

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145202.5 - phase: 0 /pseudo
(1053 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
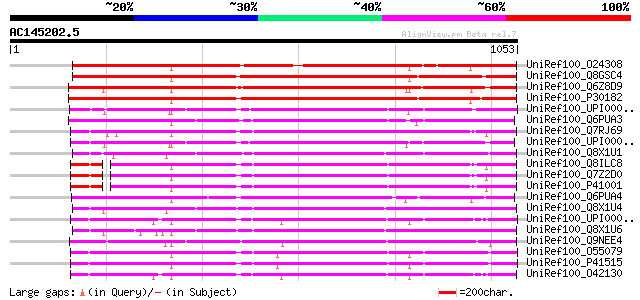
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O24308 DNA topoisomerase II [Pisum sativum] 988 0.0
UniRef100_Q8GSC4 DNA topoisomerase II [Nicotiana tabacum] 857 0.0
UniRef100_Q6Z8D9 Putative DNA topoisomerase II [Oryza sativa] 839 0.0
UniRef100_P30182 DNA topoisomerase II [Arabidopsis thaliana] 825 0.0
UniRef100_UPI0000430198 UPI0000430198 UniRef100 entry 677 0.0
UniRef100_Q6PUA3 DNA topoisomerase type 2 [Tetrahymena thermophila] 654 0.0
UniRef100_Q7RJ69 DNA topoisomerase II, putative [Plasmodium yoel... 653 0.0
UniRef100_UPI00003C1D48 UPI00003C1D48 UniRef100 entry 645 0.0
UniRef100_Q8X1U1 DNA topoisomerase II [Penicillium marneffei] 637 0.0
UniRef100_Q8ILC8 DNA topoisomerase II, putative [Plasmodium falc... 632 e-179
UniRef100_Q7Z2D0 DNA topoisomerase [Plasmodium falciparum] 632 e-179
UniRef100_P41001 DNA topoisomerase II [Plasmodium falciparum] 630 e-179
UniRef100_Q6PUA4 DNA topoisomerase type 2 [Tetrahymena thermophila] 629 e-178
UniRef100_Q8X1U4 DNA topoisomerase II [Aspergillus niger] 627 e-178
UniRef100_UPI00003AC994 UPI00003AC994 UniRef100 entry 624 e-177
UniRef100_Q8X1U6 DNA topoisomerase II [Aspergillus flavus] 624 e-177
UniRef100_Q9NEE4 DNA topoisomerase II [Leishmania major] 621 e-176
UniRef100_O55079 Topoisomerase II alpha [Homo sapiens] 621 e-176
UniRef100_P41515 DNA topoisomerase II, alpha isozyme [Cricetulus... 620 e-176
UniRef100_O42130 DNA topoisomerase II, alpha isozyme [Gallus gal... 620 e-176
>UniRef100_O24308 DNA topoisomerase II [Pisum sativum]
Length = 1462
Score = 988 bits (2553), Expect = 0.0
Identities = 560/966 (57%), Positives = 674/966 (68%), Gaps = 72/966 (7%)
Query: 130 WTNVMFKPDLDKFDMKNLEKDVIALMKKRVLDMAVCLGETVTVVLNGTVIPVKSFKDYAN 189
WT V FKPDL+KF M LE+DV+ALMKKRVLDMA C G+TV V LNGT+I KSF+DYA+
Sbjct: 203 WTKVTFKPDLEKFKMAYLEEDVVALMKKRVLDMAGCFGKTVKVELNGTLIRFKSFRDYAD 262
Query: 190 FFL---DRAKPFPLPRTHAKLGDRLEICLSLSDGKFQQVSFVNSIATIKGGTHVDYITKQ 246
FL +++KP PLPR HAK+GDR EIC+SLSDG+FQQVSFVNSIATIKGGTHVDYIT Q
Sbjct: 263 LFLKCAEKSKPMPLPRIHAKVGDRWEICISLSDGQFQQVSFVNSIATIKGGTHVDYITNQ 322
Query: 247 ITTYIKKEV-LKKKHVNVNADTVKNQLWVFVNALIDNPAFNSQTKEMLTTKPARLGLKF- 304
ITTYI +V KKK NV A TVKN LWVFVN+LIDNPAF+SQTKE LTT+ A G K
Sbjct: 323 ITTYIMNKVNKKKKDANVKAHTVKNHLWVFVNSLIDNPAFDSQTKETLTTRQASFGSKCD 382
Query: 305 FPGSMLNDVQKSQILDNLLPRIPFKHRTHV-----------------DAKLAGRPDSGKC 347
P SML DV+KS I+D LL FK + DA AG +S KC
Sbjct: 383 VPESMLKDVEKSGIVDTLLSWADFKQSKDLKKTDGTKTQRLRGCKAEDANEAGGRNSEKC 442
Query: 348 TLILTQGQYVKDLAMSGLSAVVDPDLYGVFPLSSKLLNATR-DSRKLFKKKEIQNLMAIL 406
TLILT+G K LAM+GLS VV D YGVFPL KLLN S+++ +EIQN+ IL
Sbjct: 443 TLILTEGDSAKALAMAGLS-VVGRDHYGVFPLRGKLLNVREASSKQIMDNEEIQNIKKIL 501
Query: 407 GLVRNKKYSNAKCLRYGQLMVMANQDEDGAHFKGLLINFFYSFWPSLLKVSKFMSVFTFP 466
GL +NK+Y+N K LRYG LM+MA+QD DG+H KGLLINF +SFWPSLLKV F+ FT P
Sbjct: 502 GLQQNKEYTNVKSLRYGHLMIMADQDHDGSHIKGLLINFIHSFWPSLLKVPSFLVEFTTP 561
Query: 467 IIKVSMYNASDSKKGKELSFDSMQQYEDWKKELGNTANDWEIKYCQGL--SSAQEVREYF 524
+I+ S N + + SF SM +YE WK+ LGN+A W+IKY +GL S+ QE REYF
Sbjct: 562 VIRASHPNKTIT------SFYSMPEYEAWKERLGNSATSWKIKYYKGLGTSTPQEGREYF 615
Query: 525 QDLGKHRAYFVWDDK-DETSIKLAFSKK-AEEWEAWIGNSQPGTC-NYERKVINYRNFVN 581
DLG+HR F+WDD+ D +I+LAFSKK AE + W+ N +PGTC ++E K+INY++FVN
Sbjct: 616 SDLGRHRKDFIWDDELDGNAIELAFSKKKAEGRKIWMRNFEPGTCRDHEAKLINYKDFVN 675
Query: 582 KELLFSSMKNLQRSIPSVVDGLTLGQRKTLFCSFEMNLTKQTEVCRLTAYVTEHSYCHYG 641
KEL+ S R LF SF+ L K+ +V + YV+EHS H+G
Sbjct: 676 KELILFS-------------------RADLFGSFKKKLYKEIKVAQFIGYVSEHSAYHHG 716
Query: 642 QQSIASTIIGMAQDFVGSNNINLLIPYGQFGTRDLGGKDHASSRKLYIKLNCVTRLLFPV 701
+QS+ASTIIGMAQDFVGSNNINLL P GQFGT +LGGKDHAS+R +Y +L+ VTR LF
Sbjct: 717 EQSLASTIIGMAQDFVGSNNINLLKPNGQFGTCNLGGKDHASARYIYTELSPVTRCLFHE 776
Query: 702 DDDKLLEYLNEDGRSIQPNWYIPIIPLVLVNGCHAMGTCFSSDIPKYHPYEIIKNVRRFL 761
DDKLLEYLNEDG+SI+PNWY+PIIPLVLVNG +GT +SS IP Y+P EII NVRR L
Sbjct: 777 HDDKLLEYLNEDGKSIEPNWYMPIIPLVLVNGSEGIGTGWSSYIPNYNPREIIANVRRLL 836
Query: 762 NNEEMVSMKPWYKGFGGTIEKSAKGIGYYTVDGLVERINEQTFRIKELPIRMWTEDYKKF 821
N EE+V M PWYKGF GTIEKSAK G Y V+G V I+EQTFRI ELPIR WT+DYK+F
Sbjct: 837 NGEELVPMDPWYKGFRGTIEKSAKE-GGYIVNGTVTEIDEQTFRITELPIRKWTQDYKQF 895
Query: 822 LEKITARG------LIESFRQKGDYEIVNFKVNVKQEEIATIEKDEELRKKFNLTCTIST 875
LE IT LIE FRQ GD IV+ ++ +K E+IATI E L KKF LT TIST
Sbjct: 896 LESITDGAPNVKDPLIEDFRQNGDDAIVDIEIKMKPEKIATI-LQEGLFKKFKLTSTIST 954
Query: 876 SNMYLFDAEGKIKKYNTPEQILEEFCTLRLKYYVRRKQYLVKNFTQLLRSLRIKQKFISN 935
SNM+LFDAEG KK++TPEQILEEF LRL YY + K+Y++ N +LL L K +FI
Sbjct: 955 SNMHLFDAEGN-KKFDTPEQILEEFFPLRLDYYEKSKEYILGNLNRLLLILDNKVRFILG 1013
Query: 936 IVNEKFNPINR-RAELLIEIL--------KKGKSAEPHVAGAIDYSSKEQEAGEQESVSQ 986
+VN + NR +AELLIE+ +KGKS +P VAGA D S+EQE E E+ SQ
Sbjct: 1014 VVNGEIIVSNRKKAELLIELKEKGFTPMPRKGKSTKPQVAGANDDDSEEQEDAEPETASQ 1073
Query: 987 SVSIEGATLGDYEYLSSLPFENFTLESLKKLEAELDEKENELKTLEHTSPDLMWLNDLEL 1046
SVS+EGAT GDY+ L SLP TLES++KL E EKE E + L T +WL DL+
Sbjct: 1074 SVSVEGATWGDYDDLLSLPIGTLTLESVQKLLDEKTEKEKEYEILSGTPTTSLWLKDLDE 1133
Query: 1047 FEKEFD 1052
FEK+ D
Sbjct: 1134 FEKKLD 1139
>UniRef100_Q8GSC4 DNA topoisomerase II [Nicotiana tabacum]
Length = 1482
Score = 857 bits (2215), Expect = 0.0
Identities = 480/963 (49%), Positives = 636/963 (65%), Gaps = 54/963 (5%)
Query: 130 WTNVMFKPDLDKFDMKNLEKDVIALMKKRVLDMAVCLGETVTVVLNGTVIPVKSFKDYAN 189
WT V KPDL KF+M++LE+DV+ALM+KRV+D+ CLG+TV V LN IPVKSF++Y
Sbjct: 207 WTKVSSKPDLAKFNMEHLEEDVVALMRKRVIDLGGCLGKTVKVKLNEQRIPVKSFEEYCK 266
Query: 190 FFLDR--AKPFPLPRTHAKLGDRLEICLSLSDGKFQQVSFVNSIATIKGGTHVDYITKQI 247
FLD AK L T A R EIC+SLS+G+FQQVSFVNSIATIKGGTHVDY+ QI
Sbjct: 267 LFLDSTDAKREFLKVTDADGLLRWEICVSLSEGQFQQVSFVNSIATIKGGTHVDYVANQI 326
Query: 248 TTYIKKEVLKK-KHVNVNADTVKNQLWVFVNALIDNPAFNSQTKEMLTTKPARLGLKF-F 305
+I V+KK K+ N+ A VKN LW+FVNALIDNPAF+SQTKE LT + + G K
Sbjct: 327 ANHIMGAVIKKNKNANIKAHAVKNHLWMFVNALIDNPAFDSQTKETLTLRQSSFGSKCEL 386
Query: 306 PGSMLNDVQKS-QILDNLLPRIPFKHRTHV-----------------DAKLAGRPDSGKC 347
L V+K+ I++ LL FK+ + DA AG +S KC
Sbjct: 387 QPDFLKKVEKNIGIVETLLSWADFKNSKDLKKTDGKKSEKVKVEKLEDANDAGGRNSEKC 446
Query: 348 TLILTQGQYVKDLAMSGLSAVVDPDLYGVFPLSSKLLNATRDSRK-LFKKKEIQNLMAIL 406
TLILT+G K LAM+G+S VV D YGVFPL KLLN S K + + KEI+ + IL
Sbjct: 447 TLILTEGDSAKALAMAGIS-VVGRDHYGVFPLRGKLLNVREASHKQVSENKEIEAIKKIL 505
Query: 407 GLVRNKKYSNAKCLRYGQLMVMANQDEDGAHFKGLLINFFYSFWPSLLKVSKFMSVFTFP 466
GL K+Y + K LRYG LM+M +QD DG+H KGLLINF ++FWPSLLKV F+ F P
Sbjct: 506 GLQTGKEYDSVKSLRYGHLMIMTDQDHDGSHIKGLLINFIHTFWPSLLKVPSFLIEFITP 565
Query: 467 IIKVSMYNASDSKKGKELSFDSMQQYEDWKKELGNTANDWEIKYCQGL--SSAQEVREYF 524
I+K + K GK LSF +M +YE W+K LG ++ W IKY +GL S+++E +EYF
Sbjct: 566 IVKAT------HKSGKILSFYTMPEYESWRKSLGANSSGWSIKYYKGLGTSTSKEGKEYF 619
Query: 525 QDLGKHRAYFVW-DDKDETSIKLAFSKKA-EEWEAWIGNSQPGT-CNYERKVINYRNFVN 581
QDL KHR F+W D++D SI+LAFSKK E + W+ +PGT + + K I+Y FVN
Sbjct: 620 QDLQKHRKDFIWADNQDGESIELAFSKKKIEARKNWLRQFEPGTHLDQKEKYISYTEFVN 679
Query: 582 KELLFSSMKNLQRSIPSVVDGLTLGQRKTLFCSFEMNLTKQTEVCRLTAYVTEHSYCHYG 641
KEL+ SM +LQRSIPS++DGL GQRK LFC+F+ N K+ +V + + YV+EHS H+G
Sbjct: 680 KELILFSMADLQRSIPSMLDGLKPGQRKILFCAFKRNFVKEAKVSQFSGYVSEHSAYHHG 739
Query: 642 QQSIASTIIGMAQDFVGSNNINLLIPYGQFGTRDLGGKDHASSRKLYIKLNCVTRLLFPV 701
+QS++STIIGMAQD+VGSNN+NLL P GQFGTR++GGKDHASSR +Y +L+ + R LFP
Sbjct: 740 EQSLSSTIIGMAQDYVGSNNVNLLQPNGQFGTRNMGGKDHASSRYIYTRLSPIARFLFPK 799
Query: 702 DDDKLLEYLNEDGRSIQPNWYIPIIPLVLVNGCHAMGTCFSSDIPKYHPYEIIKNVRRFL 761
+DD + +YLNEDG+ I+P WY+PI+P+VL+NG +GT +SS +P Y+P +++ NVRR L
Sbjct: 800 EDDTIHDYLNEDGQYIEPTWYVPIVPMVLINGSEGIGTGWSSYVPNYNPRDLVANVRRLL 859
Query: 762 NNEEMVSMKPWYKGFGGTIEKSA--KGIGYYTVDGLVERINEQTFRIKELPIRMWTEDYK 819
N+E M M PWYKGF GTIEK+A + YTV G++E +NE T RI ELP+R WTEDYK
Sbjct: 860 NDEPMEPMDPWYKGFKGTIEKTATKEAGATYTVTGIIEEVNETTLRISELPVRRWTEDYK 919
Query: 820 KFLEKITARG------LIESFRQKGDYEIVNFKVNVKQEEIATIEKDEELRKKFNLTCTI 873
+FLE +T I+ R GD V F+V + EE + + E L KKF L TI
Sbjct: 920 QFLESMTVSNDKAKDPFIKEVRAYGDENSVCFEV-IMSEENLILAQQEGLLKKFKLATTI 978
Query: 874 STSNMYLFDAEGKIKKYNTPEQILEEFCTLRLKYYVRRKQYLVKNFTQLLRSLRIKQKFI 933
STSNM+LFD+ GKIKKY+ PE ILEEF +RL+YY +RK+ L++ L + K KFI
Sbjct: 979 STSNMHLFDSNGKIKKYDNPEDILEEFYHVRLEYYEKRKKALLEILELELLRIENKVKFI 1038
Query: 934 SNIVNEKFNPINR-RAELLIEILKKGKSAEPH---VAGAIDYSSKEQEAGEQESVSQSVS 989
+V + NR RA+LL+E+ +KG + P V + +S + E E+E
Sbjct: 1039 LGVVKVEIIVNNRKRADLLLELKEKGFTPFPKKKAVEAVVADTSDDAEDSEEE------L 1092
Query: 990 IEGATLGDYEYLSSLPFENFTLESLKKLEAELDEKENELKTLEHTSPDLMWLNDLELFEK 1049
G GDY+YL S+P TLE +++L AE D+ E++ + + +P L+WL DL++ EK
Sbjct: 1093 NRGVRAGDYDYLLSMPIGTLTLEKVQELCAERDQLNGEVEDMRNATPKLLWLKDLDVLEK 1152
Query: 1050 EFD 1052
+ D
Sbjct: 1153 QLD 1155
>UniRef100_Q6Z8D9 Putative DNA topoisomerase II [Oryza sativa]
Length = 1525
Score = 839 bits (2167), Expect = 0.0
Identities = 482/1002 (48%), Positives = 633/1002 (63%), Gaps = 86/1002 (8%)
Query: 122 VSLCDNGK-WTNVMFKPDLDKFDMKNLEKDVIALMKKRVLDMAVCLGETVTVVLNGTVIP 180
++ C G+ WT V FKPDL KF+M +LE DV+ALM+KRV+DMA LG+TV V L+ +P
Sbjct: 203 ITKCKQGENWTRVTFKPDLAKFNMTHLENDVVALMRKRVVDMAGTLGKTVKVELDHQKVP 262
Query: 181 VKSFKDYANFFL-----DRAKPFPLPRTHAKLGDRLEICLSLSDGKFQQVSFVNSIATIK 235
V SF DY ++ DR LP K+ DR E+C+SLS+G+FQQVSFVN IATIK
Sbjct: 263 VHSFSDYVKLYIKSASKDRDDVNELPSISQKVNDRWEVCVSLSEGQFQQVSFVNRIATIK 322
Query: 236 GGTHVDYITKQITTYIKKEVLKK-KHVNVNADTVKNQLWVFVNALIDNPAFNSQTKEMLT 294
GGTHVDY+T QI T++ V K+ K+ ++ A VK+ LWVFVNALIDNPAF+SQTKE LT
Sbjct: 323 GGTHVDYVTNQIATHVMNIVNKRNKNAHMKAHNVKSHLWVFVNALIDNPAFDSQTKETLT 382
Query: 295 TKPARLGLKF-FPGSMLNDVQKSQILDNLLPRIPFKHRTHV------------------D 335
T+ A G K L V S I+ NLL FK + D
Sbjct: 383 TRQASFGSKCELSDDFLKKVGSSAIVLNLLSWAEFKLSKELQKTDGSKRSRLTGIPKLED 442
Query: 336 AKLAGRPDSGKCTLILTQGQYVKDLAMSGLSAVVDPDLYGVFPLSSKLLNATRDSRK-LF 394
A AG DS CTLILT+G K LAM+G+S VV D YGVFPL KLLN S K +
Sbjct: 443 ANGAGGKDSNNCTLILTEGDSAKALAMAGIS-VVGRDYYGVFPLRGKLLNVREASHKQIM 501
Query: 395 KKKEIQNLMAILGLVRNKKYSNAKCLRYGQLMVMANQDEDGAHFKGLLINFFYSFWPSLL 454
+ EIQN+ ILGL K+Y + K LRYG LM+M +QD DG+H KGLLINF +SFWPSL+
Sbjct: 502 ENAEIQNIKQILGLQHGKQYDSTKGLRYGHLMIMTDQDHDGSHIKGLLINFIHSFWPSLI 561
Query: 455 KVSKFMSVFTFPIIKVSMYNASDSKKGKELSFDSMQQYEDWKKELGNTANDWEIKYCQGL 514
K+ F+ F PIIK + N D K L F SM +YE WK+ LG A+ W IKY +GL
Sbjct: 562 KIPSFLVEFITPIIKAT--NKRDKKI--VLPFYSMPEYEQWKESLGGNASGWSIKYYKGL 617
Query: 515 --SSAQEVREYFQDLGKHRAYFVW-DDKDETSIKLAFSKKA-EEWEAWIGNSQPGT-CNY 569
S++ E R+YFQD+ KH+ FVW +D+D+ I+LAFSKK + + W+ N Q GT +
Sbjct: 618 GTSTSSEGRQYFQDIAKHKKDFVWKNDQDDNDIELAFSKKRITDRKEWLTNFQSGTHLDT 677
Query: 570 ERKVINYRNFVNKELLFSSMKNLQRSIPSVVDGLTLGQRKTLFCSFEMNLTKQTEVCRLT 629
E K I Y +F+NKEL+ SM +L RSIPS+VDGL GQRK LFCSF+ NL K+ +V + +
Sbjct: 678 EGKYIKYSDFINKELIQFSMADLLRSIPSMVDGLKPGQRKILFCSFKRNLVKEIKVAQFS 737
Query: 630 AYVTEHSYCHYGQQSIASTIIGMAQDFVGSNNINLLIPYGQFGTRDLGGKDHASSRKLYI 689
YV+EHS H+G+QS+ASTI GMAQDFVGSNNINLL P GQFGTRD GGKD AS+R ++
Sbjct: 738 GYVSEHSAYHHGEQSLASTITGMAQDFVGSNNINLLQPNGQFGTRDQGGKDAASARYIFT 797
Query: 690 KLNCVTRLLFPVDDDKLLEYLNEDGRSIQPNWYIPIIPLVLVNGCHAMGTCFSSDIPKYH 749
L+ +TR +FP DDD LL YL+EDG+SI+P WY+PI+P+VLVNG +GT +S+ IP Y+
Sbjct: 798 LLSPITRSIFPKDDDILLNYLDEDGQSIEPTWYVPILPMVLVNGSEGIGTGWSTFIPNYN 857
Query: 750 PYEIIKNVRRFLNNEEMVSMKPWYKGFGGTIEKSAKGIG--YYTVDGLVERINEQTFRIK 807
P +I+ N+RR LN+E + M PWY+GF G+I+K+ G YTV G++E +++ T RI
Sbjct: 858 PRDIVANLRRLLNDEPVEPMDPWYRGFKGSIQKTGTKAGGVSYTVTGIIEVVDDTTLRIT 917
Query: 808 ELPIRMWTEDYKKFL------------EKITARG----------------LIESFRQKGD 839
ELPIR W++DYK+FL +K +G IE+F D
Sbjct: 918 ELPIRRWSQDYKEFLISIGGTDKSKDKDKDKGKGKGKVKEKEKKEKDIEPFIEAFDTYSD 977
Query: 840 YEIVNFKVNVKQEEIATIEKDEELRKKFNLTCTISTSNMYLFDAEGKIKKYNTPEQILEE 899
+ V F + + +E +A I E L KKF LT TI T+NM+LFD+ GKI+KY+TPE IL+E
Sbjct: 978 DKNVEFLITLSKENMA-IALQEGLEKKFKLTTTIGTTNMHLFDSNGKIRKYDTPEDILKE 1036
Query: 900 FCTLRLKYYVRRKQYLVKNFTQLLRSLRIKQKFISNIVNEKFNPINR-RAELLIEILKKG 958
F LRL++Y +RK+ L++N L+ L K +FI +V NR RAEL +E+ +KG
Sbjct: 1037 FFGLRLEFYEKRKRVLLENIELELKKLSNKVRFILAVVEGDIIVNNRKRAELFVELKQKG 1096
Query: 959 --------KSAEPHVAGAIDYSSKEQEAGEQESVSQSVSIEGATLGDYEYLSSLPFENFT 1010
+ A P GAI+ + +E+ E +V S DYEYL S+ T
Sbjct: 1097 FDPFPRKKQRAGPSAVGAIEEDEENEESPEAANVGSS---------DYEYLLSMAIGTLT 1147
Query: 1011 LESLKKLEAELDEKENELKTLEHTSPDLMWLNDLELFEKEFD 1052
LE +++L AE ENE+ L+ T P +W+ DL+ FEKE D
Sbjct: 1148 LERVQQLIAEKGRMENEVAELKRTRPKSLWMRDLDAFEKELD 1189
>UniRef100_P30182 DNA topoisomerase II [Arabidopsis thaliana]
Length = 1473
Score = 825 bits (2132), Expect = 0.0
Identities = 475/977 (48%), Positives = 626/977 (63%), Gaps = 57/977 (5%)
Query: 122 VSLCDNGK-WTNVMFKPDLDKFDMKNLEKDVIALMKKRVLDMAVCLGETVTVVLNGTVIP 180
++ C+ + WT V FKPDL KF+M LE DV+ALM KRV D+A CLG++V V LNG IP
Sbjct: 206 ITKCNKSENWTKVTFKPDLKKFNMTELEDDVVALMSKRVFDIAGCLGKSVKVELNGKQIP 265
Query: 181 VKSFKDYANFFLDRAKPF----PLPRTHAKLGDRLEICLSLSDGKFQQVSFVNSIATIKG 236
VKSF DY + +L A PLPR K+ DR E+C+SLS+G+FQQVSFVNSIATIKG
Sbjct: 266 VKSFTDYVDLYLSAANKSRTEDPLPRLTEKVNDRWEVCVSLSEGQFQQVSFVNSIATIKG 325
Query: 237 GTHVDYITKQITTYIKKEVLKK-KHVNVNADTVKNQLWVFVNALIDNPAFNSQTKEMLTT 295
GTHVDY+T QIT +I V KK K+ NV A VKN LWVFVNALIDNPAF+SQTKE LT
Sbjct: 326 GTHVDYVTSQITNHIVAAVNKKNKNANVKAHNVKNHLWVFVNALIDNPAFDSQTKETLTL 385
Query: 296 KPARLGLKF-FPGSMLNDVQKSQILDNLLPRIPFKHRTHV-----------------DAK 337
+ + G K L V KS +++NLL FK + DA
Sbjct: 386 RQSSFGSKCELSEDFLKKVGKSGVVENLLSWADFKQNKDLKKSDGAKTGRVLVEKLDDAA 445
Query: 338 LAGRPDSGKCTLILTQGQYVKDLAMSGLSAVVDPDLYGVFPLSSKLLNATRDSR-KLFKK 396
AG +S CTLILT+G K LA++G S V+ + GVFPL KLLN S ++
Sbjct: 446 EAGGKNSRLCTLILTEGDSAKSLALAGRS-VLGNNYCGVFPLRGKLLNVREASTTQITNN 504
Query: 397 KEIQNLMAILGLVRNKKYSNAKCLRYGQLMVMANQDEDGAHFKGLLINFFYSFWPSLLKV 456
KEI+NL ILGL +N KY N LRYGQ+M+M +QD DG+H KGLLINF +SFWPSLL+V
Sbjct: 505 KEIENLKKILGLKQNMKYENVNSLRYGQMMIMTDQDHDGSHIKGLLINFIHSFWPSLLQV 564
Query: 457 SKFMSVFTFPIIKVSMYNASDSKKG--KELSFDSMQQYEDWKKELGNTANDWEIKYCQGL 514
F+ F PI+K + +KG K LSF SM +YE+WK+ L A W+IKY +GL
Sbjct: 565 PSFLVEFITPIVKAT-------RKGTKKVLSFYSMPEYEEWKESLKGNATGWDIKYYKGL 617
Query: 515 --SSAQEVREYFQDLGKHRAYFVWDDK-DETSIKLAFSKKA-EEWEAWIGNSQPGTCNYE 570
S+A+E +EYF +LG H+ FVW+D+ D +I+LAFSKK E + W+ + PG +
Sbjct: 618 GTSTAEEGKEYFSNLGLHKKDFVWEDEQDGEAIELAFSKKKIEARKNWLSSYVPGNHLDQ 677
Query: 571 RKV-INYRNFVNKELLFSSMKNLQRSIPSVVDGLTLGQRKTLFCSFEMNLTKQTEVCRLT 629
R+ + Y +FVNKEL+ SM +LQRSIPS+VDGL GQRK LF +F+ K+ +V +L
Sbjct: 678 RQPKVTYSDFVNKELILFSMADLQRSIPSMVDGLKPGQRKILFVAFKKIARKEMKVAQLV 737
Query: 630 AYVTEHSYCHYGQQSIASTIIGMAQDFVGSNNINLLIPYGQFGTRDLGGKDHASSRKLYI 689
YV+ S H+G+QS+AS IIGMAQD+VGSNNINLL+P GQFGTR GGKD AS+R ++
Sbjct: 738 GYVSLLSAYHHGEQSLASAIIGMAQDYVGSNNINLLLPNGQFGTRTSGGKDSASARYIFT 797
Query: 690 KLNCVTRLLFPVDDDKLLEYLNEDGRSIQPNWYIPIIPLVLVNGCHAMGTCFSSDIPKYH 749
KL+ VTR+LFP DDD LL+YLNEDG+ I+P WY+PIIP VLVNG +GT +S+ IP Y+
Sbjct: 798 KLSPVTRILFPKDDDLLLDYLNEDGQRIEPTWYMPIIPTVLVNGAEGIGTGWSTFIPNYN 857
Query: 750 PYEIIKNVRRFLNNEEMVSMKPWYKGFGGTIEKSAKGIG--YYTVDGLVERINEQTFRIK 807
P EI+ NVRR LN E MV M PWY+GF GTIEK+A G YT+ GL E ++E T RI
Sbjct: 858 PREIVANVRRLLNGESMVPMDPWYRGFKGTIEKTASKEGGCTYTITGLYEEVDETTIRIT 917
Query: 808 ELPIRMWTEDYKKFLEKI-TARG--LIESFRQKGDYEIVNFKVNVKQEEIATIEKDEELR 864
ELPIR W +DYK FL+ + T G + + D + V+F + + EE + E
Sbjct: 918 ELPIRRWNDDYKNFLQSLKTDNGAPFFQDVKAYNDEKSVDFDL-ILSEENMLAARQEGFL 976
Query: 865 KKFNLTCTISTSNMYLFDAEGKIKKYNTPEQILEEFCTLRLKYYVRRKQYLVKNFTQLLR 924
KKF LT TI+TSNM+LFD +G IKKY TPEQILEEF LR +YY +RK+ +VKN L
Sbjct: 977 KKFKLTTTIATSNMHLFDKKGVIKKYVTPEQILEEFFDLRFEYYEKRKETVVKNMEIELL 1036
Query: 925 SLRIKQKFISNIVNEKFNPINRRAELLIEIL---------KKGKSAEPHVAGAIDYSSKE 975
L K +FI +++ + R+ ++E L +K +S E +AGA+D + E
Sbjct: 1037 KLENKARFILAVLSGEIIVNKRKKADIVEDLRQKGFTPFPRKAESVEAAIAGAVDDDAAE 1096
Query: 976 QEAGEQESVSQSVSIEGATLGDYEYLSSLPFENFTLESLKKLEAELDEKENELKTLEHTS 1035
+ E+ V S +Y+YL ++ + T+E +++L A+ D+ + ++ T+
Sbjct: 1097 EP--EEILVDPESSSSYIPGSEYDYLLAMAIASLTIEKVEELLADRDKMIIAVADMKKTT 1154
Query: 1036 PDLMWLNDLELFEKEFD 1052
P +WL+DLE +KE +
Sbjct: 1155 PKSLWLSDLESLDKELE 1171
>UniRef100_UPI0000430198 UPI0000430198 UniRef100 entry
Length = 1272
Score = 677 bits (1748), Expect = 0.0
Identities = 405/978 (41%), Positives = 575/978 (58%), Gaps = 70/978 (7%)
Query: 124 LCDNGK---WTNVMFKPDLDKFDMKNLEKDVIALMKKRVLDMAVCLGETVTVVLNGTVIP 180
+ DN K WT + FKPDL++F M ++ D AL+ KRV DMA + + V V LN I
Sbjct: 285 ITDNKKGEEWTKITFKPDLERFGMNGIDDDTNALLMKRVYDMAGTVKD-VKVFLNDERIK 343
Query: 181 VKSFKDYANFFLDRA--------------KPFPLPRTHAKLGDRLEICLSLSDGKFQQVS 226
VK+FK Y +L+ + KP P + R E+ +LSDG+F+QVS
Sbjct: 344 VKNFKQYVEMYLNASTTATSEAAGGAAVTKP---PIIYEVANKRWEVAFALSDGEFKQVS 400
Query: 227 FVNSIATIKGGTHVDYITKQITTYIKKEVLKK-KHVNVNADTVKNQLWVFVNALIDNPAF 285
F NSIAT KGGTHVD I Q+ + ++ KK K V +KN +W+FVNALI+NPAF
Sbjct: 401 FTNSIATTKGGTHVDMIATQLANKLMDQIKKKNKAAPVKPFQIKNHMWIFVNALIENPAF 460
Query: 286 NSQTKEMLTTKPARLGLKF-FPGSMLNDVQKSQILDNLLPRIPFKH-----------RTH 333
+SQTKE LT K + G K + V K+ I+DN+L FK R+
Sbjct: 461 DSQTKENLTLKSSAFGSKCELSEEFIKKVAKTGIIDNVLNWARFKQDQILKKTDGTKRSR 520
Query: 334 V-------DAKLAGRPDSGKCTLILTQGQYVKDLAMSGLSAVVDPDLYGVFPLSSKLLNA 386
+ DA AG +S CTLILT+G K LA+SGL AVV D YGVFPL KLLN
Sbjct: 521 ISGIVKLEDANNAGGRNSKNCTLILTEGDSAKALAVSGL-AVVGRDEYGVFPLRGKLLNV 579
Query: 387 TRDSR-KLFKKKEIQNLMAILGLVRNKKYSNAKCLRYGQLMVMANQDEDGAHFKGLLINF 445
++ K EIQ++ ILGL N+ Y+ A LRYG LM+M +QD DG+H KGL+INF
Sbjct: 580 REAGHDQIVKNVEIQHIKQILGLKHNQDYATADSLRYGHLMIMTDQDHDGSHIKGLIINF 639
Query: 446 FYSFWPSLLKVSKFMSVFTFPIIKVSMYNASDSKKGKELSFDSMQQYEDWKKELGNTAND 505
F+PSLL++ F+ F PI+KV K KE+SF +M QYE+WK + N
Sbjct: 640 LDHFYPSLLRIPNFLVEFITPIVKVW-------KGKKEISFYTMPQYEEWKAQ-NNNGRG 691
Query: 506 WEIKYCQGL--SSAQEVREYFQDLGKHR-AYFVWDDKDETSIKLAFSKK-AEEWEAWIGN 561
WE KY +GL S++ + ++YF DL KHR A+ +D I +AF+KK A++ + W+
Sbjct: 692 WESKYYKGLGTSTSADAKKYFSDLDKHRLAFETMQPEDRQLIDMAFNKKKADDRKEWLRQ 751
Query: 562 SQPGT-CNYERKVINYRNFVNKELLFSSMKNLQRSIPSVVDGLTLGQRKTLFCSFEMNLT 620
+PGT +++ + + +FVNKEL+ SM + RSIPSV DGL GQRK LF F+ NLT
Sbjct: 752 FKPGTYLDHDVQSLPISDFVNKELILFSMADNIRSIPSVADGLKPGQRKVLFGCFKRNLT 811
Query: 621 KQTEVCRLTAYVTEHSYCHYGQQSIASTIIGMAQDFVGSNNINLLIPYGQFGTRDLGGKD 680
K+ +V +L YV+E + H+G+ S+ STI+G+AQ+FVGSNNINLL P GQFGTR GGKD
Sbjct: 812 KEIKVAQLAGYVSEKTAYHHGEASLTSTIVGLAQNFVGSNNINLLDPNGQFGTRLAGGKD 871
Query: 681 HASSRKLYIKLNCVTRLLFPVDDDKLLEYLNEDGRSIQPNWYIPIIPLVLVNGCHAMGTC 740
AS+R ++ L +TR +F DD LL YL EDG SI+P++++P +P+VL+NG +GT
Sbjct: 872 AASARYIFTNLPRMTRAIFHPADDGLLNYLVEDGMSIEPDYFMPTVPMVLINGADGIGTG 931
Query: 741 FSSDIPKYHPYEIIKNVRRFLNNEEMVSMKPWYKGFGGTIEKSAKGIGYYTVDGLVERIN 800
+S+ IP Y+P +I++N+RR + EE M PW++GF G IE+ + Y V G++E+IN
Sbjct: 932 WSTAIPNYNPVDIVENLRRMMRGEEPEKMTPWFRGFKGFIERIDQ--DKYKVSGIIEKIN 989
Query: 801 EQTFRIKELPIRMWTEDYKKFLEKIT-----ARGLIESFRQKGDYEIVNFKVNVKQEEIA 855
+ T I ELPIR WT+D+K+ LE++T I+ + + V+F+V++ ++ +A
Sbjct: 990 DTTLEITELPIRRWTQDFKEMLEEMTTGTDKVPQTIKDYEEHHTDTTVHFRVHMTEKNLA 1049
Query: 856 TIEKDEELRKKFNLTCTISTSNMYLFDAEGKIKKYNTPEQILEEFCTLRLKYYVRRKQYL 915
EK E K+F LT ++ T NM FD GKIKKY +PE+ILEEF RL++Y RKQ+L
Sbjct: 1050 DAEK-EGFDKRFKLTTSLGTGNMVCFDLNGKIKKYASPEEILEEFFHKRLEFYGHRKQHL 1108
Query: 916 VKNFTQLLRSLRIKQKFISNIVNEKFNPINRRAELLIEILKKGKSAEPHVAGAIDYSSKE 975
+ L + +F+ I+ ++ +R ++ L+ K +
Sbjct: 1109 ADELNRQFERLSNQARFVHMIITKELVVSGKRRPDIVAELRSLKFK------PFPKKERA 1162
Query: 976 QEAGEQESVSQSVSIEGATLGDYEYLSSLPFENFTLESLKKLEAELDEKENELKTLEHTS 1035
+E GE E+ + + DY+YL + + T E ++KL AE D KE EL L S
Sbjct: 1163 KEEGENEAAEEEEDGDEGQASDYDYLLGMAIWSLTSEKVEKLLAERDAKEQELIELLKLS 1222
Query: 1036 PDLMWLNDLELFEKEFDV 1053
P+ +W DL+ F E+ +
Sbjct: 1223 PNNIWDRDLDNFLAEWQL 1240
>UniRef100_Q6PUA3 DNA topoisomerase type 2 [Tetrahymena thermophila]
Length = 1662
Score = 654 bits (1688), Expect = 0.0
Identities = 391/972 (40%), Positives = 571/972 (58%), Gaps = 65/972 (6%)
Query: 122 VSLCDNGKWTNVMFKPDLDKFDMKNLEKDVIALMKKRVLDMAVCLGETVTVVLNGTVIPV 181
+S +T + F PDL KF MK LEKD++ LMKKRV DMA +G +V V LNG I +
Sbjct: 180 ISSYTESDYTKITFYPDLKKFHMKELEKDIVDLMKKRVYDMAGVIGGSVKVSLNGERISI 239
Query: 182 KSFKDYANFFLDR--AKPFPLPRTHAKLGDRLEICLSLSDGKFQQVSFVNSIATIKGGTH 239
SF DY +F+L A+ + R E+ SLS+G+FQQVSFVNSI T KGGTH
Sbjct: 240 SSFSDYCDFYLKNQGAEVIKVSDNKGTNNQRWEVIASLSEGQFQQVSFVNSICTSKGGTH 299
Query: 240 VDYITKQITTYIKKEVLKK-KHVNVNADTVKNQLWVFVNALIDNPAFNSQTKEMLTTKPA 298
V+YIT QIT + + + KK K +N+ VK+ LW+FVN LI+NP+F++QTKE LT +
Sbjct: 300 VNYITDQITEKLMETLKKKEKSLNIKQFQVKSHLWIFVNCLIENPSFDTQTKETLTLRQQ 359
Query: 299 RLGLKF-FPGSMLNDVQKSQILDNLLPRIPFKHRTHV-------------------DAKL 338
G + ++ KS I++++L + K + + DA
Sbjct: 360 SFGSTCEVSEKFIKELLKSNIINHILLQARAKEQAKMAKTLSAKKQSRLLGVPKLEDAND 419
Query: 339 AGRPDSGKCTLILTQGQYVKDLAMSGLSAVVDPDLYGVFPLSSKLLNATRDS--RKLFKK 396
AG ++ CTLI+T+G K LAMSG+ VV D +GVFPL K+LN RDS +++ +
Sbjct: 420 AGTRNAESCTLIITEGDSAKALAMSGIE-VVGRDKFGVFPLKGKMLNV-RDSTSKQITEN 477
Query: 397 KEIQNLMAILGLVRNKKYSNAKCLRYGQLMVMANQDEDGAHFKGLLINFFYSFWPSLLKV 456
EIQN++ ++G+ N Y N K LRYG +M+MA+QD DG+H KGL INF +FWPSL ++
Sbjct: 478 LEIQNIIKVMGMKLNTNYDNVKSLRYGSIMIMADQDHDGSHIKGLFINFIQNFWPSLFQM 537
Query: 457 SKFMSVFTFPIIKVSMYNASDSKKGKELSFDSMQQYEDWKKELGNTANDWEIKYCQGL-- 514
F+ F PI+KV N S +SF S+Q ++ WK N + W+IKY +GL
Sbjct: 538 DGFLKEFITPIVKVRKGNQS-------ISFFSIQDFKRWKSTCPNQ-HLWKIKYYKGLGT 589
Query: 515 SSAQEVREYFQDLGKHRAYFVWDDKD-ETSIKLAFSKK-AEEWEAWIGNSQPG-TCNYER 571
S+ QE +EYF ++ KH F ++ KD E +I LAF+KK AE+ + W+ P ++ +
Sbjct: 590 STDQEAQEYFGNINKHVISFKYNGKDDEDAIDLAFNKKRAEDRKKWLVKYDPEENVDHSQ 649
Query: 572 KVINYRNFVNKELLFSSMKNLQRSIPSVVDGLTLGQRKTLFCSFEMNLTKQTEVCRLTAY 631
K + Y++FVNKEL+ S + RSIPS+ DGL GQRK LF F+ NL + +V +L+ Y
Sbjct: 650 KHLTYKDFVNKELINFSNSDNARSIPSLCDGLKPGQRKILFACFKRNLKNEIKVAQLSGY 709
Query: 632 VTEHSYCHYGQQSIASTIIGMAQDFVGSNNINLLIPYGQFGTRDLGGKDHASSRKLYIKL 691
V EHS H+G+QS+ STI+GMAQ FVG+NNIN L+P GQFG+R++GGK+ AS+R ++ L
Sbjct: 710 VAEHSAYHHGEQSLESTIVGMAQSFVGTNNINYLMPIGQFGSRNMGGKEAASARYIHTCL 769
Query: 692 NCVTRLLFPVDDDKLLEYLNEDGRSIQPNWYIPIIPLVLVNGCHAMGTCFSSDIPKYHPY 751
N +TR +FP DD +L YL ++G+ ++P +Y+PIIP L+NG +GT +S+ IP Y+P
Sbjct: 770 NKITRCVFPEPDDHILYYLEDEGQQVEPIYYLPIIPTALLNGADGIGTGWSTSIPCYNPR 829
Query: 752 EIIKNVRRFLNNEEMVSMKPWYKGFGGTIEKSAKGIGYYTVDGLVERINEQTFRIKELPI 811
E++ ++ + + + PWYKGF G I + K TV G +NE T I ELPI
Sbjct: 830 ELVDSIEKRMMGVPFTDLLPWYKGFTGEISNNEK--SQITVSGKYHLVNETTIEITELPI 887
Query: 812 RMWTEDYKKFLEKITARGLIESFRQKGDYEI-----------VNFKVNVKQEEIATIEKD 860
+ W DYK FLE+ +ES + K D EI V+F V + ++++ +++
Sbjct: 888 KKWIRDYKNFLEE-----KLESTKDK-DPEIEDIKEYHAGNRVHFVVKMSEKQMEKVQR- 940
Query: 861 EELRKKFNLTCTISTSNMYLFDAEGKIKKYNTPEQILEEFCTLRLKYYVRRKQYLVKNFT 920
E + K F L TI T+NM LF+ EGK+ +Y +I+EEF +RL Y RRK+YLV
Sbjct: 941 EGIDKYFKLQSTIPTTNMVLFNHEGKLTRYANICEIMEEFYEVRLNAYKRRKEYLVSKIE 1000
Query: 921 QLLRSLRIKQKFISNIVNEKFNPIN-RRAELLIEILKKG---KSAEPHVAGAIDYSSKEQ 976
+ L L K++FI ++NE N +R +L+ E+ +G P + + S EQ
Sbjct: 1001 RDLEILSNKKRFILAVINESIRVRNAKRKDLIKELSVQGYVQMKDMPKIKSSKLTSQLEQ 1060
Query: 977 EAGEQESVSQ-SVSIEGATLGDYEYLSSLPFENFTLESLKKLEAELDEKENELKTLEHTS 1035
E + + + +Y YL SLP + T E +++++ E K++EL L T
Sbjct: 1061 EQNQDSNEENFETNPNEINAKEYNYLLSLPMWSLTYEKVEEIKQEQKNKQDELDKLVATE 1120
Query: 1036 PDLMWLNDLELF 1047
+W DL+ F
Sbjct: 1121 VIDIWKEDLKNF 1132
>UniRef100_Q7RJ69 DNA topoisomerase II, putative [Plasmodium yoelii yoelii]
Length = 1489
Score = 653 bits (1684), Expect = 0.0
Identities = 405/1033 (39%), Positives = 586/1033 (56%), Gaps = 128/1033 (12%)
Query: 127 NGK-WTNVMFKPDLDKFDMKNLEKDVIALMKKRVLDMA-VCLGETVTVVLNGTVIPVKSF 184
NGK + V FKPDLDKF M L+ D+ L+ KRV D+A C V V LNGT +PVK F
Sbjct: 193 NGKDYVKVTFKPDLDKFGMSELDDDIECLLHKRVYDLAGTC---NVRVYLNGTRLPVKDF 249
Query: 185 KDYANFFL-DRAKPFPLP---------RTHAKLG---DRLEICLSLSDG----------- 220
K Y + +L D P P T A + D L++ +S +G
Sbjct: 250 KSYVDLYLRDNNTPNTNPSVSKNALNGNTDASVNKTIDNLDVSISHENGGGAGDITPTKT 309
Query: 221 --------------------------------------KFQQVSFVNSIATIKGGTHVDY 242
+FQQVSFVNSI T KGGTHV Y
Sbjct: 310 TEENGNSNFNNNKYEEEIVKIHEKQHRWEIVISKTDGSQFQQVSFVNSICTTKGGTHVSY 369
Query: 243 ITKQITTYIKKEVLKKKH--VNVNADTVKNQLWVFVNALIDNPAFNSQTKEMLTTKPARL 300
I Q+ + K+ K + + A ++N LWVFVN LI NP F+SQTKE LTTK A+
Sbjct: 370 IVDQLLNSLSKKANAKNKGGMEIKAGHIRNHLWVFVNCLIVNPTFDSQTKETLTTKQAKF 429
Query: 301 GLK-FFPGSMLNDVQKSQILDNLLPRIPFKHRTHV----------------------DAK 337
G K +N+V KS IL N+L K + + DA
Sbjct: 430 GSKCTLTDKTINNVLKSSILSNILLWAQAKAQVELRKKMKAGSSKARERIIGIPKLEDAN 489
Query: 338 LAGRPDSGKCTLILTQGQYVKDLAMSGLSAVVDPDLYGVFPLSSKLLNATRDS--RKLFK 395
AG S +CTLILT+G K ++GLS +V D YGVFPL KLLN RD+ ++L
Sbjct: 490 DAGSKYSQECTLILTEGDSAKTSCLAGLS-IVGRDRYGVFPLKGKLLN-VRDASFKQLMD 547
Query: 396 KKEIQNLMAILGL-VRNKKYSNAKCLRYGQLMVMANQDEDGAHFKGLLINFFYSFWPSLL 454
KEIQN+ I+GL + +K + K LRYG LM+M +QD DG+H KGLLIN + FWPSLL
Sbjct: 548 NKEIQNIFKIMGLDITDKNKQDIKGLRYGSLMIMTDQDYDGSHIKGLLINMIHKFWPSLL 607
Query: 455 KVSKFMSVFTFPIIKVSMYNASDSKKGKELSFDSMQQYEDWKKELGNTANDWEIKYCQGL 514
K F+ F PI+KV K +ELSF ++ +YE WK+ W+IKY +GL
Sbjct: 608 KHKGFLCEFVTPIVKV-------QKGNQELSFFTIAEYEQWKENTNLVG--WKIKYYKGL 658
Query: 515 --SSAQEVREYFQDLGKHRAYFVW-DDKDETSIKLAFSKK-AEEWEAWIGNSQPGT-CNY 569
S+ +E ++YF D+ H+ +F+W D+D SI +AFSKK E+ + W+ N G+ ++
Sbjct: 659 GTSTDKEFKQYFSDIQSHKIFFLWTGDRDGDSIDMAFSKKRIEDRKLWLQNFILGSYVDH 718
Query: 570 ERKVINYRNFVNKELLFSSMKNLQRSIPSVVDGLTLGQRKTLFCSFEMNLTKQTEVCRLT 629
+ K ++Y +FVNKEL++ S + +RSIP+++DG GQRK L+ F+ NL + +V +L
Sbjct: 719 KEKDLSYYDFVNKELIYYSRYDTERSIPNIMDGWKPGQRKVLYGCFKRNLKNECKVAQLV 778
Query: 630 AYVTEHSYCHYGQQSIASTIIGMAQDFVGSNNINLLIPYGQFGTRDLGGKDHASSRKLYI 689
Y+ EHS H+G+ S+ TII MAQ FVGSNNIN L P GQFG+R GGKD +++R ++
Sbjct: 779 GYIAEHSAYHHGESSLQQTIINMAQTFVGSNNINFLEPCGQFGSRKEGGKDASAARYIFT 838
Query: 690 KLNCVTRLLFPVDDDKLLEYLNEDGRSIQPNWYIPIIPLVLVNGCHAMGTCFSSDIPKYH 749
KL TR +F DD +L+YLNE+G+ I+P +YIP+IP +LVNGC +GT +SS IP Y+
Sbjct: 839 KLASSTRSIFNEYDDPILKYLNEEGQKIEPQYYIPVIPTILVNGCEGIGTGYSSFIPNYN 898
Query: 750 PYEIIKNVRRFLNNEEMVSMKPWYKGFGGTIEKSAKGIGYYTVDGLVERINEQTFRIKEL 809
+II N++++++ E +V M PWYK F G IE + K GY T+ G++ +I+++T I EL
Sbjct: 899 YKDIINNIKKYIDKEPLVPMVPWYKDFKGRIEPNGKS-GYETI-GIINKIDDETLEITEL 956
Query: 810 PIRMWTEDYKKFLEKITA---RGLIESFRQKGDYEIVNFKVNVKQEEIATIEKDEELRKK 866
PI+ WT+DYK+FLE++ LI + +E V F + + ++ E +E L K
Sbjct: 957 PIKRWTQDYKEFLEELLTDEKNQLIIDYIDNSSHEDVCFTIKMDPIKLKRAE-EEGLEKV 1015
Query: 867 FNLTCTISTSNMYLFDAEGKIKKYNTPEQILEEFCTLRLKYYVRRKQYLVKNFTQLLRSL 926
F L T++T+NM LFD + K+++Y + IL++FC RL Y RK YL+ + R +
Sbjct: 1016 FKLKSTLTTTNMTLFDPKLKLQRYASELDILKDFCFHRLNAYTDRKNYLISKLEKEKRII 1075
Query: 927 RIKQKFISNIVNEKFNPINRRAELLIEILKKGKSAEPHVAGAIDYSSKEQEAGEQESVSQ 986
K KFI I+N + ++ ++L+E L + K +P+ + K++E EQE +
Sbjct: 1076 SNKTKFILAIINGELVVNKKKKKILVEELYR-KGYDPY---KDIHKLKKEEIFEQELLDA 1131
Query: 987 SVS-------IEGATLGDYEYLSSLPFENFTLESLKKLEAELDEKENELKTLEHTSPDLM 1039
+ + I G T+ DY YL ++P + TLE +++L ++L EKE EL+ L+ + + M
Sbjct: 1132 AENPEDNEEIIAGVTVKDYNYLLNMPIFSLTLEKVEELLSQLKEKEQELEILKSITVETM 1191
Query: 1040 WLNDLELFEKEFD 1052
WL D+E E+ +
Sbjct: 1192 WLKDIEKVEEAIE 1204
>UniRef100_UPI00003C1D48 UPI00003C1D48 UniRef100 entry
Length = 1518
Score = 645 bits (1664), Expect = 0.0
Identities = 389/980 (39%), Positives = 564/980 (56%), Gaps = 83/980 (8%)
Query: 127 NGKWTNVMFKPDLDKFDMKNLEKDVIALMKKRVLDMAVCLGETVTVVLNGTVIPVKSFKD 186
N ++T++ FKPDL F M ++ + ALM KRV DMA + + + V LNG ++ +K+FK
Sbjct: 279 NDEYTSITFKPDLALFGMDAIDDHMEALMLKRVYDMAGTV-KGIKVTLNGELLKIKNFKQ 337
Query: 187 YANFFLDRA-----------------------KPFPLPRTHAKLGDRLEICLSLSDGKFQ 223
Y ++ KP + + G E+ ++SDG+F+
Sbjct: 338 YVEMYVSAINELGGKADNGEEDAAGLPTSAAPKPAIIYESTVDKGRNWEVAFAVSDGEFR 397
Query: 224 QVSFVNSIATIKGGTHVDYITKQITTYIKKEVLKKKHVN-VNADTVKNQLWVFVNALIDN 282
QVSFVN+IATIKGG HVD++ Q+ + + + V K KH V + V+ QLWVFVN I N
Sbjct: 398 QVSFVNNIATIKGGKHVDHVADQVISRLIEHVKKDKHTGKVMPNQVRQQLWVFVNCQIVN 457
Query: 283 PAFNSQTKEMLTTKPARLGLKF-FPGSMLNDVQKSQILDNLLPRIPFKH----------- 330
P F+ QTKE LT K + G K+ V KS ++DN++ FK
Sbjct: 458 PTFDGQTKETLTLKQSAFGSKWTLTDEFARKVIKSGVVDNIVSFARFKQDQALKKTDGHK 517
Query: 331 RTHV-------DAKLAGRPDSGKCTLILTQGQYVKDLAMSGLSAVVDPDLYGVFPLSSKL 383
RT + DA AG + CTLILT+G K LA++G+ V D YGVFPL KL
Sbjct: 518 RTRITNIPKLEDANNAGGRKAKDCTLILTEGDSAKSLAVAGI-VEVGRDNYGVFPLRGKL 576
Query: 384 LNATRDSR-KLFKKKEIQNLMAILGLVRNKKYSNAKCLRYGQLMVMANQDEDGAHFKGLL 442
LN S ++ K EI+ + ILGL K+Y + LRYG +M+M +QD DG+H KGL+
Sbjct: 577 LNVREASHDQIMKNAEIKAIKEILGLQHGKQYLDTNSLRYGSIMIMTDQDHDGSHIKGLI 636
Query: 443 INFFYSFWPSLLKVSKFMSVFTFPIIKVSMYNASDSKKGKE-LSFDSMQQYEDWKKELGN 501
INF F+PSLLK+ F+ F PI++ +KGK+ +SF ++ +YE W+ + N
Sbjct: 637 INFLDHFYPSLLKIPGFLVEFITPIVRC--------RKGKQDISFFTIPEYETWR-DAHN 687
Query: 502 TANDWEIKYCQGL--SSAQEVREYFQDLGKHR-AYFVWDDKDETSIKLAFSKK-AEEWEA 557
W IKY +GL S +Q+ + YF+ L KH+ A+ V D D I LAF+KK A++ +
Sbjct: 688 GGKGWHIKYYKGLGTSESQDAKRYFRALEKHKLAFDVTRDGDRELIDLAFNKKKADDRKE 747
Query: 558 WIGNSQPGT-CNYERKVINYRNFVNKELLFSSMKNLQRSIPSVVDGLTLGQRKTLFCSFE 616
W+ +PGT ++ I Y +F+NKEL+ SM + RSIPS +DGL GQRK ++ F+
Sbjct: 748 WLRQFRPGTYLDHNTDRIPYADFINKELILFSMADNLRSIPSAIDGLKPGQRKVMYVMFD 807
Query: 617 MNLTKQTEVCRLTAYVTEHSYCHYGQQSIASTIIGMAQDFVGSNNINLLIPYGQFGTRDL 676
+ +V L V + H+G+ +++ TI+G+AQ+FVGSNN+NLL P GQFGTR L
Sbjct: 808 RKSNSELKVNTLVGNVLGQAAYHHGEAALSMTIVGLAQNFVGSNNVNLLEPRGQFGTRLL 867
Query: 677 GGKDHASSRKLYIKLNCVTRLLFPVDDDKLLEYLNEDGRSIQPNWYIPIIPLVLVNGCHA 736
GGKD AS R ++ L + R LFP DD LL+YL +DG+ ++P+WYIP++P VL+NG
Sbjct: 868 GGKDAASGRYIFTALPKIARALFPATDDALLDYLEDDGQKVEPHWYIPVVPQVLLNGAEG 927
Query: 737 MGTCFSSDIPKYHPYEIIKNVRRFLNNEEMVSMKPWYKGFGGTIEKSAKGIGYYTVDGLV 796
+GT +S+ +P Y+P+EI+ N+RR + E++V MKPWY+GF G +E+ G G + + G V
Sbjct: 928 IGTGWSTSVPNYNPHEIVANLRRRMAGEDVVPMKPWYRGFNGDVEEF--GPGRFKISGRV 985
Query: 797 ERINEQTFRIKELPIRMWTEDYKKFLE-------KITARGLIESFRQKGDYEIVNFKVNV 849
+I+E+T+ I ELPIR WT +YK+ LE KI A I+ +++ VNF V +
Sbjct: 986 TQIDEKTYEITELPIRTWTSNYKELLEERIVSSDKIPA--TIKDYKEYHTESTVNFIVEL 1043
Query: 850 KQEEIATIEKDEELRKKFNLTCTISTSNMYLFDAEGKIKKYNTPEQILEEFCTLRLKYYV 909
+ A I D+ + F L+C IST+NM LFD +GKIKKY + E+ILEEF LRL YY
Sbjct: 1044 TAKGQAEI-ADKGVEAFFKLSCQISTTNMVLFDQDGKIKKYTSAEEILEEFYLLRLSYYQ 1102
Query: 910 RRKQYLVKNFTQLLRSLRIKQKFISNIVNEKFNPINR-RAELLIEILKKGKSAEPHVAGA 968
+RK+YLV + L + +F+ I++ + NR RA+++ E+ +K P A
Sbjct: 1103 KRKEYLVDQLKLMYDKLSNQARFVQMIISRQLVVSNRKRADIVAELRQKDFRPFPKQGKA 1162
Query: 969 -IDYSSKEQEAGEQESVSQSVSIEGATLGDYEYLSSLPFENFTLESLKKLEAELDEKENE 1027
I ++ EA + E +S D++YL ++ N T E + KL E D KE E
Sbjct: 1163 SIAAEPEDDEAIDDEDISAD--------SDFDYLLNMAIYNLTKEKVDKLLKERDHKEEE 1214
Query: 1028 LKTLEHTSPDLMWLNDLELF 1047
LK L SP +W DL F
Sbjct: 1215 LKILIGRSPQNLWDEDLTKF 1234
>UniRef100_Q8X1U1 DNA topoisomerase II [Penicillium marneffei]
Length = 1162
Score = 637 bits (1642), Expect = 0.0
Identities = 387/966 (40%), Positives = 561/966 (58%), Gaps = 60/966 (6%)
Query: 130 WTNVMFKPDLDKFDMKNLEKDVIALMKKRVLDMAVCLGETVTVVLNGTVIPVKSFKDYAN 189
+T V FKPD KF M ++ D AL+K+RV D+A V V LNG+ IPV++FK Y
Sbjct: 203 YTKVTFKPDYQKFGMDGMDDDFEALVKRRVYDLAGTA--KVAVKLNGSRIPVRNFKKYME 260
Query: 190 FFLDRAKPFPLPRTHAKLGDRLEIC-----------LSLSDGKFQQVSFVNSIATIKGGT 238
+ K R + D+ EI ++SDG FQQVSFVNSIAT GG+
Sbjct: 261 MY---TKAIKKERGEEAVNDKSEIVTASPDPRWEVGFAVSDGSFQQVSFVNSIATTSGGS 317
Query: 239 HVDYITKQITTYIKKEVLKKKH--VNVNADTVKNQLWVFVNALIDNPAFNSQTKEMLTTK 296
HV+YI QI + + V KK + A ++N +++FVNALI NPAF SQTKE LTTK
Sbjct: 318 HVNYIADQICNRLAEAVKKKNKQGATLKASQIRNHIFIFVNALIVNPAFTSQTKEQLTTK 377
Query: 297 PARLGLK-FFPGSMLNDVQKSQILDNLLP-----------------RIPFKHRTHVDAKL 338
P++ G K V K++++ N+L R + DA
Sbjct: 378 PSQFGSKCILDDDFYKKVLKTEVMTNILHFAEKKADQLLKKTDGGRRSRMNNPKLTDANK 437
Query: 339 AGRPDSGKCTLILTQGQYVKDLAMSGLSAVVDPDLYGVFPLSSKLLNATRDSR--KLFKK 396
AG D CTLILT+G K LAM+G AVV PDL+GVFPL KLLN RD+ ++ K
Sbjct: 438 AGTKDGHHCTLILTEGDSAKGLAMAG-RAVVGPDLFGVFPLRGKLLNV-RDASVDQISKN 495
Query: 397 KEIQNLMAILGLVRNKKYSNAKCLRYGQLMVMANQDEDGAHFKGLLINFFYSFWPSLLKV 456
EIQN+ + +GL K+Y++ + LRYG LM+M +QD DG+H KGLLINF +PSLLK+
Sbjct: 496 AEIQNIKSFIGLQHKKEYTDTRGLRYGHLMIMTDQDHDGSHIKGLLINFLQVQFPSLLKI 555
Query: 457 SKFMSVFTFPIIKVSMYNASDSKKGKELSFDSMQQYEDWKKELGNTANDWEIKYCQGL-- 514
+F+ F PIIKV + + K K SF +M +YE WK+E N + W+ KY +GL
Sbjct: 556 PEFLIEFITPIIKVWKGDPKNPTKSK--SFFTMPEYEQWKEENKNDLS-WQHKYYKGLGT 612
Query: 515 SSAQEVREYFQDLGKH-RAYFVWDDKDETSIKLAFSKK-AEEWEAWIGNSQPGT-CNYER 571
S+ ++ + YF+DL +H + + D + I LAFSKK A+E + W+ +PGT ++
Sbjct: 613 STTEDAQIYFRDLDRHLKEFHTMQDHEAQLIDLAFSKKKADERKEWLRQYKPGTFLDHST 672
Query: 572 KVINYRNFVNKELLFSSMKNLQRSIPSVVDGLTLGQRKTLFCSFEMNLTKQTEVCRLTAY 631
K I Y +F+NKEL+ SM + QRSIPSVVDGL GQRK L+ F+ NL K +V L +
Sbjct: 673 KQITYTDFINKELILFSMADNQRSIPSVVDGLKPGQRKVLYTCFKRNLKKDMKVVELAGH 732
Query: 632 VTEHSYCHYGQQSIASTIIGMAQDFVGSNNINLLIPYGQFGTRDLGGKDHASSRKLYIKL 691
V+ + +G+ S+ TI+G+AQ FVGSNN+N L P G FG+R GG D AS+R +Y +L
Sbjct: 733 VSGMTAYQHGETSLQQTIVGLAQTFVGSNNLNCLEPSGNFGSRLQGGADCASARYIYTRL 792
Query: 692 NCVTRLLFPVDDDKLLEYLNEDGRSIQPNWYIPIIPLVLVNGCHAMGTCFSSDIPKYHPY 751
+ R +F D+ LL Y +DG I+P Y+P++PL+L+NG +GT +SS IP Y+P
Sbjct: 793 SPFARRVFNQQDEPLLTYNEDDGEKIEPELYVPVVPLILINGADGIGTGWSSSIPNYNPE 852
Query: 752 EIIKNVRRFLNNEEMVSMKPWYKGFGGTIEKSAKGIGYYTVDGLVERINEQTFRIKELPI 811
+I+ N++R ++ EE M+PW++GF G E + +G Y G + +N++ I ELPI
Sbjct: 853 DIVANLKRMMDGEEPEPMQPWFRGFNG--EVTPQGPDRYKFSGRIHELNDKEVEITELPI 910
Query: 812 RMWTEDYKKFLEKI----TARGLIESFRQKGDYEIVNFKVNVKQEEI-ATIEKDEELRKK 866
R WT+D+K LE I I+ ++ + V+F + + ++ + A +E E L +K
Sbjct: 911 RTWTQDFKDKLEDIIKAEKTPSFIKDYKDYNTHTKVHFVIQMDEKHLKAAVE--EGLEEK 968
Query: 867 FNLTCTISTSNMYLFDAEGKIKKYNTPEQILEEFCTLRLKYYVRRKQYLVKNFTQLLRSL 926
F L+ TI+T+N+ FD EG+I KY T IL+EF +RLKYY RRKQ+ + + L
Sbjct: 969 FKLSRTIATTNLVAFDPEGRITKYATVSDILKEFFAVRLKYYERRKQHQLSELQKQLDKF 1028
Query: 927 RIKQKFISNIVNEKF-NPINRRAELLIEILKKGKSAEPHVAGAIDYSSKEQEAGEQESVS 985
+ +FI I++ K ++A L+ E+ +KG P VA A+ EQ E+ES S
Sbjct: 1029 TNQARFIQMIIDGKLVVSKKKKATLITELKEKGFKPFPKVAEAVKAGETEQVVEEEES-S 1087
Query: 986 QSVSIEGATLGDYEYLSSLPFENFTLESLKKLEAELDEKENELKTLEHTSPDLMWLNDLE 1045
+ + E A+ Y+YL + + T E ++KL ++ +KE E+ L S + +W DL+
Sbjct: 1088 EEANTEMAS-NAYDYLLGMAIWSLTQERVEKLLRQIGDKEVEIDELIKLSKEEIWRRDLD 1146
Query: 1046 LFEKEF 1051
F E+
Sbjct: 1147 DFINEW 1152
>UniRef100_Q8ILC8 DNA topoisomerase II, putative [Plasmodium falciparum]
Length = 1472
Score = 632 bits (1630), Expect = e-179
Identities = 371/887 (41%), Positives = 536/887 (59%), Gaps = 62/887 (6%)
Query: 210 RLEICLSLSDG-KFQQVSFVNSIATIKGGTHVDYITKQITTYIKKEVLKKKH--VNVNAD 266
R EI +S SDG +FQQVSFVNSI T KGG+HV+YI +Q+ + + K+ K + + +
Sbjct: 330 RWEIVVSKSDGSQFQQVSFVNSICTTKGGSHVNYIVEQLLSSLSKKANAKNKGGMEIKSG 389
Query: 267 TVKNQLWVFVNALIDNPAFNSQTKEMLTTKPARLGLK-FFPGSMLNDVQKSQILDNLLPR 325
++N LWVFVN LI NP F+SQTKE LTTKP + G K +N+V KS IL N+L
Sbjct: 390 HIRNHLWVFVNCLIVNPTFDSQTKETLTTKPVKFGSKCILSDKTINNVLKSPILSNILLW 449
Query: 326 IPFKHRTHV----------------------DAKLAGRPDSGKCTLILTQGQYVKDLAMS 363
K + + DA AG S +CTLILT+G K ++
Sbjct: 450 AQAKAQVELKKKMKAGSSKARERIIGIPKLEDANDAGSKYSQECTLILTEGDSAKTSCLA 509
Query: 364 GLSAVVDPDLYGVFPLSSKLLNATRDS-RKLFKKKEIQNLMAILGL-VRNKKYSNAKCLR 421
GLS +V D YGVFPL KLLN S ++L KEIQN+ I+GL + +K + K LR
Sbjct: 510 GLS-IVGRDKYGVFPLKGKLLNVRDASFKQLMDNKEIQNIFRIMGLDITDKNKDDIKGLR 568
Query: 422 YGQLMVMANQDEDGAHFKGLLINFFYSFWPSLLKVSKFMSVFTFPIIKVSMYNASDSKKG 481
YG LM+M +QD DG+H KGLLIN + FWPSLLK F+S F PI+KV K
Sbjct: 569 YGSLMIMTDQDYDGSHIKGLLINMIHKFWPSLLKHKGFLSEFVTPIVKVQ-------KGS 621
Query: 482 KELSFDSMQQYEDWKKELGNTANDWEIKYCQGL--SSAQEVREYFQDLGKHRAYFVWD-D 538
+E SF ++ +YE WK+ W+IKY +GL S+ +E ++YF D+ H+ F+W D
Sbjct: 622 QEYSFFTIAEYEQWKENTNLLG--WKIKYYKGLGTSTDREFKQYFSDIKNHKIMFLWTGD 679
Query: 539 KDETSIKLAFSKKA-EEWEAWIGNSQPGT-CNYERKVINYRNFVNKELLFSSMKNLQRSI 596
+D SI +AFSKK E+ + W+ N G+ +++ K ++Y +FVNKEL++ S + +RSI
Sbjct: 680 RDGDSIDMAFSKKRIEDRKLWLQNFILGSYVDHKEKDLSYYDFVNKELIYYSRYDTERSI 739
Query: 597 PSVVDGLTLGQRKTLFCSFEMNLTKQTEVCRLTAYVTEHSYCHYGQQSIASTIIGMAQDF 656
P+++DG GQRK L+ F+ NL + +V +L Y+ EHS H+G+ S+ TII MAQ F
Sbjct: 740 PNIMDGWKPGQRKVLYGCFKRNLRNECKVAQLVGYIAEHSAYHHGESSLQQTIINMAQTF 799
Query: 657 VGSNNINLLIPYGQFGTRDLGGKDHASSRKLYIKLNCVTRLLFPVDDDKLLEYLNEDGRS 716
VGSNNIN L P GQFG+R GGKD +++R ++ KL TR +F DD +L+YLNE+G+
Sbjct: 800 VGSNNINFLEPCGQFGSRKEGGKDASAARYIFTKLASSTRSIFNEYDDPILKYLNEEGQK 859
Query: 717 IQPNWYIPIIPLVLVNGCHAMGTCFSSDIPKYHPYEIIKNVRRFLNNEEMVSMKPWYKGF 776
I+P +YIP+IP +LVNGC +GT +SS IP Y+ +II N++R++N E ++ M PWYK F
Sbjct: 860 IEPQYYIPVIPTILVNGCEGIGTGYSSFIPNYNYKDIIDNIKRYINKEPLIPMVPWYKDF 919
Query: 777 GGTIEKSAKGIGYYTVDGLVERINEQTFRIKELPIRMWTEDYKKFLEKITA---RGLIES 833
G IE + K GY T+ G++ +I+ T I ELPI+ WT+DYK+FLE++ LI
Sbjct: 920 KGRIESNGK-TGYETI-GIINKIDNDTLEITELPIKKWTQDYKEFLEELLTDEKHQLILD 977
Query: 834 FRQKGDYEIVNFKVNVKQEEIATIEKDEELRKKFNLTCTISTSNMYLFDAEGKIKKYNTP 893
+ +E + F + + ++ E +E L K F L T++T+NM LFD K+++Y+T
Sbjct: 978 YIDNSSHEDICFTIKMDPAKLQKAE-EEGLEKVFKLKSTLTTTNMTLFDPNLKLQRYSTE 1036
Query: 894 EQILEEFCTLRLKYYVRRKQYLVKNFTQLLRSLRIKQKFISNIVNEKFNPINRRAELLIE 953
IL+EFC RLK Y RK YL+ + R + K KFI IVN + ++ ++L+E
Sbjct: 1037 LDILKEFCYQRLKAYENRKSYLISKLEKEKRIISNKTKFILAIVNNELIVNKKKKKVLVE 1096
Query: 954 ILKKGKSAEPHVAGAIDYSS-KEQEAGEQESVSQSVS-------IEGATLGDYEYLSSLP 1005
L + K +P+ D + K++E EQE + + + I G T+ DY+YL S+P
Sbjct: 1097 ELYR-KGYDPYK----DINKIKKEEIFEQELLDAADNPEDNEEIIAGITVKDYDYLLSMP 1151
Query: 1006 FENFTLESLKKLEAELDEKENELKTLEHTSPDLMWLNDLELFEKEFD 1052
+ TLE ++ L +L EKE EL+ L + + + MWL D+E E+ +
Sbjct: 1152 IFSLTLEKVEDLLTQLKEKERELEILRNITVETMWLKDIEKVEEAIE 1198
Score = 54.3 bits (129), Expect = 2e-05
Identities = 31/68 (45%), Positives = 43/68 (62%), Gaps = 5/68 (7%)
Query: 127 NGK-WTNVMFKPDLDKFDMKNLEKDVIALMKKRVLDMA-VCLGETVTVVLNGTVIPVKSF 184
NGK + V FKPDL+KF M ++ D+ +L+ KRV D+A C +V V LNG + VK F
Sbjct: 193 NGKDYVKVTFKPDLNKFGMTEMDDDIESLLFKRVYDLAGTC---SVRVYLNGQRLAVKDF 249
Query: 185 KDYANFFL 192
K Y + +L
Sbjct: 250 KSYVDLYL 257
>UniRef100_Q7Z2D0 DNA topoisomerase [Plasmodium falciparum]
Length = 1397
Score = 632 bits (1630), Expect = e-179
Identities = 371/887 (41%), Positives = 536/887 (59%), Gaps = 62/887 (6%)
Query: 210 RLEICLSLSDG-KFQQVSFVNSIATIKGGTHVDYITKQITTYIKKEVLKKKH--VNVNAD 266
R EI +S SDG +FQQVSFVNSI T KGG+HV+YI +Q+ + + K+ K + + +
Sbjct: 330 RWEIVVSKSDGSQFQQVSFVNSICTTKGGSHVNYIVEQLLSSLSKKANAKNKGGMEIKSG 389
Query: 267 TVKNQLWVFVNALIDNPAFNSQTKEMLTTKPARLGLK-FFPGSMLNDVQKSQILDNLLPR 325
++N LWVFVN LI NP F+SQTKE LTTKP + G K +N+V KS IL N+L
Sbjct: 390 HIRNHLWVFVNCLIVNPTFDSQTKETLTTKPVKFGSKCILSDKTINNVLKSPILSNILLW 449
Query: 326 IPFKHRTHV----------------------DAKLAGRPDSGKCTLILTQGQYVKDLAMS 363
K + + DA AG S +CTLILT+G K ++
Sbjct: 450 AQAKAQVELKKKMKAGSSKARERIIGIPKLEDANDAGSKYSQECTLILTEGDSAKTSCLA 509
Query: 364 GLSAVVDPDLYGVFPLSSKLLNATRDS-RKLFKKKEIQNLMAILGL-VRNKKYSNAKCLR 421
GLS +V D YGVFPL KLLN S ++L KEIQN+ I+GL + +K + K LR
Sbjct: 510 GLS-IVGRDKYGVFPLKGKLLNVRDASFKQLMDNKEIQNIFRIMGLDITDKNKDDIKGLR 568
Query: 422 YGQLMVMANQDEDGAHFKGLLINFFYSFWPSLLKVSKFMSVFTFPIIKVSMYNASDSKKG 481
YG LM+M +QD DG+H KGLLIN + FWPSLLK F+S F PI+KV K
Sbjct: 569 YGSLMIMTDQDYDGSHIKGLLINMIHKFWPSLLKHKGFLSEFVTPIVKVQ-------KGS 621
Query: 482 KELSFDSMQQYEDWKKELGNTANDWEIKYCQGL--SSAQEVREYFQDLGKHRAYFVWD-D 538
+E SF ++ +YE WK+ W+IKY +GL S+ +E ++YF D+ H+ F+W D
Sbjct: 622 QEYSFFTIAEYEQWKENTNLLG--WKIKYYKGLGTSTDREFKQYFSDIKNHKIMFLWTGD 679
Query: 539 KDETSIKLAFSKKA-EEWEAWIGNSQPGT-CNYERKVINYRNFVNKELLFSSMKNLQRSI 596
+D SI +AFSKK E+ + W+ N G+ +++ K ++Y +FVNKEL++ S + +RSI
Sbjct: 680 RDGDSIDMAFSKKRIEDRKLWLQNFILGSYVDHKEKDLSYYDFVNKELIYYSRYDTERSI 739
Query: 597 PSVVDGLTLGQRKTLFCSFEMNLTKQTEVCRLTAYVTEHSYCHYGQQSIASTIIGMAQDF 656
P+++DG GQRK L+ F+ NL + +V +L Y+ EHS H+G+ S+ TII MAQ F
Sbjct: 740 PNIMDGWKPGQRKVLYGCFKRNLRNECKVAQLVGYIAEHSAYHHGESSLQQTIINMAQTF 799
Query: 657 VGSNNINLLIPYGQFGTRDLGGKDHASSRKLYIKLNCVTRLLFPVDDDKLLEYLNEDGRS 716
VGSNNIN L P GQFG+R GGKD +++R ++ KL TR +F DD +L+YLNE+G+
Sbjct: 800 VGSNNINFLEPCGQFGSRKEGGKDASAARYIFTKLASSTRSIFNEYDDPILKYLNEEGQK 859
Query: 717 IQPNWYIPIIPLVLVNGCHAMGTCFSSDIPKYHPYEIIKNVRRFLNNEEMVSMKPWYKGF 776
I+P +YIP+IP +LVNGC +GT +SS IP Y+ +II N++R++N E ++ M PWYK F
Sbjct: 860 IEPQYYIPVIPTILVNGCEGIGTGYSSFIPNYNYKDIIDNIKRYINKEPLIPMVPWYKDF 919
Query: 777 GGTIEKSAKGIGYYTVDGLVERINEQTFRIKELPIRMWTEDYKKFLEKITA---RGLIES 833
G IE + K GY T+ G++ +I+ T I ELPI+ WT+DYK+FLE++ LI
Sbjct: 920 KGRIESNGK-TGYETI-GIINKIDNDTLEITELPIKKWTQDYKEFLEELLTDEKHQLILD 977
Query: 834 FRQKGDYEIVNFKVNVKQEEIATIEKDEELRKKFNLTCTISTSNMYLFDAEGKIKKYNTP 893
+ +E + F + + ++ E +E L K F L T++T+NM LFD K+++Y+T
Sbjct: 978 YIDNSSHEDICFTIKMDPAKLQKAE-EEGLEKVFKLKSTLTTTNMTLFDPNLKLQRYSTE 1036
Query: 894 EQILEEFCTLRLKYYVRRKQYLVKNFTQLLRSLRIKQKFISNIVNEKFNPINRRAELLIE 953
IL+EFC RLK Y RK YL+ + R + K KFI IVN + ++ ++L+E
Sbjct: 1037 LDILKEFCYQRLKAYENRKSYLISKLEKEKRIISNKTKFILAIVNNELIVNKKKKKVLVE 1096
Query: 954 ILKKGKSAEPHVAGAIDYSS-KEQEAGEQESVSQSVS-------IEGATLGDYEYLSSLP 1005
L + K +P+ D + K++E EQE + + + I G T+ DY+YL S+P
Sbjct: 1097 ELYR-KGYDPYK----DINKIKKEEIFEQELLDAADNPEDNEEIIAGITVKDYDYLLSMP 1151
Query: 1006 FENFTLESLKKLEAELDEKENELKTLEHTSPDLMWLNDLELFEKEFD 1052
+ TLE ++ L +L EKE EL+ L + + + MWL D+E E+ +
Sbjct: 1152 IFSLTLEKVEDLLTQLKEKERELEILRNITVETMWLKDIEKVEEAIE 1198
Score = 54.3 bits (129), Expect = 2e-05
Identities = 31/68 (45%), Positives = 43/68 (62%), Gaps = 5/68 (7%)
Query: 127 NGK-WTNVMFKPDLDKFDMKNLEKDVIALMKKRVLDMA-VCLGETVTVVLNGTVIPVKSF 184
NGK + V FKPDL+KF M ++ D+ +L+ KRV D+A C +V V LNG + VK F
Sbjct: 193 NGKDYVKVTFKPDLNKFGMTEMDDDIESLLFKRVYDLAGTC---SVRVYLNGQRLAVKDF 249
Query: 185 KDYANFFL 192
K Y + +L
Sbjct: 250 KSYVDLYL 257
>UniRef100_P41001 DNA topoisomerase II [Plasmodium falciparum]
Length = 1398
Score = 630 bits (1626), Expect = e-179
Identities = 372/888 (41%), Positives = 537/888 (59%), Gaps = 63/888 (7%)
Query: 210 RLEICLSLSDG-KFQQVSFVNSIATIKGGTHVDYITKQITTYIKKEVLKKKH--VNVNAD 266
R EI +S SDG +FQQVSFVNSI T KGG+HV+YI +Q+ + + K+ K + + +
Sbjct: 330 RWEIVVSKSDGSQFQQVSFVNSICTTKGGSHVNYIVEQLLSSLSKKANAKNKGGMEIKSG 389
Query: 267 TVKNQLWVFVNALIDNPAFNSQTKEMLTTKPARLGLK-FFPGSMLNDVQKSQILDNLLPR 325
++N LWVFVN LI NP F+SQTKE LTTKP + G K +N+V KS IL N+L
Sbjct: 390 HIRNHLWVFVNCLIVNPTFDSQTKETLTTKPVKFGSKCILSDKTINNVLKSPILSNILLW 449
Query: 326 IPFKHRTHV----------------------DAKLAGRPDSGKCTLILTQGQYVKDLAMS 363
K + + DA AG S +CTLILT+G K ++
Sbjct: 450 AQAKAQVELKKKMKAGSSKARERIIGIPKLEDANDAGSKYSQECTLILTEGDSAKTSCLA 509
Query: 364 GLSAVVDPDLYGVFPLSSKLLNATRDS-RKLFKKKEIQNLMAILGL-VRNKKYSNAKCLR 421
GLS +V D YGVFPL KLLN S ++L KEIQN+ I+GL + +K + K LR
Sbjct: 510 GLS-IVGRDKYGVFPLKGKLLNVRDASFKQLMDNKEIQNIFRIMGLDITDKNKDDIKGLR 568
Query: 422 YGQLMVMANQDEDGAHFKGLLINFFYSFWPSLLKVSKFMSVFTFPIIKVSMYNASDSKKG 481
YG LM+M +QD DG+H KGLLIN + FWPSLLK F+S F PI+KV K
Sbjct: 569 YGSLMIMTDQDYDGSHIKGLLINMIHKFWPSLLKHKGFLSEFVTPIVKVQ-------KGS 621
Query: 482 KELSFDSMQQYEDWKKELGNTANDWEIKYCQGL--SSAQEVREYFQDLGKHRAYFVWD-D 538
+E SF ++ +YE WK+ W+IKY +GL S+ +E ++YF D+ H+ F+W D
Sbjct: 622 QEYSFFTIAEYEQWKENTNLLG--WKIKYYKGLGTSTDREFKQYFSDIKNHKIMFLWTGD 679
Query: 539 KDETSIKLAFSKKA-EEWEAWIGNSQPGT-CNYERKVINYRNFVNKELLFSSMKNLQRSI 596
+D SI +AFSKK E+ + W+ N G+ +++ K ++Y +FVNKEL++ S + +RSI
Sbjct: 680 RDGDSIDMAFSKKRIEDRKLWLQNFILGSYVDHKEKDLSYYDFVNKELIYYSRYDTERSI 739
Query: 597 PSVVDGLTLGQRKTLFCSFEMNLTKQTEVCRLTAYVTEHS-YCHYGQQSIASTIIGMAQD 655
P+++DG GQRK L+ F+ NL + +V +L Y+ EHS Y H+G+ S+ TII MAQ
Sbjct: 740 PNIMDGWKPGQRKVLYGCFKRNLRNECKVAQLVGYIAEHSAYHHHGESSLQQTIINMAQT 799
Query: 656 FVGSNNINLLIPYGQFGTRDLGGKDHASSRKLYIKLNCVTRLLFPVDDDKLLEYLNEDGR 715
FVGSNNIN L P GQFG+R GGKD +++R ++ KL TR +F DD +L+YLNE+G+
Sbjct: 800 FVGSNNINFLEPCGQFGSRKEGGKDASAARYIFTKLASSTRSIFNEYDDPILKYLNEEGQ 859
Query: 716 SIQPNWYIPIIPLVLVNGCHAMGTCFSSDIPKYHPYEIIKNVRRFLNNEEMVSMKPWYKG 775
I+P +YIP+IP +LVNGC +GT +SS IP Y+ +II N++R++N E ++ M PWYK
Sbjct: 860 KIEPQYYIPVIPTILVNGCEGIGTGYSSFIPNYNYKDIIDNIKRYINKEPLIPMVPWYKD 919
Query: 776 FGGTIEKSAKGIGYYTVDGLVERINEQTFRIKELPIRMWTEDYKKFLEKITA---RGLIE 832
F G IE + K GY T+ G++ +I+ T I ELPI+ WT+DYK+FLE++ LI
Sbjct: 920 FKGRIESNGK-TGYETI-GIINKIDNDTLEITELPIKKWTQDYKEFLEELLTDEKHQLIL 977
Query: 833 SFRQKGDYEIVNFKVNVKQEEIATIEKDEELRKKFNLTCTISTSNMYLFDAEGKIKKYNT 892
+ +E + F + + ++ E +E L K F L T++T+NM LFD K+++Y+T
Sbjct: 978 DYIDNSSHEDICFTIKMDPAKLQKAE-EEGLEKVFKLKSTLTTTNMTLFDPNLKLQRYST 1036
Query: 893 PEQILEEFCTLRLKYYVRRKQYLVKNFTQLLRSLRIKQKFISNIVNEKFNPINRRAELLI 952
IL+EFC RLK Y RK YL+ + R + K KFI IVN + ++ ++L+
Sbjct: 1037 ELDILKEFCYQRLKAYENRKSYLISKLEKEKRIISNKTKFILAIVNNELIVNKKKKKVLV 1096
Query: 953 EILKKGKSAEPHVAGAIDYSS-KEQEAGEQESVSQSVS-------IEGATLGDYEYLSSL 1004
E L + K +P+ D + K++E EQE + + + I G T+ DY+YL S+
Sbjct: 1097 EELYR-KGYDPYK----DINKIKKEEIFEQELLDAADNPEDNEEIIAGITVKDYDYLLSM 1151
Query: 1005 PFENFTLESLKKLEAELDEKENELKTLEHTSPDLMWLNDLELFEKEFD 1052
P + TLE ++ L +L EKE EL+ L + + + MWL D+E E+ +
Sbjct: 1152 PIFSLTLEKVEDLLTQLKEKERELEILRNITVETMWLKDIEKVEEAIE 1199
Score = 54.3 bits (129), Expect = 2e-05
Identities = 31/68 (45%), Positives = 43/68 (62%), Gaps = 5/68 (7%)
Query: 127 NGK-WTNVMFKPDLDKFDMKNLEKDVIALMKKRVLDMA-VCLGETVTVVLNGTVIPVKSF 184
NGK + V FKPDL+KF M ++ D+ +L+ KRV D+A C +V V LNG + VK F
Sbjct: 193 NGKDYVKVTFKPDLNKFGMTEMDDDIESLLFKRVYDLAGTC---SVRVYLNGQRLAVKDF 249
Query: 185 KDYANFFL 192
K Y + +L
Sbjct: 250 KSYVDLYL 257
>UniRef100_Q6PUA4 DNA topoisomerase type 2 [Tetrahymena thermophila]
Length = 1402
Score = 629 bits (1622), Expect = e-178
Identities = 379/967 (39%), Positives = 564/967 (58%), Gaps = 68/967 (7%)
Query: 128 GKWTNVMFKPDLDKFDMKNLEKDVIALMKKRVLDMAVCLGETVTVVLNGTVIPVKSFKDY 187
G +T V F PDL KF M+N+ D+ L++KRV DMA V V LN I +KSF Y
Sbjct: 195 GNFTCVTFYPDLSKFKMQNISDDMANLLRKRVYDMAGIFNNKVKVYLNNEQIKIKSFSQY 254
Query: 188 ANFFLDRAKPFPLPRTHAKLGDRLEICLSLSDGKFQQVSFVNSIATIKGGTHVDYITKQI 247
N F+ + DR EI ++ SDG+F+QVSFVN I T +GG HVD++ +I
Sbjct: 255 VNLFIPDIEEQYKCYDPTMTSDRWEILVTFSDGQFKQVSFVNGICTSRGGKHVDFVIDKI 314
Query: 248 TTYIKKEVLKK-KHVNVNADTVKNQLWVFVNALIDNPAFNSQTKEMLTTKPARLGLK-FF 305
+++++LKK K +++ +K L++F+N LI NP+F+SQTK+ LTTK + G K
Sbjct: 315 VERMQEQILKKNKDLDIKPHQIKQNLFIFMNCLIVNPSFDSQTKDTLTTKASAFGSKPEL 374
Query: 306 PGSMLNDVQKSQILDNLLPRIPFKHRTHV-------------------DAKLAGRPDSGK 346
+ + + I++ ++ + + + DA AG S
Sbjct: 375 SEKFIKKILGTGIIEQIVSQAKARQNAKLAKTLKGGKKGRITGIPKLDDANDAGTARSEN 434
Query: 347 CTLILTQGQYVKDLAMSGLSAVVDPDLYGVFPLSSKLLNATRDSRK-LFKKKEIQNLMAI 405
CTLILT+G K LAM+G+ VV D YGVFPL KLLN ++K + + +EIQN+ I
Sbjct: 435 CTLILTEGDSAKALAMAGIE-VVGRDNYGVFPLKGKLLNVREANKKQIIENQEIQNICKI 493
Query: 406 LGL---VRNKKYSNAKCLRYGQLMVMANQDEDGAHFKGLLINFFYSFWPSLLKVSKFMSV 462
+GL V+N Y N K LRYG LM+MA+QD DG+H KGL+INF + FWPSL K+ F+
Sbjct: 494 IGLDPAVQN--YENTKQLRYGSLMIMADQDSDGSHIKGLVINFIHHFWPSLFKMKGFLKE 551
Query: 463 FTFPIIKVSMYNASDSKKGKEL-SFDSMQQYEDWKKELGNTANDWEIKYCQGL--SSAQE 519
F PI+KV+ KG E+ SF ++Q ++++ + G+ +++KY +GL S+ +E
Sbjct: 552 FITPILKVT--------KGDEVHSFFTIQDFKNFAAQKGDRLKQYKVKYYKGLGTSTDKE 603
Query: 520 VREYFQDLGKHRAYFVW-DDKDETSIKLAFS-KKAEEWEAWIGNSQPGT-CNYERKVINY 576
+EYF + KH+ F + D+ D +I+L FS KK +E + W+ N P + +Y K +
Sbjct: 604 AKEYFSNFYKHQINFQYLDENDMEAIELGFSGKKIKERKDWLANYDPNSFVDYNIKQLRS 663
Query: 577 RNFVNKELL-FSSMKNLQRSIPSVVDGLTLGQRKTLFCSFEMNLTKQTEVCRLTAYVTEH 635
+ F+++EL+ FS+ NL RSIPS+ DGL GQRK LF F+ NL + +V +L YV EH
Sbjct: 664 KXFIDRELIHFSNADNL-RSIPSLCDGLKPGQRKILFSCFKRNLKSEIKVAQLIGYVAEH 722
Query: 636 SYCHYGQQSIASTIIGMAQDFVGSNNINLLIPYGQFGTRDLGGKDHASSRKLYIKLNCVT 695
S H+G+QS+ASTIIGMAQ+FVGSNNINL++P GQFGTR+ GGK+HAS+R ++ L+ +T
Sbjct: 723 SAYHHGEQSLASTIIGMAQNFVGSNNINLMMPIGQFGTRNQGGKEHASARYIFTSLSKIT 782
Query: 696 RLLFPVDDDKLLEYLNEDGRSIQPNWYIPIIPLVLVNGCHAMGTCFSSDIPKYHPYEIIK 755
R+LFP DD +L+YL +D + ++P WY+PIIP+ LVNG +GT +S+DIP Y+ I++
Sbjct: 783 RILFPEHDDHVLKYLEDDNQVVEPEWYMPIIPMCLVNGSSGIGTGWSTDIPNYNHKGIVE 842
Query: 756 NVRRFLNNEEMVSMKPWYKGFGGTIEKSAKG----IGYYTVDGLVERINEQTFRIKELPI 811
+R L + PWYKGF G IE + KG G Y +D + E+T +I ELP+
Sbjct: 843 QIRGKLEGRPFTELHPWYKGFQGQIEINPKGGYIVKGQYEIDEV-----EETIKITELPV 897
Query: 812 RMWTEDYK-----KFLEKITARGLIESFRQKGDYEIVNFKVNVKQEEIATIEKDEELRKK 866
WT DYK + L IE R+ V+F+V Q I++ ++ KK
Sbjct: 898 GKWTRDYKVVLIQELLSPPKGEPEIEDIREYHTNNRVHFEV---QAIPGRIQQWGDIEKK 954
Query: 867 FNLTCTISTSNMYLFDAEGKIKKYNTPEQILEEFCTLRLKYYVRRKQYLVKNFTQLLRSL 926
L+ T+ T+N+ +F+ + K+K+Y + +I+EEF LRLK Y +RK YLV + + L
Sbjct: 955 LKLSTTMQTTNIVMFNDKYKLKRYQSTVEIMEEFYALRLKCYQKRKDYLVSKIQRDVDIL 1014
Query: 927 RIKQKFISNIVNEKFNPINRRAELLIEILKK------GKSAEPHVAGAIDYSSKEQEAGE 980
+ K KFI+ +VN N R + ++ LK+ K + +D +E + E
Sbjct: 1015 QNKMKFITLVVNGTLEIRNARKKGVLNKLKELGLTPMSKITRINSTKNLDNEDEEVQQDE 1074
Query: 981 QESVSQSVSIEGATLGDYEYLSSLPFENFTLESLKKLEAELDEKENELKTLEHTSPDLMW 1040
+E Q V L +Y YL S+P + T E + +L+ +++EKE+EL L+ MW
Sbjct: 1075 EEEEEQ-VQEGEIPLSEYNYLLSMPIFSLTEEKVLQLKKQVEEKESELLILQKKEIKDMW 1133
Query: 1041 LNDLELF 1047
DL+ F
Sbjct: 1134 TEDLDKF 1140
>UniRef100_Q8X1U4 DNA topoisomerase II [Aspergillus niger]
Length = 1391
Score = 627 bits (1618), Expect = e-178
Identities = 376/962 (39%), Positives = 555/962 (57%), Gaps = 51/962 (5%)
Query: 130 WTNVMFKPDLDKFDMKNLEKDVIALMKKRVLDMAVCLGETVTVVLNGTVIPVKSFKDYAN 189
+ V FKPD KF M ++ D AL+K+RV D+A V V LNGT IP++ FK Y
Sbjct: 244 YAKVTFKPDYAKFGMDGMDDDFEALVKRRVYDLAGTA--KVNVKLNGTRIPIRGFKKYME 301
Query: 190 FFL-----DRAKPFPLPRTH---AKLGDRLEICLSLSDGKFQQVSFVNSIATIKGGTHVD 241
+ +R P P+ R E+ ++SDG FQQVSFVNSIAT GGTHV+
Sbjct: 302 MYTKAIRKERGDTGPAPKDEIITCSPDPRWEVGFAVSDGAFQQVSFVNSIATTSGGTHVN 361
Query: 242 YITKQITTYIKKEVLKKKHVNVNADT--VKNQLWVFVNALIDNPAFNSQTKEMLTTKPAR 299
YI +I + ++V KK T ++N +++FVNALI NPAF SQTKE LTTK ++
Sbjct: 362 YIADRICNRLAEQVKKKNKNGATLKTAQIRNHIFIFVNALIVNPAFTSQTKEQLTTKASQ 421
Query: 300 LGLK-FFPGSMLNDVQKSQILDNLLP-----------------RIPFKHRTHVDAKLAGR 341
G K V K+ ++DN+L R + DA AG
Sbjct: 422 FGSKCVLDEDFYKKVLKTAVMDNILHFAQQKADQMLKKTDGGRRSRMNNPKLTDANKAGT 481
Query: 342 PDSGKCTLILTQGQYVKDLAMSGLSAVVDPDLYGVFPLSSKLLNATRDSR--KLFKKKEI 399
D CTLILT+G K LAM+G AVV PDL+GVFPL KLLN RD+ ++ K +EI
Sbjct: 482 KDGHHCTLILTEGDSAKGLAMAG-RAVVGPDLFGVFPLRGKLLNV-RDASFDQISKNQEI 539
Query: 400 QNLMAILGLVRNKKYSNAKCLRYGQLMVMANQDEDGAHFKGLLINFFYSFWPSLLKVSKF 459
QN+ LGL K+Y++ + LRYG LM+M +QD DG+H KGLLINF + +PSLLK+ +F
Sbjct: 540 QNIKNFLGLQHKKEYTDTRGLRYGHLMIMTDQDHDGSHIKGLLINFLQAQFPSLLKIPEF 599
Query: 460 MSVFTFPIIKVSMYNASDSKKGKELSFDSMQQYEDWKKELGNTANDWEIKYCQGL--SSA 517
+ F PI+KV + K+ SF +M +YE W +E + W+ KY +GL S+
Sbjct: 600 LIEFITPIVKV--WKGDPKNPTKQRSFFTMPEYEAWAEEHQHERG-WDHKYYKGLGTSTT 656
Query: 518 QEVREYFQDLGKH-RAYFVWDDKDETSIKLAFSKK-AEEWEAWIGNSQPGT-CNYERKVI 574
++ + YF+DL +H + + D + I+LAFSKK A+E + W+ +PGT ++ + I
Sbjct: 657 EDAQVYFRDLDRHLKEFHTMQDNEAELIELAFSKKKADERKEWLRQFKPGTFLDHSVEKI 716
Query: 575 NYRNFVNKELLFSSMKNLQRSIPSVVDGLTLGQRKTLFCSFEMNLTKQTEVCRLTAYVTE 634
Y +F+NKEL+ SM + RSIPSVVDGL GQRK L+ F NL K +V L +V+
Sbjct: 717 TYTDFINKELILFSMADNVRSIPSVVDGLKPGQRKVLYTCFRRNLKKDMKVVELAGHVSG 776
Query: 635 HSYCHYGQQSIASTIIGMAQDFVGSNNINLLIPYGQFGTRDLGGKDHASSRKLYIKLNCV 694
+ +G S+ TI+G+AQ FVGSNNIN L P G FG+R GG D AS+R +Y +L+
Sbjct: 777 MTAYQHGDVSLQQTIVGLAQTFVGSNNINCLEPSGNFGSRLQGGSDCASARYIYTRLSPF 836
Query: 695 TRLLFPVDDDKLLEYLNEDGRSIQPNWYIPIIPLVLVNGCHAMGTCFSSDIPKYHPYEII 754
R +F D+ LL Y +DG++I+P Y+P++P++L+NG +GT +SS IP Y+P +++
Sbjct: 837 ARRIFHASDEPLLTYNVDDGKTIEPEVYMPVVPMILINGADGIGTGWSSSIPNYNPEDVV 896
Query: 755 KNVRRFLNNEEMVSMKPWYKGFGGTIEKSAKGIGYYTVDGLVERINEQTFRIKELPIRMW 814
N++R +N EE+ M+PW++GF G E ++ G + G+++ E+ I ELPIR W
Sbjct: 897 DNLKRLMNGEEIKPMQPWFRGFTG--EVTSIGGDRFKFSGIIKETGEKEVEITELPIRTW 954
Query: 815 TEDYKKFLEKI----TARGLIESFRQKGDYEIVNFKVNVKQEEIATIEKDEELRKKFNLT 870
T+D+K LE I I+ ++ + V+F + + ++ + + +E L +KF LT
Sbjct: 955 TQDFKDKLEDIIKAEKTPSFIKDYKDYNTHTKVHFVIQMDEKHLKS-AVEEGLEEKFKLT 1013
Query: 871 CTISTSNMYLFDAEGKIKKYNTPEQILEEFCTLRLKYYVRRKQYLVKNFTQLLRSLRIKQ 930
TI+T+N+ FD EG+I KY T + IL+EF +RLKYY RRKQY + + L L +
Sbjct: 1014 KTIATTNLVAFDPEGRITKYATVDDILKEFFAVRLKYYERRKQYQLNEMQKELEKLSNQA 1073
Query: 931 KFISNIVN-EKFNPINRRAELLIEILKKGKSAEPHVAGAIDYSSKEQEAGEQESVSQSVS 989
+F+ I++ E ++ L++E+ KG P VA A + + + A E E S+ V
Sbjct: 1074 RFVQMIIDGELVVSKKKKNVLVVELKDKGFKPFPKVADAAK-AGEAEPAAEDEEESEDVD 1132
Query: 990 IEGATLGDYEYLSSLPFENFTLESLKKLEAELDEKENELKTLEHTSPDLMWLNDLELFEK 1049
Y+YL + + T E +++L ++ +KE E+ TL S + +W DL+ F
Sbjct: 1133 DTSNLSNAYDYLLGMAIWSLTQERVERLRRQIGDKEMEIDTLIKLSKEDIWKRDLDDFIN 1192
Query: 1050 EF 1051
E+
Sbjct: 1193 EW 1194
>UniRef100_UPI00003AC994 UPI00003AC994 UniRef100 entry
Length = 1553
Score = 624 bits (1609), Expect = e-177
Identities = 396/989 (40%), Positives = 571/989 (57%), Gaps = 88/989 (8%)
Query: 126 DNGKWTNVMFKPDLDKFDMKNLEKDVIALMKKRVLDMAVCLGET--VTVVLNGTVIPVKS 183
D +T V F+PDL KF M L+KD++ALM +R D+A G T V V LNG +PVK
Sbjct: 211 DGEDYTCVTFQPDLSKFKMTILDKDIVALMSRRAYDIA---GSTKDVKVFLNGKRLPVKG 267
Query: 184 FKDYANFFL-DRAKPF--PLPRTHAKLGDRLEICLSLSDGKFQQVSFVNSIATIKGGTHV 240
F+ Y + +L D+ L H ++ R E+CL+LS+ FQQVSFVNSIAT KGG HV
Sbjct: 268 FRSYVDLYLKDKVDETGNALKVIHEEVNSRWEVCLTLSEKGFQQVSFVNSIATTKGGRHV 327
Query: 241 DYITKQITTYIKKEVLKKKHVN---VNADTVKNQLWVFVNALIDNPAFNSQTKEMLTTKP 297
DY+ QI T + +V+KKK+ N V VKN +W+FVN+LI+NP F+SQTKE +T +
Sbjct: 328 DYVADQIVTKLI-DVVKKKNKNGVGVKPFQVKNHMWIFVNSLIENPTFDSQTKENMTLQA 386
Query: 298 ------ARLGLKFFPGSMLNDVQKSQILDNLLPRIPFKHRTHV----------------- 334
+L KF G++ I++++L + FK +T +
Sbjct: 387 KSFGSTCKLSEKFIKGAV-----GCGIVESILNWVKFKAQTQLNKKCSAVKHTKIKGVPK 441
Query: 335 --DAKLAGRPDSGKCTLILTQGQYVKDLAMSGLSAVVDPDLYGVFPLSSKLLNATRDSRK 392
DA AG +S CTLILT+G K LA+SGL VV D YGVFPL K+LN S K
Sbjct: 442 LDDANDAGSKNSIDCTLILTEGDSAKTLAVSGLG-VVGRDKYGVFPLRGKMLNVREASHK 500
Query: 393 -LFKKKEIQNLMAILGLVRNKKYSNA---KCLRYGQLMVMANQDEDGAHFKGLLINFFYS 448
+ + EI N++ I+GL K Y + K LRYG++M+M +QD+DG+H KGLLINF +
Sbjct: 501 QIMENAEINNIIKIVGLQYKKNYEDRESLKSLRYGKIMIMTDQDQDGSHIKGLLINFIHH 560
Query: 449 FWPSLLKVSKFMSVFTFPIIKVSMYNASDSKKGKELSFDSMQQYEDWKKELGNTANDWEI 508
WPSLL+ F+ F PIIKVS K +E+ F S+ ++E+WK N N W+I
Sbjct: 561 NWPSLLR-HNFLEEFITPIIKVS-------KNKEEIPFYSIPEFEEWKSSTQNY-NSWKI 611
Query: 509 KYCQGL--SSAQEVREYFQDLGKHRAYFVWDD-KDETSIKLAFSKK-AEEWEAWIGNSQ- 563
KY +GL S+++E +EYF D+ +HR F + +D+ +I LAFSKK EE + W+ N
Sbjct: 612 KYYKGLGTSTSKEAKEYFADMARHRIGFKYSGPEDDAAITLAFSKKKVEERKEWLTNFME 671
Query: 564 ----------PGTCNY--ERKVINYRNFVNKELLFSSMKNLQRSIPSVVDGLTLGQRKTL 611
P Y + + Y +F+NKEL+ S + +RSIPS+VDGL GQRK L
Sbjct: 672 DRRQRKLHGLPEDYLYGKDTNYLTYNDFINKELVLFSNSDNERSIPSLVDGLKPGQRKVL 731
Query: 612 FCSFEMNLTKQTEVCRLTAYVTEHSYCHYGQQSIASTIIGMAQDFVGSNNINLLIPYGQF 671
F F+ N ++ +V +L V E S H+G+ S+ TII +AQ+FVGSNN+NLL P GQF
Sbjct: 732 FTCFKRNDKREVKVAQLAGSVAEMSSYHHGEASLMMTIINLAQNFVGSNNLNLLQPIGQF 791
Query: 672 GTRDLGGKDHASSRKLYIKLNCVTRLLFPVDDDKLLEYLNEDGRSIQPNWYIPIIPLVLV 731
GTR GGKD AS R ++ L+ + RLLFP DD +L +L +D + ++P WY+PIIP+VL+
Sbjct: 792 GTRLHGGKDSASPRYIFTMLSPLARLLFPPVDDNVLRFLYDDNQRVEPEWYMPIIPMVLI 851
Query: 732 NGCHAMGTCFSSDIPKYHPYEIIKNVRRFLNNEEMVSMKPWYKGFGGTIEKSAKGIGYYT 791
NG +GT +S IP + E + N+R L+ +E + M P YK F GTI++ G Y
Sbjct: 852 NGAEGIGTGWSCKIPNFDIRETVNNIRCLLDGKEPLPMLPSYKNFKGTIDE--LGPNQYV 909
Query: 792 VDGLVERINEQTFRIKELPIRMWTEDYKKFLEKITARG------LIESFRQKGDYEIVNF 845
+ G V ++ T I ELP+R WT+ YK+ + + G LI +++ V F
Sbjct: 910 ISGEVSILDSTTIEITELPVRTWTQTYKEQVLEPMLNGTEKTPPLITDYKEYHTDTTVKF 969
Query: 846 KVNVKQEEIATIEKDEELRKKFNLTCTISTSNMYLFDAEGKIKKYNTPEQILEEFCTLRL 905
V + +E++A E L K F L ++ ++M LFD G +KKY +P+ IL+EF LRL
Sbjct: 970 VVKMSEEKLAEAEA-VGLHKVFKLQTNLTCNSMVLFDHVGFLKKYESPQDILKEFFELRL 1028
Query: 906 KYYVRRKQYLVKNFTQLLRSLRIKQKFISNIVNEKFNPINRRAELLIEIL-KKGKSAEPH 964
+YY RK++L+ L + +FI ++ K N+ + LI++L ++G ++P
Sbjct: 1029 RYYGLRKEWLIGMLGAESAKLNNQARFILEKIDGKIVIENKPKKELIQVLIQRGYESDP- 1087
Query: 965 VAGAIDYSSKEQEAGEQESVSQSVSIEGATLGDYEYLSSLPFENFTLESLKKLEAELDEK 1024
V + +KE+E G+ ES +S + AT D+ YL ++P T E +L + D K
Sbjct: 1088 VKAWKELQNKEEEEGD-ESGEESAA---ATGPDFNYLLNMPLWYLTKEKKDELCKQRDNK 1143
Query: 1025 ENELKTLEHTSPDLMWLNDLELFEKEFDV 1053
+ EL+ L+H SP +W DL F +E DV
Sbjct: 1144 DKELEDLKHKSPSDLWKEDLAAFVEELDV 1172
>UniRef100_Q8X1U6 DNA topoisomerase II [Aspergillus flavus]
Length = 1219
Score = 624 bits (1608), Expect = e-177
Identities = 382/965 (39%), Positives = 555/965 (56%), Gaps = 57/965 (5%)
Query: 130 WTNVMFKPDLDKFDMKNLEKDVIALMKKRVLDMAVCLGETVTVVLNGTVIPVKSFKDYAN 189
+T V FKPD KF M ++ D AL+K+RV D+A V V LNG+ +PV++FK Y
Sbjct: 207 YTKVTFKPDYAKFGMDGMDNDFEALVKRRVYDLAGTA--KVAVKLNGSRVPVRNFKKYME 264
Query: 190 FFL-----DRAKPFPLPRTH---AKLGDRLEICLSLSDGKFQQVSFVNSIATIKGGTHVD 241
+ +R P + R E+ ++SDG FQQVSFVNSIAT GGTHV+
Sbjct: 265 MYTKAIRRERGDDGPAAKDEIITCSPDPRWEVGFAVSDGSFQQVSFVNSIATTSGGTHVN 324
Query: 242 YITKQITTYIKKEVLKKKHVNVNADTVK-----NQLWVFVNALIDNPAFNSQTKEMLTTK 296
YI QI + + +V KK N N T+K N +++FVNA I NPAF SQTKE LTTK
Sbjct: 325 YIADQICSKLADQVKKK---NKNGATLKPAQIRNHIFIFVNAQIVNPAFTSQTKEQLTTK 381
Query: 297 PARLGLK------FFPGSMLNDVQKS------QILDNLLPRIPFKHRTHV------DAKL 338
++ G K F+ + DV + Q D +L + R + DA
Sbjct: 382 SSQFGSKCVLEEDFYKKILKTDVMSNILHFAQQKADQMLKKTDGGRRARMNNPKLTDANK 441
Query: 339 AGRPDSGKCTLILTQGQYVKDLAMSGLSAVVDPDLYGVFPLSSKLLNATRDS--RKLFKK 396
AG D CTLILT+G K LAM+G AVV PDL+GVFPL KLLN RD+ ++ K
Sbjct: 442 AGTKDGHHCTLILTEGDSAKGLAMAG-RAVVGPDLFGVFPLRGKLLNV-RDATFEQISKN 499
Query: 397 KEIQNLMAILGLVRNKKYSNAKCLRYGQLMVMANQDEDGAHFKGLLINFFYSFWPSLLKV 456
EIQN+ LGL K+Y++ + LRYG LM+M +QD DG+H KGLLINF + +PSLL++
Sbjct: 500 AEIQNIKNFLGLQHKKEYTDTRGLRYGHLMIMTDQDHDGSHIKGLLINFLQAQFPSLLRI 559
Query: 457 SKFMSVFTFPIIKVSMYNASDSKKGKELSFDSMQQYEDWKKELGNTANDWEIKYCQGL-- 514
+F+ F PIIKV + K+ SF +M +YE WK+E + WE KY +GL
Sbjct: 560 PEFLIEFITPIIKV--WKGDPKNPTKQRSFFTMPEYEAWKEEHKHERG-WEHKYYKGLGT 616
Query: 515 SSAQEVREYFQDLGKH-RAYFVWDDKDETSIKLAFSKK-AEEWEAWIGNSQPGT-CNYER 571
S+ ++ + YF+DL +H + + D + I+LAFSKK A+E + W+ +PGT ++
Sbjct: 617 STTEDAQVYFRDLDRHLKEFHTMQDNEVGLIELAFSKKKADERKEWLRQFKPGTFLDHSV 676
Query: 572 KVINYRNFVNKELLFSSMKNLQRSIPSVVDGLTLGQRKTLFCSFEMNLTKQTEVCRLTAY 631
I+Y +F+NKEL+ SM + RSIPSVVDGL GQRK L+ F NL K +V L +
Sbjct: 677 SKISYTDFINKELILFSMADNVRSIPSVVDGLKPGQRKVLYTCFRRNLKKDMKVVELAGH 736
Query: 632 VTEHSYCHYGQQSIASTIIGMAQDFVGSNNINLLIPYGQFGTRDLGGKDHASSRKLYIKL 691
V+ + +G S+ TI+G+AQ FVGSNN+N L P G FG+R GG D AS+R +Y +L
Sbjct: 737 VSGMTAYQHGDISLQQTIVGLAQSFVGSNNVNCLEPSGNFGSRLQGGSDCASARYIYTRL 796
Query: 692 NCVTRLLFPVDDDKLLEYLNEDGRSIQPNWYIPIIPLVLVNGCHAMGTCFSSDIPKYHPY 751
+ R +F DD LL Y +DG I+P Y+P++P++LVNG +GT +SS IP Y+P
Sbjct: 797 SPFARRVFHAHDDPLLTYNEDDGEKIEPEIYVPVVPMILVNGADGIGTGWSSSIPNYNPE 856
Query: 752 EIIKNVRRFLNNEEMVSMKPWYKGFGGTIEKSAKGIGYYTVDGLVERINEQTFRIKELPI 811
+I+ N++R ++ EE M+PW++GF G E + G + G+++ E+ I ELPI
Sbjct: 857 DIVDNLKRLMDGEETRPMQPWFRGFTG--EVTDVGGDRFKFSGIIKETGEKEVEITELPI 914
Query: 812 RMWTEDYKKFLEKI----TARGLIESFRQKGDYEIVNFKVNVKQEEIATIEKDEELRKKF 867
R WT+D+K LE I I+ ++ + V+F + + ++ + + E L +KF
Sbjct: 915 RTWTQDFKDKLEDIIKAEKTPSFIKDYKDYNTHTKVHFVIQLDEKNMKS-ALSEGLEEKF 973
Query: 868 NLTCTISTSNMYLFDAEGKIKKYNTPEQILEEFCTLRLKYYVRRKQYLVKNFTQLLRSLR 927
L+ T++T+N+ FD EG+I KY T + IL+EF T RLK+Y +RKQY + + L L
Sbjct: 974 KLSKTVATTNLVAFDPEGRITKYATVDDILKEFYTYRLKFYEKRKQYQLSQLQKELDKLS 1033
Query: 928 IKQKFISNIVNEKFNPINRRAELLIEILK-KGKSAEPHVAGAIDYSSKEQEAGEQESVSQ 986
+ +F+ I++ K ++ +L++ LK KG A P VA A+ +E+ E + S
Sbjct: 1034 NQARFVQEIIDGKLVVSKKKKNVLVQELKDKGYKAFPKVAEAVKAGEEERVVEEDDDESD 1093
Query: 987 SVSIEGATLGDYEYLSSLPFENFTLESLKKLEAELDEKENELKTLEHTSPDLMWLNDLEL 1046
S E Y+YL + + T E ++KL ++ EKE E+ L S + +W DL+
Sbjct: 1094 SQDTE-VLSSAYDYLLGMAIWSLTQERVEKLRRQIGEKEVEVDALIKMSKEDIWKRDLDD 1152
Query: 1047 FEKEF 1051
F E+
Sbjct: 1153 FIAEW 1157
>UniRef100_Q9NEE4 DNA topoisomerase II [Leishmania major]
Length = 1497
Score = 621 bits (1602), Expect = e-176
Identities = 365/984 (37%), Positives = 559/984 (56%), Gaps = 64/984 (6%)
Query: 125 CDNGKWTNVMFKPDLDKFDMKNLEKDVIALMKKRVLDMAVCLGETVTVVLNGTVIPVKSF 184
CD+ +T V F PD F + +D++ +MK+RV D+A C +T+ LN I +F
Sbjct: 184 CDSADYTMVTFYPDFALFHVDKFTEDMLLMMKRRVYDIAGCTDKTLKCFLNDERIACTTF 243
Query: 185 KDYANFFLDRAKPFPLPRTHAKLGDRLEICLSLSDGKFQQVSFVNSIATIKGGTHVDYIT 244
+Y + + + +++++ DR E+C+ +S+ QQVSFVNSIAT +GGTHV YI
Sbjct: 244 PEYVDLYPTMGEERKAA-SYSRVNDRWEVCVRVSNIGPQQVSFVNSIATTRGGTHVKYIM 302
Query: 245 KQITTYIKKEVLKKKHVNVNADTVKNQLWVFVNALIDNPAFNSQTKEMLTTKPARLGLKF 304
QIT+ + ++ KKK +V ++ +++FVN LI+NP F+SQTKE L T ++ G
Sbjct: 303 DQITSKVMEQAKKKKKTDVKPHMIRPHIFLFVNCLIENPGFDSQTKETLNTVKSKFGSTC 362
Query: 305 -FPGSMLNDVQKSQILDN------------LLPRIPFKHRTHV-------DAKLAGRPDS 344
P S+++ + S I++ + ++ R + DA AG S
Sbjct: 363 DLPNSIVDYIMSSGIVERSVEMANSKIAREMASKMRSSDRKQIMGIPKLDDANEAGGKHS 422
Query: 345 GKCTLILTQGQYVKDLAMSGLSAVVDPDLYGVFPLSSKLLNATRDS-RKLFKKKEIQNLM 403
+CTLILT+G K L +GL AV D D +GVFPL K LN S +KL +EIQ +M
Sbjct: 423 HRCTLILTEGDSAKTLCTAGL-AVKDRDYFGVFPLRGKPLNVREASLKKLAACEEIQCVM 481
Query: 404 AILGLVRNKKYSNAKCLRYGQLMVMANQDEDGAHFKGLLINFFYSFWPSLLKVSKFMSVF 463
I+GL + Y N+ LRYG LM+M++QD DG+H KGLLINF + FWP+LL+V F+ F
Sbjct: 482 KIMGLDIRQTYENSDGLRYGHLMIMSDQDHDGSHIKGLLINFIHCFWPNLLRVPGFLQQF 541
Query: 464 TFPIIKVSMYNASDSKKGKELSFDSMQQYEDWKKELGNTANDWEIKYCQGL--SSAQEVR 521
PI+K + GK +SF SM Y +WKK +G+ ++++I+Y +GL S A+E R
Sbjct: 542 ITPIVKARPKGRGGA--GKAISFFSMPDYFEWKKAIGDNLSNYQIRYYKGLGTSGAEEGR 599
Query: 522 EYFQDLGKHRAYFVWDDK-DETSIKLAFSK-KAEEWEAWIGNSQPG-----TCNYERKVI 574
EYF+++ +HR FV ++ +E I +AF K + E+ + WI N + + +Y + +
Sbjct: 600 EYFENIDRHRLNFVEQEQSEEDRIVMAFGKDRVEDRKEWITNFKTNVNVNESMDYNVRQV 659
Query: 575 NYRNFVNKELLFSSMKNLQRSIPSVVDGLTLGQRKTLFCSFEMNLTKQTEVCRLTAYVTE 634
+YR+FV+KEL+ S+ + +RSIPSV+DG GQRK LF F+ NL +V +L YV+E
Sbjct: 660 SYRDFVDKELILFSIADCERSIPSVIDGFKPGQRKILFSCFKRNLVNSIKVVQLAGYVSE 719
Query: 635 HSYCHYGQQSIASTIIGMAQDFVGSNNINLLIPYGQFGTRDLGGKDHASSRKLYIKLNCV 694
HS H+G+QS+ TI+GMAQDFVGSNN+ LL GQFGTR GGKDHA+ R ++ +L V
Sbjct: 720 HSAYHHGEQSLVQTIVGMAQDFVGSNNVPLLRKDGQFGTRLHGGKDHAAGRYIFTRLMQV 779
Query: 695 TRLLFPVDDDKLLEYLNEDGRSIQPNWYIPIIPLVLVNGCHAMGTCFSSDIPKYHPYEII 754
R +F DD ++EY ++DG S++P +YIP+IP+VLVNG +GT FS+ IP Y+P EII
Sbjct: 780 ARAIFHPADDFVVEYKDDDGLSVEPFFYIPVIPMVLVNGTTGIGTGFSTSIPNYNPLEII 839
Query: 755 KNVRRFLNNEEMVSMKPWYKGFGGTIEKSAKGIGYYTVDGLVERINEQTFRIKELPIRMW 814
N+ R + EE+ M+PWY GF GT+E+ K G + G E + I ELPI +W
Sbjct: 840 SNLERLMRGEEVHKMQPWYFGFTGTMEE--KEPGKFISHGRYEIRPDGVVCITELPIGVW 897
Query: 815 TEDYKKFLEKITARGLIESFRQKGDYEIVNFKVNVKQEEIATIEKDEELRKKFNLTCTIS 874
T+ YKKFLE + + L+ +R+ V+F+V + + + + + ++ L +
Sbjct: 898 TQAYKKFLEDMREKELVVQYREHTTDVTVDFEVFLHPQVLEQWVAQDVVEERLQLKEYVH 957
Query: 875 TSNMYLFDAEGKIKKYNTPEQILEEFCTLRLKYYVRRKQYLVKNFTQLLRSLRIKQKFIS 934
+N+ F+ EG+I KY+ E +L+EF +RL YY +RKQYL+ +L L +F+
Sbjct: 958 ATNIIAFNKEGQITKYSDAESVLKEFYLVRLDYYAKRKQYLLGELRRLTSKLENMVRFVR 1017
Query: 935 NIVNEKFNPINRRAELLIEILKKGK--SAEPHVAGAIDYSSKEQEAGEQESVSQSVSIEG 992
+V R+ + L+E L++ PH + SS E G+ E + + E
Sbjct: 1018 EVVEGTLVVTKRKKKELLEDLQRRMYLPFPPHTKKKL--SSTTVEEGDDEGAPDAQNSEE 1075
Query: 993 ATLG------------------------DYEYLSSLPFENFTLESLKKLEAELDEKENEL 1028
LG DY+YL + + T E + +LEA++ +
Sbjct: 1076 DVLGVGNAADSGADDGAAEPPEMRRATRDYDYLLGMKLWSLTAEMIARLEAQMARARRDT 1135
Query: 1029 KTLEHTSPDLMWLNDLELFEKEFD 1052
+E +P MW+ DL++ + +
Sbjct: 1136 AAMEARTPKDMWMEDLKVLRTKLE 1159
>UniRef100_O55079 Topoisomerase II alpha [Homo sapiens]
Length = 1526
Score = 621 bits (1601), Expect = e-176
Identities = 390/987 (39%), Positives = 569/987 (57%), Gaps = 80/987 (8%)
Query: 127 NGK-WTNVMFKPDLDKFDMKNLEKDVIALMKKRVLDMAVCLGET--VTVVLNGTVIPVKS 183
NG+ +T + F+PDL KF M++L+KD++ALM +R D+A G T V V LNG +PVK
Sbjct: 209 NGEDYTCITFQPDLSKFKMQSLDKDIVALMVRRAYDIA---GSTKDVKVFLNGNKLPVKG 265
Query: 184 FKDYANFFL-DRAKPF--PLPRTHAKLGDRLEICLSLSDGKFQQVSFVNSIATIKGGTHV 240
F+ Y + +L D+ L H ++ R E+CL++S+ FQQ+SFVNSIAT KGG HV
Sbjct: 266 FRSYVDMYLKDKLDETGNALKVVHEQVNPRWEVCLTMSEKGFQQISFVNSIATSKGGRHV 325
Query: 241 DYITKQITTYIKKEVLKKKH---VNVNADTVKNQLWVFVNALIDNPAFNSQTKEMLTTKP 297
DY+ QI + + +V+KKK+ V V A VKN +W+FVNALI+NP F+SQTKE +T +
Sbjct: 326 DYVADQIVSKLV-DVVKKKNKGGVAVKAHQVKNHMWIFVNALIENPTFDSQTKENMTLQA 384
Query: 298 ARLGLKF-FPGSMLNDVQKSQILDNLLPRIPFKHRTHV-------------------DAK 337
G + I++++L + FK + + DA
Sbjct: 385 KSFGSTCQLSEKFIKAAIGCGIVESILNWVKFKAQIQLNKKCSAVKHNRIKGIPKLDDAN 444
Query: 338 LAGRPDSGKCTLILTQGQYVKDLAMSGLSAVVDPDLYGVFPLSSKLLNATRDSRK-LFKK 396
AG +S +CTLILT+G K LA+SGL VV D YGVFPL K+LN S K + +
Sbjct: 445 DAGSRNSTECTLILTEGDSAKTLAVSGLG-VVGRDKYGVFPLGGKILNVREASHKQIMEN 503
Query: 397 KEIQNLMAILGLVRNKKYSNA---KCLRYGQLMVMANQDEDGAHFKGLLINFFYSFWPSL 453
EI N++ I+GL K Y + K LRYG++M+M +QD+DG+H KGLLINF + WPSL
Sbjct: 504 AEINNIIKIVGLQYKKNYEDEDSLKTLRYGKIMIMTDQDQDGSHIKGLLINFIHHNWPSL 563
Query: 454 LKVSKFMSVFTFPIIKVSMYNASDSKKGKELSFDSMQQYEDWKKELGNTANDWEIKYCQG 513
L+ +F+ F PI+KVS K +EL+F S+ ++E+WK N W++KY +G
Sbjct: 564 LR-HRFLEEFITPIVKVS-------KNKQELAFYSLPEFEEWKSSTPNHKK-WKVKYYKG 614
Query: 514 L--SSAQEVREYFQDLGKHRAYFVWDD-KDETSIKLAFSKKA----EEW----------E 556
L S+++E +EYF D+ +HR F + +D+ +I LAFSKK +EW
Sbjct: 615 LGTSTSKEAKEYFADMKRHRIQFKYSGPEDDAAISLAFSKKQVDDRKEWLTHFMEDRRQR 674
Query: 557 AWIGNSQPGTCNYERKVINYRNFVNKELLFSSMKNLQRSIPSVVDGLTLGQRKTLFCSFE 616
+G + + Y +F+NKEL+ S + +RSIPS+VDGL GQRK LF F+
Sbjct: 675 KLLGLPEDYLYGQTTTYLTYNDFINKELILFSNSDNERSIPSMVDGLKPGQRKVLFTCFK 734
Query: 617 MNLTKQTEVCRLTAYVTEHSYCHYGQQSIASTIIGMAQDFVGSNNINLLIPYGQFGTRDL 676
N ++ +V +L V E S H+G+ S+ TII +AQ+FVGSNN+NLL P GQFGTR
Sbjct: 735 RNDKREVKVAQLAGSVAEMSSYHHGEMSLMMTIINLAQNFVGSNNLNLLQPIGQFGTRLH 794
Query: 677 GGKDHASSRKLYIKLNCVTRLLFPVDDDKLLEYLNEDGRSIQPNWYIPIIPLVLVNGCHA 736
GGKD AS R ++ L+ +TRLLFP DD L++L +D + ++P WYIPIIP+VL+NG
Sbjct: 795 GGKDSASPRYIFTMLSPLTRLLFPPKDDHTLKFLYDDNQRVEPEWYIPIIPMVLINGAEG 854
Query: 737 MGTCFSSDIPKYHPYEIIKNVRRFLNNEEMVSMKPWYKGFGGTIEKSAKGIGYYTVDGLV 796
+GT +S IP + E++ N+RR L+ EE + M P YK F GTIE+ A Y ++G V
Sbjct: 855 IGTGWSCKIPNFDIREVVNNIRRLLDGEEPLPMLPSYKNFKGTIEELAS--NQYVINGEV 912
Query: 797 ERINEQTFRIKELPIRMWTEDYKKFLEKITARG------LIESFRQKGDYEIVNFKVNVK 850
+N T I ELPIR WT+ YK+ + + G LI +R+ V F + +
Sbjct: 913 AILNSTTIEISELPIRTWTQTYKEQVLEPMLNGTEKTPPLITDYREYHTDTTVKFVIKMT 972
Query: 851 QEEIATIEKDEELRKKFNLTCTISTSNMYLFDAEGKIKKYNTPEQILEEFCTLRLKYYVR 910
+E++A E+ L K F L +++ ++M LFD G +KKY+T IL++F LRLKYY
Sbjct: 973 EEKLAEAER-VGLHKVFKLQTSLTCNSMVLFDHVGCLKKYDTVLDILKDFFELRLKYYGL 1031
Query: 911 RKQYLVKNFTQLLRSLRIKQKFISNIVNEKFNPINR-RAELLIEILKKGKSAEPHVAGAI 969
RK++L+ L + +FI ++ K N+ + EL+ ++++G ++P A
Sbjct: 1032 RKEWLLGMLGAESAKLNNQARFILEKIDGKIIIENKPKKELIKVLIQRGYDSDPVKAWKE 1091
Query: 970 DYSSKEQEAGEQES---VSQSVSIEGATLGDYEYLSSLPFENFTLESLKKLEAELDEKEN 1026
E +ES S SV+ G T + YL +P T E +L + +EKE
Sbjct: 1092 AQQKVPDEEENEESDNENSDSVAESGPT---FNYLLDMPLWYLTKEKKDELCKQRNEKEQ 1148
Query: 1027 ELKTLEHTSPDLMWLNDLELFEKEFDV 1053
EL TL++ SP +W DL +F +E +V
Sbjct: 1149 ELNTLKNKSPSDLWKEDLAVFIEELEV 1175
>UniRef100_P41515 DNA topoisomerase II, alpha isozyme [Cricetulus griseus]
Length = 1526
Score = 620 bits (1600), Expect = e-176
Identities = 390/987 (39%), Positives = 569/987 (57%), Gaps = 80/987 (8%)
Query: 127 NGK-WTNVMFKPDLDKFDMKNLEKDVIALMKKRVLDMAVCLGET--VTVVLNGTVIPVKS 183
NG+ +T + F+PDL KF M++L+KD++ALM +R D+A G T V V LNG +PVK
Sbjct: 209 NGEDYTCITFQPDLSKFKMQSLDKDIVALMVRRAYDIA---GSTKDVKVFLNGNKLPVKG 265
Query: 184 FKDYANFFL-DRAKPF--PLPRTHAKLGDRLEICLSLSDGKFQQVSFVNSIATIKGGTHV 240
F+ Y + +L D+ L H ++ R E+CL++S+ FQQ+SFVNSIAT KGG HV
Sbjct: 266 FRSYVDMYLKDKLDETGNALKVVHEQVNPRWEVCLTMSEKGFQQISFVNSIATSKGGRHV 325
Query: 241 DYITKQITTYIKKEVLKKKH---VNVNADTVKNQLWVFVNALIDNPAFNSQTKEMLTTKP 297
DY+ QI + + +V+KKK+ V V A VKN +W+FVNALI+NP F+SQTKE +T +
Sbjct: 326 DYVADQIVSKLV-DVVKKKNKGGVAVKAHQVKNHMWIFVNALIENPTFDSQTKENMTLQA 384
Query: 298 ARLGLKF-FPGSMLNDVQKSQILDNLLPRIPFKHRTHV-------------------DAK 337
G + I++++L + FK + + DA
Sbjct: 385 KSFGSTCQLSEKFIKAAIGCGIVESILNWVKFKAQIQLNKKCSAVKHNRIKGIPKLDDAN 444
Query: 338 LAGRPDSGKCTLILTQGQYVKDLAMSGLSAVVDPDLYGVFPLSSKLLNATRDSRK-LFKK 396
AG +S +CTLILT+G K LA+SGL VV D YGVFPL K+LN S K + +
Sbjct: 445 DAGSRNSTECTLILTEGDSAKTLAVSGLG-VVGRDKYGVFPLRGKILNVREASHKQIMEN 503
Query: 397 KEIQNLMAILGLVRNKKYSNA---KCLRYGQLMVMANQDEDGAHFKGLLINFFYSFWPSL 453
EI N++ I+GL K Y + K LRYG++M+M +QD+DG+H KGLLINF + WPSL
Sbjct: 504 AEINNIIKIVGLQYKKNYEDEDSLKTLRYGKIMIMTDQDQDGSHIKGLLINFIHHNWPSL 563
Query: 454 LKVSKFMSVFTFPIIKVSMYNASDSKKGKELSFDSMQQYEDWKKELGNTANDWEIKYCQG 513
L+ +F+ F PI+KVS K +EL+F S+ ++E+WK N W++KY +G
Sbjct: 564 LR-HRFLEEFITPIVKVS-------KNKQELAFYSLPEFEEWKSSTPNHKK-WKVKYYKG 614
Query: 514 L--SSAQEVREYFQDLGKHRAYFVWDD-KDETSIKLAFSKKA----EEW----------E 556
L S+++E +EYF D+ +HR F + +D+ +I LAFSKK +EW
Sbjct: 615 LGTSTSKEAKEYFADMKRHRIQFKYSGPEDDAAISLAFSKKQVDDRKEWLTHFMEDRRQR 674
Query: 557 AWIGNSQPGTCNYERKVINYRNFVNKELLFSSMKNLQRSIPSVVDGLTLGQRKTLFCSFE 616
+G + + Y +F+NKEL+ S + +RSIPS+VDGL GQRK LF F+
Sbjct: 675 KLLGLPEDYLYGQTTTYLTYNDFINKELILFSNSDNERSIPSMVDGLKPGQRKVLFTCFK 734
Query: 617 MNLTKQTEVCRLTAYVTEHSYCHYGQQSIASTIIGMAQDFVGSNNINLLIPYGQFGTRDL 676
N ++ +V +L V E S H+G+ S+ TII +AQ+FVGSNN+NLL P GQFGTR
Sbjct: 735 RNDKREVKVAQLAGSVAEMSSYHHGEMSLMMTIINLAQNFVGSNNLNLLQPIGQFGTRLH 794
Query: 677 GGKDHASSRKLYIKLNCVTRLLFPVDDDKLLEYLNEDGRSIQPNWYIPIIPLVLVNGCHA 736
GGKD AS R ++ L+ +TRLLFP DD L++L +D + ++P WYIPIIP+VL+NG
Sbjct: 795 GGKDSASPRYIFTMLSPLTRLLFPPKDDHTLKFLYDDNQRVEPEWYIPIIPMVLINGAEG 854
Query: 737 MGTCFSSDIPKYHPYEIIKNVRRFLNNEEMVSMKPWYKGFGGTIEKSAKGIGYYTVDGLV 796
+GT +S IP + E++ N+RR L+ EE + M P YK F GTIE+ A Y ++G V
Sbjct: 855 IGTGWSCKIPNFDIREVVNNIRRLLDGEEPLPMLPSYKNFKGTIEELAS--NQYVINGEV 912
Query: 797 ERINEQTFRIKELPIRMWTEDYKKFLEKITARG------LIESFRQKGDYEIVNFKVNVK 850
+N T I ELPIR WT+ YK+ + + G LI +R+ V F + +
Sbjct: 913 AILNSTTIEISELPIRTWTQTYKEQVLEPMLNGTEKTPPLITDYREYHTDTTVKFVIKMT 972
Query: 851 QEEIATIEKDEELRKKFNLTCTISTSNMYLFDAEGKIKKYNTPEQILEEFCTLRLKYYVR 910
+E++A E+ L K F L +++ ++M LFD G +KKY+T IL++F LRLKYY
Sbjct: 973 EEKLAEAER-VGLHKVFKLQTSLTCNSMVLFDHVGCLKKYDTVLDILKDFFELRLKYYGL 1031
Query: 911 RKQYLVKNFTQLLRSLRIKQKFISNIVNEKFNPINR-RAELLIEILKKGKSAEPHVAGAI 969
RK++L+ L + +FI ++ K N+ + EL+ ++++G ++P A
Sbjct: 1032 RKEWLLGMLGAESAKLNNQARFILEKIDGKIIIENKPKKELIKVLIQRGYDSDPVKAWKE 1091
Query: 970 DYSSKEQEAGEQES---VSQSVSIEGATLGDYEYLSSLPFENFTLESLKKLEAELDEKEN 1026
E +ES S SV+ G T + YL +P T E +L + +EKE
Sbjct: 1092 AQQKVPDEEENEESDNENSDSVAESGPT---FNYLLDMPLWYLTKEKKDELCKQRNEKEQ 1148
Query: 1027 ELKTLEHTSPDLMWLNDLELFEKEFDV 1053
EL TL++ SP +W DL +F +E +V
Sbjct: 1149 ELNTLKNKSPSDLWKEDLAVFIEELEV 1175
>UniRef100_O42130 DNA topoisomerase II, alpha isozyme [Gallus gallus]
Length = 1553
Score = 620 bits (1598), Expect = e-176
Identities = 394/988 (39%), Positives = 569/988 (56%), Gaps = 88/988 (8%)
Query: 126 DNGKWTNVMFKPDLDKFDMKNLEKDVIALMKKRVLDMAVCLGET--VTVVLNGTVIPVKS 183
D +T V F+PDL KF M L+KD++ALM +R D+A G T V V LNG +PVK
Sbjct: 211 DGEDYTCVTFQPDLSKFKMTILDKDIVALMSRRAYDIA---GSTKDVKVFLNGKRLPVKG 267
Query: 184 FKDYANFFL-DRAKPF--PLPRTHAKLGDRLEICLSLSDGKFQQVSFVNSIATIKGGTHV 240
F+ Y + +L D+ L H ++ R E+CL+LS+ FQQVSFVNSIAT KGG HV
Sbjct: 268 FRSYVDLYLKDKVDETGNALKVIHEEVNSRWEVCLTLSEKGFQQVSFVNSIATTKGGRHV 327
Query: 241 DYITKQITTYIKKEVLKKKHVN---VNADTVKNQLWVFVNALIDNPAFNSQTKEMLTTKP 297
DY+ QI T + +V+KKK+ N V VKN +W+FVN+LI+NP F+SQTKE +T +
Sbjct: 328 DYVADQIVTKLI-DVVKKKNKNGVGVKPFQVKNHMWIFVNSLIENPTFDSQTKENMTLQA 386
Query: 298 ------ARLGLKFFPGSMLNDVQKSQILDNLLPRIPFKHRTHV----------------- 334
+L KF G++ I++++L + FK +T +
Sbjct: 387 KSFGSTCKLSEKFIKGAV-----GCGIVESILNWVKFKAQTQLNKKCSAVKHTKIKGVPK 441
Query: 335 --DAKLAGRPDSGKCTLILTQGQYVKDLAMSGLSAVVDPDLYGVFPLSSKLLNATRDSRK 392
DA AG +S CTLILT+G K LA+SGL VV D YGVFPL K+LN S K
Sbjct: 442 LDDANDAGSKNSIDCTLILTEGDSAKTLAVSGLG-VVGRDKYGVFPLRGKMLNVREASHK 500
Query: 393 -LFKKKEIQNLMAILGLVRNKKYSNA---KCLRYGQLMVMANQDEDGAHFKGLLINFFYS 448
+ + EI N++ I+GL K Y + K LRYG++M+M +QD+DG+H KGLLINF +
Sbjct: 501 QIMENAEINNIIKIVGLQYKKNYEDRESLKSLRYGKIMIMTDQDQDGSHIKGLLINFIHH 560
Query: 449 FWPSLLKVSKFMSVFTFPIIKVSMYNASDSKKGKELSFDSMQQYEDWKKELGNTANDWEI 508
WPSLL+ F+ F PIIKVS K +E+ F S+ ++E+WK N N W+I
Sbjct: 561 NWPSLLR-HNFLEEFITPIIKVS-------KNKEEIPFYSIPEFEEWKSSTQNY-NSWKI 611
Query: 509 KYCQGL--SSAQEVREYFQDLGKHRAYFVWDD-KDETSIKLAFSKK-AEEWEAWIGNSQ- 563
KY +GL S+++E +EYF D+ +HR F + +D+ +I LAFSKK EE + W+ N
Sbjct: 612 KYYKGLGTSTSKEAKEYFADMARHRIGFKYSGPEDDAAITLAFSKKKVEERKEWLTNFME 671
Query: 564 ----------PGTCNY--ERKVINYRNFVNKELLFSSMKNLQRSIPSVVDGLTLGQRKTL 611
P Y + + Y +F+NKEL+ S + +RSIPS+VDGL GQRK L
Sbjct: 672 DRRQRNVHGLPEDYLYGKDTNYLTYNDFINKELVLFSNSDNERSIPSLVDGLKPGQRKVL 731
Query: 612 FCSFEMNLTKQTEVCRLTAYVTEHSYCHYGQQSIASTIIGMAQDFVGSNNINLLIPYGQF 671
F F+ N ++ + +L V E S H+G+ S+ TII +AQ+FVGSNN+NLL P GQF
Sbjct: 732 FTCFKRNDKREVKGAQLAGSVAEMSSYHHGEASLMMTIINLAQNFVGSNNLNLLQPIGQF 791
Query: 672 GTRDLGGKDHASSRKLYIKLNCVTRLLFPVDDDKLLEYLNEDGRSIQPNWYIPIIPLVLV 731
GTR GGKD AS R ++ L+ + RLLFP DD +L +L +D + ++P WY+PIIP+VL+
Sbjct: 792 GTRLHGGKDSASPRYIFTMLSPLARLLFPPVDDNVLRFLYDDNQRVEPEWYMPIIPMVLI 851
Query: 732 NGCHAMGTCFSSDIPKYHPYEIIKNVRRFLNNEEMVSMKPWYKGFGGTIEKSAKGIGYYT 791
NG +GT +S IP + E + N+R L+ +E + M P YK F GTI++ G Y
Sbjct: 852 NGAEGIGTGWSCKIPNFDIRETVNNIRCLLDGKEPLPMLPSYKNFKGTIDE--LGPNQYV 909
Query: 792 VDGLVERINEQTFRIKELPIRMWTEDYKKFLEKITARG------LIESFRQKGDYEIVNF 845
+ G V ++ T I ELP+R WT+ YK+ + + G LI +++ V F
Sbjct: 910 ISGEVSILDSTTIEITELPVRTWTQTYKEQVLEPMLNGTEKTPPLITDYKEYHTDTTVKF 969
Query: 846 KVNVKQEEIATIEKDEELRKKFNLTCTISTSNMYLFDAEGKIKKYNTPEQILEEFCTLRL 905
V + +E++A E L K F L ++ ++M LFD G +KKY +P+ IL+EF LRL
Sbjct: 970 VVKMSEEKLAEAEA-VGLHKVFKLQTNLTCNSMVLFDHVGFLKKYESPQDILKEFFELRL 1028
Query: 906 KYYVRRKQYLVKNFTQLLRSLRIKQKFISNIVNEKFNPINRRAELLIEIL-KKGKSAEPH 964
+YY RK++L+ L + +FI ++ K N+ + LI++L ++G ++P
Sbjct: 1029 RYYGLRKEWLIGMLGAESAKLNNQARFILEKIDGKIVIENKPKKELIQVLIQRGYESDP- 1087
Query: 965 VAGAIDYSSKEQEAGEQESVSQSVSIEGATLGDYEYLSSLPFENFTLESLKKLEAELDEK 1024
V + +KE+E G+ ES +S + AT D+ YL ++P T E +L + D K
Sbjct: 1088 VKAWKELQNKEEEEGD-ESGEESAA---ATGPDFNYLLNMPLWYLTKEKKDELCKQRDNK 1143
Query: 1025 ENELKTLEHTSPDLMWLNDLELFEKEFD 1052
+ EL+ L+H SP +W DL F +E D
Sbjct: 1144 DKELEDLKHKSPSDLWKEDLAAFVEELD 1171
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.323 0.139 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,723,804,566
Number of Sequences: 2790947
Number of extensions: 73689616
Number of successful extensions: 213593
Number of sequences better than 10.0: 3461
Number of HSP's better than 10.0 without gapping: 1054
Number of HSP's successfully gapped in prelim test: 2409
Number of HSP's that attempted gapping in prelim test: 207206
Number of HSP's gapped (non-prelim): 4399
length of query: 1053
length of database: 848,049,833
effective HSP length: 138
effective length of query: 915
effective length of database: 462,899,147
effective search space: 423552719505
effective search space used: 423552719505
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 80 (35.4 bits)
Medicago: description of AC145202.5


