: 675 GVKNHGLRSGSHVVLVFWKPPKCSKSLVGGG--VPLTQLVGFERVEVGRSMTEKFTVDFD 732
Query: 486 VCKLLSVVDKSGIRRIPMGEHSLHIG 511
VCK LS+VD G R++ G H L IG
Sbjct: 733 VCKALSLVDTHGKRKLVTGHHKLVIG 758
>UniRef100_Q9LXA8 Beta-xylosidase-like protein [Arabidopsis thaliana]
Length = 792
Score = 274 bits (700), Expect = 5e-72
Identities = 149/323 (46%), Positives = 202/323 (62%), Gaps = 13/323 (4%)
Query: 208 KTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETVDRTSLLLPGHQQD 267
KT + GCS+V+C D FG A+ A+ AD I+V GLD S E E DR SL LPG Q+D
Sbjct: 472 KTSYASGCSDVSCDSDTGFGEAVAIAKGADFVIVVAGLDLSQETEDKDRVSLSLPGKQKD 531
Query: 268 LVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAIADILFGTAS 327
LVS VAA SK P ILVL GGPVD+TFAKNDP++ I+W GYPG+ GG A+A+I+FG +
Sbjct: 532 LVSHVAAVSKKPVILVLTGGGPVDVTFAKNDPRIGSIIWIGYPGETGGQALAEIIFGDFN 591
Query: 328 PGGKLPVTWYPQEYLKNLAMTNMAMRP-SKIGYPGRTYRFYKGPVVYPFGHGLTYTHFVH 386
PGG+LP TWYP+ + ++AM++M MR S GYPGRTYRFY GP VY FG GL+YT F +
Sbjct: 592 PGGRLPTTWYPESF-TDVAMSDMHMRANSSRGYPGRTYRFYTGPQVYSFGTGLSYTKFEY 650
Query: 387 ELSSAPTVVSV-----PVHGHR----HGNNTNISNKAIRVTHARCGKLSIALHVDVKNVG 437
++ SAP +S+ H+ HG + ++ C L + V V N G
Sbjct: 651 KILSAPIRLSLSELLPQQSSHKKQLQHGEELRYLQLDDVIVNS-CESLRFNVRVHVSNTG 709
Query: 438 SRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVVDKSG 497
DG+H +++FS P + VP+K L+ +++VHV + I CK LSV + G
Sbjct: 710 EIDGSHVVMLFSKMPPVLS-GVPEKQLIGYDRVHVRSNEMMETVFVIDPCKQLSVANDVG 768
Query: 498 IRRIPMGEHSLHIGDVKHSVSLQ 520
R IP+G H L +GD++HS+S++
Sbjct: 769 KRVIPLGSHVLFLGDLQHSLSVE 791
>UniRef100_Q8GZ32 Putative beta-xylosidase [Arabidopsis thaliana]
Length = 732
Score = 274 bits (700), Expect = 5e-72
Identities = 149/323 (46%), Positives = 202/323 (62%), Gaps = 13/323 (4%)
Query: 208 KTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETVDRTSLLLPGHQQD 267
KT + GCS+V+C D FG A+ A+ AD I+V GLD S E E DR SL LPG Q+D
Sbjct: 412 KTSYASGCSDVSCDSDTGFGEAVAIAKGADFVIVVAGLDLSQETEDKDRVSLSLPGKQKD 471
Query: 268 LVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAIADILFGTAS 327
LVS VAA SK P ILVL GGPVD+TFAKNDP++ I+W GYPG+ GG A+A+I+FG +
Sbjct: 472 LVSHVAAVSKKPVILVLTGGGPVDVTFAKNDPRIGSIIWIGYPGETGGQALAEIIFGDFN 531
Query: 328 PGGKLPVTWYPQEYLKNLAMTNMAMRP-SKIGYPGRTYRFYKGPVVYPFGHGLTYTHFVH 386
PGG+LP TWYP+ + ++AM++M MR S GYPGRTYRFY GP VY FG GL+YT F +
Sbjct: 532 PGGRLPTTWYPESF-TDVAMSDMHMRANSSRGYPGRTYRFYTGPQVYSFGTGLSYTKFEY 590
Query: 387 ELSSAPTVVSV-----PVHGHR----HGNNTNISNKAIRVTHARCGKLSIALHVDVKNVG 437
++ SAP +S+ H+ HG + ++ C L + V V N G
Sbjct: 591 KILSAPIRLSLSELLPQQSSHKKQLQHGEELRYLQLDDVIVNS-CESLRFNVRVHVSNTG 649
Query: 438 SRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVVDKSG 497
DG+H +++FS P + VP+K L+ +++VHV + I CK LSV + G
Sbjct: 650 EIDGSHVVMLFSKMPPVLS-GVPEKQLIGYDRVHVRSNEMMETVFVIDPCKQLSVANDVG 708
Query: 498 IRRIPMGEHSLHIGDVKHSVSLQ 520
R IP+G H L +GD++HS+S++
Sbjct: 709 KRVIPLGSHVLFLGDLQHSLSVE 731
>UniRef100_Q7X7M4 OSJNBa0074L08.23 protein [Oryza sativa]
Length = 770
Score = 261 bits (666), Expect = 5e-68
Identities = 141/339 (41%), Positives = 202/339 (58%), Gaps = 27/339 (7%)
Query: 200 LQGIGRYA-KTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETVDRTS 258
++G+ Y KT GC +V C FG A++AA+ AD +L+ GL+ + E E DR S
Sbjct: 442 VKGMQAYVPKTTFAAGCKDVPCNSTDGFGEAIEAAKRADVVVLIAGLNLTEETEDHDRVS 501
Query: 259 LLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAI 318
LLLPG Q DL+ VA+ +K P +LVLM GGPVD++FAK+DP++A ILW GYPG+ GG +
Sbjct: 502 LLLPGRQMDLIHTVASVTKKPVVLVLMGGGPVDVSFAKHDPRIASILWIGYPGEVGGNVL 561
Query: 319 ADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMR-PSKIGYPGRTYRFYKGPVVYPFGH 377
+ILFG +PGGKLP+TWYP+ + + M +M MR + GYPGRTYRFY G VVY FG+
Sbjct: 562 PEILFGKYNPGGKLPITWYPESFTA-VPMDDMNMRADASRGYPGRTYRFYTGDVVYGFGY 620
Query: 378 GLTYTHFVHELSSAPTVVSV------------PVHGHRHGNNTNISNKAIRVTH-ARCGK 424
GL+Y+ + + + AP +S+ P + R G + ++V A C
Sbjct: 621 GLSYSKYSYSILQAPKKISLSRSSVPDLISRKPAYTRRDGVD------YVQVEDIASCEA 674
Query: 425 LSIALHVDVKNVGSRDGTHTLLVF--SAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRV 482
L +H+ V N G+ DG+H +L+F S P G+ P K LV FE+VH A V +
Sbjct: 675 LQFPVHISVSNDGAMDGSHAVLLFASSKPSFPGS---PIKQLVGFERVHTAAGRSTDVEI 731
Query: 483 NIHVCKLLSVVDKSGIRRIPMGEHSLHIGDVKHSVSLQA 521
+ CKL+S + G R + +G H L +GD +H + ++A
Sbjct: 732 TVDPCKLMSFANTEGTRVLFLGTHVLMVGDEEHELLIEA 770
>UniRef100_Q75RZ3 Putative beta-xylosidase [Triticum aestivum]
Length = 573
Score = 245 bits (626), Expect = 2e-63
Identities = 140/329 (42%), Positives = 186/329 (55%), Gaps = 9/329 (2%)
Query: 199 PLQGIGRYAK-TIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETVDRT 257
P Q + Y K QGC+ C + G A+ AA AD +L +GLDQ+ E E VDR
Sbjct: 248 PFQALQGYVKDATFVQGCNAAVC-NVSNIGEAVHAASSADYVVLFMGLDQNQEREEVDRL 306
Query: 258 SLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAA 317
L LPG Q+ LV+KVA A+K P ILVL+ GGPVD+TFAKN+PK+ I+WAGYPGQAGG A
Sbjct: 307 ELGLPGMQESLVNKVADAAKKPVILVLLCGGPVDVTFAKNNPKIGAIVWAGYPGQAGGIA 366
Query: 318 IADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPS-KIGYPGRTYRFYKGPVVYPFG 376
IA +LFG +PGG+LPVTWYP+E+ + MT+M MR GYPGRTYRFYKG VY FG
Sbjct: 367 IAQVLFGEHNPGGRLPVTWYPKEFTA-VPMTDMRMRADPSTGYPGRTYRFYKGKTVYNFG 425
Query: 377 HGLTYTHFVHELSS----APTVVSVPVHGHRHGNNTNISNKAIRVTHARCGKLSIALHVD 432
+GL+Y+ + H +S P++ + +S + C +L V
Sbjct: 426 YGLSYSKYSHRFASEGTKPPSMSGIEGLKATASAAGTVSYDVEEMGAEACDRLRFPAVVR 485
Query: 433 VKNVGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSV 492
V+N G DG H +L+F PN P L+ F+ VH+ A V + CK S
Sbjct: 486 VQNHGPMDGRHPVLLFLRWPN-ATDGRPASQLIGFQSVHLRADEAAHVEFEVSPCKHFSR 544
Query: 493 VDKSGIRRIPMGEHSLHIGDVKHSVSLQA 521
+ G + I G H + +GD + +S A
Sbjct: 545 AAEDGRKVIDQGSHFVKVGDDEFELSFMA 573
>UniRef100_Q94IY5 Putative beta-xylosidase [Oryza sativa]
Length = 818
Score = 245 bits (626), Expect = 2e-63
Identities = 135/341 (39%), Positives = 202/341 (58%), Gaps = 21/341 (6%)
Query: 195 KLCR---PLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEA 251
K CR P G+ + + C +C A AA+ DATI+V GL+ S+E
Sbjct: 479 KPCRVVTPYDGVRKVVSSTSVHACDKGSC------DTAAAAAKTVDATIVVAGLNMSVER 532
Query: 252 ETVDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPG 311
E+ DR LLLP Q ++ VA AS P +LV+MS G VD++FA+++PK+ ++WAGYPG
Sbjct: 533 ESNDREDLLLPWSQASWINAVAEASPSPIVLVIMSAGGVDVSFAQDNPKIGAVVWAGYPG 592
Query: 312 QAGGAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRP-SKIGYPGRTYRFYKG- 369
+ GG AIAD+LFG +PGG+LP+TWY EY+ + MT+MA+RP ++ GYPGRTY+FY G
Sbjct: 593 EEGGTAIADVLFGKYNPGGRLPLTWYKNEYVSKIPMTSMALRPDAEHGYPGRTYKFYGGA 652
Query: 370 PVVYPFGHGLTYTHFVHELSSAPTVVSVPVHG--------HRHGNNTNISNKAIRVTHAR 421
V+YPFGHGL+YT+F + ++A V+V V ++ G ++ + A+ V
Sbjct: 653 DVLYPFGHGLSYTNFTYASATAAAPVTVKVGAWEYCKQLTYKAGVSSPPACPAVNVASHA 712
Query: 422 CGKLSIALHVDVKNVGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVR 481
C + ++ V V N G RDGTH + +++APP P+K LVAF +V V A V
Sbjct: 713 CQE-EVSFAVTVANTGGRDGTHVVPMYTAPP-AEVDGAPRKQLVAFRRVRVAAGAAVEVA 770
Query: 482 VNIHVCKLLSVVDKSGIRRIPMGEHSLHIGDVKHSVSLQAE 522
++VCK ++V+++ +P G + +GD S+S +
Sbjct: 771 FALNVCKAFAIVEETAYTVVPSGVSRVLVGDDALSLSFPVQ 811
>UniRef100_Q8W011 Beta-D-xylosidase [Hordeum vulgare]
Length = 777
Score = 245 bits (626), Expect = 2e-63
Identities = 142/332 (42%), Positives = 187/332 (55%), Gaps = 13/332 (3%)
Query: 199 PLQGIGRYAKTIH-QQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETVDRT 257
PLQ + Y K QGC+ C + G A+ AA AD +L +GLDQ+ E E VDR
Sbjct: 450 PLQALQGYVKDARFVQGCNAAVC-NVSNIGEAVHAAGSADYVVLFMGLDQNQEREEVDRL 508
Query: 258 SLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAA 317
L LPG Q+ LV+ VA A+K P ILVL+ GGPVD+TFAKN+PK+ I+WAGYPGQAGG A
Sbjct: 509 ELGLPGMQESLVNSVADAAKKPVILVLLCGGPVDVTFAKNNPKIGAIVWAGYPGQAGGIA 568
Query: 318 IADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPS-KIGYPGRTYRFYKGPVVYPFG 376
IA +LFG +PGG+LPVTWYP+E+ + MT+M MR GYPGRTYRFYKG VY FG
Sbjct: 569 IAQVLFGDHNPGGRLPVTWYPKEFTA-VPMTDMRMRADPSTGYPGRTYRFYKGKTVYNFG 627
Query: 377 HGLTYTHFVHELSSAPT-------VVSVPVHGHRHGNNTNISNKAIRVTHARCGKLSIAL 429
+GL+Y+ + H +S T + + T +S + C +L
Sbjct: 628 YGLSYSKYSHRFASKGTKPPSMSGIEGLKATARASAAGT-VSYDVEEMGAEACDRLRFPA 686
Query: 430 HVDVKNVGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKL 489
V V+N G DG H +L+F PN P L+ F+ VH+ A V + CK
Sbjct: 687 VVRVQNHGPMDGGHLVLLFLRWPN-ATDGRPASQLIGFQSVHLRADEAAHVEFEVSPCKH 745
Query: 490 LSVVDKSGIRRIPMGEHSLHIGDVKHSVSLQA 521
LS + G + I G H + +GD + +S A
Sbjct: 746 LSRAAEDGRKVIDQGSHFVRVGDDEFELSFMA 777
>UniRef100_Q9SGZ5 F28K19.27 [Arabidopsis thaliana]
Length = 696
Score = 239 bits (610), Expect = 1e-61
Identities = 132/326 (40%), Positives = 197/326 (59%), Gaps = 16/326 (4%)
Query: 195 KLCRPLQGIGRYAKT-IHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAET 253
K PL + Y K ++ QGC +VAC + A+ A++AD +L++GLDQ+ E E
Sbjct: 370 KTVTPLDALRSYVKNAVYHQGCDSVAC-SNAAIDQAVAIAKNADHVVLIMGLDQTQEKED 428
Query: 254 VDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQA 313
DR L LPG QQ+L++ VA A+K P +LVL+ GGPVDI+FA N+ K+ I+WAGYPG+A
Sbjct: 429 FDRVDLSLPGKQQELITSVANAAKKPVVLVLICGGPVDISFAANNNKIGSIIWAGYPGEA 488
Query: 314 GGAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPSKIGYPGRTYRFYKGPVVY 373
GG AI++I+FG +PGG+LPVTWYPQ ++ N+ MT+M MR S GYPGRTY+FYKGP VY
Sbjct: 489 GGIAISEIIFGDHNPGGRLPVTWYPQSFV-NIQMTDMRMR-SATGYPGRTYKFYKGPKVY 546
Query: 374 PFGHGLTYTHFVHELSSAPTVVSVPVHGHRHGNNTNISNKAIRVT------HARCGKLSI 427
FGHGL+Y+ + + T+ ++ ++ TN + ++R T C
Sbjct: 547 EFGHGLSYSAYSYRFK---TLAETNLYLNQSKAQTN--SDSVRYTLVSEMGKEGCDVAKT 601
Query: 428 ALHVDVKNVGSRDGTHTLLVFSAPPNGGNHWV-PQKSLVAFEKVHVPAKTKQRVRVNIHV 486
+ V+V+N G G H +L+F+ GG +K LV F+ + + K + I +
Sbjct: 602 KVTVEVENQGEMAGKHPVLMFARHERGGEDGKRAEKQLVGFKSIVLSNGEKAEMEFEIGL 661
Query: 487 CKLLSVVDKSGIRRIPMGEHSLHIGD 512
C+ LS ++ G+ + G++ L +GD
Sbjct: 662 CEHLSRANEFGVMVLEEGKYFLTVGD 687
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.337 0.146 0.508
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 873,432,277
Number of Sequences: 2790947
Number of extensions: 36033775
Number of successful extensions: 106490
Number of sequences better than 10.0: 386
Number of HSP's better than 10.0 without gapping: 308
Number of HSP's successfully gapped in prelim test: 78
Number of HSP's that attempted gapping in prelim test: 105471
Number of HSP's gapped (non-prelim): 465
length of query: 528
length of database: 848,049,833
effective HSP length: 132
effective length of query: 396
effective length of database: 479,644,829
effective search space: 189939352284
effective search space used: 189939352284
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 77 (34.3 bits)
Medicago: description of AC145329.13


