

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144931.14 - phase: 0 /pseudo
(122 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
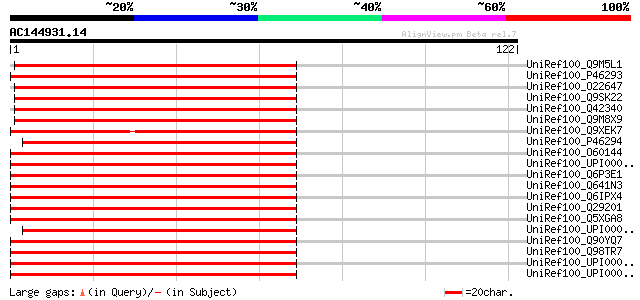
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9M5L1 40S ribosomal protein S16 [Euphorbia esula] 138 3e-32
UniRef100_P46293 40S ribosomal protein S16 [Gossypium hirsutum] 130 5e-30
UniRef100_O22647 40S ribosomal protein S16 [Fritillaria agrestis] 127 8e-29
UniRef100_Q9SK22 40S ribosomal protein S16 [Arabidopsis thaliana] 123 8e-28
UniRef100_Q42340 40S ribosomal protein S16 [Arabidopsis thaliana] 123 1e-27
UniRef100_Q9M8X9 Putative 40S ribosomal protein S16 [Arabidopsis... 114 4e-25
UniRef100_Q9XEK7 40S ribosomal protein S16 [Tortula ruralis] 112 2e-24
UniRef100_P46294 40S ribosomal protein S16 [Oryza sativa] 105 2e-22
UniRef100_O60144 40S ribosomal protein S16 [Schizosaccharomyces ... 104 5e-22
UniRef100_UPI00003A96FE UPI00003A96FE UniRef100 entry 103 9e-22
UniRef100_Q6P3E1 Hypothetical protein [Rattus norvegicus] 103 9e-22
UniRef100_Q641N3 40S ribosomal protein S16 [Homo sapiens] 103 9e-22
UniRef100_Q6IPX4 Hypothetical protein [Homo sapiens] 103 9e-22
UniRef100_Q29201 40S ribosomal protein S16 [Sus scrofa] 103 9e-22
UniRef100_Q5XGA8 Hypothetical protein [Xenopus tropicalis] 103 1e-21
UniRef100_UPI000023EF76 UPI000023EF76 UniRef100 entry 102 3e-21
UniRef100_Q90YQ7 40S ribosomal protein S16 [Ictalurus punctatus] 102 3e-21
UniRef100_Q98TR7 40S ribosomal protein S16 [Heteropneustes fossi... 102 3e-21
UniRef100_UPI00002BB43E UPI00002BB43E UniRef100 entry 101 3e-21
UniRef100_UPI0000366D72 UPI0000366D72 UniRef100 entry 100 8e-21
>UniRef100_Q9M5L1 40S ribosomal protein S16 [Euphorbia esula]
Length = 144
Score = 138 bits (347), Expect = 3e-32
Identities = 66/68 (97%), Positives = 66/68 (97%)
Query: 2 EQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGVD 61
E VQCFGRKK AVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGVD
Sbjct: 6 ESVQCFGRKKTAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGVD 65
Query: 62 MRIRVKGG 69
MRIRVKGG
Sbjct: 66 MRIRVKGG 73
>UniRef100_P46293 40S ribosomal protein S16 [Gossypium hirsutum]
Length = 145
Score = 130 bits (328), Expect = 5e-30
Identities = 63/69 (91%), Positives = 64/69 (92%)
Query: 1 MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 60
+E VQCFGRKK AVAVTHCKRGRGLIKINGCPIELVEPEILRFKA EPILLLGR RF GV
Sbjct: 6 IESVQCFGRKKTAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAVEPILLLGRQRFTGV 65
Query: 61 DMRIRVKGG 69
DMRIRVKGG
Sbjct: 66 DMRIRVKGG 74
>UniRef100_O22647 40S ribosomal protein S16 [Fritillaria agrestis]
Length = 145
Score = 127 bits (318), Expect = 8e-29
Identities = 59/68 (86%), Positives = 64/68 (93%)
Query: 2 EQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGVD 61
E VQCFGRKK AVAVTHCKRGRGLIK+NG PIELV+PEILR+KA+EPILLLGRHRF GVD
Sbjct: 7 ESVQCFGRKKTAVAVTHCKRGRGLIKVNGSPIELVKPEILRYKAFEPILLLGRHRFVGVD 66
Query: 62 MRIRVKGG 69
MRIRV+GG
Sbjct: 67 MRIRVRGG 74
>UniRef100_Q9SK22 40S ribosomal protein S16 [Arabidopsis thaliana]
Length = 146
Score = 123 bits (309), Expect = 8e-28
Identities = 57/68 (83%), Positives = 62/68 (90%)
Query: 2 EQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGVD 61
E VQCFGRKK AVAVTHCKRG GLIK+NGCPIEL +PEILRFK +EPILLLG+HRFAGV+
Sbjct: 8 ESVQCFGRKKTAVAVTHCKRGSGLIKLNGCPIELFQPEILRFKIFEPILLLGKHRFAGVN 67
Query: 62 MRIRVKGG 69
MRIRV GG
Sbjct: 68 MRIRVNGG 75
>UniRef100_Q42340 40S ribosomal protein S16 [Arabidopsis thaliana]
Length = 146
Score = 123 bits (308), Expect = 1e-27
Identities = 56/68 (82%), Positives = 62/68 (90%)
Query: 2 EQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGVD 61
E VQCFGRKK AVAVTHCKRG GLIK+NGCPIEL +PEILRFK +EP+LLLG+HRFAGV+
Sbjct: 8 ESVQCFGRKKTAVAVTHCKRGSGLIKLNGCPIELFQPEILRFKIFEPVLLLGKHRFAGVN 67
Query: 62 MRIRVKGG 69
MRIRV GG
Sbjct: 68 MRIRVNGG 75
>UniRef100_Q9M8X9 Putative 40S ribosomal protein S16 [Arabidopsis thaliana]
Length = 146
Score = 114 bits (286), Expect = 4e-25
Identities = 52/68 (76%), Positives = 59/68 (86%)
Query: 2 EQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGVD 61
E VQCFGRKK A AVT+CKRG G+IK+NG PIEL +PEILRFK +EP+LLLG+HRFAGVD
Sbjct: 8 ESVQCFGRKKTATAVTYCKRGSGMIKLNGSPIELYQPEILRFKIFEPVLLLGKHRFAGVD 67
Query: 62 MRIRVKGG 69
MRIR GG
Sbjct: 68 MRIRATGG 75
>UniRef100_Q9XEK7 40S ribosomal protein S16 [Tortula ruralis]
Length = 142
Score = 112 bits (280), Expect = 2e-24
Identities = 54/69 (78%), Positives = 63/69 (91%), Gaps = 1/69 (1%)
Query: 1 MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 60
+E VQCFGRKK AVAVT+C+RGRGLI++ PIELVEPEILR+KA+EP+LLLGR +FAGV
Sbjct: 4 VESVQCFGRKKTAVAVTYCRRGRGLIRLT-VPIELVEPEILRYKAFEPVLLLGRSKFAGV 62
Query: 61 DMRIRVKGG 69
DMRIRVKGG
Sbjct: 63 DMRIRVKGG 71
>UniRef100_P46294 40S ribosomal protein S16 [Oryza sativa]
Length = 149
Score = 105 bits (263), Expect = 2e-22
Identities = 49/66 (74%), Positives = 57/66 (86%)
Query: 4 VQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGVDMR 63
VQCFGRKK AVAV++CK GRGLIK+NG PIEL+ PE+LR KA+EPILL GR RF +DMR
Sbjct: 13 VQCFGRKKAAVAVSYCKPGRGLIKVNGVPIELIRPEMLRLKAFEPILLAGRSRFKDIDMR 72
Query: 64 IRVKGG 69
IRV+GG
Sbjct: 73 IRVRGG 78
>UniRef100_O60144 40S ribosomal protein S16 [Schizosaccharomyces pombe]
Length = 140
Score = 104 bits (259), Expect = 5e-22
Identities = 46/69 (66%), Positives = 56/69 (80%)
Query: 1 MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 60
M+ VQCFG+K NA AV HCK G+GLIK+NG P+ LV+PEILR K YEPIL+ G +FAGV
Sbjct: 1 MQSVQCFGKKGNATAVAHCKVGKGLIKVNGAPLSLVQPEILRMKVYEPILVAGADKFAGV 60
Query: 61 DMRIRVKGG 69
D+R+RV GG
Sbjct: 61 DIRVRVSGG 69
>UniRef100_UPI00003A96FE UPI00003A96FE UniRef100 entry
Length = 160
Score = 103 bits (257), Expect = 9e-22
Identities = 45/69 (65%), Positives = 57/69 (82%)
Query: 1 MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 60
++ VQ FGRKK A AV HCKRG GLIK+NG P+E++EP L++K EP+LLLG+ RFAGV
Sbjct: 21 LQSVQVFGRKKTATAVAHCKRGNGLIKVNGRPLEMIEPRTLQYKLLEPVLLLGKERFAGV 80
Query: 61 DMRIRVKGG 69
D+R+RVKGG
Sbjct: 81 DIRVRVKGG 89
>UniRef100_Q6P3E1 Hypothetical protein [Rattus norvegicus]
Length = 161
Score = 103 bits (257), Expect = 9e-22
Identities = 45/69 (65%), Positives = 57/69 (82%)
Query: 1 MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 60
++ VQ FGRKK A AV HCKRG GLIK+NG P+E++EP L++K EP+LLLG+ RFAGV
Sbjct: 22 LQSVQVFGRKKTATAVAHCKRGNGLIKVNGRPLEMIEPRTLQYKLLEPVLLLGKERFAGV 81
Query: 61 DMRIRVKGG 69
D+R+RVKGG
Sbjct: 82 DIRVRVKGG 90
>UniRef100_Q641N3 40S ribosomal protein S16 [Homo sapiens]
Length = 157
Score = 103 bits (257), Expect = 9e-22
Identities = 45/69 (65%), Positives = 57/69 (82%)
Query: 1 MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 60
++ VQ FGRKK A AV HCKRG GLIK+NG P+E++EP L++K EP+LLLG+ RFAGV
Sbjct: 18 LQSVQVFGRKKTATAVAHCKRGNGLIKVNGRPLEMIEPRTLQYKLLEPVLLLGKERFAGV 77
Query: 61 DMRIRVKGG 69
D+R+RVKGG
Sbjct: 78 DIRVRVKGG 86
>UniRef100_Q6IPX4 Hypothetical protein [Homo sapiens]
Length = 152
Score = 103 bits (257), Expect = 9e-22
Identities = 45/69 (65%), Positives = 57/69 (82%)
Query: 1 MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 60
++ VQ FGRKK A AV HCKRG GLIK+NG P+E++EP L++K EP+LLLG+ RFAGV
Sbjct: 7 LQSVQVFGRKKTATAVAHCKRGNGLIKVNGRPLEMIEPRTLQYKLLEPVLLLGKERFAGV 66
Query: 61 DMRIRVKGG 69
D+R+RVKGG
Sbjct: 67 DIRVRVKGG 75
>UniRef100_Q29201 40S ribosomal protein S16 [Sus scrofa]
Length = 135
Score = 103 bits (257), Expect = 9e-22
Identities = 45/69 (65%), Positives = 57/69 (82%)
Query: 1 MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 60
++ VQ FGRKK A AV HCKRG GLIK+NG P+E++EP L++K EP+LLLG+ RFAGV
Sbjct: 6 LQSVQVFGRKKTATAVAHCKRGNGLIKVNGRPLEMIEPRTLQYKLLEPVLLLGKERFAGV 65
Query: 61 DMRIRVKGG 69
D+R+RVKGG
Sbjct: 66 DIRVRVKGG 74
>UniRef100_Q5XGA8 Hypothetical protein [Xenopus tropicalis]
Length = 146
Score = 103 bits (256), Expect = 1e-21
Identities = 45/69 (65%), Positives = 57/69 (82%)
Query: 1 MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 60
++ VQ FGRKK A AV HCKRG GLIK+NG P+E++EP L++K EP+LLLG+ RFAGV
Sbjct: 7 LQSVQVFGRKKTATAVAHCKRGNGLIKVNGRPLEMIEPATLQYKLLEPVLLLGKERFAGV 66
Query: 61 DMRIRVKGG 69
D+R+RVKGG
Sbjct: 67 DIRVRVKGG 75
>UniRef100_UPI000023EF76 UPI000023EF76 UniRef100 entry
Length = 143
Score = 102 bits (253), Expect = 3e-21
Identities = 44/66 (66%), Positives = 56/66 (84%)
Query: 4 VQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGVDMR 63
VQCFG+KK A AV HCK G+GLIK+NG P++LV+PEILRFK YEP+L++G +FA VD+R
Sbjct: 7 VQCFGKKKTATAVAHCKAGKGLIKVNGRPLQLVQPEILRFKVYEPLLVVGLDKFANVDVR 66
Query: 64 IRVKGG 69
+RV GG
Sbjct: 67 VRVSGG 72
>UniRef100_Q90YQ7 40S ribosomal protein S16 [Ictalurus punctatus]
Length = 146
Score = 102 bits (253), Expect = 3e-21
Identities = 45/69 (65%), Positives = 56/69 (80%)
Query: 1 MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 60
++ VQ FGRKK A AV HCKRG GLIK+NG P+E +EP L++K EP+LLLG+ RFAGV
Sbjct: 7 LQSVQVFGRKKTATAVAHCKRGNGLIKVNGRPLETIEPATLQYKLLEPVLLLGKERFAGV 66
Query: 61 DMRIRVKGG 69
D+R+RVKGG
Sbjct: 67 DIRVRVKGG 75
>UniRef100_Q98TR7 40S ribosomal protein S16 [Heteropneustes fossilis]
Length = 146
Score = 102 bits (253), Expect = 3e-21
Identities = 45/69 (65%), Positives = 56/69 (80%)
Query: 1 MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 60
++ VQ FGRKK A AV HCKRG GLIK+NG P+E +EP L++K EP+LLLG+ RFAGV
Sbjct: 7 LQSVQVFGRKKTATAVAHCKRGNGLIKVNGRPLETIEPATLQYKLLEPVLLLGKERFAGV 66
Query: 61 DMRIRVKGG 69
D+R+RVKGG
Sbjct: 67 DIRVRVKGG 75
>UniRef100_UPI00002BB43E UPI00002BB43E UniRef100 entry
Length = 146
Score = 101 bits (252), Expect = 3e-21
Identities = 44/69 (63%), Positives = 57/69 (81%)
Query: 1 MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 60
++ VQ FGRKK A AV HCKRG GL+K+NG P+E++EP L++K EP+LLLG+ RFAGV
Sbjct: 7 LQSVQVFGRKKTATAVAHCKRGNGLLKVNGKPLEMLEPATLQYKLLEPVLLLGKERFAGV 66
Query: 61 DMRIRVKGG 69
D+R+RVKGG
Sbjct: 67 DIRVRVKGG 75
>UniRef100_UPI0000366D72 UPI0000366D72 UniRef100 entry
Length = 149
Score = 100 bits (249), Expect = 8e-21
Identities = 43/69 (62%), Positives = 57/69 (82%)
Query: 1 MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 60
++ VQ FGRKK A AV HC+RG GLIK+NG P+E+ EP +L+++ EP+LLLG+ RFAGV
Sbjct: 10 LQSVQVFGRKKTATAVAHCRRGNGLIKVNGQPLEMTEPRMLQYELLEPVLLLGKERFAGV 69
Query: 61 DMRIRVKGG 69
D+R+RVKGG
Sbjct: 70 DIRVRVKGG 78
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.325 0.147 0.448
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 150,222,378
Number of Sequences: 2790947
Number of extensions: 4360851
Number of successful extensions: 8029
Number of sequences better than 10.0: 194
Number of HSP's better than 10.0 without gapping: 124
Number of HSP's successfully gapped in prelim test: 70
Number of HSP's that attempted gapping in prelim test: 7899
Number of HSP's gapped (non-prelim): 194
length of query: 122
length of database: 848,049,833
effective HSP length: 98
effective length of query: 24
effective length of database: 574,537,027
effective search space: 13788888648
effective search space used: 13788888648
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 67 (30.4 bits)
Medicago: description of AC144931.14


