

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144726.3 - phase: 0
(151 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
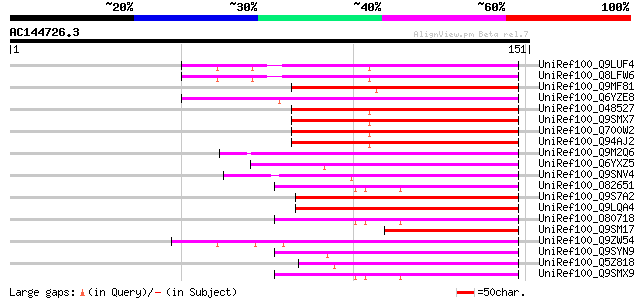
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LUF4 Similarity to squamosa promoter binding protein... 84 8e-16
UniRef100_Q8LFW6 Hypothetical protein [Arabidopsis thaliana] 84 8e-16
UniRef100_Q9MF81 Orf102a protein [Beta vulgaris subsp. vulgaris] 62 3e-09
UniRef100_Q6YZE8 SBP-domain protein-like [Oryza sativa] 60 1e-08
UniRef100_O48527 Putative squamosa-promoter binding protein [Ara... 60 1e-08
UniRef100_Q9SMX7 Squamosa promoter binding protein-like 9 [Arabi... 60 1e-08
UniRef100_Q700W2 Putative squamosa-promoter binding protein [Ara... 60 1e-08
UniRef100_Q94AJ2 Putative squamosa-promoter binding protein [Ara... 60 1e-08
UniRef100_Q9M2Q6 Squamosa promoter-binding protein homolog [Arab... 58 5e-08
UniRef100_Q6YXZ5 Putative squamosa promoter binding protein 2 [O... 57 8e-08
UniRef100_Q9SNV4 Squamosa promoter binding protein-homologue 4 [... 57 8e-08
UniRef100_O82651 Squamosa-promoter binding protein-like 1 [Arabi... 57 1e-07
UniRef100_Q9S7A2 Squamosa promoter binding protein-like 6 [Arabi... 57 1e-07
UniRef100_Q9LQA4 F4N2.13 [Arabidopsis thaliana] 57 1e-07
UniRef100_O80718 Putative squamosa-promoter binding protein [Ara... 57 1e-07
UniRef100_Q9SM17 SBP-domain protein 3 [Zea mays] 55 4e-07
UniRef100_Q9ZW54 Squamosa promoter binding protein-like 11 [Arab... 54 5e-07
UniRef100_Q9SYN9 F9H16.3 protein [Arabidopsis thaliana] 54 7e-07
UniRef100_Q5Z818 Squamosa promoter binding protein 2-like [Oryza... 54 7e-07
UniRef100_Q9SMX9 Squamosa promoter binding protein-like 1 [Arabi... 54 9e-07
>UniRef100_Q9LUF4 Similarity to squamosa promoter binding protein [Arabidopsis
thaliana]
Length = 359
Score = 83.6 bits (205), Expect = 8e-16
Identities = 51/107 (47%), Positives = 63/107 (58%), Gaps = 13/107 (12%)
Query: 51 EFFVDLKLG-HVGNSSAESG-------LTKSKDVGGFTKITSSSSSTSGSSKKARAINNG 102
+F DLKLG ++GNSS+ G L+K KD + + S S SSK+ R G
Sbjct: 39 DFSFDLKLGRNIGNSSSVFGDTEQVISLSKWKD----SALAKPEGSRSSSSKRTRGNGVG 94
Query: 103 T-QIVSSLADGCQADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
T Q+ L DGC +D SN R YH RHKVC +HSKT VT+ GHKQRF
Sbjct: 95 TNQMPICLVDGCDSDFSNCREYHKRHKVCDVHSKTPVVTINGHKQRF 141
>UniRef100_Q8LFW6 Hypothetical protein [Arabidopsis thaliana]
Length = 336
Score = 83.6 bits (205), Expect = 8e-16
Identities = 51/107 (47%), Positives = 63/107 (58%), Gaps = 13/107 (12%)
Query: 51 EFFVDLKLG-HVGNSSAESG-------LTKSKDVGGFTKITSSSSSTSGSSKKARAINNG 102
+F DLKLG ++GNSS+ G L+K KD + + S S SSK+ R G
Sbjct: 16 DFSFDLKLGRNIGNSSSVFGDTEQVISLSKWKD----SALAKPEGSRSSSSKRTRGNGVG 71
Query: 103 T-QIVSSLADGCQADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
T Q+ L DGC +D SN R YH RHKVC +HSKT VT+ GHKQRF
Sbjct: 72 TNQMPICLVDGCDSDFSNCREYHKRHKVCDVHSKTPVVTINGHKQRF 118
>UniRef100_Q9MF81 Orf102a protein [Beta vulgaris subsp. vulgaris]
Length = 102
Score = 61.6 bits (148), Expect = 3e-09
Identities = 33/67 (49%), Positives = 42/67 (62%), Gaps = 1/67 (1%)
Query: 83 TSSSSSTSGSSKKARAINNGTQI-VSSLADGCQADLSNYRGYHMRHKVCKLHSKTSQVTL 141
+SSSSS SK+ R N+G + V DGC +DL+ R YH RHKVC+ HSKT V +
Sbjct: 3 SSSSSSPVALSKRGRPRNSGAKNPVFCSVDGCASDLNQCREYHRRHKVCERHSKTPVVLV 62
Query: 142 GGHKQRF 148
GG K +F
Sbjct: 63 GGKKPQF 69
>UniRef100_Q6YZE8 SBP-domain protein-like [Oryza sativa]
Length = 455
Score = 60.1 bits (144), Expect = 1e-08
Identities = 37/108 (34%), Positives = 47/108 (43%), Gaps = 10/108 (9%)
Query: 51 EFFVDLKLGHVGNSSAESGLTKSKDVG----------GFTKITSSSSSTSGSSKKARAIN 100
E VDLKLG +G + G T +++SS + A
Sbjct: 51 ECSVDLKLGGLGEFGGGGAQPRVAVAGEPAKGKGPAAAATGAAAAASSAPAKRPRGAAAA 110
Query: 101 NGTQIVSSLADGCQADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
Q S DGC+ DLS R YH RHKVC+ HSKT V + G + RF
Sbjct: 111 GQQQCPSCAVDGCKEDLSKCRDYHRRHKVCEAHSKTPLVVVSGREMRF 158
>UniRef100_O48527 Putative squamosa-promoter binding protein [Arabidopsis thaliana]
Length = 375
Score = 59.7 bits (143), Expect = 1e-08
Identities = 30/68 (44%), Positives = 44/68 (64%), Gaps = 2/68 (2%)
Query: 83 TSSSSSTSGSSKKARAINNGT--QIVSSLADGCQADLSNYRGYHMRHKVCKLHSKTSQVT 140
+ SSSS S+++ R +G QI +GC DL+N +GY+ RH+VC +HSKT +VT
Sbjct: 47 SGSSSSGGRSNRRVRGGGSGQSGQIPRCQVEGCGMDLTNAKGYYSRHRVCGVHSKTPKVT 106
Query: 141 LGGHKQRF 148
+ G +QRF
Sbjct: 107 VAGIEQRF 114
>UniRef100_Q9SMX7 Squamosa promoter binding protein-like 9 [Arabidopsis thaliana]
Length = 373
Score = 59.7 bits (143), Expect = 1e-08
Identities = 30/68 (44%), Positives = 44/68 (64%), Gaps = 2/68 (2%)
Query: 83 TSSSSSTSGSSKKARAINNGT--QIVSSLADGCQADLSNYRGYHMRHKVCKLHSKTSQVT 140
+ SSSS S+++ R +G QI +GC DL+N +GY+ RH+VC +HSKT +VT
Sbjct: 47 SGSSSSGGRSNRRVRGGGSGQSGQIPRCQVEGCGMDLTNAKGYYSRHRVCGVHSKTPKVT 106
Query: 141 LGGHKQRF 148
+ G +QRF
Sbjct: 107 VAGIEQRF 114
>UniRef100_Q700W2 Putative squamosa-promoter binding protein [Arabidopsis thaliana]
Length = 378
Score = 59.7 bits (143), Expect = 1e-08
Identities = 30/68 (44%), Positives = 44/68 (64%), Gaps = 2/68 (2%)
Query: 83 TSSSSSTSGSSKKARAINNGT--QIVSSLADGCQADLSNYRGYHMRHKVCKLHSKTSQVT 140
+ SSSS S+++ R +G QI +GC DL+N +GY+ RH+VC +HSKT +VT
Sbjct: 47 SGSSSSGGRSNRRVRGGGSGQSGQIPRCQVEGCGMDLTNAKGYYSRHRVCGVHSKTPKVT 106
Query: 141 LGGHKQRF 148
+ G +QRF
Sbjct: 107 VAGIEQRF 114
>UniRef100_Q94AJ2 Putative squamosa-promoter binding protein [Arabidopsis thaliana]
Length = 374
Score = 59.7 bits (143), Expect = 1e-08
Identities = 30/68 (44%), Positives = 44/68 (64%), Gaps = 2/68 (2%)
Query: 83 TSSSSSTSGSSKKARAINNGT--QIVSSLADGCQADLSNYRGYHMRHKVCKLHSKTSQVT 140
+ SSSS S+++ R +G QI +GC DL+N +GY+ RH+VC +HSKT +VT
Sbjct: 47 SGSSSSGGRSNRRVRGGGSGQSGQIPRCQVEGCGMDLTNAKGYYSRHRVCGVHSKTPKVT 106
Query: 141 LGGHKQRF 148
+ G +QRF
Sbjct: 107 VAGIEQRF 114
>UniRef100_Q9M2Q6 Squamosa promoter-binding protein homolog [Arabidopsis thaliana]
Length = 354
Score = 57.8 bits (138), Expect = 5e-08
Identities = 32/87 (36%), Positives = 46/87 (52%), Gaps = 1/87 (1%)
Query: 62 GNSSAESGLTKSKDVGGFTKITSSSSSTSGSSKKARAINNGTQIVSSLADGCQADLSNYR 121
G+SS ES + S + KI S S S + + + +GC+ DLSN +
Sbjct: 14 GSSSTESS-SLSGGLRFGQKIYFEDGSGSRSKNRVNTVRKSSTTARCQVEGCRMDLSNVK 72
Query: 122 GYHMRHKVCKLHSKTSQVTLGGHKQRF 148
Y+ RHKVC +HSK+S+V + G QRF
Sbjct: 73 AYYSRHKVCCIHSKSSKVIVSGLHQRF 99
>UniRef100_Q6YXZ5 Putative squamosa promoter binding protein 2 [Oryza sativa]
Length = 387
Score = 57.0 bits (136), Expect = 8e-08
Identities = 31/84 (36%), Positives = 45/84 (52%), Gaps = 6/84 (7%)
Query: 71 TKSKDVGGFTKITSSSSSTS------GSSKKARAINNGTQIVSSLADGCQADLSNYRGYH 124
T +DV G + SS S S G +KK + TQ +GC DLS+ + YH
Sbjct: 134 TYFEDVCGGQNVKSSPSGVSVATPSPGLAKKVKVAQQNTQNPHCQVEGCNVDLSSAKPYH 193
Query: 125 MRHKVCKLHSKTSQVTLGGHKQRF 148
+H+VC+ HSKT +V + G ++RF
Sbjct: 194 RKHRVCEPHSKTLKVIVAGLERRF 217
>UniRef100_Q9SNV4 Squamosa promoter binding protein-homologue 4 [Antirrhinum majus]
Length = 257
Score = 57.0 bits (136), Expect = 8e-08
Identities = 33/88 (37%), Positives = 51/88 (57%), Gaps = 4/88 (4%)
Query: 63 NSSAESGLTKSKDVGGFTKITSSSSSTSGSSKKARA--INNGTQIVSSLADGCQADLSNY 120
+SS+ +GL K + + + SS S SKK R+ + G Q +GC+ DLS+
Sbjct: 4 SSSSLNGLNFGKKI--YFENVGSSGLQSSPSKKGRSGGVVQGGQPPRCQVEGCKIDLSDA 61
Query: 121 RGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
+ Y+ RHKVC +HSK+ +V + G +QRF
Sbjct: 62 KAYYSRHKVCGMHSKSPKVIVAGIEQRF 89
>UniRef100_O82651 Squamosa-promoter binding protein-like 1 [Arabidopsis thaliana]
Length = 881
Score = 56.6 bits (135), Expect = 1e-07
Identities = 36/95 (37%), Positives = 49/95 (50%), Gaps = 24/95 (25%)
Query: 78 GFTKITSSSSSTSGSSKKARAI-----NNG--TQIVSSLADG-----------------C 113
G + +SSS S G+ KK RA+ NG T ++ +DG C
Sbjct: 52 GNSSNSSSSCSDEGNDKKRRAVAIQGDTNGALTLNLNGESDGLFPAKKTKSGAVCQVENC 111
Query: 114 QADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
+ADLS + YH RHKVC++HSK + T+GG QRF
Sbjct: 112 EADLSKVKDYHRRHKVCEMHSKATSATVGGILQRF 146
>UniRef100_Q9S7A2 Squamosa promoter binding protein-like 6 [Arabidopsis thaliana]
Length = 405
Score = 56.6 bits (135), Expect = 1e-07
Identities = 30/65 (46%), Positives = 40/65 (61%)
Query: 84 SSSSSTSGSSKKARAINNGTQIVSSLADGCQADLSNYRGYHMRHKVCKLHSKTSQVTLGG 143
+SS + SKK+RA N +Q GC DLS+ + YH RH+VC+ HSKTS V + G
Sbjct: 100 TSSRGFALPSKKSRASNLCSQNPLCQVYGCSKDLSSSKDYHKRHRVCEAHSKTSVVIVNG 159
Query: 144 HKQRF 148
+QRF
Sbjct: 160 LEQRF 164
>UniRef100_Q9LQA4 F4N2.13 [Arabidopsis thaliana]
Length = 378
Score = 56.6 bits (135), Expect = 1e-07
Identities = 30/65 (46%), Positives = 40/65 (61%)
Query: 84 SSSSSTSGSSKKARAINNGTQIVSSLADGCQADLSNYRGYHMRHKVCKLHSKTSQVTLGG 143
+SS + SKK+RA N +Q GC DLS+ + YH RH+VC+ HSKTS V + G
Sbjct: 89 TSSRGFALPSKKSRASNLCSQNPLCQVYGCSKDLSSSKDYHKRHRVCEAHSKTSVVIVNG 148
Query: 144 HKQRF 148
+QRF
Sbjct: 149 LEQRF 153
>UniRef100_O80718 Putative squamosa-promoter binding protein [Arabidopsis thaliana]
Length = 240
Score = 56.6 bits (135), Expect = 1e-07
Identities = 36/95 (37%), Positives = 49/95 (50%), Gaps = 24/95 (25%)
Query: 78 GFTKITSSSSSTSGSSKKARAI-----NNG--TQIVSSLADG-----------------C 113
G + +SSS S G+ KK RA+ NG T ++ +DG C
Sbjct: 52 GNSSNSSSSCSDEGNDKKRRAVAIQGDTNGALTLNLNGESDGLFPAKKTKSGAVCQVENC 111
Query: 114 QADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
+ADLS + YH RHKVC++HSK + T+GG QRF
Sbjct: 112 EADLSKVKDYHRRHKVCEMHSKATSATVGGILQRF 146
>UniRef100_Q9SM17 SBP-domain protein 3 [Zea mays]
Length = 435
Score = 54.7 bits (130), Expect = 4e-07
Identities = 23/39 (58%), Positives = 29/39 (73%)
Query: 110 ADGCQADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
A+GC+ADLS + YH RHKVC+ H+K S V GG +QRF
Sbjct: 197 AEGCKADLSGAKHYHRRHKVCEYHAKASVVATGGKQQRF 235
>UniRef100_Q9ZW54 Squamosa promoter binding protein-like 11 [Arabidopsis thaliana]
Length = 393
Score = 54.3 bits (129), Expect = 5e-07
Identities = 36/116 (31%), Positives = 59/116 (50%), Gaps = 15/116 (12%)
Query: 48 CANEFFVDLKLG---HVGNSSAESGL-------TKSKDVGG-----FTKITSSSSSTSGS 92
C+N FV +K V +SAES L T S++ G + ++ + S
Sbjct: 100 CSNIDFVQVKAPTALEVSVASAESDLCLKLGKRTYSEEYWGRNNNEISAVSMKLLTPSVV 159
Query: 93 SKKARAINNGTQIVSSLADGCQADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
+ K++ + DGC+ DLS+ +GYH +HKVC+ HSK +V++ G ++RF
Sbjct: 160 AGKSKLCGQSMPVPRCQIDGCELDLSSAKGYHRKHKVCEKHSKCPKVSVSGLERRF 215
>UniRef100_Q9SYN9 F9H16.3 protein [Arabidopsis thaliana]
Length = 1035
Score = 53.9 bits (128), Expect = 7e-07
Identities = 26/73 (35%), Positives = 41/73 (55%), Gaps = 2/73 (2%)
Query: 78 GFTKITSSSSSTSG--SSKKARAINNGTQIVSSLADGCQADLSNYRGYHMRHKVCKLHSK 135
G T + ++++T +KK R+ + G D C DLS+ + YH RHKVC++HSK
Sbjct: 88 GLTAVEETTTTTQNVRPNKKVRSGSPGGNYPMCQVDNCTEDLSHAKDYHRRHKVCEVHSK 147
Query: 136 TSQVTLGGHKQRF 148
++ +G QRF
Sbjct: 148 ATKALVGKQMQRF 160
>UniRef100_Q5Z818 Squamosa promoter binding protein 2-like [Oryza sativa]
Length = 475
Score = 53.9 bits (128), Expect = 7e-07
Identities = 28/70 (40%), Positives = 40/70 (57%), Gaps = 6/70 (8%)
Query: 85 SSSSTSGSS------KKARAINNGTQIVSSLADGCQADLSNYRGYHMRHKVCKLHSKTSQ 138
SSS+ SG + KK + TQ +GC+ DLS+ R YH +HKVC+ HSK +
Sbjct: 151 SSSAASGVTCPSTVVKKMKVSQQSTQSSYCQVEGCKVDLSSAREYHRKHKVCEAHSKAPK 210
Query: 139 VTLGGHKQRF 148
V + G ++RF
Sbjct: 211 VIVSGLERRF 220
>UniRef100_Q9SMX9 Squamosa promoter binding protein-like 1 [Arabidopsis thaliana]
Length = 881
Score = 53.5 bits (127), Expect = 9e-07
Identities = 35/95 (36%), Positives = 48/95 (49%), Gaps = 24/95 (25%)
Query: 78 GFTKITSSSSSTSGSSKKARAI-----NNG--TQIVSSLADG-----------------C 113
G + +SSS S G+ KK RA+ NG T ++ +DG C
Sbjct: 52 GNSSNSSSSCSDEGNDKKRRAVAIQGDTNGALTLNLNGESDGLFPAKKTKSGAVCQVENC 111
Query: 114 QADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
+ADLS + YH RHKVC++HSK + T+ G QRF
Sbjct: 112 EADLSKVKDYHRRHKVCEMHSKATSATVEGILQRF 146
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.324 0.137 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 248,336,917
Number of Sequences: 2790947
Number of extensions: 9039239
Number of successful extensions: 45404
Number of sequences better than 10.0: 220
Number of HSP's better than 10.0 without gapping: 99
Number of HSP's successfully gapped in prelim test: 121
Number of HSP's that attempted gapping in prelim test: 45147
Number of HSP's gapped (non-prelim): 318
length of query: 151
length of database: 848,049,833
effective HSP length: 127
effective length of query: 24
effective length of database: 493,599,564
effective search space: 11846389536
effective search space used: 11846389536
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 67 (30.4 bits)
Medicago: description of AC144726.3


