

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144726.5 - phase: 0 /pseudo
(293 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
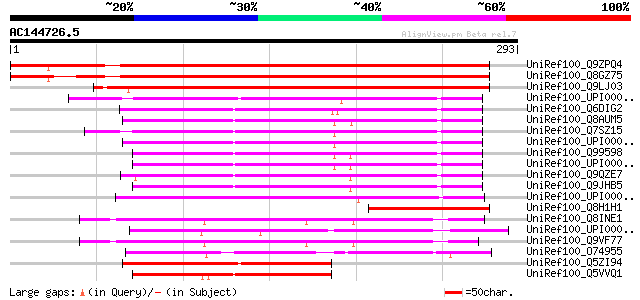
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9ZPQ4 Hypothetical protein At2g03780 [Arabidopsis tha... 355 8e-97
UniRef100_Q8GZ75 Hypothetical protein At2g03780/F19B11.23 [Arabi... 344 1e-93
UniRef100_Q9LJ03 Similar to putative translin [Oryza sativa] 326 5e-88
UniRef100_UPI00003ACDBC UPI00003ACDBC UniRef100 entry 137 3e-31
UniRef100_Q6DIG2 Translin-associated factor X [Xenopus tropicalis] 134 2e-30
UniRef100_Q8AUM5 Translin associated factor X [Fugu rubripes] 134 3e-30
UniRef100_Q7SZ15 MGC64311 protein [Xenopus laevis] 131 2e-29
UniRef100_UPI00002BB7FE UPI00002BB7FE UniRef100 entry 130 3e-29
UniRef100_Q99598 Translin-associated protein X [Homo sapiens] 127 5e-28
UniRef100_UPI0000368713 UPI0000368713 UniRef100 entry 126 6e-28
UniRef100_Q9QZE7 Translin associated protein X [Mus musculus] 126 8e-28
UniRef100_Q9JHB5 Trax [Rattus norvegicus] 125 1e-27
UniRef100_UPI0000431EF8 UPI0000431EF8 UniRef100 entry 125 2e-27
UniRef100_Q8H1H1 Translin-associated factor X [Lycopersicon escu... 118 2e-25
UniRef100_Q8INE1 CG5063-PB, isoform B [Drosophila melanogaster] 108 1e-22
UniRef100_UPI000042C634 UPI000042C634 UniRef100 entry 107 3e-22
UniRef100_Q9VF77 CG5063-PA, isoform A [Drosophila melanogaster] 100 6e-20
UniRef100_O74955 SPCC736.09c protein [Schizosaccharomyces pombe] 100 6e-20
UniRef100_Q5ZI94 Hypothetical protein [Gallus gallus] 94 6e-18
UniRef100_Q5VVQ1 Translin-associated factor X [Homo sapiens] 85 2e-15
>UniRef100_Q9ZPQ4 Hypothetical protein At2g03780 [Arabidopsis thaliana]
Length = 299
Score = 355 bits (911), Expect = 8e-97
Identities = 185/279 (66%), Positives = 219/279 (78%), Gaps = 10/279 (3%)
Query: 1 MLRSFSSSFHRLSLFMASEKT--QRLHQSILLFFDFGFSTVTGTNFQNTSKRPKTMSIAT 58
ML SS+F R++ + + K QRLHQ ++ FG V + ++ K+ +TMS
Sbjct: 1 MLSCSSSAFQRVAFMLMAPKLKPQRLHQMLISNDGFGVCVVAESGVEHLVKKARTMS--- 57
Query: 59 DTATVTDSAMKEPFTKYTEYLNNLNDKRERVVKASRDITMNSKKVIFQVHRMSKYNKDEV 118
T+S+MK+ F+ Y +YLNN N+KRERVVK SRDITMNSKKVIFQVHR+SK NK+EV
Sbjct: 58 -----TESSMKDAFSTYADYLNNFNEKRERVVKVSRDITMNSKKVIFQVHRLSKDNKEEV 112
Query: 119 LEKAEKDLAAVTNQHVSRLVKELQGTDFWKLRRAYSPGIQEYVEAATFCSFCKNGTLLKL 178
LEKA KDL AV +QH +RL+KELQGTDFWKLRRAYSPG+QEYVEAATF FC +GTL L
Sbjct: 113 LEKAGKDLEAVRDQHFARLMKELQGTDFWKLRRAYSPGVQEYVEAATFYKFCLSGTLCTL 172
Query: 179 DEINKTLLPLSDPSLQPLQINILDYILGLADLTGELMRLAIGRISDGELEFAEKICSFAR 238
DEIN TL+PLSDPSL+PLQINILDYILGLADLTGELMR+AIGRISDGE+EFA++IC F R
Sbjct: 173 DEINTTLVPLSDPSLEPLQINILDYILGLADLTGELMRMAIGRISDGEIEFAQRICQFVR 232
Query: 239 DIYRELTLVVPHMDDSSDMKTKMETMLQSVMKIENVFMS 277
I+REL LVVP MDDS DMK+KME MLQSV+KIEN S
Sbjct: 233 QIHRELMLVVPKMDDSYDMKSKMEVMLQSVIKIENACFS 271
>UniRef100_Q8GZ75 Hypothetical protein At2g03780/F19B11.23 [Arabidopsis thaliana]
Length = 287
Score = 344 bits (883), Expect = 1e-93
Identities = 182/279 (65%), Positives = 215/279 (76%), Gaps = 22/279 (7%)
Query: 1 MLRSFSSSFHRLSLFMASEKT--QRLHQSILLFFDFGFSTVTGTNFQNTSKRPKTMSIAT 58
ML SS+F R++ + + K QRLHQ + + ++ K+ +TMS
Sbjct: 1 MLSCSSSAFQRVAFMLMAPKLKPQRLHQ------------IAESGVEHLVKKARTMS--- 45
Query: 59 DTATVTDSAMKEPFTKYTEYLNNLNDKRERVVKASRDITMNSKKVIFQVHRMSKYNKDEV 118
T+S+MK+ F+ Y +YLNN N+KRERVVK SRDITMNSKKVIFQVHR+SK NK+EV
Sbjct: 46 -----TESSMKDAFSTYADYLNNFNEKRERVVKVSRDITMNSKKVIFQVHRLSKDNKEEV 100
Query: 119 LEKAEKDLAAVTNQHVSRLVKELQGTDFWKLRRAYSPGIQEYVEAATFCSFCKNGTLLKL 178
LEKA KDL AV +QH +RL+KELQGTDFWKLRRAYSPG+QEYVEAATF FC +GTL L
Sbjct: 101 LEKAGKDLEAVRDQHFARLMKELQGTDFWKLRRAYSPGVQEYVEAATFYKFCLSGTLCTL 160
Query: 179 DEINKTLLPLSDPSLQPLQINILDYILGLADLTGELMRLAIGRISDGELEFAEKICSFAR 238
DEIN TL+PLSDPSL+PLQINILDYILGLADLTGELMR+AIGRISDGE+EFA++IC F R
Sbjct: 161 DEINTTLVPLSDPSLEPLQINILDYILGLADLTGELMRMAIGRISDGEIEFAQRICQFVR 220
Query: 239 DIYRELTLVVPHMDDSSDMKTKMETMLQSVMKIENVFMS 277
I+REL LVVP MDDS DMK+KME MLQSV+KIEN S
Sbjct: 221 QIHRELMLVVPKMDDSYDMKSKMEVMLQSVIKIENACFS 259
>UniRef100_Q9LJ03 Similar to putative translin [Oryza sativa]
Length = 324
Score = 326 bits (835), Expect = 5e-88
Identities = 167/235 (71%), Positives = 192/235 (81%), Gaps = 8/235 (3%)
Query: 49 KRPKTMSIATDTATVTDSA------MKEPFTKYTEYLNNLNDKRERVVKASRDITMNSKK 102
KR +TM ATD A SA MK F K+ EYLN LNDKRER+VKASRD+TMNSKK
Sbjct: 62 KRSRTM--ATDAAATAHSASAGCSAMKAEFAKHAEYLNTLNDKRERLVKASRDLTMNSKK 119
Query: 103 VIFQVHRMSKYNKDEVLEKAEKDLAAVTNQHVSRLVKELQGTDFWKLRRAYSPGIQEYVE 162
IFQVHR+SK NK+EVL KAE DL V NQ++ +LVKELQGTDFWKLRRAY+ G+QEYVE
Sbjct: 120 AIFQVHRISKNNKEEVLSKAENDLTVVVNQYIGKLVKELQGTDFWKLRRAYTFGVQEYVE 179
Query: 163 AATFCSFCKNGTLLKLDEINKTLLPLSDPSLQPLQINILDYILGLADLTGELMRLAIGRI 222
AATFC FCK GTLL L EIN +LL L D S++PLQIN+LDY+LG+ADL+GELMRLAIGRI
Sbjct: 180 AATFCRFCKTGTLLSLAEINDSLLELGDKSVEPLQINVLDYVLGVADLSGELMRLAIGRI 239
Query: 223 SDGELEFAEKICSFARDIYRELTLVVPHMDDSSDMKTKMETMLQSVMKIENVFMS 277
SDGE+E+A+ IC+F RDIYRELTLVVP MDD+S+MK KMETMLQSV+KIEN S
Sbjct: 240 SDGEVEYAKNICAFVRDIYRELTLVVPLMDDNSEMKKKMETMLQSVVKIENACFS 294
>UniRef100_UPI00003ACDBC UPI00003ACDBC UniRef100 entry
Length = 285
Score = 137 bits (346), Expect = 3e-31
Identities = 87/256 (33%), Positives = 141/256 (54%), Gaps = 26/256 (10%)
Query: 35 GFSTVTGTNFQNTSKRPKTMSIATDTATVTDSAMKEPFTKYTEYLNNLNDKRERVVKASR 94
GF NF + +R + ++ + +T F + L+ +DK ER+VK SR
Sbjct: 4 GFRKRKHDNFPHGQRREEKENVNPSSPLMTS------FKSFQLELDTRHDKYERLVKLSR 57
Query: 95 DITMNSKKVIFQVHR-MSKYNKDEVLEKAEKDLAAVTNQHVSRLVKELQGTDFWKLRRAY 153
DIT+ SK+ IF +HR +S N +EVL ++E L AV + + ++ +EL G D ++ RA
Sbjct: 58 DITIESKRTIFLLHRYISAPNGEEVLNESEVKLDAVRRK-IKQVAQELIGEDMYQFHRAI 116
Query: 154 SPGIQEYVEAATFCSFCKNGTLLKLDEINKTLLPLSD----------------PSLQPLQ 197
SPG+QEY+EA +F F K +L+ ++EINK L+ ++ P L+
Sbjct: 117 SPGLQEYIEAVSFQYFIKTRSLISIEEINKQLVFTAEDREETTNMTSTSQDKQPHTWSLK 176
Query: 198 INILDYILGLADLTGELMRLAIGRISDGELEFAEKICSFARDIYRELTLVVPHMDDSSDM 257
+ +DY+LG+ADLTGELMRL I + +G+++ ++ F R IY T + ++
Sbjct: 177 VTPVDYLLGVADLTGELMRLCISSVGNGDIDTPFELSQFLRQIYDGFTFI--GNTGPYEV 234
Query: 258 KTKMETMLQSVMKIEN 273
K+ T+ QS+ K+EN
Sbjct: 235 SKKLYTLKQSLAKVEN 250
>UniRef100_Q6DIG2 Translin-associated factor X [Xenopus tropicalis]
Length = 297
Score = 134 bits (338), Expect = 2e-30
Identities = 83/236 (35%), Positives = 133/236 (56%), Gaps = 29/236 (12%)
Query: 64 TDSAMKEPFTKYTEYLNNLNDKRERVVKASRDITMNSKKVIFQVHRM-SKYNKDEVLEKA 122
+ SA+ F + L+ +DK ER+VK RDIT+ SK+ IF +HRM S +NK++VL +A
Sbjct: 31 SSSAVLMSFKAFQHDLDARHDKYERLVKLGRDITIESKRTIFLLHRMISDHNKEDVLSEA 90
Query: 123 EKDLAAVTNQHVSRLVKELQGTDFWKLRRAYSPGIQEYVEAATFCSFCKNGTLLKLDEIN 182
E L AV Q + + +EL G D ++ RA++PG+QEYVEA TF F ++ TL+ ++EIN
Sbjct: 91 ETKLLAV-RQKIKEIAEELVGEDMYQYHRAFTPGLQEYVEAITFKHFIESRTLVTINEIN 149
Query: 183 KTL-------LPL------------------SDPSLQPLQINILDYILGLADLTGELMRL 217
K L +P+ S S +Q+ +DY+LG+ADLTGELMR
Sbjct: 150 KQLIFEDLENMPMITTESFCGNLSSSPDNRHSKISALSIQVTPVDYLLGVADLTGELMRY 209
Query: 218 AIGRISDGELEFAEKICSFARDIYRELTLVVPHMDDSSDMKTKMETMLQSVMKIEN 273
I + +G+++ ++ F R ++ + + ++ K+ + QS+ K+EN
Sbjct: 210 CISSVGNGDIDTPFELSCFLRQVFDGFSYI--GNTGPYEISRKIHVLKQSLSKVEN 263
>UniRef100_Q8AUM5 Translin associated factor X [Fugu rubripes]
Length = 280
Score = 134 bits (337), Expect = 3e-30
Identities = 80/222 (36%), Positives = 131/222 (58%), Gaps = 17/222 (7%)
Query: 66 SAMKEPFTKYTEYLNNLNDKRERVVKASRDITMNSKKVIFQVHRMSKYNKDE-VLEKAEK 124
SA+ F + + L+ +DK ER+VK SRD+T+ SK+ IF +HR++ E VL +A+
Sbjct: 27 SAIMSVFRVFQQELDTKHDKYERLVKISRDVTIESKRTIFLLHRVTSVQDAEAVLNEADS 86
Query: 125 DLAAVTNQHVSRLVKELQGTDFWKLRRAYSPGIQEYVEAATFCSFCKNGTLLKLDEINKT 184
L AV Q + ++ KELQG D ++ RA++PGIQE+VEAA+F + ++ +L+ L+EIN
Sbjct: 87 KLDAV-RQKIGQIAKELQGEDIYQFHRAFTPGIQEFVEAASFLHYIRHRSLVSLEEINAR 145
Query: 185 LL---PLSDPSLQPL----------QINILDYILGLADLTGELMRLAIGRISDGELEFAE 231
L+ P PS+ + Q+ DY+LG+ADLTGELMRL I + +G+++
Sbjct: 146 LVFVRPEEPPSMDSVEAGPAGALTFQVTPSDYLLGVADLTGELMRLCISSVGNGDIDTPF 205
Query: 232 KICSFARDIYRELTLVVPHMDDSSDMKTKMETMLQSVMKIEN 273
++ F R I+ + ++ K+ + QS+ K+E+
Sbjct: 206 QLSQFLRQIHDGFFYI--GNTGPYEVSKKLHVLRQSLGKVED 245
>UniRef100_Q7SZ15 MGC64311 protein [Xenopus laevis]
Length = 297
Score = 131 bits (330), Expect = 2e-29
Identities = 83/256 (32%), Positives = 137/256 (53%), Gaps = 35/256 (13%)
Query: 44 FQNTSKRPKTMSIATDTATVTDSAMKEPFTKYTEYLNNLNDKRERVVKASRDITMNSKKV 103
FQ + ++ + S+ + +A V F + L+ +DK ER+VK RDIT+ SK+
Sbjct: 17 FQRSQRKDEKGSVHSSSAVVM------AFKDFQSELDARHDKYERLVKLGRDITIESKRT 70
Query: 104 IFQVHR-MSKYNKDEVLEKAEKDLAAVTNQHVSRLVKELQGTDFWKLRRAYSPGIQEYVE 162
IF +HR MS +NK++VL +AE L V Q + + +EL G D ++ RA++PG+QEYVE
Sbjct: 71 IFLLHRIMSDHNKEDVLSEAETKLLTV-RQKIREIAEELVGEDMYQYHRAFTPGLQEYVE 129
Query: 163 AATFCSFCKNGTLLKLDEINKTLL-------------------------PLSDPSLQPLQ 197
A TF F ++ TL+ ++EINK L+ S + +Q
Sbjct: 130 AITFKHFIESRTLVTINEINKQLIFEGLENMPTITRESFCSNLSCSTENDHSKITALRIQ 189
Query: 198 INILDYILGLADLTGELMRLAIGRISDGELEFAEKICSFARDIYRELTLVVPHMDDSSDM 257
+ +DY+LG+ADLTGELMR I + +G+++ ++ F R ++ + ++
Sbjct: 190 VTPVDYLLGVADLTGELMRYCISSVGNGDIDTPFELSCFLRQVFDGFAYI--GNTGPYEI 247
Query: 258 KTKMETMLQSVMKIEN 273
K+ + QS+ K+EN
Sbjct: 248 SRKIHVLKQSLSKVEN 263
>UniRef100_UPI00002BB7FE UPI00002BB7FE UniRef100 entry
Length = 278
Score = 130 bits (328), Expect = 3e-29
Identities = 75/220 (34%), Positives = 130/220 (59%), Gaps = 15/220 (6%)
Query: 66 SAMKEPFTKYTEYLNNLNDKRERVVKASRDITMNSKKVIFQVHRMSKY-NKDEVLEKAEK 124
SA+ F + + L+ +DK ER+VK SRD+T+ SK+ IF +HR++ + + +L +A+
Sbjct: 28 SAVLSVFKVFQQELDIKHDKYERLVKISRDVTIESKRTIFLLHRVTSVPDAEALLSEADT 87
Query: 125 DLAAVTNQHVSRLVKELQGTDFWKLRRAYSPGIQEYVEAATFCSFCKNGTLLKLDEINKT 184
L AV Q + ++ +EL+G D ++ RA++PGIQE+VEAA+F + ++ +L+ L+EIN
Sbjct: 88 KLEAV-RQKIGQIAEELRGEDIYQFHRAFTPGIQEFVEAASFLHYIRHRSLISLEEINAR 146
Query: 185 LL-----------PLSDPSLQPLQINILDYILGLADLTGELMRLAIGRISDGELEFAEKI 233
L+ P Q+ DY+LG+ADLTGELMRL I + +G+++ ++
Sbjct: 147 LVFVGSKELDNKDSAGSPEALTFQVTPSDYLLGVADLTGELMRLCISSVGNGDIDTPFQL 206
Query: 234 CSFARDIYRELTLVVPHMDDSSDMKTKMETMLQSVMKIEN 273
F R I+ + + ++ K+ T+ QS+ K+E+
Sbjct: 207 SQFLRQIHDGFSYI--GNTGPYEVSKKLHTLRQSLGKVED 244
>UniRef100_Q99598 Translin-associated protein X [Homo sapiens]
Length = 290
Score = 127 bits (318), Expect = 5e-28
Identities = 77/220 (35%), Positives = 127/220 (57%), Gaps = 21/220 (9%)
Query: 72 FTKYTEYLNNLNDKRERVVKASRDITMNSKKVIFQVHRM-SKYNKDEVLEKAEKDLAAVT 130
F + + L+ +DK ER+VK SRDIT+ SK+ IF +HR+ S + +++L ++E L V
Sbjct: 39 FKSFQQELDARHDKYERLVKLSRDITVESKRTIFLLHRITSAPDMEDILTESEIKLDGV- 97
Query: 131 NQHVSRLVKELQGTDFWKLRRAYSPGIQEYVEAATFCSFCKNGTLLKLDEINKTLL---- 186
Q + ++ +EL G D + RA + G+QEYVEA +F F K +L+ +DEINK L+
Sbjct: 98 RQKIFQVAQELSGEDMHQFHRAITTGLQEYVEAVSFQHFIKTRSLISMDEINKQLIFTTE 157
Query: 187 --------PLSDPSLQP-----LQINILDYILGLADLTGELMRLAIGRISDGELEFAEKI 233
P SD + L++ +DY+LG+ADLTGELMR+ I + +G+++ ++
Sbjct: 158 DNGKENKTPSSDAQDKQFGTWRLRVTPVDYLLGVADLTGELMRMCINSVGNGDIDTPFEV 217
Query: 234 CSFARDIYRELTLVVPHMDDSSDMKTKMETMLQSVMKIEN 273
F R +Y + + ++ K+ T+ QS+ K+EN
Sbjct: 218 SQFLRQVYDGFSFI--GNTGPYEVSKKLYTLKQSLAKVEN 255
>UniRef100_UPI0000368713 UPI0000368713 UniRef100 entry
Length = 344
Score = 126 bits (317), Expect = 6e-28
Identities = 76/220 (34%), Positives = 127/220 (57%), Gaps = 21/220 (9%)
Query: 72 FTKYTEYLNNLNDKRERVVKASRDITMNSKKVIFQVHRM-SKYNKDEVLEKAEKDLAAVT 130
F + + L+ +DK ER+VK SRD+T+ SK+ IF +HR+ S + +++L ++E L V
Sbjct: 93 FKSFQQELDARHDKYERLVKLSRDVTVESKRTIFLLHRITSAPDMEDILTESEIKLDGV- 151
Query: 131 NQHVSRLVKELQGTDFWKLRRAYSPGIQEYVEAATFCSFCKNGTLLKLDEINKTLL---- 186
Q + ++ +EL G D + RA + G+QEYVEA +F F K +L+ +DEINK L+
Sbjct: 152 RQKIFQVAQELSGEDMHQFHRAITTGLQEYVEAVSFQHFIKTRSLISMDEINKQLIFTTE 211
Query: 187 --------PLSDPSLQP-----LQINILDYILGLADLTGELMRLAIGRISDGELEFAEKI 233
P SD + L++ +DY+LG+ADLTGELMR+ I + +G+++ ++
Sbjct: 212 DNGKENKTPSSDAQDKQFGTWRLRVTPVDYLLGVADLTGELMRMCINSVGNGDIDTPFEV 271
Query: 234 CSFARDIYRELTLVVPHMDDSSDMKTKMETMLQSVMKIEN 273
F R +Y + + ++ K+ T+ QS+ K+EN
Sbjct: 272 SQFLRQVYDGFSFI--GNTGPYEVSKKLYTLKQSLAKVEN 309
>UniRef100_Q9QZE7 Translin associated protein X [Mus musculus]
Length = 290
Score = 126 bits (316), Expect = 8e-28
Identities = 77/231 (33%), Positives = 131/231 (56%), Gaps = 25/231 (10%)
Query: 65 DSAMKEP----FTKYTEYLNNLNDKRERVVKASRDITMNSKKVIFQVHRM-SKYNKDEVL 119
D+++ P F + + L+ +DK ER+VK SRDIT+ SK+ IF +HR+ S + +E+L
Sbjct: 28 DASLSSPVMLAFKSFQQELDARHDKYERLVKLSRDITVESKRTIFLLHRITSAPDMEEIL 87
Query: 120 EKAEKDLAAVTNQHVSRLVKELQGTDFWKLRRAYSPGIQEYVEAATFCSFCKNGTLLKLD 179
++E L V Q + ++ +EL G D + RA + G+QEYVEA +F F K +L+ ++
Sbjct: 88 TESESKLDGV-RQKILQVAQELSGEDMHQFHRAVTTGLQEYVEAVSFQHFIKTRSLISME 146
Query: 180 EINKTLLPLSDPSLQP-----------------LQINILDYILGLADLTGELMRLAIGRI 222
EINK L ++ S + L++ +DY+LG+ADLTGELMR+ I +
Sbjct: 147 EINKQLTFTAEDSGKESKTPPAEGQEKQLVTWRLKLTPVDYLLGVADLTGELMRMCINSV 206
Query: 223 SDGELEFAEKICSFARDIYRELTLVVPHMDDSSDMKTKMETMLQSVMKIEN 273
+G+++ ++ F R +Y + + ++ K+ T+ QS+ K+EN
Sbjct: 207 GNGDIDTPFEVSQFLRQVYDGFSFI--GNTGPYEVSKKLYTLKQSLAKVEN 255
>UniRef100_Q9JHB5 Trax [Rattus norvegicus]
Length = 290
Score = 125 bits (314), Expect = 1e-27
Identities = 75/220 (34%), Positives = 126/220 (57%), Gaps = 21/220 (9%)
Query: 72 FTKYTEYLNNLNDKRERVVKASRDITMNSKKVIFQVHRM-SKYNKDEVLEKAEKDLAAVT 130
F + + L+ +DK ER+VK SRDIT+ SK+ IF +HR+ S + +E+L ++E L V
Sbjct: 39 FKSFQQELDTRHDKYERLVKLSRDITVESKRTIFLLHRITSAPDMEEILTESESKLDGV- 97
Query: 131 NQHVSRLVKELQGTDFWKLRRAYSPGIQEYVEAATFCSFCKNGTLLKLDEINKTLLPLSD 190
Q + ++ +EL G D + RA + G+QEYVEA +F F + +L+ ++EIN+ L +D
Sbjct: 98 RQKMLQVAQELSGEDMHQFHRAVTTGLQEYVEAVSFQHFIRTRSLISMEEINRQLTFTTD 157
Query: 191 PSLQP-----------------LQINILDYILGLADLTGELMRLAIGRISDGELEFAEKI 233
S + L+I +DY+LG+ADLTGELMR+ I + +G+++ ++
Sbjct: 158 DSGKESKAPPADGQDKQLVTWRLKITPVDYLLGVADLTGELMRMCINSVGNGDIDTPFEV 217
Query: 234 CSFARDIYRELTLVVPHMDDSSDMKTKMETMLQSVMKIEN 273
F R +Y + + ++ K+ T+ QS+ K+EN
Sbjct: 218 SQFLRQVYDGFSFI--GNTGPYEVSKKLYTLKQSLSKVEN 255
>UniRef100_UPI0000431EF8 UPI0000431EF8 UniRef100 entry
Length = 264
Score = 125 bits (313), Expect = 2e-27
Identities = 75/216 (34%), Positives = 123/216 (56%), Gaps = 5/216 (2%)
Query: 62 TVTDSAMKEPFTKYTEYLNNLNDKRERVVKASRDITMNSKKVIFQVHRMSKYNKDE-VLE 120
T +S + + F Y L++ +D+ ER+VK RDIT+ SK++IF +H + K +K E VL
Sbjct: 21 TNENSFVLQQFRAYATELDDKHDRFERIVKTGRDITIESKRIIFLLHTIDKKSKQESVLC 80
Query: 121 KAEKDLAAVTNQHVSRLVKELQGTDFWKLRRAYSPGIQEYVEAATFCSFCKNGTLLKLDE 180
+A+ L V H + +EL+ D + +AY G++EY+EA TF + +G + E
Sbjct: 81 EADLRLQNVAQNHFKAISRELENQDPYLYLKAYRNGLEEYIEAVTFYQYLSSGNMKNWLE 140
Query: 181 INKTLLPLSDPSLQPLQINI--LDYILGLADLTGELMRLAIGRISDGELEFAEKICSFAR 238
+ K L S + +QI I +YILG++DLTGELMR I ++ G+ + C+F R
Sbjct: 141 LGKALTYTSSEKSKFIQILITPFEYILGISDLTGELMRQCINNLATGDSASCYETCNFVR 200
Query: 239 DIYRELTLVVPHMDDSSDMKTKMETMLQSVMKIENV 274
++Y+ V + ++ K+ T+ QS+ K+ENV
Sbjct: 201 NMYKGFLGCV--SISNKEINRKLCTLKQSLHKMENV 234
>UniRef100_Q8H1H1 Translin-associated factor X [Lycopersicon esculentum]
Length = 98
Score = 118 bits (296), Expect = 2e-25
Identities = 59/70 (84%), Positives = 63/70 (89%)
Query: 208 ADLTGELMRLAIGRISDGELEFAEKICSFARDIYRELTLVVPHMDDSSDMKTKMETMLQS 267
ADLTGELMRLAIGRIS+GEL+FAEKICSFAR+IYR LTL+ P MDDSSDMK KMETMLQS
Sbjct: 1 ADLTGELMRLAIGRISEGELDFAEKICSFAREIYRNLTLIAPEMDDSSDMKQKMETMLQS 60
Query: 268 VMKIENVFMS 277
VMKIEN S
Sbjct: 61 VMKIENACFS 70
>UniRef100_Q8INE1 CG5063-PB, isoform B [Drosophila melanogaster]
Length = 292
Score = 108 bits (271), Expect = 1e-22
Identities = 80/267 (29%), Positives = 131/267 (48%), Gaps = 44/267 (16%)
Query: 41 GTNFQNTSKRPKTMSIATDTATVTDSAMKEPFTKYTEYLNNLNDKRERVVKASRDITMNS 100
G +NT+ R + + A DS + + F Y+ L +D+ ER+VK SRDIT+ S
Sbjct: 6 GAGHRNTAPRKRQIPAAQ---LDEDSPIVQQFRIYSNELIMKHDRHERIVKLSRDITIES 62
Query: 101 KKVIFQVHRMS--KYNKDEVLEKAEKDLAAVTNQHVSRLVKELQGTDFWKLRRAYSPGIQ 158
K++IF +H + K NK++VLE+A + L + + + EL+ D ++ R +YSPG+Q
Sbjct: 63 KRIIFLLHSIDSRKQNKEKVLEEARQRLNKLIAVNFRAVALELRDQDVYQFRSSYSPGLQ 122
Query: 159 EYVEAATFCSFC-------KNGTLLKLD-EINKTLLPLSDPSLQPLQ------------- 197
E++EA T+ + +N T D + + ++ + S QP +
Sbjct: 123 EFIEAYTYMEYLCHEDAEGENETKSVSDWQAIQAVMQYVEESSQPKEEPTEGEDVQAIAQ 182
Query: 198 ----------INILDYILGLADLTGELMRLAIGRISDGELEFAEKICSFARDIYRELTLV 247
++ +YILGL+DLTGELMR I + G+ + C + Y L
Sbjct: 183 VESPKKFQFFVDPTEYILGLSDLTGELMRRCINSLGSGDTDTCLDTCKALQHFYSGLR-- 240
Query: 248 VPHMDDSSDMKTKMETMLQSVMKIENV 274
+ ++ K+ TM QSV+K ENV
Sbjct: 241 ------ARELWRKITTMKQSVLKAENV 261
>UniRef100_UPI000042C634 UPI000042C634 UniRef100 entry
Length = 270
Score = 107 bits (268), Expect = 3e-22
Identities = 74/230 (32%), Positives = 115/230 (49%), Gaps = 23/230 (10%)
Query: 70 EPFTKYTEYLNNLNDKRERVVKASRDITMNSKKVIFQVHR------MSKYNKDEVLEKAE 123
+ F Y L++ N RE+++ SR IT SKK+IF +HR + + EK E
Sbjct: 28 QTFEAYRAELDDENALREKLIILSRSITQLSKKLIFHLHRGATSQPAQRQKNNNEAEKKE 87
Query: 124 KDLAAVTNQHVSRLVKELQG----TDFWKLRRAYSPGIQEYVEAATFCSFCKNGTLLKLD 179
+++AAV L G + FWK R++ +PG++EY+E +F + ++G L+ LD
Sbjct: 88 REIAAVFKNIRQELSDARPGESWESGFWKWRKSITPGLEEYIEGLSFMWYLQHGGLVPLD 147
Query: 180 EINKTLLPLSDPSLQPL-QINILDYILGLADLTGELMRLAIGRISDGELEFAEKICSFAR 238
++ K LSD + +PL + DYILG++DLTGELMR A + G+ E IC F R
Sbjct: 148 QVQKA---LSDENGEPLIFVTPEDYILGMSDLTGELMRYATNALGTGDHETPLSICDFVR 204
Query: 239 DIYRELTLVVPHMDDSSDMKTKMETMLQSVMKIENVFMSEDQSIYHFWDQ 288
+ + K E +S+ KIE V + + F D+
Sbjct: 205 TVKTHAI---------RQLSKKQEETQRSLEKIEKVCYALRLRLIEFADR 245
>UniRef100_Q9VF77 CG5063-PA, isoform A [Drosophila melanogaster]
Length = 258
Score = 100 bits (248), Expect = 6e-20
Identities = 75/264 (28%), Positives = 126/264 (47%), Gaps = 44/264 (16%)
Query: 41 GTNFQNTSKRPKTMSIATDTATVTDSAMKEPFTKYTEYLNNLNDKRERVVKASRDITMNS 100
G +NT+ R + + A DS + + F Y+ L +D+ ER+VK SRDIT+ S
Sbjct: 6 GAGHRNTAPRKRQIPAAQ---LDEDSPIVQQFRIYSNELIMKHDRHERIVKLSRDITIES 62
Query: 101 KKVIFQVHRMS--KYNKDEVLEKAEKDLAAVTNQHVSRLVKELQGTDFWKLRRAYSPGIQ 158
K++IF +H + K NK++VLE+A + L + + + EL+ D ++ R +YSPG+Q
Sbjct: 63 KRIIFLLHSIDSRKQNKEKVLEEARQRLNKLIAVNFRAVALELRDQDVYQFRSSYSPGLQ 122
Query: 159 EYVEAATFCSFC-------KNGTLLKLD-EINKTLLPLSDPSLQPLQ------------- 197
E++EA T+ + +N T D + + ++ + S QP +
Sbjct: 123 EFIEAYTYMEYLCHEDAEGENETKSVSDWQAIQAVMQYVEESSQPKEEPTEGEDVQAIAQ 182
Query: 198 ----------INILDYILGLADLTGELMRLAIGRISDGELEFAEKICSFARDIYRELTLV 247
++ +YILGL+DLTGELMR I + G+ + C + Y L
Sbjct: 183 VESPKKFQFFVDPTEYILGLSDLTGELMRRCINSLGSGDTDTCLDTCKALQHFYSGLV-- 240
Query: 248 VPHMDDSSDMKTKMETMLQSVMKI 271
SS ++ + +L SV +
Sbjct: 241 ------SSPLRNSINFLLFSVTSV 258
>UniRef100_O74955 SPCC736.09c protein [Schizosaccharomyces pombe]
Length = 231
Score = 100 bits (248), Expect = 6e-20
Identities = 64/222 (28%), Positives = 115/222 (50%), Gaps = 30/222 (13%)
Query: 68 MKEPFTKYTEYLNNLNDKRERVVKASRDITMNSKKVIFQVHRMSK---------YNKDEV 118
M+E F + +L DKRE++++ SR+IT+ SK++IF +H+ S +++ +
Sbjct: 1 MEEEFLSFKNFLQEDQDKREKIIRLSREITIQSKRMIFLLHQTSSSDGFPLPKDFDRTSI 60
Query: 119 LEKAEKDLAAVTNQHVSRLVKELQGTDFWKLRRAYSPGIQEYVEAATFCSFCKNGTLLKL 178
EK ++ + L +EL G + K A + G+QEYVEA TF + + GTLL
Sbjct: 61 FEKK-------IHKELESLKRELAGLNADKFSSACTHGLQEYVEAVTFKFWLQTGTLLS- 112
Query: 179 DEINKTLLPLSDPSLQPLQINILDYILGLADLTGELMRLAIGRISDGELEFAEKICSFAR 238
D S + + IN +DY+LG+ D+TGE+MR + S ++ + F R
Sbjct: 113 ---------CKDSSFR-ISINFIDYVLGVCDMTGEIMRFLVTNGSKFSVQQLTQQVKFLR 162
Query: 239 DIYRELTLVVPHMDD--SSDMKTKMETMLQSVMKIENVFMSE 278
+++ + + H+ S+++ K+ M S+ K+E + S+
Sbjct: 163 GLHKNCS-EIEHLPSKVKSELQQKLSVMENSISKVEGICYSK 203
>UniRef100_Q5ZI94 Hypothetical protein [Gallus gallus]
Length = 166
Score = 93.6 bits (231), Expect = 6e-18
Identities = 52/122 (42%), Positives = 79/122 (64%), Gaps = 2/122 (1%)
Query: 66 SAMKEPFTKYTEYLNNLNDKRERVVKASRDITMNSKKVIFQVHR-MSKYNKDEVLEKAEK 124
S + F + L+ +DK ER+VK SRDIT+ SK+ IF +HR +S N +EVL ++E
Sbjct: 34 SPLMTSFKSFQLELDTRHDKYERLVKLSRDITIESKRTIFLLHRYISAPNGEEVLNESEV 93
Query: 125 DLAAVTNQHVSRLVKELQGTDFWKLRRAYSPGIQEYVEAATFCSFCKNGTLLKLDEINKT 184
L AV + + ++ +EL G D ++ RA SPG+QEY+EA +F F K +L+ ++EINK
Sbjct: 94 KLDAVRRK-IKQVAQELIGEDMYQFHRAISPGLQEYIEAVSFQYFIKTRSLISIEEINKQ 152
Query: 185 LL 186
L+
Sbjct: 153 LV 154
>UniRef100_Q5VVQ1 Translin-associated factor X [Homo sapiens]
Length = 163
Score = 85.1 bits (209), Expect = 2e-15
Identities = 49/123 (39%), Positives = 77/123 (61%), Gaps = 9/123 (7%)
Query: 72 FTKYTEYLNNLNDKRERVVKASRDITMNSKKVIFQVHRM--SKY------NKDEVLEKAE 123
F + + L+ +DK ER+VK SRDIT+ SK+ IF +HR+ S+Y + +++L ++E
Sbjct: 39 FKSFQQELDARHDKYERLVKLSRDITVESKRTIFLLHRITSSQYLDHSAPDMEDILTESE 98
Query: 124 KDLAAVTNQHVSRLVKELQGTDFWKLRRAYSPGIQEYVEAATFCSFCKNGTLLKLDEINK 183
L V Q + ++ +EL G D + RA + G+QEYVEA +F F K +L+ +DEINK
Sbjct: 99 IKLDGV-RQKIFQVAQELSGEDMHQFHRAITTGLQEYVEAVSFQHFIKTRSLISMDEINK 157
Query: 184 TLL 186
L+
Sbjct: 158 QLI 160
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.133 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 433,527,004
Number of Sequences: 2790947
Number of extensions: 16832891
Number of successful extensions: 56916
Number of sequences better than 10.0: 144
Number of HSP's better than 10.0 without gapping: 60
Number of HSP's successfully gapped in prelim test: 84
Number of HSP's that attempted gapping in prelim test: 56757
Number of HSP's gapped (non-prelim): 190
length of query: 293
length of database: 848,049,833
effective HSP length: 126
effective length of query: 167
effective length of database: 496,390,511
effective search space: 82897215337
effective search space used: 82897215337
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 74 (33.1 bits)
Medicago: description of AC144726.5


