

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144931.10 + phase: 0 /pseudo
(416 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
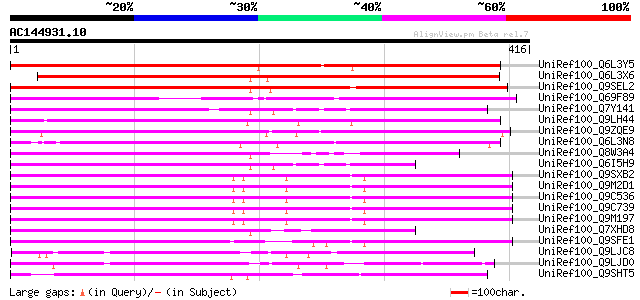
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6L3Y5 Putative gag-pol polyprotein [Solanum demissum] 422 e-117
UniRef100_Q6L3X6 Putative polyprotein [Solanum demissum] 372 e-101
UniRef100_Q9SEL2 Gag-pol polyprotein [Vitis vinifera] 351 2e-95
UniRef100_Q69F89 Gag-pol polyprotein [Phaseolus vulgaris] 313 6e-84
UniRef100_Q7Y141 Putative polyprotein [Oryza sativa] 261 3e-68
UniRef100_Q9LH44 Copia-like retrotransposable element [Arabidops... 254 4e-66
UniRef100_Q9ZQE9 Putative retroelement pol polyprotein [Arabidop... 232 1e-59
UniRef100_Q6L3N8 Putative gag-pol polyprotein [Solanum demissum] 229 8e-59
UniRef100_Q8W3A4 Putative gag-pol polyprotein [Oryza sativa] 225 2e-57
UniRef100_Q6I5H9 Putative polyprotein [Oryza sativa] 218 3e-55
UniRef100_Q9SXB2 T28P6.8 protein [Arabidopsis thaliana] 204 5e-51
UniRef100_Q9M2D1 Copia-type polyprotein [Arabidopsis thaliana] 204 5e-51
UniRef100_Q9C536 Copia-type polyprotein, putative [Arabidopsis t... 202 1e-50
UniRef100_Q9C739 Copia-type polyprotein, putative [Arabidopsis t... 202 1e-50
UniRef100_Q9M197 Copia-type reverse transcriptase-like protein [... 201 4e-50
UniRef100_Q7XHD8 Putative copia-type polyprotein [Oryza sativa] 196 1e-48
UniRef100_Q9SFE1 T26F17.17 [Arabidopsis thaliana] 189 2e-46
UniRef100_Q9LJC8 Gag-pol polyprotein-like [Arabidopsis thaliana] 151 3e-35
UniRef100_Q9LJD0 Gag-pol polyprotein-like [Arabidopsis thaliana] 150 5e-35
UniRef100_Q9SHT5 Putative retroelement pol polyprotein [Arabidop... 142 2e-32
>UniRef100_Q6L3Y5 Putative gag-pol polyprotein [Solanum demissum]
Length = 1133
Score = 422 bits (1086), Expect = e-117
Identities = 213/420 (50%), Positives = 292/420 (68%), Gaps = 28/420 (6%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
MKTYL+ LW VE+ N P PL NPTL QIR+H +E+AK +AL+ IHAA+ D IF +
Sbjct: 1 MKTYLKGLGLWKYVEEDNAPEPLRANPTLQQIRSHEEEIAKGPKALSFIHAAVSDSIFTR 60
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
I+ E AKE+W+ L +E+QGS+RT++M++LNLR+EFE L+MKETE V+++ DRI K+ Q
Sbjct: 61 IMTCESAKESWDMLKQEFQGSDRTRQMQILNLRKEFEMLRMKETEKVKDYIDRIMKIFNQ 120
Query: 121 IRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRSLR 180
IRLLGE + +QR+VEK++V LPE FEAKISS+E+ + S +T+ EL NAL A E+R++ R
Sbjct: 121 IRLLGEKVEEQRIVEKVIVTLPEKFEAKISSIEDTWDMSRVTLNELSNALLAVEKRKAFR 180
Query: 181 MEENV-EGAFLASNKGKNQ----------------------SFKSFGEKKFPPCPHCKKD 217
EE+ E A +A+ + K Q + + K+PPCP+CKK
Sbjct: 181 KEESTSEIALVAAQRSKAQLDGEPKRQQNGRYGKEKKEQSNNRGRYRRSKYPPCPYCKKT 240
Query: 218 THLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQEQQARVVEHHQEDEEQLFKAS---- 273
H DKFCWYRPGV+C+ C Q GHV+KVCK N Q QA+V E+ ++ EE+LF AS
Sbjct: 241 NHTDKFCWYRPGVQCKLCKQFGHVDKVCKINQN-QPAQAQVTENVEDPEEKLFAASIAGE 299
Query: 274 CYLACSSKETWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVETL 333
C +A ++ WL+DSGCT+HMT N++ F+ LD+ ++SKV G+G+ V GKG VAV+T
Sbjct: 300 CNVAAQDQDVWLVDSGCTHHMTANLNIFEWLDKKYFSKVRLGDGRLVDAAGKGAVAVQTP 359
Query: 334 LGIKYILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLMIVEMRGKSFP 393
G+K I +VLFVPEI+QSLLSVGQ+++KNY+L FK+ C I DP G KL+ V+M +SFP
Sbjct: 360 SGMKIISNVLFVPEISQSLLSVGQLLDKNYALLFKDKTCEIMDPTGIKLLSVKMNNRSFP 419
>UniRef100_Q6L3X6 Putative polyprotein [Solanum demissum]
Length = 1758
Score = 372 bits (954), Expect = e-101
Identities = 178/387 (45%), Positives = 270/387 (68%), Gaps = 17/387 (4%)
Query: 23 LPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIKILNLEIAKEAWNKLMEEYQGSE 82
LP NPTLAQI+++++E K+ +A +++ A+ D +F +I+ + AKEAW++L EEYQGS+
Sbjct: 22 LPENPTLAQIKSNNEEKTKKSKAKSLMQNAVADSVFYRIMACKTAKEAWDRLKEEYQGSD 81
Query: 83 RTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCLP 142
RT++M+VLNL+REFE L M++ ET+ +++DRIS ++ IRLLGE+ +D+R+VEK+LV LP
Sbjct: 82 RTRQMQVLNLKREFECLNMQDDETISKYADRISLIVNNIRLLGEEFTDKRIVEKVLVTLP 141
Query: 143 EMFEAKISSLEENKNFSEITVPELVNALQASEQRRSLRMEENVEGAFLA----------- 191
E FE+KISS E++K+ ++++ EL+ ALQA EQRR++R ++ EGA
Sbjct: 142 ERFESKISSFEKSKDLGKLSLGELMGALQAQEQRRNMRRDKFTEGAVSVQKQIFGKGKQQ 201
Query: 192 ---SNKGKNQSFKSFGE--KKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCK 246
+NK K+ + G+ KKFPPC +CK+ THL+K+CW+R C C Q GH+ KVCK
Sbjct: 202 VNQNNKVKHDGGNNSGDVKKKFPPCKYCKRTTHLEKYCWWRVDAICGNCKQTGHISKVCK 261
Query: 247 NKTNQQ-EQQARVVEHHQEDEEQLFKASCYLACSSKETWLIDSGCTNHMTNNVSFFKELD 305
++ N QA+V + E+QLF S + S ++W++DSGCT+H+ N+ FK LD
Sbjct: 262 SRANASGSLQAQVADAADAHEDQLFAVSYFSINESSDSWILDSGCTHHLCNDAEMFKFLD 321
Query: 306 ESFYSKVVFGNGQHVKVKGKGVVAVETLLGIKYILDVLFVPEINQSLLSVGQMMEKNYSL 365
+++ SKV GNG+ V+VKG+G +++ + GIK I D+L+ P+++Q+LLSVGQM+E NYSL
Sbjct: 322 DTYKSKVKVGNGEAVEVKGRGTMSISIISGIKTIPDILYTPDMSQNLLSVGQMLENNYSL 381
Query: 366 HFKNMKCTIFDPYGSKLMIVEMRGKSF 392
HFKN +C + DP G +L V+M F
Sbjct: 382 HFKNHECVVSDPSGVELFYVKMSNIMF 408
>UniRef100_Q9SEL2 Gag-pol polyprotein [Vitis vinifera]
Length = 581
Score = 351 bits (900), Expect = 2e-95
Identities = 174/410 (42%), Positives = 267/410 (64%), Gaps = 15/410 (3%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
M +L+A +W+ VE+ PPL +PT+AQ++ H + ++ +A A + AA+ IFIK
Sbjct: 25 MTVHLQALDVWEAVEENYEVPPLGADPTVAQMKLHKERRTRKAKAKACLFAAVSPSIFIK 84
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
I+ ++ A E W L EEY+G ER K M+V+NL REFE KM+E++ V++++ ++ + +
Sbjct: 85 IMKIDSAAEIWEYLKEEYKGDERIKNMQVMNLIREFEMKKMRESDAVKDYAAQLLSIANK 144
Query: 121 IRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRSLR 180
+RLLG++ S++++V+KILV LP+ +EA ISSLE +K+ S I++ EL+++L+A EQRR +R
Sbjct: 145 VRLLGKEFSNEKIVQKILVTLPKKYEATISSLENSKDLSTISLTELLHSLEAVEQRRLMR 204
Query: 181 MEENVEGAFLA-------SNKGKNQSFKSFGEKK----FPPCPHCKKDTHLDKFCWYRPG 229
+ EGAF A GK + KS G + FPPCPHC+K H + CW+
Sbjct: 205 QGDTAEGAFQARMHKNAGHKNGKVNNNKSCGNNQKNGVFPPCPHCQKTNHSPQKCWWETR 264
Query: 230 VKCRACNQLGHVEKVCKNKTNQQEQQARVVEHHQEDEEQLFKASCYLACSSKETWLIDSG 289
+C C + GHVE++CK + ++ Q + +Q A+ S+ E+WL+DSG
Sbjct: 265 CECNKCGKQGHVERICKINSRKKLVQQLTIARRSNCLQQRVLAN----KSTSESWLVDSG 320
Query: 290 CTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVETLLGIKYILDVLFVPEIN 349
CTNHMTNN F+ELD + SKV GNG+++ VKGKG VA+E+ G+K I DVLFVP+IN
Sbjct: 321 CTNHMTNNQDLFRELDRTTISKVRIGNGEYIPVKGKGTVAIESQTGLKLIYDVLFVPDIN 380
Query: 350 QSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLMIVEMRGKSFPRGVEED 399
Q+LLSVGQ++EK + ++F++ C I D G ++ ++M+GKSF + ED
Sbjct: 381 QNLLSVGQLVEKEFKVYFEDRNCIIKDAEGKEVFNIKMKGKSFALDLLED 430
>UniRef100_Q69F89 Gag-pol polyprotein [Phaseolus vulgaris]
Length = 529
Score = 313 bits (802), Expect = 6e-84
Identities = 170/407 (41%), Positives = 245/407 (59%), Gaps = 41/407 (10%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
M+TYL A LW+ VE+ LP NPT+AQI++ D+ K +A A + AA+ +F +
Sbjct: 18 METYLEALDLWEAVEEDYEIQSLPENPTVAQIKSQKDKKMKRSKAKACLFAAVSPMVFTR 77
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
I++L+ AKE W+ L EY+G ER + M+ LNL REFE KMKE+E+++E+SDR+ + +
Sbjct: 78 IMSLKSAKEIWDYLKAEYEGDERIRGMQALNLIREFELQKMKESESIKEYSDRLLTIANK 137
Query: 121 IRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRSLR 180
K+ S+I++ EL+NALQA EQRR++R
Sbjct: 138 ---------------------------------NTKDLSKISLAELLNALQAQEQRRAMR 164
Query: 181 MEENVEGAFLASNKGKNQSFKSFGEKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLGH 240
+ +EG L ++G +++ G + +PPC HC K H CW RP KC CNQLGH
Sbjct: 165 QDGAIEGVLLVKHEG---GWRNKG-RNYPPCKHCNKLGHPPFKCWRRPDAKCNKCNQLGH 220
Query: 241 VEKVCKNKTNQQEQQARVVEHHQEDEEQLFKASCYLACSSKETWLIDSGCTNHMTNNVSF 300
+C+N++ QQ+ A++ E+E+ LF A+ + + S +WLIDSGCTNHMT++
Sbjct: 221 EAIICRNQSEQQDVDAQIAN---EEEDVLFVATGFCSNISSASWLIDSGCTNHMTHDKEL 277
Query: 301 FKELDESFYSKVVFGNGQHVKVKGKGVVAVETLLGIKYILDVLFVPEINQSLLSVGQMME 360
FKEL + +V GNG+H+ VKGK +A+ + GIK I DVLFVPEI+Q+LLSVGQ++E
Sbjct: 278 FKELRTTDVKRVRIGNGEHLTVKGKDTIAITSYKGIKIITDVLFVPEIDQNLLSVGQLLE 337
Query: 361 KNYSLHFKNMKCTIFDPYGSKLMIVEMRGKSFP-RGVEEDILACFSK 406
K Y + F+N C I D G L VEM+GKSF +EE+ +A SK
Sbjct: 338 KGYKVMFENKHCLIKDVDGRDLFKVEMKGKSFALNPMEEEHIAFKSK 384
>UniRef100_Q7Y141 Putative polyprotein [Oryza sativa]
Length = 1335
Score = 261 bits (667), Expect = 3e-68
Identities = 141/419 (33%), Positives = 244/419 (57%), Gaps = 50/419 (11%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
M+T L +Q LWD+VE G T Q ++ +++ + +AL +I + + +F +
Sbjct: 21 MRTLLLSQGLWDIVENGYQEYSAGETLTAEQKKSLAEDRMSDAKALFLIQQGVAESLFPR 80
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
I+ + +KEAW+KL EE+QGS++ +K+ LRR+F+ L MKE+E V+++ R+ +++ Q
Sbjct: 81 IIGAKKSKEAWDKLKEEFQGSQKVLAVKLQTLRRQFQNLLMKESEKVKDYFSRVIEIVNQ 140
Query: 121 IRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRSLR 180
+RL GED++DQ+VVEKIL+ LPE +E +++ EE+K+ S+ ++L++ E+R+ R
Sbjct: 141 MRLYGEDINDQKVVEKILISLPEKYEYIVAATEESKDLSK-------DSLESHEERKLQR 193
Query: 181 MEENVEGAFLA--SNKGKNQSFK-SFGEKKFPP--------------------------- 210
++E AF + S + +N F+ +F + FP
Sbjct: 194 EGSSIENAFQSKLSFRPQNSRFRGNFQKNGFPMRDRGYFQKNGFSRQKEDGQERREKGTS 253
Query: 211 -----CPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQEQQARVVEHHQED 265
C C+K +H CW + + C C + GH+ K C+ + + +A + ++
Sbjct: 254 SSNLWCDICQKSSHTTDMCWKK--MTCNKCKRKGHIAKYCRTR---EINRANFSQEKEKS 308
Query: 266 EEQLFKASCYLACSSK-ETWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKG 324
EE +F SC+ A K + W+IDSGCTNHM + + F+E+D S+++K+ GNG + +G
Sbjct: 309 EEMVF--SCHTAQEEKDDVWVIDSGCTNHMAADPNLFREMDSSYHAKIHMGNGSIAQSEG 366
Query: 325 KGVVAVETLLGIKYILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLM 383
KG VAV+T G K+I DVL VP++ Q+LLS+GQ++E Y+++F++ C I D ++L+
Sbjct: 367 KGTVAVQTADGPKFIKDVLLVPDLKQNLLSIGQLLEHGYAVYFEDFSCKILDRKNNRLV 425
>UniRef100_Q9LH44 Copia-like retrotransposable element [Arabidopsis thaliana]
Length = 1499
Score = 254 bits (648), Expect = 4e-66
Identities = 151/411 (36%), Positives = 231/411 (55%), Gaps = 19/411 (4%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
M T L+ + LWDV+E G P A R D+V K+ AL I+ +A+ D IF +
Sbjct: 24 MITILKTRKLWDVIENGVTSNSSPETSP-ALTRERDDQVMKDMMALQILQSAVSDSIFPR 82
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
I A EAWN L E+QGS + K + + LRRE+E LKM+E ET+ +F+ ++ + Q
Sbjct: 83 IAPASSATEAWNALEMEFQGSSQVKMINLQTLRREYENLKMEEGETINDFTTKLINLSNQ 142
Query: 121 IRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRSLR 180
+R+ GE+ SD +VV+KIL+ +P+ F++ + LE+ K+ S ++V EL+ L+A E+R +LR
Sbjct: 143 LRVHGEEKSDYQVVQKILISVPQQFDSIVGVLEQTKDLSTLSVTELIGTLKAHERRLNLR 202
Query: 181 MEENVEGAF---LASNKGKNQSFKSFGEKKFPPCPHCKKDTHLDKFCWYRPGV------- 230
+ EGAF ++G+N+ K K C CK++ H + C+ +
Sbjct: 203 EDRINEGAFNGEKLGSRGENKQNKIRHGKTNMWCGVCKRNNHNEVDCFRKKSESISQRGG 262
Query: 231 ----KCRACNQLGHVEKVCK-NKTNQQEQQARVVEHHQEDE-EQLFKA--SCYLACSSKE 282
+C C++ GH+ + CK K + E +EDE LF A ++ +E
Sbjct: 263 SYERRCYVCDKQGHIARDCKLRKGERAHLSIEESEDEKEDECHMLFSAVEEKEISTIGEE 322
Query: 283 TWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVETLLGIKYILDV 342
TWL+DSGCTNHM+ +V F LD S + GNG V +GKG + V T G I DV
Sbjct: 323 TWLVDSGCTNHMSKDVRHFIALDRSKKIIIRIGNGGKVVSEGKGDIRVSTNKGDHVIKDV 382
Query: 343 LFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLMIVEMRGKSFP 393
L+VPE+ ++LLSV QM+ Y + F++ KC I D G K++ ++M+ +SFP
Sbjct: 383 LYVPELARNLLSVSQMISNGYRVIFEDNKCVIQDLKGRKILDIKMKDRSFP 433
>UniRef100_Q9ZQE9 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1347
Score = 232 bits (592), Expect = 1e-59
Identities = 139/426 (32%), Positives = 228/426 (52%), Gaps = 28/426 (6%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLP--NNPTLAQIRNHSDE-VAKEGRALAIIHAALHDDI 57
M T R + LW VVE+G P+ P A+ + +E V + AL I+ A+ D I
Sbjct: 24 MATIFRTRKLWSVVEEGVPVEPVQAEETPETARAKTLREEAVTNDTMALQILQTAVTDQI 83
Query: 58 FIKILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKV 117
F +I +KEAW+ L +EYQGS + + +K+ +LRRE+E LKM + + ++ F+D++ +
Sbjct: 84 FSRIAAASSSKEAWDVLKDEYQGSPQVRLVKLQSLRREYENLKMYDNDNIKTFTDKLIVL 143
Query: 118 ITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRR 177
Q+ GE ++ ++++KIL+ LP F++ +S LE+ ++ +T+ EL+ L+A E R
Sbjct: 144 EIQLTYHGEKKTNTQLIQKILISLPAKFDSIVSVLEQTRDLDALTMSELLGILKAQEARV 203
Query: 178 SLRMEENVEGAFLASNKGKNQSFKSFG-------EKKFPPCPHCKKDTHLDKFCWYRP-- 228
+ R E EGAF +KG+ FK +KK+ C K H ++ C +P
Sbjct: 204 TAREESTKEGAFYVRSKGRESGFKQDNTNNRVNQDKKW--CGFHKSSKHTEEECREKPKN 261
Query: 229 ---------GVKCRACNQLGHVEKVCKNKTNQQEQQARVVEHHQEDEEQLFKASCYLACS 279
+KC C ++GH C++K N++ + E ++ LF AS + +
Sbjct: 262 DDHGKNKRSNIKCYKCGKIGHYANECRSK-NKERAHVTLEEEDVNEDHMLFSASEEESTT 320
Query: 280 SKE-TWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVETLLGIKY 338
+E WL+DSGCTNHMT +F +++S + NG V GKG + V T G +
Sbjct: 321 LREDVWLVDSGCTNHMTKEERYFSNINKSIKVPIRVRNGDIVMTAGKGDITVMTRHGKRI 380
Query: 339 ILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLMIVEMRGKSFP---RG 395
I +V VP + ++LLSV Q++ Y + F++ +C I D G ++M +EM KSF
Sbjct: 381 IKNVFLVPGLEKNLLSVPQIISSGYWVRFQDKRCIIQDANGKEIMNIEMTDKSFKIKLSS 440
Query: 396 VEEDIL 401
VEE+ +
Sbjct: 441 VEEEAM 446
>UniRef100_Q6L3N8 Putative gag-pol polyprotein [Solanum demissum]
Length = 1333
Score = 229 bits (585), Expect = 8e-59
Identities = 137/405 (33%), Positives = 225/405 (54%), Gaps = 22/405 (5%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
MKT ++Q LWD+VE G +P Q+R H ++ +AL I AL D+IF +
Sbjct: 29 MKTLFKSQELWDIVETG-----IPEG-NANQMREHRK---RDSKALFTIQQALDDEIFPR 79
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
I +E +K+AW L +EY G ++ +K+ LRR+FE L M E E+V+ + R S ++ +
Sbjct: 80 ISAVETSKQAWEILKQEYFGDDKVITVKLQTLRRDFETLFMNENESVQGYLSRTSAIVNR 139
Query: 121 IRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRSLR 180
+R GE + +Q VV K+L L FE ++++EE+K+ S + EL+++L A E R +
Sbjct: 140 MRSYGEKIDNQIVVSKVLRSLTTKFEHVVTAIEESKDLSTYSFDELMSSLLAHEDRLNRS 199
Query: 181 MEE------NVEGAFLASNKGKNQSFKSFGEKKFPPCPH---CKKDTHLDKFCWYRPGVK 231
E+ V+G F K +N + + G F + + +F Y+ ++
Sbjct: 200 REKVQEKAFQVKGEFSYKGKAENSAGRGHGRGNFRGRGRGGSGRGRNQVGEFRQYKSNIQ 259
Query: 232 CRACNQLGHVEKVCKNKTNQQEQQARVVEHHQEDEEQLFKASCYLACSSKETWLIDSGCT 291
CR C + GH E C K +++ A + + E+E +LF AS + S+ W IDSGC+
Sbjct: 260 CRYCKKFGHKEVDCWTKQKDEQKDANFTQ-NVEEESKLFMASSQITESANAVWFIDSGCS 318
Query: 292 NHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVETLLG-IKYILDVLFVPEINQ 350
NHM+++ S F++LDES S+V G+ + V ++GKG V ++T+ G +K++ DV +VP +
Sbjct: 319 NHMSSSKSLFRDLDESQKSEVRLGDDKQVHIEGKGTVEIKTVQGNVKFLYDVQYVPTLAH 378
Query: 351 SLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLM--IVEMRGKSFP 393
+LLSVGQ+M YS+ F + C I D + + + + K FP
Sbjct: 379 NLLSVGQLMTSGYSVVFYDNACDIKDKESGRTIARVPMTQNKMFP 423
>UniRef100_Q8W3A4 Putative gag-pol polyprotein [Oryza sativa]
Length = 1167
Score = 225 bits (574), Expect = 2e-57
Identities = 124/363 (34%), Positives = 217/363 (59%), Gaps = 46/363 (12%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
M+T L +Q LWD+VE G T Q ++ +++ + +AL +I + + +F +
Sbjct: 21 MRTLLLSQGLWDIVENGYQEYSAGETLTAEQKKSLAEDRMSDAKALFLIQQGVAESLFPR 80
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
I+ + +KEAW+KL EE+QGS++ +K+ LRR+F+ L MKE+E V+++ R+ +++ Q
Sbjct: 81 IIGAKKSKEAWDKLKEEFQGSQKVLAVKLQTLRRQFQNLLMKESEKVKDYFSRVIEIVNQ 140
Query: 121 IRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRSLR 180
IRL GED++DQ+VVE+IL+ LPE +E ++++EE+K+ S +T+ +L+++L++ E+R+ R
Sbjct: 141 IRLYGEDINDQKVVEEILISLPEKYEYIVAAIEESKDLSTLTIQQLMSSLESHEERKLQR 200
Query: 181 MEENVEGAFLA--SNKGKNQSFK-SFGEKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQ 237
++E AF + S + +N F+ +F + FP
Sbjct: 201 EGSSIENAFQSKLSFRPQNSRFRGNFQKNGFP-------------------------MRD 235
Query: 238 LGHVEKVCKNKTNQQEQQARVVEHHQEDEEQLFKASCYLACSSKETWLIDSGCTNHMTNN 297
G+ + KN ++Q++ + +E EE+ + W+IDSGCTNHM +
Sbjct: 236 RGYFQ---KNGFSRQKEDG---QERREKEEK------------DDVWVIDSGCTNHMAAD 277
Query: 298 VSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVETLLGIKYILDVLFVPEINQSLLSVGQ 357
+ F+E+D S+++K+ GNG + +GKG VAV+T G K+I DVL VP++ Q+LLS+GQ
Sbjct: 278 PNLFREMDSSYHAKIHMGNGSIAQSEGKGTVAVQTADGPKFIKDVLLVPDLKQNLLSIGQ 337
Query: 358 MME 360
++E
Sbjct: 338 LLE 340
>UniRef100_Q6I5H9 Putative polyprotein [Oryza sativa]
Length = 1136
Score = 218 bits (554), Expect = 3e-55
Identities = 117/360 (32%), Positives = 207/360 (57%), Gaps = 42/360 (11%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
M+T L +Q LWD+VE G T Q ++ ++ + +AL +I + + +F +
Sbjct: 21 MRTLLLSQGLWDIVENGYQEYSAGETLTAEQKKSLVEDRMSDAKALFLIQQGVAESLFPR 80
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
I+ + +KEAW+KL EE+Q S++ +K+ LRR+F+ L+MKE+E V+++ R+ +++ Q
Sbjct: 81 IIGAKKSKEAWDKLKEEFQASQKVLAVKLQTLRRQFQNLQMKESEKVKDYFSRVIEIVNQ 140
Query: 121 IRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRSLR 180
+RL GED++DQ+VV+KIL+ LPE +E ++++EE+K+ S +T+ +L+++L++ E+R+ R
Sbjct: 141 MRLYGEDINDQKVVKKILISLPEKYEYIVAAIEESKDLSTLTIQQLMSSLESHEERKLQR 200
Query: 181 MEENVEGAFLA--SNKGKNQSFK-SFGEKKFPP--------------------------- 210
+VE AF + S + +N F+ +F + FP
Sbjct: 201 EGSSVENAFQSKLSFRPQNSRFRGNFQKNGFPMRDRGFQKNGFSRQNEYGQERREKGTSS 260
Query: 211 ----CPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQEQQARVVEHHQEDE 266
C C+K +H CW + + C C + GH+ K C+ + + +A + ++ E
Sbjct: 261 SNLWCDICQKSSHTTYMCWKK--MTCNKCKRKGHIAKYCRTR---EINRANFSQEKEKSE 315
Query: 267 EQLFKASCYLACSSK-ETWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGK 325
E +F SC+ A K + W+IDSGCTNHM + + F+E+D S+++K+ GNG + KGK
Sbjct: 316 EMVF--SCHTAQEEKDDVWVIDSGCTNHMAADPNLFREMDSSYHAKIHMGNGSIAQSKGK 373
>UniRef100_Q9SXB2 T28P6.8 protein [Arabidopsis thaliana]
Length = 1352
Score = 204 bits (518), Expect = 5e-51
Identities = 132/433 (30%), Positives = 218/433 (49%), Gaps = 31/433 (7%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
MK L A +W++VEKG P + + Q D ++ +AL +I+ L +D F K
Sbjct: 25 MKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRDKKALCLIYQGLDEDTFEK 84
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
++ AKEAW KL Y+G+++ KK+++ LR EFEAL+MKE E V ++ R+ V
Sbjct: 85 VVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVSDYFSRVLTVTNN 144
Query: 121 IRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRS-- 178
++ GE L D R++EK+L L FE ++ +EE K+ +T+ +L+ +LQA E+++
Sbjct: 145 LKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSLQAYEEKKKKK 204
Query: 179 -----------LRMEENVE-----GAFLASNKGKNQSFKSFGEKKFPPCPHCKKDTHL-- 220
+ EEN + G +G+ G + + + +
Sbjct: 205 EDIVEQVLNMQITKEENGQSYQRRGGGQVRGRGRGGYGNGRGWRPHEDNTNQRGENSSRG 264
Query: 221 -----DKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQ-EQQARVVEHHQEDEEQLFKASC 274
K + + VKC C + GH CK +N++ E++A VE ++E+ L AS
Sbjct: 265 RGKGHPKSRYDKSSVKCYNCGKFGHYASECKAPSNKKFEEKANYVEEKIQEEDMLLMAS- 323
Query: 275 YLACSSKET--WLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVET 332
Y KE W +DSG +NHM S F ELDES V G+ ++VKGKG + +
Sbjct: 324 YKKDEQKENHKWYLDSGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRL 383
Query: 333 LLG-IKYILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLMIVEM-RGK 390
G ++I +V ++P + ++LS+GQ++EK Y + K+ +I D + + V M + +
Sbjct: 384 KNGDHQFISNVYYIPSMKTNILSLGQLLEKGYDIRLKDNNLSIRDQESNLITKVPMSKNR 443
Query: 391 SFPRGVEEDILAC 403
F + DI C
Sbjct: 444 MFVLNIRNDIAQC 456
>UniRef100_Q9M2D1 Copia-type polyprotein [Arabidopsis thaliana]
Length = 1352
Score = 204 bits (518), Expect = 5e-51
Identities = 132/433 (30%), Positives = 218/433 (49%), Gaps = 31/433 (7%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
MK L A +W++VEKG P + + Q D ++ +AL +I+ L +D F K
Sbjct: 25 MKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRDKKALCLIYQGLDEDTFEK 84
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
++ AKEAW KL Y+G+++ KK+++ LR EFEAL+MKE E V ++ R+ V
Sbjct: 85 VVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVSDYFSRVLTVTNN 144
Query: 121 IRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRS-- 178
++ GE L D R++EK+L L FE ++ +EE K+ +T+ +L+ +LQA E+++
Sbjct: 145 LKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSLQAYEEKKKKK 204
Query: 179 -----------LRMEENVE-----GAFLASNKGKNQSFKSFGEKKFPPCPHCKKDTHL-- 220
+ EEN + G +G+ G + + + +
Sbjct: 205 EDIAEQVLNMQITKEENGQSYQRRGGGQVRGRGRGGYGNGRGWRPHEDNTNQRGENSSRG 264
Query: 221 -----DKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQ-EQQARVVEHHQEDEEQLFKASC 274
K + + VKC C + GH CK +N++ E++A VE ++E+ L AS
Sbjct: 265 RGKGHPKSRYDKSSVKCYNCGKFGHYASECKAPSNKKFEEKAHYVEEKIQEEDMLLMAS- 323
Query: 275 YLACSSKET--WLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVET 332
Y KE W +DSG +NHM S F ELDES V G+ ++VKGKG + +
Sbjct: 324 YKKDEQKENHKWYLDSGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRL 383
Query: 333 LLG-IKYILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLMIVEM-RGK 390
G ++I +V ++P + ++LS+GQ++EK Y + K+ +I D + + V M + +
Sbjct: 384 KNGDHQFISNVYYIPSMKTNILSLGQLLEKGYDIRLKDNNLSIRDQESNLITKVPMSKNR 443
Query: 391 SFPRGVEEDILAC 403
F + DI C
Sbjct: 444 MFVLNIRNDIAQC 456
>UniRef100_Q9C536 Copia-type polyprotein, putative [Arabidopsis thaliana]
Length = 1320
Score = 202 bits (514), Expect = 1e-50
Identities = 131/433 (30%), Positives = 218/433 (50%), Gaps = 31/433 (7%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
MK L A +W++VEKG P + + Q D ++ +AL +I+ L +D F K
Sbjct: 25 MKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRDKKALCLIYQGLDEDTFEK 84
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
++ AKEAW KL Y+G+++ KK+++ LR EFEAL+MKE E V ++ R+ V
Sbjct: 85 VVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVSDYFSRVLTVTNN 144
Query: 121 IRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRS-- 178
++ GE L D R++EK+L L FE ++ +EE K+ +T+ +L+ +LQA E+++
Sbjct: 145 LKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSLQAYEEKKKKK 204
Query: 179 -----------LRMEENVE-----GAFLASNKGKNQSFKSFGEKKFPPCPHCKKDTHL-- 220
+ EEN + G +G+ G + + + +
Sbjct: 205 EDIVEQVLNMQITKEENGQSYQRRGGGQVRGRGRGGYGNGRGWRPHEDNTNQRGENSSRG 264
Query: 221 -----DKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQ-EQQARVVEHHQEDEEQLFKASC 274
K + + VKC C + GH CK +N++ E++A VE ++E+ L AS
Sbjct: 265 RGKGHPKSRYDKSSVKCYNCGKFGHYASECKAPSNKKFEEKANYVEEKIQEEDMLLMAS- 323
Query: 275 YLACSSKET--WLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVET 332
Y +E W +DSG +NHM S F ELDES V G+ ++VKGKG + +
Sbjct: 324 YKKDEQEENHKWYLDSGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRL 383
Query: 333 LLG-IKYILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLMIVEM-RGK 390
G ++I +V ++P + ++LS+GQ++EK Y + K+ +I D + + V M + +
Sbjct: 384 KNGDHQFISNVYYIPSMKTNILSLGQLLEKGYDIRLKDNNLSIRDQESNLITKVPMSKNR 443
Query: 391 SFPRGVEEDILAC 403
F + DI C
Sbjct: 444 MFVLNIRNDIAQC 456
>UniRef100_Q9C739 Copia-type polyprotein, putative [Arabidopsis thaliana]
Length = 1352
Score = 202 bits (514), Expect = 1e-50
Identities = 131/433 (30%), Positives = 218/433 (50%), Gaps = 31/433 (7%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
MK L A +W++VEKG P + + Q D ++ +AL +I+ L +D F K
Sbjct: 25 MKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRDKKALCLIYQGLDEDTFEK 84
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
++ AKEAW KL Y+G+++ KK+++ LR EFEAL+MKE E V ++ R+ V
Sbjct: 85 VVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVSDYFSRVLTVTNN 144
Query: 121 IRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRS-- 178
++ GE L D R++EK+L L FE ++ +EE K+ +T+ +L+ +LQA E+++
Sbjct: 145 LKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSLQAYEEKKKKK 204
Query: 179 -----------LRMEENVE-----GAFLASNKGKNQSFKSFGEKKFPPCPHCKKDTHL-- 220
+ EEN + G +G+ G + + + +
Sbjct: 205 EDIIEQVLNMQITKEENGQSYQRRGGGQVRGRGRGGYGNGRGWRPHEDNTNQRGENSSRG 264
Query: 221 -----DKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQ-EQQARVVEHHQEDEEQLFKASC 274
K + + VKC C + GH CK +N++ E++A VE ++E+ L AS
Sbjct: 265 RGKGHPKSRYDKSSVKCYNCGKFGHYASECKAPSNKKFEEKANYVEEKIQEEDMLLMAS- 323
Query: 275 YLACSSKET--WLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVET 332
Y +E W +DSG +NHM S F ELDES V G+ ++VKGKG + +
Sbjct: 324 YKKDEQEENHKWYLDSGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRL 383
Query: 333 LLG-IKYILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLMIVEM-RGK 390
G ++I +V ++P + ++LS+GQ++EK Y + K+ +I D + + V M + +
Sbjct: 384 KNGDHQFISNVYYIPSMKTNILSLGQLLEKGYDIRLKDNNLSIRDQESNLITKVPMSKNR 443
Query: 391 SFPRGVEEDILAC 403
F + DI C
Sbjct: 444 MFVLNIRNDIAQC 456
>UniRef100_Q9M197 Copia-type reverse transcriptase-like protein [Arabidopsis
thaliana]
Length = 1272
Score = 201 bits (510), Expect = 4e-50
Identities = 130/433 (30%), Positives = 218/433 (50%), Gaps = 31/433 (7%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
MK L A +W++VEKG P + + Q D ++ +AL +I+ L +D F K
Sbjct: 25 MKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRDKKALCLIYQGLDEDTFEK 84
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
++ AKEAW KL Y+G+++ KK+++ LR EFEAL+MKE E V ++ R+ V
Sbjct: 85 VVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVSDYFSRVLTVTNN 144
Query: 121 IRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRS-- 178
++ GE L D R++EK+L L FE ++ +EE K+ +T+ +L+ +LQA E+++
Sbjct: 145 LKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSLQAYEEKKKKK 204
Query: 179 -----------LRMEENVE-----GAFLASNKGKNQSFKSFGEKKFPPCPHCKKDTHL-- 220
+ EEN + G +G+ G + + + +
Sbjct: 205 EDIVEQVLNMQITKEENGQSYQRRGGGQVRGRGRGGYGNGRGWRPHEDNTNQRGENSSRG 264
Query: 221 -----DKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQ-EQQARVVEHHQEDEEQLFKASC 274
K + + VKC C + GH CK +N++ +++A VE ++E+ L AS
Sbjct: 265 RGKGHPKSRYDKSSVKCYNCGKFGHYASECKAPSNKKFKEKANYVEEKIQEEDMLLMAS- 323
Query: 275 YLACSSKET--WLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVET 332
Y +E W +DSG +NHM S F ELDES V G+ ++VKGKG + +
Sbjct: 324 YKKDEQEENHKWYLDSGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRL 383
Query: 333 LLG-IKYILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLMIVEM-RGK 390
G ++I +V ++P + ++LS+GQ++EK Y + K+ +I D + + V M + +
Sbjct: 384 KNGDHQFISNVYYIPSMKTNILSLGQLLEKGYDIRLKDNNLSIRDKESNLITKVPMSKNR 443
Query: 391 SFPRGVEEDILAC 403
F + DI C
Sbjct: 444 MFVLNIRNDIAQC 456
>UniRef100_Q7XHD8 Putative copia-type polyprotein [Oryza sativa]
Length = 1350
Score = 196 bits (497), Expect = 1e-48
Identities = 106/329 (32%), Positives = 197/329 (59%), Gaps = 20/329 (6%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
M+T L +Q LWD+VE G T Q ++ +++ + +AL +I + + +F +
Sbjct: 21 MRTLLLSQGLWDIVENGYQEYSAGETLTAEQKKSLAEDRMSDAKALFLIQQGVAESLFPR 80
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
I+ + +KEAW+KL EE+QGS++ +K+ LRR+F+ L MKE+E V+++ R+ +++ Q
Sbjct: 81 IIGAKKSKEAWDKLKEEFQGSQKVLAVKLQTLRRQFQNLLMKESEKVKDYFPRVIEIVNQ 140
Query: 121 IRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRSLR 180
+RL GED++DQ+VVEKIL+ LPE +E ++++EE+K+ S +T+ +L+++L++ E+R+ R
Sbjct: 141 MRLYGEDINDQKVVEKILISLPEKYEYIVAAIEESKDLSTLTIQQLMSSLESHEERKLQR 200
Query: 181 MEENVEGAFLA--SNKGKNQSFK-SFGEKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQ 237
++E AF + S + +N F+ +F + +FP D+ + + G +
Sbjct: 201 EGSSIENAFQSKLSFRPQNSRFRGNFQKNEFPM---------RDRGYFQKNGFSRQK--- 248
Query: 238 LGHVEKVCKNKTNQQEQQARVVEHHQEDEEQLFKASCYLACSSKE-TWLIDSGCTNHMTN 296
E + + + E+ ++ H + + + SC+ A K+ W+IDSGCTNHM
Sbjct: 249 ----EDGQERREKELEKLIGLIFHKKRKKSEEMVFSCHTAQEEKDDVWVIDSGCTNHMAA 304
Query: 297 NVSFFKELDESFYSKVVFGNGQHVKVKGK 325
+ + F+E+D S+++K+ GNG + +GK
Sbjct: 305 DPNLFREMDSSYHAKIHMGNGSIAQSEGK 333
>UniRef100_Q9SFE1 T26F17.17 [Arabidopsis thaliana]
Length = 1291
Score = 189 bits (479), Expect = 2e-46
Identities = 126/418 (30%), Positives = 213/418 (50%), Gaps = 39/418 (9%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
MK L A +W++VEKG P + + Q D ++ +AL +I+ L +D F K
Sbjct: 25 MKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRDKKALCLIYQGLDEDTFEK 84
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
++ AKEAW KL Y+G ++ KK+++ LR EFEAL+MKE E V ++ R+ V
Sbjct: 85 VVEATSAKEAWEKLRTSYKGVDQVKKVRLQTLRGEFEALQMKEGELVSDYFSRVLTVTNN 144
Query: 121 IRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRSLR 180
++ GE L D R++EK+L L FE ++ +EE K+ +T+ +L+ +LQA E+++ +
Sbjct: 145 LKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLGSLQAYEEKK--K 202
Query: 181 MEENVEGAFLASNKGKNQSFKSFGEKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLGH 240
+E++ L K ++ +S+ R G + R + G+
Sbjct: 203 KKEDIVEQVLNMRITKEENGQSYQR---------------------RGGGEVRGRGRGGY 241
Query: 241 VE----KVCKNKTNQQ-------EQQARVVEHHQEDEEQLFKASCYLACSSKET--WLID 287
+ ++ TNQ+ E++A VE ++E+ L AS Y +E W +D
Sbjct: 242 GNGRGWRPHEDNTNQRAPSNKKFEEKANYVEEKIQEEDMLLMAS-YKKDEQEENHKWYLD 300
Query: 288 SGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVETLLG-IKYILDVLFVP 346
SG +NHM S F ELDES V G+ ++VKGKG + + G ++I +V ++P
Sbjct: 301 SGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRLKNGDHQFISNVYYIP 360
Query: 347 EINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLMIVEM-RGKSFPRGVEEDILAC 403
+ ++LS+GQ++EK Y + K+ +I D + + V M + + F + DI C
Sbjct: 361 SMKTNILSLGQLLEKGYDIRLKDNNLSIRDQESNLITKVPMSKNRMFVLNIRNDIAQC 418
>UniRef100_Q9LJC8 Gag-pol polyprotein-like [Arabidopsis thaliana]
Length = 420
Score = 151 bits (382), Expect = 3e-35
Identities = 116/386 (30%), Positives = 187/386 (48%), Gaps = 36/386 (9%)
Query: 2 KTYLRAQSLWDVVEKGNNPPP--LPNNPT------LAQIRNHSDEVAKEGRALAIIHAAL 53
K L +WDVV+ G +P P +P LAQ RN V K+ +AL I+ ++L
Sbjct: 23 KKRLVENGVWDVVQNGVSPNPTKIPQLAATIQAVDLAQWRNR---VIKDNKALKIMQSSL 79
Query: 54 HDDIFIKILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDR 113
D +F K +++ AKE W+ L + TK+ K+ L ++FE L M E E + + R
Sbjct: 80 PDSVFRKTISIASAKELWDLLKK----GNDTKEAKLRRLEKQFEKLMMYEGEPMDLYLKR 135
Query: 114 ISKVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQAS 173
+ ++ + +LG +SD +V+ K+L L ++ I L+E ++T+ +L+ A +
Sbjct: 136 VEEITERFEVLGNPISDDKVITKLLTSLSWPYDDSIPVLKEFMTLPDLTLRDLLKAFELF 195
Query: 174 EQRRSLRMEENVEGAFLASNKGKNQSFKSFGEKKFPPCPHCKKDTHL---DKFCWYRPGV 230
+E + K N K+ E+ PC C K+ H D + +RP V
Sbjct: 196 GSHPETMPQELM--------KFINILRKAHSERM--PCGICVKNNHNQEEDLYYNHRPRV 245
Query: 231 -KCRACNQL---GHVEKVCKNKTNQQEQQARVVEHHQEDEEQLFKASCYLACSSKETWLI 286
NQ GH + C N N Q+ + R+ Q+ E + + + W++
Sbjct: 246 VNLSGANQFRGQGHYARDCSNTRNLQQAEKRI----QKPEHLMLGVTVGGITFDEGMWMV 301
Query: 287 DSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVETLLGIKYILDVLFVP 346
+ T+HMT FF LD S+ ++V +G+ V +GKG V + T G K I +VLFVP
Sbjct: 302 HTTTTDHMTPYEKFFTTLDRSYRARVGLADGKVVMAEGKGDVMIMTREGKKRIKNVLFVP 361
Query: 347 EINQSLLSVGQMMEKNYSLHFKNMKC 372
IN++ LSV QM ++ S+ F KC
Sbjct: 362 GINKNALSVAQMTDQGCSVTFGGGKC 387
>UniRef100_Q9LJD0 Gag-pol polyprotein-like [Arabidopsis thaliana]
Length = 411
Score = 150 bits (380), Expect = 5e-35
Identities = 111/404 (27%), Positives = 195/404 (47%), Gaps = 47/404 (11%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIR-----NHSDEVAKEGRALAIIHAALHD 55
+KT L +WDVV+ G +P P N A I+ + V K+ +AL I+ ++L D
Sbjct: 22 VKTRLVENGVWDVVQNGVSPNPTTNPELAATIKAVDLAQWRNLVVKDNKALKIMQSSLPD 81
Query: 56 DIFIKILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRIS 115
+F K +++ AKE W+ L + TK+ K+ L ++FE L M E E ++ + R+
Sbjct: 82 SVFRKTISIASAKELWDLLKK----GNDTKEAKLRRLEKQFENLMMYEGERMKLYLKRLE 137
Query: 116 KVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQ 175
K+I ++ +LG +SD +V+ K+L LP ++ + L+E E+T +L+ A +
Sbjct: 138 KIIEELFVLGNPISDDKVIAKLLTSLPRSYDDSVPVLKEFMTLPELTHRDLLKAFEMFGS 197
Query: 176 RRSLRMEENVEGAFLASNKGKNQSFKSFGEKKFPPCPHCKKDTHLDKFCWYRPGV----- 230
+E ++ + +++ E+ + C C+ H + C+Y P V
Sbjct: 198 DPKTMPQELMKFIMILR--------RAWSERMW--CSACETYNHNQEDCYYNPRVVYLGG 247
Query: 231 ---KCRACNQLGHVEKVCKNKTNQQ--EQQARVVEHHQEDEEQLFKASCYLACSSKETWL 285
+C C + GH + C N N Q +++ V + DE+ W+
Sbjct: 248 GRGQCFQCGERGHFARDCCNTRNLQPIDRKREVKDDFTYDEDM---------------WM 292
Query: 286 IDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVETLLGIKY-ILDVLF 344
+ + NHM+ FF LD ++ + V NG V V+G G V + G++ I DVLF
Sbjct: 293 LYTTSENHMSPYEKFFTSLDRTYKAGVQLPNGNVVMVEGIGDVKF-MMEGVRMTIKDVLF 351
Query: 345 VPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLMIVEMR 388
VP IN+++LS +M ++ YS+ + + I D G K+ + +R
Sbjct: 352 VPWINKNILSFSRMKKRGYSMEVDDGEWIIRDRTG-KVFVKTLR 394
>UniRef100_Q9SHT5 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1307
Score = 142 bits (358), Expect = 2e-32
Identities = 101/406 (24%), Positives = 189/406 (45%), Gaps = 49/406 (12%)
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
M+ LR +W+ ++ G SD++ K A A++ ++ + ++
Sbjct: 1 MEATLRVHKVWETIDPG------------------SDDMEKNDMARALLFQSVPESTILQ 42
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
+ + +K W + G+ER K+ K+ L EF+ L MK+ ET+ EF RIS++ T+
Sbjct: 43 VGKHKTSKAMWEAIKTRNLGAERVKEAKLQTLMAEFDRLNMKDNETIDEFVGRISEISTK 102
Query: 121 IRLLGEDLSDQRVVEKILVCLP-EMFEAKISSLEENKNFSEITVPELVNALQASEQR--- 176
LGE++ + ++V+K L LP + + I++LE+ + + ++V ++ E R
Sbjct: 103 SESLGEEIEESKIVKKFLKSLPRKKYIHIIAALEQILDLNTTGFEDIVGRMKTYEDRVCD 162
Query: 177 -------RSLRMEENVEGAF-----------LASNKGKNQSFKSFGEKKFPPCPHCKKDT 218
+ M N E ++ +S +G+ +K C C K
Sbjct: 163 EDDSPEEQGKLMYANSESSYDTRGGRGRGRGRSSGRGRGGYGYQQRDKSKVICYRCDKTG 222
Query: 219 HLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQEQQARVVEHHQEDEEQLFKASCYLAC 278
H C R +L ++ +N + E ++ ++ E+ K + AC
Sbjct: 223 HYASECLDR-------LLKLIKAQEQQQNNEDDDEIESLMMHEVVYLNERSVKPKEFEAC 275
Query: 279 SSKETWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVETLLGI-K 337
S +W +D+G +NHMT N+ +F +L+E KV FG+ + +KGKG + + T GI K
Sbjct: 276 SD-NSWYLDNGASNHMTGNLQWFSKLNEMITGKVRFGDDSRIDIKGKGSIVLITKGGIRK 334
Query: 338 YILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLM 383
+ DV F+P++ +++S+GQ E + K+ + T+ D G L+
Sbjct: 335 TLTDVYFIPDLKSNIISLGQATEAGCDVRMKDDQLTLHDREGCLLL 380
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.135 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 686,641,473
Number of Sequences: 2790947
Number of extensions: 28541880
Number of successful extensions: 101257
Number of sequences better than 10.0: 948
Number of HSP's better than 10.0 without gapping: 405
Number of HSP's successfully gapped in prelim test: 544
Number of HSP's that attempted gapping in prelim test: 99498
Number of HSP's gapped (non-prelim): 1571
length of query: 416
length of database: 848,049,833
effective HSP length: 130
effective length of query: 286
effective length of database: 485,226,723
effective search space: 138774842778
effective search space used: 138774842778
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 76 (33.9 bits)
Medicago: description of AC144931.10


