

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144807.5 + phase: 0
(112 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
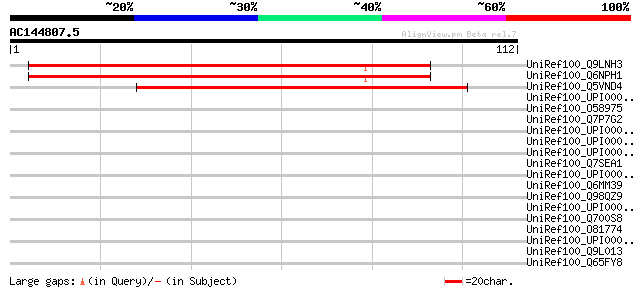
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LNH3 F21D18.7 [Arabidopsis thaliana] 92 2e-18
UniRef100_Q6NPH1 At1g48200 [Arabidopsis thaliana] 92 2e-18
UniRef100_Q5VND4 Hypothetical protein P0710H01.6 [Oryza sativa] 68 6e-11
UniRef100_UPI0000321A69 UPI0000321A69 UniRef100 entry 35 0.31
UniRef100_O58975 Hypothetical protein PH1208 [Pyrococcus horikos... 34 0.91
UniRef100_Q7P7G2 ATP-dependent DNA helicase pcrA [Fusobacterium ... 33 1.6
UniRef100_UPI000033C8EF UPI000033C8EF UniRef100 entry 32 2.7
UniRef100_UPI00003615E0 UPI00003615E0 UniRef100 entry 32 3.5
UniRef100_UPI00003615DD UPI00003615DD UniRef100 entry 32 3.5
UniRef100_Q7SEA1 Predicted protein [Neurospora crassa] 32 3.5
UniRef100_UPI000032A5D2 UPI000032A5D2 UniRef100 entry 32 4.5
UniRef100_Q6MM39 ThrS protein [Bdellovibrio bacteriovorus] 32 4.5
UniRef100_Q98QZ9 Hypothetical protein MYPU_2110 [Mycoplasma pulm... 32 4.5
UniRef100_UPI000042D07C UPI000042D07C UniRef100 entry 31 5.9
UniRef100_Q700S8 Variable membrane protein precursor [Mycoplasma... 31 5.9
UniRef100_O81774 Hypothetical protein F28M20.70 [Arabidopsis tha... 31 5.9
UniRef100_UPI00002ED111 UPI00002ED111 UniRef100 entry 31 7.7
UniRef100_Q9L013 Hypothetical protein SCO2300 [Streptomyces coel... 31 7.7
UniRef100_Q65FY8 YteR [Bacillus licheniformis] 31 7.7
>UniRef100_Q9LNH3 F21D18.7 [Arabidopsis thaliana]
Length = 254
Score = 92.4 bits (228), Expect = 2e-18
Identities = 48/91 (52%), Positives = 63/91 (68%), Gaps = 2/91 (2%)
Query: 5 HKIVDIAKRASENNTVINVGLGAVFAILGARSYNQQKIIEALEAEKDSLTKTNKSIRNTL 64
+KI A NN VIN LG F +LG RS QQK +EAL +K+SL K+NK+++ T+
Sbjct: 3 NKIAMFLSEAMNNNAVINTCLGVSFVVLGLRSDKQQKYVEALAEQKESLFKSNKAMKLTM 62
Query: 65 WDWKQQLYAEAAS--DSAPVPLARLYEIYGE 93
W+WKQQL+AEAAS ++A VPL+ L IYGE
Sbjct: 63 WEWKQQLFAEAASAGNAAVVPLSTLKAIYGE 93
>UniRef100_Q6NPH1 At1g48200 [Arabidopsis thaliana]
Length = 118
Score = 92.4 bits (228), Expect = 2e-18
Identities = 48/91 (52%), Positives = 63/91 (68%), Gaps = 2/91 (2%)
Query: 5 HKIVDIAKRASENNTVINVGLGAVFAILGARSYNQQKIIEALEAEKDSLTKTNKSIRNTL 64
+KI A NN VIN LG F +LG RS QQK +EAL +K+SL K+NK+++ T+
Sbjct: 3 NKIAMFLSEAMNNNAVINTCLGVSFVVLGLRSDKQQKYVEALAEQKESLFKSNKAMKLTM 62
Query: 65 WDWKQQLYAEAAS--DSAPVPLARLYEIYGE 93
W+WKQQL+AEAAS ++A VPL+ L IYGE
Sbjct: 63 WEWKQQLFAEAASAGNAAVVPLSTLKAIYGE 93
>UniRef100_Q5VND4 Hypothetical protein P0710H01.6 [Oryza sativa]
Length = 117
Score = 67.8 bits (164), Expect = 6e-11
Identities = 32/73 (43%), Positives = 46/73 (62%)
Query: 29 FAILGARSYNQQKIIEALEAEKDSLTKTNKSIRNTLWDWKQQLYAEAASDSAPVPLARLY 88
F +LG RS+ QQ I+ LEA K SL N ++ + +W W+++L+A AA+ S P+ +RL
Sbjct: 29 FVVLGWRSWEQQHEIDELEARKASLRAANTAMSSAMWAWREELFALAAAPSPPISASRLR 88
Query: 89 EIYGEAAPPQQSA 101
IYGE PP A
Sbjct: 89 VIYGEEQPPASPA 101
>UniRef100_UPI0000321A69 UPI0000321A69 UniRef100 entry
Length = 277
Score = 35.4 bits (80), Expect = 0.31
Identities = 25/81 (30%), Positives = 40/81 (48%), Gaps = 8/81 (9%)
Query: 17 NNTVINVGLGAVFAILGARSYNQQKIIEALEAE----KDSLTKTNKSIRNTLWDWKQQL- 71
NN + G V I G Q +++EAL E K+ + NKSI N+ ++++QL
Sbjct: 47 NNINFTIDYGQVLGIAGIAGNGQVELMEALSGETLGKKEEIIFENKSIGNSKIEFRRQLG 106
Query: 72 ---YAEAASDSAPVPLARLYE 89
E ++ A VP +L+E
Sbjct: 107 IEFVPEERNNHATVPDLKLHE 127
>UniRef100_O58975 Hypothetical protein PH1208 [Pyrococcus horikoshii]
Length = 250
Score = 33.9 bits (76), Expect = 0.91
Identities = 29/84 (34%), Positives = 39/84 (45%), Gaps = 4/84 (4%)
Query: 8 VDIAKRASENNTVINVGLGAVFAILGARSYNQQKIIEALEAEKDS-LTKTNKSIRNTLWD 66
V++AK SEN I + G VF I R ++ IE L AE DS K I+N W
Sbjct: 159 VELAKEISENGHFIGIVTGIVF-IPEVRKVAEKMDIEGLLAETDSPYMSPYKGIKNKPWF 217
Query: 67 WKQQLYAEAASDSAPVPLARLYEI 90
K + E S +P+ + EI
Sbjct: 218 VK--IVLEELSKIKEIPIEEVEEI 239
>UniRef100_Q7P7G2 ATP-dependent DNA helicase pcrA [Fusobacterium nucleatum subsp.
vincentii ATCC 49256]
Length = 693
Score = 33.1 bits (74), Expect = 1.6
Identities = 32/110 (29%), Positives = 49/110 (44%), Gaps = 21/110 (19%)
Query: 5 HKIVDIAKRASENNTVINVGLGAVFAILGAR---------SYNQQKIIEALEAEKDSLT- 54
+KI+D+ R S N V+ +++ GA +YN KII+ E + + T
Sbjct: 182 YKIIDLIARKSSNLCVVGDENQSIYGFRGANILNILNFETNYNNAKIIKLEENYRSTTTI 241
Query: 55 --------KTNKSIRN-TLW--DWKQQLYAEAASDSAPVPLARLYEIYGE 93
K NKS R+ LW + K L A D+A ++R+ EI E
Sbjct: 242 LDAANELIKNNKSSRDKKLWTQNGKGDLIKVLACDNARDEVSRIIEIIKE 291
>UniRef100_UPI000033C8EF UPI000033C8EF UniRef100 entry
Length = 289
Score = 32.3 bits (72), Expect = 2.7
Identities = 22/64 (34%), Positives = 32/64 (49%), Gaps = 3/64 (4%)
Query: 32 LGARS---YNQQKIIEALEAEKDSLTKTNKSIRNTLWDWKQQLYAEAASDSAPVPLARLY 88
LGA S Y + AL +D + + SI+ W Q L+A A+ +AP LA +
Sbjct: 149 LGAHSFGLYIHNDTMRALGRPQDMFSDSAISIQPIFAQWIQGLHAAASGSTAPNALAGVS 208
Query: 89 EIYG 92
EI+G
Sbjct: 209 EIFG 212
>UniRef100_UPI00003615E0 UPI00003615E0 UniRef100 entry
Length = 675
Score = 32.0 bits (71), Expect = 3.5
Identities = 29/98 (29%), Positives = 47/98 (47%), Gaps = 11/98 (11%)
Query: 3 FVHKIVDIAKRASENNTVINVGL--GAVFAIL----GARSYNQQKIIEALEAEKDSLTKT 56
F VD A V+ +G G V ++ G+ + ++ ++E LE KD+ +
Sbjct: 394 FTQIAVDRVGAADGQYDVMFIGTDKGTVLKVINVPKGSWNNMEELLLEELEVFKDASSII 453
Query: 57 NKSIRNTLWDWKQQLYAEAASDSAPVPLARLYEIYGEA 94
N I + +QQLY +A+ A VPL R +YG+A
Sbjct: 454 NMQISSK----RQQLYVGSATGIAQVPLHRC-SVYGKA 486
>UniRef100_UPI00003615DD UPI00003615DD UniRef100 entry
Length = 729
Score = 32.0 bits (71), Expect = 3.5
Identities = 29/98 (29%), Positives = 47/98 (47%), Gaps = 11/98 (11%)
Query: 3 FVHKIVDIAKRASENNTVINVGL--GAVFAIL----GARSYNQQKIIEALEAEKDSLTKT 56
F VD A V+ +G G V ++ G+ + ++ ++E LE KD+ +
Sbjct: 400 FTQIAVDRVGAADGQYDVMFIGTDKGTVLKVINVPKGSWNNMEELLLEELEVFKDASSII 459
Query: 57 NKSIRNTLWDWKQQLYAEAASDSAPVPLARLYEIYGEA 94
N I + +QQLY +A+ A VPL R +YG+A
Sbjct: 460 NMQISSK----RQQLYVGSATGIAQVPLHRC-SVYGKA 492
>UniRef100_Q7SEA1 Predicted protein [Neurospora crassa]
Length = 2202
Score = 32.0 bits (71), Expect = 3.5
Identities = 21/80 (26%), Positives = 37/80 (46%), Gaps = 18/80 (22%)
Query: 37 YNQQKIIEALEAEKDSLTKTNKS-------------IRNTLWDWKQQLYAEAASDSAPV- 82
Y+ +++EA + + D+LT T +S + ++ W + + A D APV
Sbjct: 1495 YSYDQVVEATKRQWDALTSTARSNYSNKAFAAAQHPLDKGIFTWLEDTFKRVAKDKAPVS 1554
Query: 83 --PLARLYEIYGEAA--PPQ 98
P A+ Y G ++ PPQ
Sbjct: 1555 STPAAQNYSASGNSSSEPPQ 1574
>UniRef100_UPI000032A5D2 UPI000032A5D2 UniRef100 entry
Length = 226
Score = 31.6 bits (70), Expect = 4.5
Identities = 23/93 (24%), Positives = 46/93 (48%), Gaps = 4/93 (4%)
Query: 9 DIAKRASENNTVINVGLGAVFAILGARSYNQQKIIEALEAEKDSLTK---TNKSIRNTLW 65
D A A E + + +GA+ + L A+ YN + +I+ +E E++++T+ K+I+ +
Sbjct: 73 DTAGSAKELSKSNDTTIGAIASTLSAKIYNLKVLIKNIEDERNNVTRFLIMGKNIKQPKY 132
Query: 66 DWKQQLYAEAASDSAPVPLARLYEIYGEAAPPQ 98
+ ++ +P A LY+ G A Q
Sbjct: 133 ETNKKYITSCIFRLKSIP-AALYKCLGGFATNQ 164
>UniRef100_Q6MM39 ThrS protein [Bdellovibrio bacteriovorus]
Length = 416
Score = 31.6 bits (70), Expect = 4.5
Identities = 24/92 (26%), Positives = 42/92 (45%), Gaps = 23/92 (25%)
Query: 4 VHKIVDIAKRASENNTVINVGLGAVFAILGARSYNQQKIIEALEAEKDSLTKTNKSIRNT 63
+HK++++ ++A + ILG Y + ++ +EAE+ S K + N
Sbjct: 173 IHKVLEMYQKA--------------YRILGISDY-RLRLSRGVEAEEAS----QKYVANP 213
Query: 64 LWDWKQQLYAEAASDSAPVPLARLYEIYGEAA 95
LWDW + + E +S +E GEAA
Sbjct: 214 LWDWAEGILREVLVESG----LPFFEAPGEAA 241
>UniRef100_Q98QZ9 Hypothetical protein MYPU_2110 [Mycoplasma pulmonis]
Length = 3216
Score = 31.6 bits (70), Expect = 4.5
Identities = 17/55 (30%), Positives = 34/55 (60%), Gaps = 3/55 (5%)
Query: 18 NTVINVGLGAVF-AILGARSYNQQKIIEALEAEKDSLTKTNKSIRNTLWDWKQQL 71
N+++ +G A ++L + S +Q + AL+ ++ K N SI+ T+W+WK++L
Sbjct: 2741 NSLVEIGPNAFKNSVLTSLSGLEQTKLSALK--ENVFEKANDSIKTTIWNWKKEL 2793
>UniRef100_UPI000042D07C UPI000042D07C UniRef100 entry
Length = 157
Score = 31.2 bits (69), Expect = 5.9
Identities = 29/107 (27%), Positives = 46/107 (42%), Gaps = 25/107 (23%)
Query: 8 VDIAKRASENNTVINVGLGAVFAILGARSYNQQKIIEALEAEKDSLTKTNKSIRNTLWDW 67
VD AK ++ G+G V ++ G + +Q+K T+ N WD+
Sbjct: 56 VDAAKAQKAGAATVDSGIGKVKSVFGYVTGDQEK-----------QTEGNTQAEKAQWDY 104
Query: 68 KQQLYAEAASDS-APVPL-------ARLYEIYGEAA-PPQQSAPGNL 105
KQ A SDS A +P+ +L + G AA P++ GN+
Sbjct: 105 KQ-----ATSDSIAAIPIPSVEGAKGKLESVVGMAAEDPEKQKEGNV 146
>UniRef100_Q700S8 Variable membrane protein precursor [Mycoplasma hominis]
Length = 1828
Score = 31.2 bits (69), Expect = 5.9
Identities = 20/70 (28%), Positives = 34/70 (48%)
Query: 10 IAKRASENNTVINVGLGAVFAILGARSYNQQKIIEALEAEKDSLTKTNKSIRNTLWDWKQ 69
I K ++ N+ + A A+ A+S Q+I EAE ++LTK + + +
Sbjct: 1317 ITKNKADKNSTLEEITNATKALEDAKSKLDQEIKTKKEAEFNNLTKAKTELSDLITSSSN 1376
Query: 70 QLYAEAASDS 79
Q A+A SD+
Sbjct: 1377 QAPADAISDA 1386
>UniRef100_O81774 Hypothetical protein F28M20.70 [Arabidopsis thaliana]
Length = 171
Score = 31.2 bits (69), Expect = 5.9
Identities = 21/81 (25%), Positives = 39/81 (47%), Gaps = 15/81 (18%)
Query: 27 AVFAILGARSYNQQKIIEALEAEKDSLTKTNKSIRNTLWDWKQQLYAEAASDSAP----- 81
A+ +L + NQ ++ EA+EA +L + + R+ + DW ++LY ++ S P
Sbjct: 7 AIMYLLSLETINQSEV-EAVEA---ALPSADSASRSNIVDWAEKLYGQSISAVTPGVKNL 62
Query: 82 ------VPLARLYEIYGEAAP 96
+P+AR E + P
Sbjct: 63 LSSDQQLPVARTVEALTDGKP 83
>UniRef100_UPI00002ED111 UPI00002ED111 UniRef100 entry
Length = 228
Score = 30.8 bits (68), Expect = 7.7
Identities = 25/92 (27%), Positives = 36/92 (38%), Gaps = 6/92 (6%)
Query: 17 NNTVIN----VGLGAVFAILGARSY--NQQKIIEALEAEKDSLTKTNKSIRNTLWDWKQQ 70
NN V++ + LG F +L Y + II L L N LW W Q
Sbjct: 9 NNVVVSAPSGLALGGGFEVLVQSDYVVSHTNIITGLVETLVGLIPAGGGTTNMLWRWMQT 68
Query: 71 LYAEAASDSAPVPLARLYEIYGEAAPPQQSAP 102
A+ SD AP + + A+ P ++ P
Sbjct: 69 QEAKKDSDFAPQKVFNIIGYGKTASSPIEAIP 100
>UniRef100_Q9L013 Hypothetical protein SCO2300 [Streptomyces coelicolor]
Length = 196
Score = 30.8 bits (68), Expect = 7.7
Identities = 20/88 (22%), Positives = 41/88 (45%)
Query: 2 EFVHKIVDIAKRASENNTVINVGLGAVFAILGARSYNQQKIIEALEAEKDSLTKTNKSIR 61
+ V ++++ + A E + +GAV + + + +E L+AE S+TK + +
Sbjct: 65 DVVLEVMERRESAQERVAELTERVGAVQGKIDDATARRDAAVEELDAEITSVTKEREVVA 124
Query: 62 NTLWDWKQQLYAEAASDSAPVPLARLYE 89
++ D LY + V A+LY+
Sbjct: 125 GSVPDDLLTLYGKLREQQGGVGAAKLYQ 152
>UniRef100_Q65FY8 YteR [Bacillus licheniformis]
Length = 373
Score = 30.8 bits (68), Expect = 7.7
Identities = 18/62 (29%), Positives = 34/62 (54%), Gaps = 5/62 (8%)
Query: 25 LGAVFAILGARSYNQQKIIEALEAEKDSLTKTNKSIRNTL----WDWKQQL-YAEAASDS 79
+G FA+ AR++N+ ++ + + E+ + K K R L WD K+Q+ +A+ S
Sbjct: 144 MGGPFALKYARAFNEPELYDKVILEEKLMRKHTKDCRTGLYYHAWDEKRQMPWADPVSGC 203
Query: 80 AP 81
+P
Sbjct: 204 SP 205
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.316 0.131 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 172,025,577
Number of Sequences: 2790947
Number of extensions: 5776172
Number of successful extensions: 20596
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 20588
Number of HSP's gapped (non-prelim): 19
length of query: 112
length of database: 848,049,833
effective HSP length: 88
effective length of query: 24
effective length of database: 602,446,497
effective search space: 14458715928
effective search space used: 14458715928
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC144807.5


