

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145219.5 + phase: 0
(130 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
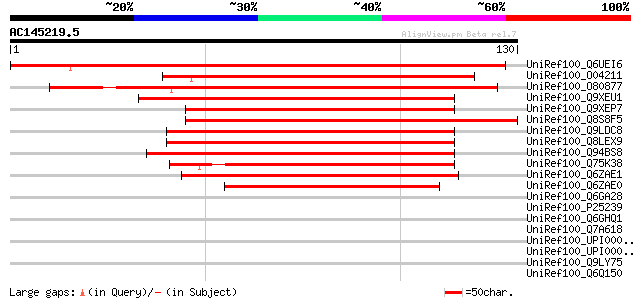
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6UEI6 Early flowering 4 [Mesembryanthemum crystallinum] 122 1e-27
UniRef100_O04211 Expressed protein [Arabidopsis thaliana] 113 1e-24
UniRef100_O80877 Hypothetical protein At2g29950 [Arabidopsis tha... 94 7e-19
UniRef100_Q9XEU1 Hypothetical protein [Oryza sativa] 86 1e-16
UniRef100_Q9XEP7 Hypothetical protein [Sorghum bicolor] 84 5e-16
UniRef100_Q8S8F5 Expressed protein [Arabidopsis thaliana] 84 7e-16
UniRef100_Q9LDC8 F28G4.6 protein [Arabidopsis thaliana] 81 5e-15
UniRef100_Q8LEX9 Hypothetical protein [Arabidopsis thaliana] 79 3e-14
UniRef100_Q94BS8 Hypothetical protein At1g72630 [Arabidopsis tha... 77 1e-13
UniRef100_Q75K38 Expressed protein [Oryza sativa] 75 4e-13
UniRef100_Q6ZAE1 Hypothetical protein P0410E02.4 [Oryza sativa] 67 9e-11
UniRef100_Q6ZAE0 Hypothetical protein P0410E02.6 [Oryza sativa] 53 1e-06
UniRef100_Q6GA28 Putative cell division protein [Staphylococcus ... 37 0.076
UniRef100_P25239 Type IIS restriction enzyme Eco57I (EC 3.1.21.4... 37 0.099
UniRef100_Q6GHQ1 Putative cell division protein [Staphylococcus ... 37 0.13
UniRef100_Q7A618 Div1b protein [Staphylococcus aureus] 37 0.13
UniRef100_UPI0000432CD5 UPI0000432CD5 UniRef100 entry 36 0.17
UniRef100_UPI00002E9F02 UPI00002E9F02 UniRef100 entry 36 0.17
UniRef100_Q9LY75 Peptidyl-prolyl cis-trans isomerase [Arabidopsi... 36 0.17
UniRef100_Q6Q150 Multidomain cyclophilin type peptidyl-prolyl ci... 36 0.17
>UniRef100_Q6UEI6 Early flowering 4 [Mesembryanthemum crystallinum]
Length = 139
Score = 122 bits (307), Expect = 1e-27
Identities = 65/128 (50%), Positives = 82/128 (63%), Gaps = 1/128 (0%)
Query: 1 MSTLYSPSSAMEDT-NHRSKKNQRRRRPSATPTNNDYSDVEDAEDGDPEAWQTLNKSFRQ 59
M + S+MED NH + RR + NN+ + E GDP W +SF +
Sbjct: 1 MMSYSGAESSMEDGGNHHRQTLAGNRRRRSNNNNNNNNGEEYFSGGDPAEWNAFAESFSK 60
Query: 60 VQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNISKVASLYSDLNSDFTNICH 119
VQSVLDRNR +IQQ NEN QSR+PDNMVKNV +IQELNGNISKVAS+YSDL+ +FT +
Sbjct: 61 VQSVLDRNRMLIQQANENHQSRVPDNMVKNVAIIQELNGNISKVASIYSDLSVNFTGVVG 120
Query: 120 QQQRSSHN 127
Q ++ N
Sbjct: 121 QHRQQGGN 128
>UniRef100_O04211 Expressed protein [Arabidopsis thaliana]
Length = 111
Score = 113 bits (282), Expect = 1e-24
Identities = 55/81 (67%), Positives = 68/81 (83%), Gaps = 1/81 (1%)
Query: 40 EDAEDG-DPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNG 98
E+AE G DP W+ L+++FRQVQSVLDRNR++IQQVN+N QSRM DNM KNV LIQELNG
Sbjct: 15 EEAEQGEDPAMWENLDRNFRQVQSVLDRNRSLIQQVNDNHQSRMADNMSKNVALIQELNG 74
Query: 99 NISKVASLYSDLNSDFTNICH 119
NISKV ++YSDLN+ F++ H
Sbjct: 75 NISKVVNMYSDLNTSFSSGFH 95
>UniRef100_O80877 Hypothetical protein At2g29950 [Arabidopsis thaliana]
Length = 125
Score = 94.0 bits (232), Expect = 7e-19
Identities = 52/116 (44%), Positives = 72/116 (61%), Gaps = 4/116 (3%)
Query: 11 MEDTNHRSKKNQRRRRPSATPTNNDYSDVE-DAEDGDPEAWQTLNKSFRQVQSVLDRNRA 69
ME + +RS R S ND DV A D E W TL+ F++ Q LD+NR
Sbjct: 1 MEASRNRSLVGNNR---SPEMNENDGEDVAASAAVEDVEVWDTLSNGFKRAQLYLDQNRD 57
Query: 70 IIQQVNENQQSRMPDNMVKNVGLIQELNGNISKVASLYSDLNSDFTNICHQQQRSS 125
+IQ+VNEN SR+PDN+ +NVGLI E+NGNIS+V +YSDL+ +F Q++R++
Sbjct: 58 LIQRVNENHMSRIPDNVSRNVGLINEINGNISQVMEIYSDLSLNFAKKFDQRRRTT 113
>UniRef100_Q9XEU1 Hypothetical protein [Oryza sativa]
Length = 117
Score = 86.3 bits (212), Expect = 1e-16
Identities = 39/81 (48%), Positives = 59/81 (72%)
Query: 34 NDYSDVEDAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLI 93
+ +S + + D + QT +KSF QVQS+LD+NR +I ++N+N +SR PDN+ +NVGLI
Sbjct: 4 DSFSGMANGGQVDNKLIQTFHKSFVQVQSILDQNRMLINEINQNHESRAPDNLTRNVGLI 63
Query: 94 QELNGNISKVASLYSDLNSDF 114
+ELN NI +V LY+DL++ F
Sbjct: 64 RELNNNIRRVVGLYADLSASF 84
>UniRef100_Q9XEP7 Hypothetical protein [Sorghum bicolor]
Length = 438
Score = 84.3 bits (207), Expect = 5e-16
Identities = 38/69 (55%), Positives = 54/69 (78%)
Query: 46 DPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNISKVAS 105
D + QT +KSF QVQS+LD+NR +I ++N+N +SR PDN+ +NVGLI+ELN NI +V
Sbjct: 10 DGKLIQTFHKSFVQVQSLLDQNRMLISEINQNHESRAPDNLTRNVGLIRELNNNIRRVVG 69
Query: 106 LYSDLNSDF 114
LY+DL++ F
Sbjct: 70 LYADLSASF 78
>UniRef100_Q8S8F5 Expressed protein [Arabidopsis thaliana]
Length = 109
Score = 84.0 bits (206), Expect = 7e-16
Identities = 37/85 (43%), Positives = 58/85 (67%)
Query: 46 DPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNISKVAS 105
D + QT KSF QVQ++LD NR +I ++N+N +S++PDN+ +NVGLI+ELN N+ +VA
Sbjct: 17 DGKILQTFEKSFVQVQNILDHNRLLINEINQNHESKIPDNLGRNVGLIRELNNNVRRVAH 76
Query: 106 LYSDLNSDFTNICHQQQRSSHNRGK 130
LY DL+++F+ + G+
Sbjct: 77 LYVDLSNNFSKSMEASSEGDSSEGR 101
>UniRef100_Q9LDC8 F28G4.6 protein [Arabidopsis thaliana]
Length = 141
Score = 81.3 bits (199), Expect = 5e-15
Identities = 37/74 (50%), Positives = 53/74 (71%)
Query: 41 DAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNI 100
D + D + Q+ KSF VQ +LD+NR +I ++N+N +S+ PDN+ +NVGLI+ELN NI
Sbjct: 11 DRHNMDGKLLQSFQKSFVDVQDILDQNRLLINEINQNHESKQPDNLGRNVGLIKELNNNI 70
Query: 101 SKVASLYSDLNSDF 114
+VASLY DL+ F
Sbjct: 71 RRVASLYGDLSHSF 84
>UniRef100_Q8LEX9 Hypothetical protein [Arabidopsis thaliana]
Length = 114
Score = 78.6 bits (192), Expect = 3e-14
Identities = 36/74 (48%), Positives = 52/74 (69%)
Query: 41 DAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNI 100
D + D + Q+ KSF VQ +LD+NR +I ++N+ +S+ PDN+ +NVGLI+ELN NI
Sbjct: 11 DRHNMDGKLLQSFQKSFVDVQDILDQNRLLINEINQXHESKQPDNLGRNVGLIKELNNNI 70
Query: 101 SKVASLYSDLNSDF 114
+VASLY DL+ F
Sbjct: 71 RRVASLYGDLSHSF 84
>UniRef100_Q94BS8 Hypothetical protein At1g72630 [Arabidopsis thaliana]
Length = 119
Score = 76.6 bits (187), Expect = 1e-13
Identities = 37/79 (46%), Positives = 52/79 (64%)
Query: 36 YSDVEDAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQE 95
YS + D + Q KSF QVQ +LD+NR +I ++N+N +S+ D++ +NVGLI+E
Sbjct: 10 YSGFGERYQMDGKLLQNFQKSFVQVQDILDQNRLLINEINQNHESKQADHLGRNVGLIRE 69
Query: 96 LNGNISKVASLYSDLNSDF 114
LN NI VASLY DL+ F
Sbjct: 70 LNNNIRTVASLYGDLSHSF 88
>UniRef100_Q75K38 Expressed protein [Oryza sativa]
Length = 114
Score = 74.7 bits (182), Expect = 4e-13
Identities = 39/74 (52%), Positives = 56/74 (74%), Gaps = 4/74 (5%)
Query: 42 AEDGDP-EAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNI 100
AEDG A+QT SF QVQS+LD+NR +I ++N+N +S++P ++ +NVGLI+ELN NI
Sbjct: 11 AEDGKVLHAFQT---SFVQVQSLLDQNRVLINEINQNHESKVPGDLSRNVGLIRELNNNI 67
Query: 101 SKVASLYSDLNSDF 114
+V LY+DL+S F
Sbjct: 68 RRVVDLYADLSSLF 81
>UniRef100_Q6ZAE1 Hypothetical protein P0410E02.4 [Oryza sativa]
Length = 144
Score = 67.0 bits (162), Expect = 9e-11
Identities = 28/71 (39%), Positives = 51/71 (71%)
Query: 45 GDPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNISKVA 104
G + Q L ++F +VQ +L++NR +IQ++++N ++R D + +NV LI+ELN NI++V
Sbjct: 36 GGGKVVQVLQRNFGEVQGILEQNRVLIQEISQNHEARDADGLTRNVALIRELNTNIARVV 95
Query: 105 SLYSDLNSDFT 115
LY++L+ F+
Sbjct: 96 DLYANLSGSFS 106
>UniRef100_Q6ZAE0 Hypothetical protein P0410E02.6 [Oryza sativa]
Length = 357
Score = 53.1 bits (126), Expect = 1e-06
Identities = 21/55 (38%), Positives = 41/55 (74%)
Query: 56 SFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNISKVASLYSDL 110
S +V+ +L++N +IQ++++N ++R D + +NV LI++LN NI++V LY++L
Sbjct: 256 SLGEVRGILEQNHTLIQEISQNHKARDADRLTRNVALIRDLNTNIARVVDLYANL 310
>UniRef100_Q6GA28 Putative cell division protein [Staphylococcus aureus]
Length = 440
Score = 37.4 bits (85), Expect = 0.076
Identities = 23/77 (29%), Positives = 39/77 (49%), Gaps = 2/77 (2%)
Query: 14 TNHRSKKNQRRRRPSATP-TNNDYSDVEDAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQ 72
T + SKK ++R+R AT +N + D D D E + +NK F++ QS + N +
Sbjct: 24 TRNTSKKRRQRKRSKATHFSNQNKDDTSQQADFDEEIY-LINKDFKKEQSNDENNDSASS 82
Query: 73 QVNENQQSRMPDNMVKN 89
+ N N D+ ++N
Sbjct: 83 RANNNNIDDSTDSNIEN 99
>UniRef100_P25239 Type IIS restriction enzyme Eco57I (EC 3.1.21.4) (Endonuclease
Eco57I) [Includes: Adenine-specific methyltransferase
activity Eco57IA (EC 2.1.1.72) (M.Eco57IA)] [Escherichia
coli]
Length = 998
Score = 37.0 bits (84), Expect = 0.099
Identities = 24/78 (30%), Positives = 41/78 (51%), Gaps = 3/78 (3%)
Query: 35 DYSDVEDAEDGDP--EAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGL 92
D+ D + E D Q LNK + Q Q DR++ I ++ E ++ +M D ++KN+
Sbjct: 922 DFDDPKQKEIHDTISSKQQYLNKLYSQTQKSADRDKIIFERQFEQEKIQM-DYLIKNLFD 980
Query: 93 IQELNGNISKVASLYSDL 110
+ +L+ I V LY +L
Sbjct: 981 LGDLDSEIPTVEDLYKNL 998
>UniRef100_Q6GHQ1 Putative cell division protein [Staphylococcus aureus]
Length = 440
Score = 36.6 bits (83), Expect = 0.13
Identities = 22/77 (28%), Positives = 38/77 (48%), Gaps = 2/77 (2%)
Query: 14 TNHRSKKNQRRRRPSATP-TNNDYSDVEDAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQ 72
T + SKK ++R+R AT +N + D D D E + +NK F++ Q + N +
Sbjct: 24 TRNTSKKRRQRKRSKATHFSNQNKDDTSQQADFDEEIY-LINKDFKKEQGNEENNDSASS 82
Query: 73 QVNENQQSRMPDNMVKN 89
N+N D+ ++N
Sbjct: 83 HANDNNIDDSTDSNIEN 99
>UniRef100_Q7A618 Div1b protein [Staphylococcus aureus]
Length = 439
Score = 36.6 bits (83), Expect = 0.13
Identities = 22/77 (28%), Positives = 39/77 (50%), Gaps = 2/77 (2%)
Query: 14 TNHRSKKNQRRRRPSATP-TNNDYSDVEDAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQ 72
T + SKK ++R+R AT +N + D D D E + +NK F++ +S + N +
Sbjct: 23 TRNTSKKRRQRKRSKATHFSNQNKDDTSQQADFDEEIY-LINKDFKKEESNDENNGSASS 81
Query: 73 QVNENQQSRMPDNMVKN 89
N+N D+ ++N
Sbjct: 82 HANDNNIDDSTDSNIEN 98
>UniRef100_UPI0000432CD5 UPI0000432CD5 UniRef100 entry
Length = 142
Score = 36.2 bits (82), Expect = 0.17
Identities = 29/123 (23%), Positives = 53/123 (42%), Gaps = 2/123 (1%)
Query: 5 YSPSSAMEDTNHRSKKNQRRRRPSATPTNNDYSDVEDAEDGDPEAWQTLNKSFRQVQSVL 64
Y + + + SK QRRR+ T N Y + E+ D E+ + + S QS L
Sbjct: 22 YQSLTQTQQNVNNSKMLQRRRKKKELSTMNSYINYENISKSDSESLTSSSGSPLSNQS-L 80
Query: 65 DRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNISKVASLYSDLNSDFTNICHQQQRS 124
+ + I Q++E ++ + K +L +I + A L D N +C+ +
Sbjct: 81 NSSNTSISQISEKSITKSQTLLSKKKANRSQLKVSIKETAELIEDQNKS-KKLCNNEYLI 139
Query: 125 SHN 127
S++
Sbjct: 140 SND 142
>UniRef100_UPI00002E9F02 UPI00002E9F02 UniRef100 entry
Length = 197
Score = 36.2 bits (82), Expect = 0.17
Identities = 27/78 (34%), Positives = 41/78 (51%), Gaps = 12/78 (15%)
Query: 40 EDAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQQVNE-NQQSRMPDNMVKNVGLIQELNG 98
ED + D E +T+NK +Q++S+ VNE + +RMP + K LI+ +N
Sbjct: 8 EDKKSFD-ENLKTINKQIKQIESL----------VNEFSDFARMPKPIFKKYDLIKIINE 56
Query: 99 NISKVASLYSDLNSDFTN 116
NIS + L S +N F N
Sbjct: 57 NISLMNELDSSINISFKN 74
>UniRef100_Q9LY75 Peptidyl-prolyl cis-trans isomerase [Arabidopsis thaliana]
Length = 570
Score = 36.2 bits (82), Expect = 0.17
Identities = 29/109 (26%), Positives = 52/109 (47%), Gaps = 20/109 (18%)
Query: 17 RSKKNQRRRRPSATPTNNDYSDVEDAEDGDPEAWQTLNKSFRQVQ------------SVL 64
+ + +++ RP P+ N SD E + D E + +K+ + V+ SV
Sbjct: 282 KGRSGKKKARPDRKPSTNSSSDTESSSSSDDE--KVGHKAIKSVKVDNADQHANLDDSVK 339
Query: 65 DRNRAIIQQVNENQQSRMPD-NMVKNVGLIQELNGNISKVASLYSDLNS 112
R+R+ I++ N+N +S+ P + V+ +G NGN S S DL +
Sbjct: 340 SRSRSPIRRRNQNSRSKSPSRSPVRVLG-----NGNRSPSRSPVRDLGN 383
>UniRef100_Q6Q150 Multidomain cyclophilin type peptidyl-prolyl cis-trans isomerase
[Arabidopsis thaliana]
Length = 566
Score = 36.2 bits (82), Expect = 0.17
Identities = 29/109 (26%), Positives = 52/109 (47%), Gaps = 20/109 (18%)
Query: 17 RSKKNQRRRRPSATPTNNDYSDVEDAEDGDPEAWQTLNKSFRQVQ------------SVL 64
+ + +++ RP P+ N SD E + D E + +K+ + V+ SV
Sbjct: 282 KGRSGKKKARPDRKPSTNSSSDTESSSSSDDE--KVGHKAIKSVKVDNADQHANLDDSVK 339
Query: 65 DRNRAIIQQVNENQQSRMPD-NMVKNVGLIQELNGNISKVASLYSDLNS 112
R+R+ I++ N+N +S+ P + V+ +G NGN S S DL +
Sbjct: 340 SRSRSPIRRRNQNSRSKSPSRSPVRVLG-----NGNRSPSRSPVRDLGN 383
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.309 0.123 0.340
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 221,221,391
Number of Sequences: 2790947
Number of extensions: 8515425
Number of successful extensions: 28885
Number of sequences better than 10.0: 189
Number of HSP's better than 10.0 without gapping: 52
Number of HSP's successfully gapped in prelim test: 137
Number of HSP's that attempted gapping in prelim test: 28793
Number of HSP's gapped (non-prelim): 225
length of query: 130
length of database: 848,049,833
effective HSP length: 106
effective length of query: 24
effective length of database: 552,209,451
effective search space: 13253026824
effective search space used: 13253026824
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 67 (30.4 bits)
Medicago: description of AC145219.5


