

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144731.10 + phase: 0 /pseudo
(222 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
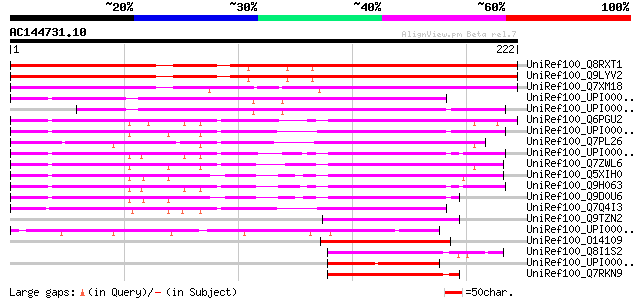
Sequences producing significant alignments: (bits) Value
UniRef100_Q8RXT1 Hypothetical protein At5g13240 [Arabidopsis tha... 201 1e-50
UniRef100_Q9LYV2 Hypothetical protein T31B5_60 [Arabidopsis thal... 201 1e-50
UniRef100_Q7XM18 OSJNBa0084K01.13 protein [Oryza sativa] 162 5e-39
UniRef100_UPI000035FA4E UPI000035FA4E UniRef100 entry 89 6e-17
UniRef100_UPI000029BC72 UPI000029BC72 UniRef100 entry 87 4e-16
UniRef100_Q6PGU2 Repressor of RNA polymerase III transcription M... 85 1e-15
UniRef100_UPI0000249370 UPI0000249370 UniRef100 entry 84 3e-15
UniRef100_Q7PL26 CG40196-PA.3 [Drosophila melanogaster] 82 1e-14
UniRef100_UPI000036E4D0 UPI000036E4D0 UniRef100 entry 80 5e-14
UniRef100_Q7ZWL6 Maf1-pending-prov protein [Xenopus laevis] 79 7e-14
UniRef100_Q5XIH0 Hypothetical protein [Rattus norvegicus] 79 7e-14
UniRef100_Q9H063 Repressor of RNA polymerase III transcription M... 79 7e-14
UniRef100_Q9D0U6 Repressor of RNA polymerase III transcription M... 79 9e-14
UniRef100_Q7Q4I3 ENSANGP00000019317 [Anopheles gambiae str. PEST] 79 1e-13
UniRef100_Q9TZN2 Hypothetical protein C43H8.2 [Caenorhabditis el... 57 5e-07
UniRef100_UPI00002349F9 UPI00002349F9 UniRef100 entry 52 1e-05
UniRef100_O14109 ORF N150 [Schizosaccharomyces pombe] 51 2e-05
UniRef100_Q8I1S2 Hypothetical protein PFD0800c [Plasmodium falci... 47 3e-04
UniRef100_UPI000021AB1D UPI000021AB1D UniRef100 entry 46 8e-04
UniRef100_Q7RKN9 Hypothetical protein [Plasmodium yoelii yoelii] 46 8e-04
>UniRef100_Q8RXT1 Hypothetical protein At5g13240 [Arabidopsis thaliana]
Length = 224
Score = 201 bits (510), Expect = 1e-50
Identities = 120/235 (51%), Positives = 147/235 (62%), Gaps = 25/235 (10%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEILDYLGKSS 60
MKFLE LDRL+ FL +LNLGERTIKGC+EAYSCKH+G+DK+LS+SL NE+LDYLGKSS
Sbjct: 1 MKFLEYTNLDRLNVFLGHLNLGERTIKGCLEAYSCKHAGSDKRLSLSLENEMLDYLGKSS 60
Query: 61 DNDSPSPVDSLSSPVDSLSARTSPAGRHWFT*FLPSITCIQIM----ISAQLKLTSFSQR 116
D DS SSPVD L +R+S + +T Q+ SA FS+
Sbjct: 61 DTDS-------SSPVDLLLSRSSRKALIYLV-----LTLYQMYPDYDFSAVKAHQFFSEE 108
Query: 117 NHGT---VLSRFLKPTCL------KPQSLLDALFKAVDEVVSLDDCEIYGYLPDSEADPL 167
+ T + + ++ + SLL+ ++KA+DEVV L +CEIY Y P+ ADP
Sbjct: 109 SWDTFKQIFNNYMFEASKEWTERNEDGSLLEVIYKALDEVVKLAECEIYVYNPNPNADPF 168
Query: 168 PERGAIWSFNFLFYNRKLKRIVTFRLSCFSNLIADGFSLDEICDEYDEEIFADMD 222
E GAIWSF FLFYNRKLKR+ FR SC SNL D F D E DEEIFADMD
Sbjct: 169 LEEGAIWSFCFLFYNRKLKRVAGFRFSCTSNLANDAFLTDSPPYEEDEEIFADMD 223
>UniRef100_Q9LYV2 Hypothetical protein T31B5_60 [Arabidopsis thaliana]
Length = 283
Score = 201 bits (510), Expect = 1e-50
Identities = 120/235 (51%), Positives = 147/235 (62%), Gaps = 25/235 (10%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEILDYLGKSS 60
MKFLE LDRL+ FL +LNLGERTIKGC+EAYSCKH+G+DK+LS+SL NE+LDYLGKSS
Sbjct: 60 MKFLEYTNLDRLNVFLGHLNLGERTIKGCLEAYSCKHAGSDKRLSLSLENEMLDYLGKSS 119
Query: 61 DNDSPSPVDSLSSPVDSLSARTSPAGRHWFT*FLPSITCIQIM----ISAQLKLTSFSQR 116
D DS SSPVD L +R+S + +T Q+ SA FS+
Sbjct: 120 DTDS-------SSPVDLLLSRSSRKALIYLV-----LTLYQMYPDYDFSAVKAHQFFSEE 167
Query: 117 NHGT---VLSRFLKPTCL------KPQSLLDALFKAVDEVVSLDDCEIYGYLPDSEADPL 167
+ T + + ++ + SLL+ ++KA+DEVV L +CEIY Y P+ ADP
Sbjct: 168 SWDTFKQIFNNYMFEASKEWTERNEDGSLLEVIYKALDEVVKLAECEIYVYNPNPNADPF 227
Query: 168 PERGAIWSFNFLFYNRKLKRIVTFRLSCFSNLIADGFSLDEICDEYDEEIFADMD 222
E GAIWSF FLFYNRKLKR+ FR SC SNL D F D E DEEIFADMD
Sbjct: 228 LEEGAIWSFCFLFYNRKLKRVAGFRFSCTSNLANDAFLTDSPPYEEDEEIFADMD 282
>UniRef100_Q7XM18 OSJNBa0084K01.13 protein [Oryza sativa]
Length = 224
Score = 162 bits (411), Expect = 5e-39
Identities = 97/232 (41%), Positives = 132/232 (56%), Gaps = 19/232 (8%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEILDYLGKSS 60
MKFLE P D ++ FL NL+LG+ TI+G +EA+SCKH+G D++LSISL +EILD L KSS
Sbjct: 1 MKFLEYTPFDSINLFLDNLDLGDCTIRGNLEAFSCKHTGNDRRLSISLEHEILDCLSKSS 60
Query: 61 DNDSPSPVDSLSSPVDSLSARTSPAG--------RHWFT*FLPSITCIQIMISAQLKLTS 112
D+D SSPV+ LS R+S H + + S + + + S
Sbjct: 61 DSDH-------SSPVEHLSCRSSRKTLIYLVLTLSHMYPDYDFSAVRAHLFFKEE-EWES 112
Query: 113 FSQRNHGTVLSRFLKPTCLKPQ--SLLDALFKAVDEVVSLDDCEIYGYLPDSEADPLPER 170
F + T LS K + SLLD++ +DEV+ + + +IY Y PD + DP E+
Sbjct: 113 FKEMTD-TYLSEASKQWAATNEGTSLLDSMTNVIDEVIKIGESDIYSYNPDQDGDPFLEK 171
Query: 171 GAIWSFNFLFYNRKLKRIVTFRLSCFSNLIADGFSLDEICDEYDEEIFADMD 222
G IWS NF FYNRKLKR+V+FR SC S + D F D +E+ DMD
Sbjct: 172 GVIWSINFFFYNRKLKRVVSFRCSCISKISGDDFLTSAPSDGEEEDALIDMD 223
>UniRef100_UPI000035FA4E UPI000035FA4E UniRef100 entry
Length = 208
Score = 89.4 bits (220), Expect = 6e-17
Identities = 65/207 (31%), Positives = 100/207 (47%), Gaps = 22/207 (10%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEILDYLGKSS 60
MK LE + + LS L + +GE I G IE+YSCK +G DK + E G+
Sbjct: 3 MKLLENSSFEALSSRLC-VEVGESRILGRIESYSCKMAGDDKHMFKQFCQE-----GEPH 56
Query: 61 DNDSPSPVDSLSSPVDSLSARTSPAGRHWFT*FLPSITCIQIMIS-----------AQLK 109
++ SP S S+ SL ++S G + + T ++ + + +
Sbjct: 57 VLEALSPPQSSSAASPSLLGKSSEDGENPLSDKCCRKTLFYLITTLNESFRPDYDFSAAR 116
Query: 110 LTSFSQRNH-----GTVLSRFLKPTCLKPQSLLDALFKAVDEVVSLDDCEIYGYLPDSEA 164
FS+ + V S + SL L+ A+D+ ++L C+IY Y PD ++
Sbjct: 117 AHEFSRESSLNWVANAVNSSLFSAVGEEFNSLRPELWNAIDQEINLQSCDIYSYNPDLDS 176
Query: 165 DPLPERGAIWSFNFLFYNRKLKRIVTF 191
DP E G++WSFN+ FYN+KLKRIV F
Sbjct: 177 DPFGEEGSLWSFNYFFYNKKLKRIVFF 203
>UniRef100_UPI000029BC72 UPI000029BC72 UniRef100 entry
Length = 243
Score = 86.7 bits (213), Expect = 4e-16
Identities = 61/204 (29%), Positives = 97/204 (46%), Gaps = 23/204 (11%)
Query: 30 IEAYSCKHSGADKKLSISLSNEILDYLGKSSDNDSPSPVDSLSSPVDSLSARTSPAGRHW 89
IE+YSCK +G DK + E G+ ++ SP S S SL ++S G +
Sbjct: 27 IESYSCKMAGDDKHMFKQFCQE-----GEPHVLEALSPPQSSSGASPSLLGKSSEDGENP 81
Query: 90 FT*FLPSITCIQIMIS-----------AQLKLTSFSQRNH-----GTVLSRFLKPTCLKP 133
+ T ++ + + + FS+ + V S +
Sbjct: 82 LSDKCCRKTLFYLITTLNESFRPDYDFSAARAHEFSRESSLNWVANAVNSSLFSAVGEEF 141
Query: 134 QSLLDALFKAVDEVVSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRL 193
SL L+ A+D+ ++L C+IY Y PD ++DP E G++WSFN+ FYN+KLKRIV F
Sbjct: 142 NSLRPELWNAIDQEINLQGCDIYSYNPDLDSDPFGEEGSLWSFNYFFYNKKLKRIVFF-- 199
Query: 194 SCFSNLIADGFSLDEICDEYDEEI 217
+C S + G+ D + +E D E+
Sbjct: 200 TCRSVSVLSGYGCDSLDNELDMEL 223
>UniRef100_Q6PGU2 Repressor of RNA polymerase III transcription MAF1 homolog
[Brachydanio rerio]
Length = 247
Score = 85.1 bits (209), Expect = 1e-15
Identities = 77/257 (29%), Positives = 118/257 (44%), Gaps = 54/257 (21%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNE----ILDYL 56
MK LE + + ++ L+ + G+ I G IE+YSCK +G DK++ E +L+ L
Sbjct: 1 MKLLENSSFEAINSLLT-IETGDCKIIGRIESYSCKMAGDDKQMFKQFCQEGQPHVLEAL 59
Query: 57 GKS-----SDNDSPSPVDSLSSPV-DSLSART------------------SPAGRHWFT* 92
S N +D P+ D S +T S H F+
Sbjct: 60 SPPQSSGISPNKLSHSLDDGEGPLSDKCSRKTLFYLIATLNESFRPDYDFSRTKGHDFS- 118
Query: 93 FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDEVVSLDD 152
PS+ + +++ L + +H L+PQ L++A+D+ +SL +
Sbjct: 119 KEPSVNWVFNAVNSSLSAAAGEAYSH------------LQPQ-----LWEALDKEISLAE 161
Query: 153 CEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFSNLIAD-----GFSLD 207
C+IY Y PD ++DP E G +WSFN+ FYN++LKRIV F S +A G LD
Sbjct: 162 CDIYSYNPDLDSDPYGEEGNLWSFNYFFYNKRLKRIVFFTCRSVSLFMAPRDSGIGTELD 221
Query: 208 EICDE--YDEEIFADMD 222
DE Y+EE +MD
Sbjct: 222 LELDEGSYEEEEEGNMD 238
>UniRef100_UPI0000249370 UPI0000249370 UniRef100 entry
Length = 237
Score = 84.0 bits (206), Expect = 3e-15
Identities = 69/246 (28%), Positives = 108/246 (43%), Gaps = 50/246 (20%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNE----ILDYL 56
MK LE + + LS L + G+ I G IE+YSCK +G DK + E +L+ L
Sbjct: 1 MKLLENSRFEALSSQLC-VETGDAQIIGRIESYSCKMAGDDKHMFKQFCQEGEPHVLEAL 59
Query: 57 GKSSDNDSPSPV------DSLSSPVDSLSART-------------------SPAGRHWFT 91
+ +PSP + +P+ R S A H F+
Sbjct: 60 SPPQSSSAPSPNLLGKSGEDAENPLSDKCCRKTLFYLITTLNESFRPDYDFSAARAHEFS 119
Query: 92 *FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDEVVSLD 151
PS + +++ L Q N +L L+ A+D+ ++L
Sbjct: 120 -REPSANWVANSVNSSLYSAVGEQFN-----------------TLGPELWTAIDQEINLQ 161
Query: 152 DCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFSNLIADGFSLDEICD 211
C+IY Y PD ++DP E G++WSFN+ FYN+KLKRI+ F +C S + + + +
Sbjct: 162 GCDIYSYNPDLDSDPFGEEGSLWSFNYFFYNKKLKRILFF--TCRSVSVLSSYGRSGLDN 219
Query: 212 EYDEEI 217
E D E+
Sbjct: 220 ELDMEL 225
>UniRef100_Q7PL26 CG40196-PA.3 [Drosophila melanogaster]
Length = 226
Score = 81.6 bits (200), Expect = 1e-14
Identities = 73/243 (30%), Positives = 108/243 (44%), Gaps = 55/243 (22%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKL--------------SI 46
MK LE + + +++ LS G TI G IE+YSCK A+K L ++
Sbjct: 1 MKLLESSRFEAINNALSIQTSGI-TIFGRIESYSCKMVAAEKVLYKRFTADSHGHDLQAL 59
Query: 47 SLSNEILDYLGKSSDNDSPSPVDSLSSPVDSLSART------------------SPAGRH 88
S + D+ N+S S + ++ D++S +T S A H
Sbjct: 60 SPPQTLADFSPNFRRNNSQSGDEGITL-CDTISRKTLFYLIATLNASFEPDYDFSEAKSH 118
Query: 89 WFT*FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDEVV 148
F+ PS+ + I A L + Q Q++ L+ AVD+ V
Sbjct: 119 EFS-KEPSLQWVMNSIHANLSALAGDQY-----------------QAIRQPLWSAVDDEV 160
Query: 149 SLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFSNLIAD---GFS 205
L +C+IY Y PD DP E G +WSFN+ FYN+KLKRIV F ++L A FS
Sbjct: 161 ILSECDIYSYNPDLSCDPFGEPGCLWSFNYFFYNKKLKRIVFFTSRAVNSLYAGETLPFS 220
Query: 206 LDE 208
++E
Sbjct: 221 MEE 223
>UniRef100_UPI000036E4D0 UPI000036E4D0 UniRef100 entry
Length = 256
Score = 79.7 bits (195), Expect = 5e-14
Identities = 77/246 (31%), Positives = 114/246 (46%), Gaps = 51/246 (20%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNE----ILDYL 56
MK LE + + ++ L+ + G+ I G IE+YSCK +G DK + E +L+ L
Sbjct: 1 MKLLENSSFEAINSQLT-VETGDAHIIGRIESYSCKMAGDDKHMFKQFCQEGQPHVLEAL 59
Query: 57 ------GKSSDNDSPSPVDSLSSPV-DSLSART------------------SPAGRHWFT 91
G S S S P+ D S +T S A H F+
Sbjct: 60 SPPQTSGLSPSRLSKSQGGEEEGPLSDKCSRKTLFYLIATLNESFRPDYDFSTARSHEFS 119
Query: 92 *FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDEVVSLD 151
PS++ + ++ L T + K LKPQ L+ AVDE + L
Sbjct: 120 -REPSLSWVVNAVNCSLF----------TAVREDFKD--LKPQ-----LWNAVDEEICLA 161
Query: 152 DCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFSNLIADGFSLDEICD 211
+C+IY Y PD ++DP E G++WSFN+ FYN++LKRIV F SC S + ++ E +
Sbjct: 162 ECDIYSYNPDLDSDPFGEDGSLWSFNYFFYNKRLKRIVFF--SCRS-ISGSTYTPSEAGN 218
Query: 212 EYDEEI 217
E D E+
Sbjct: 219 ELDMEL 224
>UniRef100_Q7ZWL6 Maf1-pending-prov protein [Xenopus laevis]
Length = 245
Score = 79.3 bits (194), Expect = 7e-14
Identities = 75/248 (30%), Positives = 107/248 (42%), Gaps = 51/248 (20%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNE----ILDYL 56
MK LE + + ++ L + G+ I G IE+YSCK +G DK + E +L+ L
Sbjct: 1 MKLLENSHFEAVNSQLC-VETGDAHIIGRIESYSCKMAGDDKHMFKQFCQEGQPHVLEAL 59
Query: 57 GKSSDNDSPSPV------DSLSSPVDSLSART------------------SPAGRHWFT* 92
SPS + D D S +T S A H F+
Sbjct: 60 SPPQTGISPSRLGKSQGADEEGPLSDKCSRKTLFYLIATLNESFRPDYDFSAARAHEFS- 118
Query: 93 FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDEVVSLDD 152
PS+ + +++ L V + T L+P L+ AVDE ++L +
Sbjct: 119 REPSLNWVVNAVNSSL------------VSALGDDFTSLRPN-----LWDAVDEEINLSE 161
Query: 153 CEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFSNLIAD----GFSLDE 208
C+IY Y PD + DP E G +WSFN+ FYN+KLKRIV F S G LD
Sbjct: 162 CDIYSYNPDLDCDPFGEDGNLWSFNYFFYNKKLKRIVFFSCRSISGYTYTHCDAGAELDM 221
Query: 209 ICDEYDEE 216
+E D+E
Sbjct: 222 DLEEEDDE 229
>UniRef100_Q5XIH0 Hypothetical protein [Rattus norvegicus]
Length = 260
Score = 79.3 bits (194), Expect = 7e-14
Identities = 73/256 (28%), Positives = 109/256 (42%), Gaps = 64/256 (25%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNE----ILDYL 56
MK LE + + ++ L+ + G+ I G IE+YSCK +G DK + E +L+ L
Sbjct: 1 MKLLENSSFEAINSQLT-VETGDAHIIGRIESYSCKMAGDDKHMFKQFCQEGQPHVLEAL 59
Query: 57 G-------------KSSDNDSPSPV-----------------DSLSSPVDSLSARTSPAG 86
KS + SP+ +S D +AR+
Sbjct: 60 SPPQTSGLSPSRLSKSQGGEDESPLSDKCSRKTLFYLIATLNESFRPDYDFSTARSHEFS 119
Query: 87 RHWFT*FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDE 146
R PS+ + ++ L V F LKPQ L+ AVDE
Sbjct: 120 RE------PSLRWVVNAVNCSL---------FSAVREDF---KALKPQ-----LWNAVDE 156
Query: 147 VVSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFS------NLI 200
+ L +C+IY Y PD ++DP E G++WSFN+ FYN++LKRIV F S +
Sbjct: 157 EICLAECDIYSYNPDLDSDPFGEDGSLWSFNYFFYNKRLKRIVFFSCRSISGSTYTPSEA 216
Query: 201 ADGFSLDEICDEYDEE 216
+ L+ +E DEE
Sbjct: 217 GNALDLELGAEEVDEE 232
>UniRef100_Q9H063 Repressor of RNA polymerase III transcription MAF1 homolog [Homo
sapiens]
Length = 256
Score = 79.3 bits (194), Expect = 7e-14
Identities = 77/246 (31%), Positives = 113/246 (45%), Gaps = 51/246 (20%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNE----ILDYL 56
MK LE + + ++ L+ + G+ I G IE+YSCK +G DK + E +L+ L
Sbjct: 1 MKLLENSSFEAINSQLT-VETGDAHIIGRIESYSCKMAGDDKHMFKQFCQEGQPHVLEAL 59
Query: 57 ------GKSSDNDSPSPVDSLSSPV-DSLSART------------------SPAGRHWFT 91
G S S S P+ D S +T S A H F+
Sbjct: 60 SPPQTSGLSPSRLSKSQGGEEEGPLSDKCSRKTLFYLIATLNESFRPDYDFSTARSHEFS 119
Query: 92 *FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDEVVSLD 151
PS++ + ++ L V F LKPQ L+ AVDE + L
Sbjct: 120 -REPSLSWVVNAVNCSL---------FSAVREDFKD---LKPQ-----LWNAVDEEICLA 161
Query: 152 DCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFSNLIADGFSLDEICD 211
+C+IY Y PD ++DP E G++WSFN+ FYN++LKRIV F SC S + ++ E +
Sbjct: 162 ECDIYSYNPDLDSDPFGEDGSLWSFNYFFYNKRLKRIVFF--SCRS-ISGSTYTPSEAGN 218
Query: 212 EYDEEI 217
E D E+
Sbjct: 219 ELDMEL 224
>UniRef100_Q9D0U6 Repressor of RNA polymerase III transcription MAF1 homolog [Mus
musculus]
Length = 258
Score = 79.0 bits (193), Expect = 9e-14
Identities = 70/231 (30%), Positives = 102/231 (43%), Gaps = 60/231 (25%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNE----ILDYL 56
MK LE + + ++ L+ + G+ I G IE+YSCK +G DK + E +L+ L
Sbjct: 1 MKLLENSSFEAINSQLT-VETGDAHIIGRIESYSCKMAGDDKHMFKQFCQEGQPHVLEAL 59
Query: 57 G-------------KSSDNDSPSPV-----------------DSLSSPVDSLSARTSPAG 86
KS + SP+ +S D +AR+
Sbjct: 60 SPPQTSGLSPSRLSKSQGGEDESPLSDKCSRKTLFYLIATLNESFRPDYDFSTARSHEFS 119
Query: 87 RHWFT*FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDE 146
R PS+ + ++ L V F LKPQ L+ AVDE
Sbjct: 120 RE------PSLRWVVNAVNCSL---------FSAVREDF---KALKPQ-----LWNAVDE 156
Query: 147 VVSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFS 197
+ L +C+IY Y PD ++DP E G++WSFN+ FYN++LKRIV F SC S
Sbjct: 157 EICLAECDIYSYNPDLDSDPFGEDGSLWSFNYFFYNKRLKRIVFF--SCRS 205
>UniRef100_Q7Q4I3 ENSANGP00000019317 [Anopheles gambiae str. PEST]
Length = 207
Score = 78.6 bits (192), Expect = 1e-13
Identities = 68/221 (30%), Positives = 98/221 (43%), Gaps = 49/221 (22%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEI----LDYL 56
MK LE + +++ L ++ G+ TI G IE+YSCK +G DK L ++E L L
Sbjct: 1 MKLLESTRFEAVNNKL-HIQTGDSTIYGRIESYSCKMAGNDKALYKRFTSEQAPTDLQAL 59
Query: 57 GKSSDNDSPSPV---DSLSSP-----VDSLSART------------------SPAGRHWF 90
SP SLS D++S +T S A H F
Sbjct: 60 SPPQTLQDMSPQILRSSLSGDEGVTLCDTISRKTLFYLIATLNAAFEPDYDFSDARSHEF 119
Query: 91 T*FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDEVVSL 150
+ PS+ + I L + Q + + AL+ +++ +SL
Sbjct: 120 S-KEPSLQWVLNSIEGNLGPVAGEQYS-----------------KIRHALWTTIEDEISL 161
Query: 151 DDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTF 191
DC+IY Y PD +DP E G +WSFN+ FYN+KLKRIV F
Sbjct: 162 KDCDIYSYNPDLNSDPFGEPGCLWSFNYFFYNKKLKRIVFF 202
>UniRef100_Q9TZN2 Hypothetical protein C43H8.2 [Caenorhabditis elegans]
Length = 245
Score = 56.6 bits (135), Expect = 5e-07
Identities = 24/60 (40%), Positives = 34/60 (56%)
Query: 138 DALFKAVDEVVSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFS 197
+ L+ +DE + DC+IY + E DP E G IW+ F+FYN+ LKR V + C S
Sbjct: 163 EELWGIIDEAIVPGDCQIYSFKSQFEDDPFTEDGCIWALAFIFYNKGLKRFVLLTIRCLS 222
>UniRef100_UPI00002349F9 UPI00002349F9 UniRef100 entry
Length = 314
Score = 52.0 bits (123), Expect = 1e-05
Identities = 62/228 (27%), Positives = 92/228 (40%), Gaps = 47/228 (20%)
Query: 1 MKFLECAPLDRLSDFLSNLNL--GERTIKGCIEAYSCKHSGADKKL----SISLSNEILD 54
MKFL PL D S+LN + I G E Y K +D+KL SL +
Sbjct: 1 MKFL---PLPEFEDVTSSLNFDTADCHIVGGCELYITKAPRSDRKLYKNIEQSLEAQYES 57
Query: 55 YLGKSSDNDSPSPVD-------SLSSPVDSLSARTSPAGRHWFT*FLPSITCIQ--IMIS 105
L S+ P+ D S SSP LS +S R F + ++ S
Sbjct: 58 VLRLSASLSPPTASDAAASLNLSRSSPFGPLSDHSS---RRTFAYLIATLNASHPDYDFS 114
Query: 106 AQLKLTSFSQRNHGTVLSRFLKPTC--LKPQSLLDA-----------------------L 140
L+ + F + + + + T L+P+ LD +
Sbjct: 115 HVLRPSDFHREKNLKRVMNTIDSTLFNLRPRETLDLTPPSPSAVSGSYNSEASARWGRRM 174
Query: 141 FKAVDEVVSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRI 188
+K +DE +SL +C IY Y PD + + GAIWS ++ F+NR KR+
Sbjct: 175 WKILDEQLSLKECNIYSYSPDEDPSDADD-GAIWSLHYFFFNRIRKRV 221
>UniRef100_O14109 ORF N150 [Schizosaccharomyces pombe]
Length = 150
Score = 51.2 bits (121), Expect = 2e-05
Identities = 17/57 (29%), Positives = 36/57 (62%)
Query: 137 LDALFKAVDEVVSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRL 193
++ +++ +D ++L DC +Y Y PDS++ P + IW ++ F+N+ +KR++ L
Sbjct: 51 VNGIWEIIDRHINLSDCSVYSYTPDSDSVPYGDDALIWGMSYFFFNKNMKRMLYLSL 107
>UniRef100_Q8I1S2 Hypothetical protein PFD0800c [Plasmodium falciparum]
Length = 389
Score = 47.4 bits (111), Expect = 3e-04
Identities = 31/94 (32%), Positives = 53/94 (55%), Gaps = 20/94 (21%)
Query: 140 LFKAVDEVVSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSC---- 195
++K + E++ C++Y YL D++ DP ++ +I SFN+ F+ +K KRI+ +SC
Sbjct: 294 IWKILKELIDFKYCDVYTYLNDTDNDPYVDKESISSFNYFFFAKKNKRILF--ISCITKP 351
Query: 196 ----------FSNL---IADGFSLDEICDEYDEE 216
F+NL I D S++E +EYD+E
Sbjct: 352 KYKNQKNDEDFNNLYMGIQDDPSINE-QNEYDDE 384
>UniRef100_UPI000021AB1D UPI000021AB1D UniRef100 entry
Length = 611
Score = 45.8 bits (107), Expect = 8e-04
Identities = 18/49 (36%), Positives = 34/49 (68%), Gaps = 1/49 (2%)
Query: 140 LFKAVDEVVSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRI 188
++ +D+ ++LDDC ++ Y P +E E GAIW+ ++ F+N++LKR+
Sbjct: 407 MWATIDKEMALDDCNVFSYQP-AENPFDEEEGAIWALHYFFFNKQLKRV 454
>UniRef100_Q7RKN9 Hypothetical protein [Plasmodium yoelii yoelii]
Length = 232
Score = 45.8 bits (107), Expect = 8e-04
Identities = 20/58 (34%), Positives = 38/58 (65%), Gaps = 2/58 (3%)
Query: 140 LFKAVDEVVSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFS 197
++K + E++ C+IY YL +++ DP ++ +I SFN+ F+ +K KRI+ +SC +
Sbjct: 138 VWKLLKELIDFKSCDIYTYLNNTDKDPYIDKQSISSFNYFFFAKKTKRILF--ISCIT 193
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.323 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 363,763,630
Number of Sequences: 2790947
Number of extensions: 14614935
Number of successful extensions: 33357
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 25
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 33301
Number of HSP's gapped (non-prelim): 67
length of query: 222
length of database: 848,049,833
effective HSP length: 123
effective length of query: 99
effective length of database: 504,763,352
effective search space: 49971571848
effective search space used: 49971571848
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 72 (32.3 bits)
Medicago: description of AC144731.10


