

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145218.1 - phase: 0
(63 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
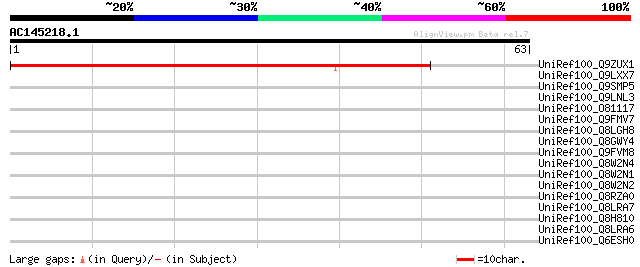
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9ZUX1 Putative cytochrome P450 [Arabidopsis thaliana] 47 1e-04
UniRef100_Q9LXX7 Cytochrome P450-like protein [Arabidopsis thali... 37 0.10
UniRef100_Q9SMP5 Cytochrome P450-like protein [Arabidopsis thali... 37 0.13
UniRef100_Q9LNL3 F12K21.15 [Arabidopsis thaliana] 37 0.13
UniRef100_O81117 Cytochrome P450 94A1 [Vicia sativa] 35 0.66
UniRef100_Q9FMV7 Cytochrome P450-like protein [Arabidopsis thali... 34 1.1
UniRef100_Q8LGH8 Cytochrome P450-like protein [Arabidopsis thali... 34 1.1
UniRef100_Q8GWY4 Putative cytochrome P450 [Arabidopsis thaliana] 34 1.1
UniRef100_Q9FVM8 Cytochrome P450 [Triticum aestivum] 33 2.5
UniRef100_Q8W2N4 Cytochrome P450-dependent fatty acid hydroxylas... 32 3.3
UniRef100_Q8W2N1 Cytochrome P450-dependent fatty acid hydroxylas... 32 3.3
UniRef100_Q8W2N2 Cytochrome P450-dependent fatty acid hydroxylas... 32 4.3
UniRef100_Q8RZA0 Putative cytochrome P450-like protein [Oryza sa... 32 4.3
UniRef100_Q8LRA7 Putative cytochrome P450-dependent fatty acid h... 32 4.3
UniRef100_Q8H810 Putative cytochrome P450 [Oryza sativa] 32 5.6
UniRef100_Q8LRA6 Putative cytochrome P450-dependent fatty acid h... 31 7.3
UniRef100_Q6ESH0 Putative cytochrome P450 [Oryza sativa] 31 7.3
>UniRef100_Q9ZUX1 Putative cytochrome P450 [Arabidopsis thaliana]
Length = 495
Score = 47.0 bits (110), Expect = 1e-04
Identities = 25/52 (48%), Positives = 36/52 (69%), Gaps = 1/52 (1%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDE-KIFDLRDIMRSSS 51
+ASLEL SV+VR +A EI+ EI TRL+P + SF+ + + DL+D+ R S
Sbjct: 127 LASLELGSVSVRVFAHEIVKTEIETRLLPILTSFSDNPGSVLDLQDVFRRFS 178
>UniRef100_Q9LXX7 Cytochrome P450-like protein [Arabidopsis thaliana]
Length = 499
Score = 37.4 bits (85), Expect = 0.10
Identities = 16/46 (34%), Positives = 30/46 (64%)
Query: 2 ASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
AS E ++ ++R + + + EI+TRL+P + A + K+ DL+DI+
Sbjct: 129 ASYEFSTKSLRDFVMSNVTVEINTRLVPVLAEAATNGKLIDLQDIL 174
>UniRef100_Q9SMP5 Cytochrome P450-like protein [Arabidopsis thaliana]
Length = 506
Score = 37.0 bits (84), Expect = 0.13
Identities = 15/48 (31%), Positives = 32/48 (66%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIMR 48
+AS E ++ ++RS+A E++ +E+ RL+P + + A DL+D+++
Sbjct: 133 LASHEFSTRSLRSFAFEVLKDEVENRLVPVLSTAADVGTTVDLQDVLK 180
>UniRef100_Q9LNL3 F12K21.15 [Arabidopsis thaliana]
Length = 498
Score = 37.0 bits (84), Expect = 0.13
Identities = 16/46 (34%), Positives = 29/46 (62%)
Query: 2 ASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
AS E ++ ++R + + + EI+TRL+P + A K+ DL+DI+
Sbjct: 129 ASYEFSTKSLRDFVMSNVTVEINTRLVPVLVEAATTGKLIDLQDIL 174
>UniRef100_O81117 Cytochrome P450 94A1 [Vicia sativa]
Length = 514
Score = 34.7 bits (78), Expect = 0.66
Identities = 15/48 (31%), Positives = 29/48 (60%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIMR 48
+AS E + ++R++ I++ E+ RLIP + S + I D +DI++
Sbjct: 145 VASHEFNTKSIRNFVEHIVDTELTNRLIPILTSSTQTNNILDFQDILQ 192
>UniRef100_Q9FMV7 Cytochrome P450-like protein [Arabidopsis thaliana]
Length = 510
Score = 33.9 bits (76), Expect = 1.1
Identities = 15/48 (31%), Positives = 28/48 (58%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIMR 48
+AS E T ++R + EI+ EE+ RLIP + S + D +++++
Sbjct: 135 LASHEFTMRSLREFTFEILREEVQNRLIPVLSSAVDCGETVDFQEVLK 182
>UniRef100_Q8LGH8 Cytochrome P450-like protein [Arabidopsis thaliana]
Length = 508
Score = 33.9 bits (76), Expect = 1.1
Identities = 15/48 (31%), Positives = 28/48 (58%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIMR 48
+AS E T ++R + EI+ EE+ RLIP + S + D +++++
Sbjct: 133 LASHEFTMRSLREFTFEILREEVQNRLIPVLSSAVDCGETVDFQEVLK 180
>UniRef100_Q8GWY4 Putative cytochrome P450 [Arabidopsis thaliana]
Length = 510
Score = 33.9 bits (76), Expect = 1.1
Identities = 15/48 (31%), Positives = 28/48 (58%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIMR 48
+AS E T ++R + EI+ EE+ RLIP + S + D +++++
Sbjct: 135 LASHEFTMRSLREFTFEILREEVQNRLIPVLSSAVDCGETVDFQEVLK 182
>UniRef100_Q9FVM8 Cytochrome P450 [Triticum aestivum]
Length = 541
Score = 32.7 bits (73), Expect = 2.5
Identities = 15/46 (32%), Positives = 28/46 (60%)
Query: 2 ASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
A+LE T+ T+R+ ++ IH RL+P + A+ + DL+D++
Sbjct: 134 AALEFTTRTLRTAMSRWVSRSIHGRLLPILADAAKGKAQVDLQDLL 179
>UniRef100_Q8W2N4 Cytochrome P450-dependent fatty acid hydroxylase [Nicotiana
tabacum]
Length = 510
Score = 32.3 bits (72), Expect = 3.3
Identities = 13/48 (27%), Positives = 30/48 (62%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIMR 48
+AS E + ++R + +++ E+ RLIP + + A ++ + D +DI++
Sbjct: 141 VASHEFNTRSLRKFVETVVDTELSERLIPILATAAANKTVLDFQDILQ 188
>UniRef100_Q8W2N1 Cytochrome P450-dependent fatty acid hydroxylase [Nicotiana
tabacum]
Length = 511
Score = 32.3 bits (72), Expect = 3.3
Identities = 13/48 (27%), Positives = 30/48 (62%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIMR 48
+AS E + ++R + +++ E+ RLIP + + A ++ + D +DI++
Sbjct: 142 VASHEFNTRSLRKFVETVVDTELSERLIPILATAAANKTVLDFQDILQ 189
>UniRef100_Q8W2N2 Cytochrome P450-dependent fatty acid hydroxylase [Nicotiana
tabacum]
Length = 511
Score = 32.0 bits (71), Expect = 4.3
Identities = 12/48 (25%), Positives = 31/48 (64%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIMR 48
++S E + ++R + +++ E++ RLIP + + A ++ + D +DI++
Sbjct: 139 VSSHEFNTKSLRKFVETVVDTELNERLIPILATAAVEKTVLDFQDILQ 186
>UniRef100_Q8RZA0 Putative cytochrome P450-like protein [Oryza sativa]
Length = 544
Score = 32.0 bits (71), Expect = 4.3
Identities = 12/47 (25%), Positives = 28/47 (59%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
+AS +S ++R ++ ++ +H RL+P + + A + DL+D++
Sbjct: 137 LASYSFSSRSLRRFSARVLRAHLHRRLVPLLDAAAGSGEAVDLQDVL 183
>UniRef100_Q8LRA7 Putative cytochrome P450-dependent fatty acid hydroxylase [Oryza
sativa]
Length = 508
Score = 32.0 bits (71), Expect = 4.3
Identities = 12/46 (26%), Positives = 27/46 (58%)
Query: 2 ASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
AS E ++R++ ++ + E+ RL+P + RD + D++D++
Sbjct: 136 ASYEFNKRSLRNFVVDTVRSEVVERLLPLLERAERDGRTLDVQDVL 181
>UniRef100_Q8H810 Putative cytochrome P450 [Oryza sativa]
Length = 523
Score = 31.6 bits (70), Expect = 5.6
Identities = 13/46 (28%), Positives = 27/46 (58%)
Query: 2 ASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
AS E + ++R++ ++ E+H RL+P + A + + DL+D +
Sbjct: 137 ASYEFNTRSLRAFVARCVHGELHGRLLPLLRRAAAEGRAIDLQDAL 182
>UniRef100_Q8LRA6 Putative cytochrome P450-dependent fatty acid hydroxylase [Oryza
sativa]
Length = 512
Score = 31.2 bits (69), Expect = 7.3
Identities = 13/46 (28%), Positives = 27/46 (58%)
Query: 2 ASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
AS E ++RS+ ++ + E+ RL+P + RD + D++D++
Sbjct: 138 ASYEFNQRSLRSFVVDTVRFEVVERLLPLLEWARRDGRTLDVQDVL 183
>UniRef100_Q6ESH0 Putative cytochrome P450 [Oryza sativa]
Length = 532
Score = 31.2 bits (69), Expect = 7.3
Identities = 15/46 (32%), Positives = 27/46 (58%)
Query: 2 ASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
A+LE T+ T+R+ ++ IH RL+P + A + DL+D++
Sbjct: 134 AALEFTTRTLRTAMSRWVSRSIHHRLLPILDDAAAGKAHVDLQDLL 179
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.325 0.135 0.352
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 78,221,026
Number of Sequences: 2790947
Number of extensions: 2062280
Number of successful extensions: 6810
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 17
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 6793
Number of HSP's gapped (non-prelim): 17
length of query: 63
length of database: 848,049,833
effective HSP length: 39
effective length of query: 24
effective length of database: 739,202,900
effective search space: 17740869600
effective search space used: 17740869600
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC145218.1


