

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144807.8 - phase: 0
(361 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
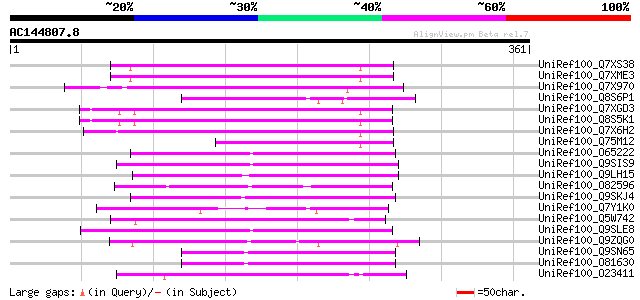
Sequences producing significant alignments: (bits) Value
UniRef100_Q7XS38 OSJNBa0081G05.2 protein [Oryza sativa] 90 1e-16
UniRef100_Q7XME3 OSJNBa0061G20.16 protein [Oryza sativa] 90 1e-16
UniRef100_Q7X970 BZIP-like protein [Oryza sativa] 87 9e-16
UniRef100_Q8S6P1 Putative reverse transcriptase [Oryza sativa] 80 7e-14
UniRef100_Q7XGD3 Putative retroelement [Oryza sativa] 80 9e-14
UniRef100_Q8S5K1 Putative retroelement [Oryza sativa] 80 9e-14
UniRef100_Q7X6H2 OSJNBb0003A12.9 protein [Oryza sativa] 78 3e-13
UniRef100_Q75M12 Hypothetical protein P0676G05.2 [Oryza sativa] 72 2e-11
UniRef100_O65222 F7N22.17 protein [Arabidopsis thaliana] 70 1e-10
UniRef100_Q9SIS9 Putative non-LTR retroelement reverse transcrip... 69 2e-10
UniRef100_Q9LH15 Ta11 non-LTR retroelement protein-like [Arabido... 69 2e-10
UniRef100_O82596 F11O4.11 protein [Arabidopsis thaliana] 67 1e-09
UniRef100_Q9SKJ4 Putative non-LTR retroelement reverse transcrip... 67 1e-09
UniRef100_Q7Y1K0 Hypothetical protein OSJNBb0056B16.21 [Oryza sa... 66 1e-09
UniRef100_Q5W742 Hypothetical protein OJ1005_D04.4 [Oryza sativa] 66 2e-09
UniRef100_Q9SLE8 Putative Ta11-like non-LTR retroelement protein... 64 5e-09
UniRef100_Q9ZQG0 Putative Ta11-like non-LTR retroelement protein... 64 9e-09
UniRef100_Q9SN65 Hypothetical protein F25I24.40 [Arabidopsis tha... 63 1e-08
UniRef100_O81630 F8M12.22 protein [Arabidopsis thaliana] 63 1e-08
UniRef100_O23411 Hypothetical protein AT4g15600 [Arabidopsis tha... 63 1e-08
>UniRef100_Q7XS38 OSJNBa0081G05.2 protein [Oryza sativa]
Length = 1711
Score = 89.7 bits (221), Expect = 1e-16
Identities = 58/207 (28%), Positives = 97/207 (46%), Gaps = 10/207 (4%)
Query: 71 DDSDIQERLEEC------QNSILGKIVSEKAIHRNSIQNALSNIWCNPKGFRVEHIGDKL 124
DD ++ LEE Q ++L + S K ++ ++ + W + + D L
Sbjct: 149 DDDAVEIELEEDEVKKAGQWTVLARFYSLKFPNQVALFEDMRRAWRLRAEMSYKSLKDNL 208
Query: 125 FHFFMDEQEDTKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCR 184
F + D K +++G PW R L+V + + + PIWV++ LP
Sbjct: 209 FIVSFSAEGDHKFVLQGGPWLHRGDALLVADFNGLVSPSMVPLETMPIWVRIYDLPLVMM 268
Query: 185 TKQMGIKIGSSIGTVLASELYEYP-DKKLIIKIKVNLAVSTPIKAGIYIGSAKDGAHWI- 242
K+ G+ GS +G V ++ E +K +I+V+L V+ P+K + I GA I
Sbjct: 269 NKERGVIYGSKLGKVREVDVQEDGCNKHDFFRIRVDLPVNRPLKRMLAIKIKVKGAEEIR 328
Query: 243 --DFRYENLPQFCFACGLIGHTEANCK 267
+ RYE +P FCF CG IGH++ C+
Sbjct: 329 RFNLRYERIPHFCFFCGFIGHSDKECE 355
>UniRef100_Q7XME3 OSJNBa0061G20.16 protein [Oryza sativa]
Length = 1285
Score = 89.7 bits (221), Expect = 1e-16
Identities = 58/207 (28%), Positives = 97/207 (46%), Gaps = 10/207 (4%)
Query: 71 DDSDIQERLEEC------QNSILGKIVSEKAIHRNSIQNALSNIWCNPKGFRVEHIGDKL 124
DD ++ LEE Q ++L + S K ++ ++ + W + + D L
Sbjct: 149 DDDAVEIELEEDEVKKAGQWTVLARFYSLKFPNQVALFEDMRRAWRLRAEMSYKSLKDNL 208
Query: 125 FHFFMDEQEDTKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCR 184
F + D K +++G PW R L+V + + + PIWV++ LP
Sbjct: 209 FIVSFSAEGDHKFVLQGGPWLHRGDALLVADFNGLVSPSMVPLETMPIWVRIYDLPLVMM 268
Query: 185 TKQMGIKIGSSIGTVLASELYEYP-DKKLIIKIKVNLAVSTPIKAGIYIGSAKDGAHWI- 242
K+ G+ GS +G V ++ E +K +I+V+L V+ P+K + I GA I
Sbjct: 269 NKERGVIYGSKLGKVREVDVQEDGCNKHDFFRIRVDLPVNRPLKRMLAIKIKVKGAEEIR 328
Query: 243 --DFRYENLPQFCFACGLIGHTEANCK 267
+ RYE +P FCF CG IGH++ C+
Sbjct: 329 RFNLRYERIPHFCFFCGFIGHSDKECE 355
>UniRef100_Q7X970 BZIP-like protein [Oryza sativa]
Length = 2367
Score = 86.7 bits (213), Expect = 9e-16
Identities = 60/240 (25%), Positives = 110/240 (45%), Gaps = 11/240 (4%)
Query: 39 NAITSQDPAIENDPTEETVTEDTAETDSFIYYDDSDIQERLEECQNSILGKIVSEKAIHR 98
N ++SQ P + + ED E I +++++Q+ Q +IL + S K ++
Sbjct: 318 NNLSSQGPTVFTEGEVVDEEEDAVE----IEIEENEVQKA---GQWTILARFYSLKFPNQ 370
Query: 99 NSIQNALSNIWCNPKGFRVEHIGDKLFHFFMDEQEDTKRIIRGNPWFFRNSWLIVHPWRR 158
+++ + W + + D LF + + D +++G+PW R L+V +
Sbjct: 371 SALFEDMRRAWRLRAEMSYKSLKDNLFIVSFNAEGDYNFVLQGDPWLHRGDALLVAEFNG 430
Query: 159 DIDARSLEFRHAPIWVQLRGLPTQCRTKQMGIKIGSSIGTVLASELYEYP-DKKLIIKIK 217
++ + PIWV++ LP + G GS +G V ++ E +K +I+
Sbjct: 431 LLNPSMVNLDTVPIWVRIYDLPLVMMNRARGELYGSKMGKVREVDVQEDGCNKHDFFRIR 490
Query: 218 VNLAVSTPIKAGIYIG---SAKDGAHWIDFRYENLPQFCFACGLIGHTEANCKLHDTRTN 274
V+L V+ P++ + I K+ RYE +P FCF CG IGH++ C+ T N
Sbjct: 491 VDLPVNRPLRRMLAIKIKVQGKEETRRFHLRYERVPHFCFFCGFIGHSDKECEKRLTNEN 550
>UniRef100_Q8S6P1 Putative reverse transcriptase [Oryza sativa]
Length = 1509
Score = 80.5 bits (197), Expect = 7e-14
Identities = 46/168 (27%), Positives = 78/168 (46%), Gaps = 8/168 (4%)
Query: 120 IGDKLFHFFMDEQEDTKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGL 179
+G+ F F + +R + PW F +++ + F PIWV+ L
Sbjct: 40 LGENHFLFTFHQASGKRRALEDGPWMFNKDLVVMIDLDETKLIEDMIFAFVPIWVRAMKL 99
Query: 180 PTQCRTKQMGIKIGSSIGTVLASELYEYPDKKLI---IKIKVNLAVSTPIKAGI--YIGS 234
P TK+ G+ IG +G + +L E D + ++IK+ + + P+ G+ ++G
Sbjct: 100 PLGLMTKETGMAIGREVGEFMTMDLEE--DGSAVGQFLRIKIRIDIRKPLMRGVTLFVG- 156
Query: 235 AKDGAHWIDFRYENLPQFCFACGLIGHTEANCKLHDTRTNAESRNKNV 282
A + W YE LP FC+ CG++GHTE C+ A NK++
Sbjct: 157 ADERPLWCPLVYEFLPDFCYICGIVGHTEKLCEKKLAEGEAPLFNKSL 204
>UniRef100_Q7XGD3 Putative retroelement [Oryza sativa]
Length = 1694
Score = 80.1 bits (196), Expect = 9e-14
Identities = 60/229 (26%), Positives = 101/229 (43%), Gaps = 12/229 (5%)
Query: 49 ENDPTEETVTEDTAETDSFIYYDDSDI---QERLEECQNS----ILGKIVSEKAIHRNSI 101
EN+ + T T T+ + DD D+ +E Q + IL + S + ++ ++
Sbjct: 230 ENNQVDAT-QGPTVFTEGDLVEDDEDVVVVNIEDDELQTAGLWTILARFYSLRTPNQVAL 288
Query: 102 QNALSNIWCNPKGFRVEHIGDKLFHFFMDEQEDTKRIIRGNPWFFRNSWLIVHPWRRDID 161
+ + W + + D +F + D +I+G PW R L+V +
Sbjct: 289 FDDMRRAWRLRADISYKSLRDNMFIITFSAEGDYDFVIKGGPWIHRGDALLVASFDGITC 348
Query: 162 ARSLEFRHAPIWVQLRGLPTQCRTKQMGIKIGSSIGTVLASELYEYP-DKKLIIKIKVNL 220
++ PIWV++ LP TK G GS +G V ++ + +K I+V+L
Sbjct: 349 PSNVPLDVVPIWVRIYDLPLVLMTKVRGEMFGSKLGKVREVDVGDDGRNKHDFFHIRVDL 408
Query: 221 AVSTPIKAGIYIGSAKDGAHWI---DFRYENLPQFCFACGLIGHTEANC 266
V P+K+ I I G + D YE +P FCF CG IGH++ +C
Sbjct: 409 PVKRPLKSQIAIRMTVQGKEVLRRFDLHYERVPHFCFICGFIGHSDRDC 457
>UniRef100_Q8S5K1 Putative retroelement [Oryza sativa]
Length = 1888
Score = 80.1 bits (196), Expect = 9e-14
Identities = 60/229 (26%), Positives = 101/229 (43%), Gaps = 12/229 (5%)
Query: 49 ENDPTEETVTEDTAETDSFIYYDDSDI---QERLEECQNS----ILGKIVSEKAIHRNSI 101
EN+ + T T T+ + DD D+ +E Q + IL + S + ++ ++
Sbjct: 204 ENNQVDAT-QGPTVFTEGDLVEDDEDVVVVNIEDDELQTAGLWTILARFYSLRTPNQVAL 262
Query: 102 QNALSNIWCNPKGFRVEHIGDKLFHFFMDEQEDTKRIIRGNPWFFRNSWLIVHPWRRDID 161
+ + W + + D +F + D +I+G PW R L+V +
Sbjct: 263 FDDMRRAWRLRADISYKSLRDNMFIITFSAEGDYDFVIKGGPWIHRGDALLVASFDGITC 322
Query: 162 ARSLEFRHAPIWVQLRGLPTQCRTKQMGIKIGSSIGTVLASELYEYP-DKKLIIKIKVNL 220
++ PIWV++ LP TK G GS +G V ++ + +K I+V+L
Sbjct: 323 PSNVPLDVVPIWVRIYDLPLVLMTKVRGEMFGSKLGKVREVDVGDDGRNKHDFFHIRVDL 382
Query: 221 AVSTPIKAGIYIGSAKDGAHWI---DFRYENLPQFCFACGLIGHTEANC 266
V P+K+ I I G + D YE +P FCF CG IGH++ +C
Sbjct: 383 PVKRPLKSQIAIRMTVQGKEVLRRFDLHYERVPHFCFICGFIGHSDRDC 431
>UniRef100_Q7X6H2 OSJNBb0003A12.9 protein [Oryza sativa]
Length = 884
Score = 78.2 bits (191), Expect = 3e-13
Identities = 58/220 (26%), Positives = 97/220 (43%), Gaps = 5/220 (2%)
Query: 52 PTEETVTEDTAETDSFIYYDDSDIQERLEECQNSILGKIVSEKAIHRNSIQNALSNIWCN 111
P+ T E E + + D +D E + Q +IL + S + + +++ +S
Sbjct: 101 PSAFTEGEIPEEEEEVVEVDIAD-DEMKQVVQWTILARFYSMRVPNHSALFEDISKASRR 159
Query: 112 PKGFRVEHIGDKLFHFFMDEQEDTKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRHAP 171
+ + D LF + D K +I+G PW + L+V + +
Sbjct: 160 RSDMSYKSLHDNLFIINFKAEGDYKFVIQGGPWLHKGDALLVAEFDGLTCPSEVSLDVVT 219
Query: 172 IWVQLRGLPTQCRTKQMGIKIGSSIGTVLASELYEYP-DKKLIIKIKVNLAVSTPIKAGI 230
IW+++ LP TK +G GS G V ++ E ++ +I+V L V P K+ I
Sbjct: 220 IWIRIYDLPLVLMTKAIGELYGSKFGKVREVDVEEDGRNRHDFFRIRVELPVKKPPKSKI 279
Query: 231 YIGSAKDGAHWI---DFRYENLPQFCFACGLIGHTEANCK 267
I + G + D RYE +P FCF CG IGH+ +C+
Sbjct: 280 AIKTTVHGVEAVRRFDVRYERVPFFCFICGYIGHSVKDCE 319
>UniRef100_Q75M12 Hypothetical protein P0676G05.2 [Oryza sativa]
Length = 1936
Score = 72.4 bits (176), Expect = 2e-11
Identities = 41/128 (32%), Positives = 64/128 (49%), Gaps = 4/128 (3%)
Query: 144 WFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGIKIGSSIGTVLASE 203
W R L+V + + + PIWV++ LP K+ G+ GS +G V +
Sbjct: 287 WLHRGDALLVADFNGLVSPSMVPLETVPIWVRIYDLPLVMMNKERGVIYGSKLGKVREVD 346
Query: 204 LYEYP-DKKLIIKIKVNLAVSTPIKAGIYIGSAKDGAHWI---DFRYENLPQFCFACGLI 259
+ E +K +I+V+L V+ P+K + I GA I + RYE +P FCF CG I
Sbjct: 347 VQEDGCNKHDFFRIRVDLPVNRPLKRMLAIKIKVKGAEEIRRFNLRYERIPHFCFFCGFI 406
Query: 260 GHTEANCK 267
GH++ C+
Sbjct: 407 GHSDKECE 414
>UniRef100_O65222 F7N22.17 protein [Arabidopsis thaliana]
Length = 416
Score = 69.7 bits (169), Expect = 1e-10
Identities = 43/184 (23%), Positives = 78/184 (42%), Gaps = 2/184 (1%)
Query: 85 SILGKIVSEKAIHRNSIQNALSNIWCNPKGFRVEHIGDKLFHFFMDEQEDTKRIIRGNPW 144
S++ V+ + + ++ +S +W P + F +E ++R PW
Sbjct: 59 SVVVTTVNPRKQNLRALIGQMSRVWGFPDSCVGRILERGRVQFKFQTEEAMNLVLRRGPW 118
Query: 145 FFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGIKIGSSIGTVLASEL 204
F + L +H W +I L+ P WVQ+ G+P T M +G+ +G V +
Sbjct: 119 SFNDWMLSIHRWYPNISETELKI--IPFWVQITGIPLLFLTNAMARCVGNRLGIVSTVDF 176
Query: 205 YEYPDKKLIIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRYENLPQFCFACGLIGHTEA 264
E + +++K++ + P++ A D I +R+E L FC CG + H
Sbjct: 177 DENSNHVGFVRVKIDWNLDDPLRFQRNFQFAVDENTVIKYRFERLRNFCSKCGSLKHDVK 236
Query: 265 NCKL 268
C L
Sbjct: 237 ECVL 240
>UniRef100_Q9SIS9 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1715
Score = 69.3 bits (168), Expect = 2e-10
Identities = 45/197 (22%), Positives = 86/197 (42%), Gaps = 3/197 (1%)
Query: 75 IQERLEECQNSILGKIVSEKAIHRNSIQNALSNIWCNPKGFRVEHIGDKLFHFFMDEQED 134
+Q +++ + +LG+ + + + I + W R I ++ F F +E
Sbjct: 28 VQRAVDDNRFCLLGRPLMPRNQNLRQILTTVPRTWGLVGFVRGRIIQNRRFQFIFPSEES 87
Query: 135 TKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGIKIGS 194
++R PW F + L+V W D+D L F P W+Q+R +P Q ++ I
Sbjct: 88 LDTVLRRGPWSFADRMLVVERWTPDMDPLVLNF--IPFWIQVREIPLQFLNLEVIDNIAG 145
Query: 195 SIGTVLASELYEYPDKKL-IIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRYENLPQFC 253
S+G A + + ++ ++I++ V+ P++ + + F YE L FC
Sbjct: 146 SLGERKAVDFDPFTTTRVEFVRIQIKWDVNHPLRFQRNYQFSLGVNTVLSFYYERLRGFC 205
Query: 254 FACGLIGHTEANCKLHD 270
CG + H +C L +
Sbjct: 206 DVCGRMTHDAGSCVLQN 222
>UniRef100_Q9LH15 Ta11 non-LTR retroelement protein-like [Arabidopsis thaliana]
Length = 241
Score = 68.9 bits (167), Expect = 2e-10
Identities = 43/186 (23%), Positives = 82/186 (43%), Gaps = 5/186 (2%)
Query: 86 ILGKIVSEKAIHRNSIQNALSNIWCNPKGFRVEHIGDKLFHFFMDEQEDTKRIIRGNPWF 145
+ G+ V + + SI ++ IW + + FHF +E + ++R PW
Sbjct: 39 LFGRPVMPRRQNLRSIVASMPRIWGQSGLVHGRIMEGRQFHFIFTLEESLETVLRRGPWA 98
Query: 146 FRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGIKIGSSIGTVLASEL- 204
F + +++ W I F P WVQ+RG+P Q + + IG ++G VL ++
Sbjct: 99 FNDWMILLQRWEPQIPL----FPFIPFWVQIRGIPFQFLNRGVVEHIGRALGQVLDTDFN 154
Query: 205 YEYPDKKLIIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRYENLPQFCFACGLIGHTEA 264
E + ++ ++ ++ P++ + + FRYE L FC CG++ H
Sbjct: 155 VEVVARMDFARVLLHWDITHPLRFQRHFQFTAGVNTLLRFRYERLRGFCEVCGMLTHDFG 214
Query: 265 NCKLHD 270
C + +
Sbjct: 215 ACLIQN 220
>UniRef100_O82596 F11O4.11 protein [Arabidopsis thaliana]
Length = 577
Score = 66.6 bits (161), Expect = 1e-09
Identities = 51/194 (26%), Positives = 82/194 (41%), Gaps = 9/194 (4%)
Query: 74 DIQERLEECQNSILGKIVSEKAIHRNSIQNALSNIWCNPKGFRV-EHIGDKLFHFFMDEQ 132
D E ++E +++G + + A S+ LSN W N KG +G F F + +
Sbjct: 28 DNSELIQENSLTLMGILTNPSAQRLWSLIPFLSNRW-NLKGKATGSDLGRGCFQFRFEYE 86
Query: 133 EDTKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGIKI 192
ED ++++ P+ F +I+ W+ I P W++L+GLP K+M I
Sbjct: 87 EDLQKVLDNRPYHFGQWMVILQRWKPVISPSFPS--EIPFWIELQGLPMHYWIKEMLYAI 144
Query: 193 GSSIGTVLASELYEYPDKKLIIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRYENLPQF 252
G G V+ E+ KIKV + PI + + Y+NL
Sbjct: 145 GKEAGEVVDHEI-----SPAAAKIKVLINGLQPITKETVVEFPDGSEALVYLEYKNLKSH 199
Query: 253 CFACGLIGHTEANC 266
C C + H EA+C
Sbjct: 200 CHHCQRLSHAEADC 213
>UniRef100_Q9SKJ4 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1750
Score = 66.6 bits (161), Expect = 1e-09
Identities = 42/184 (22%), Positives = 76/184 (40%), Gaps = 2/184 (1%)
Query: 85 SILGKIVSEKAIHRNSIQNALSNIWCNPKGFRVEHIGDKLFHFFMDEQEDTKRIIRGNPW 144
S++ V+ + + ++ + +W P I F +E ++R PW
Sbjct: 38 SLVVTTVNPRKQNLRALIGQMPRVWGFPDSCVGRIIDKGRVQFKFQSEEAMNLVLRRGPW 97
Query: 145 FFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGIKIGSSIGTVLASEL 204
F + L +H W ++ E + P WVQ+ G+P T M +G+ +G V +
Sbjct: 98 SFNDWMLSIHRWYPNLS--EAEMKIIPFWVQITGIPLLFLTNAMARCVGNRLGHVADVDF 155
Query: 205 YEYPDKKLIIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRYENLPQFCFACGLIGHTEA 264
E + +++++N + P++ A I FR+E L FC CG + H
Sbjct: 156 DENSNHTGFVRVRINWNLDDPLRFQRNFQFADGENTVIKFRFERLRNFCTKCGSLKHDIK 215
Query: 265 NCKL 268
C L
Sbjct: 216 VCTL 219
>UniRef100_Q7Y1K0 Hypothetical protein OSJNBb0056B16.21 [Oryza sativa]
Length = 813
Score = 66.2 bits (160), Expect = 1e-09
Identities = 50/210 (23%), Positives = 91/210 (42%), Gaps = 39/210 (18%)
Query: 61 TAETDSFIYYDDSDIQERLEECQNSILGKIVSEKAIHRNSIQNALSNIWCNPKGFRVEHI 120
T E ++ + D D +E + ++GK++S +H ++ A+ NP G ++ I
Sbjct: 16 TTEEENVAEFSDDDETGDPQEVEWMLVGKVLSPAIVHATTVFRAMKPARGNPYGLKIRSI 75
Query: 121 GDKLFHFFMDE---QEDTKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLR 177
+K + F+ + + D +R + G+PW +LE
Sbjct: 76 DEKADNIFVTKFKVERDLQRALGGSPWM------------------NLE----------- 106
Query: 178 GLPTQCRTKQMGIKIGSSIGTVLASELYEYPDKKL---IIKIKVNLAVSTPIKAGIYIGS 234
PT+ K G +G V E+ D K ++ +V + V+ ++ G+ + +
Sbjct: 107 -PPTRLDDKHRGELAMGLVGVVRKLEVDS--DGKASGPYLRGRVAIVVAKSLRRGMLLKT 163
Query: 235 AKDGA-HWIDFRYENLPQFCFACGLIGHTE 263
KD A W +YE LP +C ACG++GH E
Sbjct: 164 KKDAAPEWFYLQYEKLPYYCLACGIVGHLE 193
>UniRef100_Q5W742 Hypothetical protein OJ1005_D04.4 [Oryza sativa]
Length = 796
Score = 65.9 bits (159), Expect = 2e-09
Identities = 42/195 (21%), Positives = 86/195 (43%), Gaps = 7/195 (3%)
Query: 71 DDSDIQERLEECQNSI----LGKIVSEKAIHRNSIQNALSNIWCNPKGFRVEHIGDKLFH 126
+D +++E L+E + L ++ +EK+ ++ ++ + W K + D LF
Sbjct: 46 EDVELEEDLKELEADARWLALARVHTEKSFSHGALFASMRSAWNCAKEVDFRSMKDNLFS 105
Query: 127 FFMDEQEDTKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTK 186
++ D +R++ PW FR +++ + + S+ P WVQ+ L + RT
Sbjct: 106 IQLNCLADWERVMEEGPWIFRGCPVLLAEYDGWSEIESVVLETYPTWVQIHDLKEKIRTG 165
Query: 187 QMGIKIGSSIGTVLASELYEYPDKKLIIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRY 246
+ ++ GTV+ + ++++V + V +P + K + Y
Sbjct: 166 SIATQLVRKAGTVIKLDEASVKGSGDGVRVRVKVDVPSPASRSTTLNKKK---WFFRLMY 222
Query: 247 ENLPQFCFACGLIGH 261
E + FC CGL+GH
Sbjct: 223 EKMLVFCGICGLVGH 237
>UniRef100_Q9SLE8 Putative Ta11-like non-LTR retroelement protein [Arabidopsis
thaliana]
Length = 240
Score = 64.3 bits (155), Expect = 5e-09
Identities = 48/218 (22%), Positives = 90/218 (41%), Gaps = 3/218 (1%)
Query: 50 NDPTEETVTEDTAETDSFIYYDDSDIQERLEECQNSILGKIVSEKAIHRNSIQNALSNIW 109
N ++ + E T DS + + +E S++G+ ++ ++ + IW
Sbjct: 2 NMELDKAIFEMTINDDSPLILSNQPQLSSIERNSCSLIGRFLNPSHQRMSNWILDMPRIW 61
Query: 110 CNPKGFRVEHIGDKLFHFFMDEQEDTKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRH 169
R + + F FF +D I++ W +I+ W L
Sbjct: 62 RIYSRVRGVALSQERFQFFYKSNDDLLEILKTGVWTQDEWCVIMDRWVEKPPEEYLMI-- 119
Query: 170 APIWVQLRGLPTQCRTKQMGIKIGSSIGTVLASEL-YEYPDKKLIIKIKVNLAVSTPIKA 228
PIW++LR +P T++ KI +G V+ EL E + ++++VNL V P++
Sbjct: 120 LPIWIRLRNIPVNYYTEETIKKIAGCVGQVVKVELDLEKSQAQDYVRVQVNLDVRNPLRN 179
Query: 229 GIYIGSAKDGAHWIDFRYENLPQFCFACGLIGHTEANC 266
+ + F YE + + CF C + H +A+C
Sbjct: 180 SKSVQVPSGEVVSVTFDYERIRKRCFFCQRLTHDKADC 217
>UniRef100_Q9ZQG0 Putative Ta11-like non-LTR retroelement protein [Arabidopsis
thaliana]
Length = 483
Score = 63.5 bits (153), Expect = 9e-09
Identities = 47/223 (21%), Positives = 99/223 (44%), Gaps = 11/223 (4%)
Query: 70 YDDSDIQERLEECQN--SILGKIVSEKAIHRNSIQNALSNIWCNPKGFRVEHIGDKLFHF 127
++ D+ E L +N S++G++++ ++ + W + + ++ F F
Sbjct: 17 FEMPDLPEYLSSERNKFSLIGRVLNPACQPMKNLLRNMPRKWQKEGKVKGIALNNERFQF 76
Query: 128 FMDEQEDTKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQ 187
D + D + ++ F + ++V W + L R+ +WVQ+ +P T +
Sbjct: 77 IFDSEHDLEEVLAKGVHTFNDWSIVVDRWYENPPDDYL--RYMLLWVQIWNIPVNYNTAK 134
Query: 188 MGIKIGSSIGTVLASELYEYPDKKLI---IKIKVNLAVSTPIKAGIYIGSAKDGAHWIDF 244
K+G IG V E+ P+ K I +++KV VS P++ + G+ ++F
Sbjct: 135 AITKLGDLIGEV--KEVVFNPELKQIREFVRVKVLFDVSRPLRRSKVVNFRNGGSATVNF 192
Query: 245 RYENLPQFCFACGLIGHTEANCKL--HDTRTNAESRNKNVLGP 285
YE + + C+ C + H + C + + A +R K ++ P
Sbjct: 193 HYERIQKRCYECQRLTHEKDVCPILVKARQDLATARRKGIIIP 235
>UniRef100_Q9SN65 Hypothetical protein F25I24.40 [Arabidopsis thaliana]
Length = 1294
Score = 63.2 bits (152), Expect = 1e-08
Identities = 38/149 (25%), Positives = 65/149 (43%), Gaps = 2/149 (1%)
Query: 120 IGDKLFHFFMDEQEDTKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGL 179
+G+ F +E +++ PW F + L VH W +I E + P WVQ++G+
Sbjct: 73 VGNGRVQFKFRNEESMNLVLQRGPWSFNDWMLSVHRWFPNITEG--EMKIIPFWVQIQGI 130
Query: 180 PTQCRTKQMGIKIGSSIGTVLASELYEYPDKKLIIKIKVNLAVSTPIKAGIYIGSAKDGA 239
P T M +GS +G V + E ++ +++K+ ++ ++
Sbjct: 131 PILYLTNAMARVVGSRLGYVTDVDFDENANQMGFVRVKLAWNFDDHLRFQRNFQFQENEN 190
Query: 240 HWIDFRYENLPQFCFACGLIGHTEANCKL 268
I FR+E L FC CG + H C L
Sbjct: 191 TIIKFRFERLRNFCTKCGSLKHDVKECNL 219
>UniRef100_O81630 F8M12.22 protein [Arabidopsis thaliana]
Length = 1662
Score = 63.2 bits (152), Expect = 1e-08
Identities = 38/149 (25%), Positives = 65/149 (43%), Gaps = 2/149 (1%)
Query: 120 IGDKLFHFFMDEQEDTKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGL 179
+G+ F +E +++ PW F + L VH W +I E + P WVQ++G+
Sbjct: 73 VGNGRVQFKFRNEESMNLVLQRGPWSFNDWMLSVHRWFPNITEG--EMKIIPFWVQIQGI 130
Query: 180 PTQCRTKQMGIKIGSSIGTVLASELYEYPDKKLIIKIKVNLAVSTPIKAGIYIGSAKDGA 239
P T M +GS +G V + E ++ +++K+ ++ ++
Sbjct: 131 PILYLTNAMARVVGSRLGYVTDVDFDENANQMGFVRVKLAWNFDDHLRFQRNFQFQENEN 190
Query: 240 HWIDFRYENLPQFCFACGLIGHTEANCKL 268
I FR+E L FC CG + H C L
Sbjct: 191 TIIKFRFERLRNFCTKCGSLKHDVKECNL 219
>UniRef100_O23411 Hypothetical protein AT4g15600 [Arabidopsis thaliana]
Length = 655
Score = 62.8 bits (151), Expect = 1e-08
Identities = 42/207 (20%), Positives = 89/207 (42%), Gaps = 10/207 (4%)
Query: 75 IQERLEECQNSILGKIVSEKAIHRNSIQNALS----NIWCNPKGFRVEHIGDKLFHFFMD 130
I++ + + N + + + K + R++ LS ++W G V + + F +
Sbjct: 80 IEKEVLDAMNGLYKQCMLVKILGRHTTIEVLSRKLRDLWRPTGGMSVLDLPRQFFMVRFE 139
Query: 131 EQEDTKRIIRGNPWFFRNSWLIVHPWRRDIDARSLEFRHAPIWVQLRGLPTQCRTKQMGI 190
+ED + G PW S L+V W + + P+WV++ LP ++ +
Sbjct: 140 VEEDYMMALTGGPWRVLGSILMVQAWSPEFNPLRDVIETTPVWVRVANLPVTFYHNEILL 199
Query: 191 KIGSSIGTVLASELYEY-PDKKLIIKIKVNLAVSTPIKAGIYIGSAKDGAHWIDFRYENL 249
I + +G + +L ++ ++ V + + P+K + + + +++ YE L
Sbjct: 200 GIAAGLGKPIKVDLTTLRKERGRFARVCVEVNLKNPLKGTLVVNGER---YFVS--YEGL 254
Query: 250 PQFCFACGLIGHTEANCKLHDTRTNAE 276
C CG+ GHT NC + T ++
Sbjct: 255 QTICSLCGIYGHTVNNCPMGRTTQGSQ 281
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.135 0.425
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 655,748,773
Number of Sequences: 2790947
Number of extensions: 27698287
Number of successful extensions: 98716
Number of sequences better than 10.0: 122
Number of HSP's better than 10.0 without gapping: 31
Number of HSP's successfully gapped in prelim test: 91
Number of HSP's that attempted gapping in prelim test: 98569
Number of HSP's gapped (non-prelim): 136
length of query: 361
length of database: 848,049,833
effective HSP length: 128
effective length of query: 233
effective length of database: 490,808,617
effective search space: 114358407761
effective search space used: 114358407761
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 75 (33.5 bits)
Medicago: description of AC144807.8


