

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144721.3 - phase: 0 /pseudo
(750 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
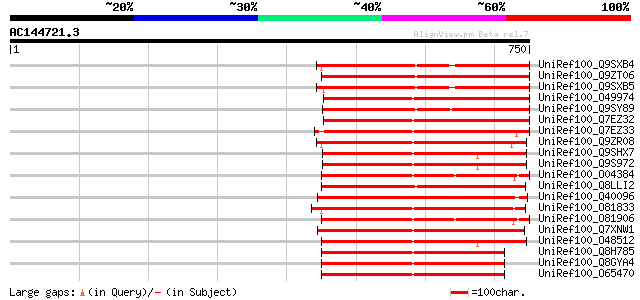
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SXB4 T28P6.6 protein [Arabidopsis thaliana] 382 e-104
UniRef100_Q9ZT06 Receptor-like protein kinase [Arabidopsis thali... 380 e-104
UniRef100_Q9SXB5 T28P6.5 protein [Arabidopsis thaliana] 357 6e-97
UniRef100_O49974 KI domain interacting kinase 1 [Zea mays] 351 4e-95
UniRef100_Q9SY89 T25B24.4 protein [Arabidopsis thaliana] 348 3e-94
UniRef100_Q7EZ32 Putative S-receptor kinase KIK1 [Oryza sativa] 348 4e-94
UniRef100_Q7EZ33 Putative S-receptor kinase KIK1 [Oryza sativa] 339 2e-91
UniRef100_Q9ZR08 Putative receptor kinase [Arabidopsis thaliana] 337 6e-91
UniRef100_Q9SHX7 F1E22.15 [Arabidopsis thaliana] 337 8e-91
UniRef100_Q9S972 ARK2 product/receptor-like serine/threonine pro... 337 8e-91
UniRef100_O04384 Serine/threonine kinase [Brassica oleracea] 337 1e-90
UniRef100_Q8LLI2 S-locus receptor-like kinase RLK11 [Oryza sativa] 335 2e-90
UniRef100_Q40096 Receptor protein kinase [Ipomoea trifida] 335 2e-90
UniRef100_O81833 Putative receptor protein kinase [Arabidopsis t... 335 2e-90
UniRef100_O81906 Serine/threonine kinase-like protein [Arabidops... 335 4e-90
UniRef100_Q7XNW1 OSJNBb0015G09.5 protein [Oryza sativa] 334 7e-90
UniRef100_O48512 Receptor kinase 1 [Brassica campestris] 334 7e-90
UniRef100_Q8H785 Hypothetical protein [Arabidopsis thaliana] 332 2e-89
UniRef100_Q8GYA4 Putative receptor-like protein kinase 4 RLK4 [A... 332 2e-89
UniRef100_O65470 Serine/threonine kinase-like protein [Arabidops... 332 2e-89
>UniRef100_Q9SXB4 T28P6.6 protein [Arabidopsis thaliana]
Length = 820
Score = 382 bits (981), Expect = e-104
Identities = 195/311 (62%), Positives = 237/311 (75%), Gaps = 12/311 (3%)
Query: 444 QGRFWP----GIQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERG 499
QG F P +Q+GQEIAVKRLS+ASGQG+EE +NEVVVISKLQHRNLV+LLGCC+
Sbjct: 517 QGGFGPVYKGKLQEGQEIAVKRLSRASGQGLEELVNEVVVISKLQHRNLVKLLGCCIAGE 576
Query: 500 EKMLVYEFMPNKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLK 559
E+MLVYEFMP KSLD +LFD + K LDW+ R NI+ GI RG++YLHRDSRL+IIHRDLK
Sbjct: 577 ERMLVYEFMPKKSLDYYLFDSRRAKLLDWKTRFNIINGICRGLLYLHRDSRLRIIHRDLK 636
Query: 560 ASNILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVY 619
ASNILLD+ +IPKISDFGLARI G E DEANT+RVVGTYGYM PEYAMGGLFSEKSDV+
Sbjct: 637 ASNILLDENLIPKISDFGLARIFPGNE-DEANTRRVVGTYGYMAPEYAMGGLFSEKSDVF 695
Query: 620 SFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRC 679
S GV+LLEI+SGRRN+ + +L+ + W +W E SL+D E++D FE + +C
Sbjct: 696 SLGVILLEIISGRRNS-------NSTLLAYVWSIWNEGEINSLVDPEIFDLLFEKEIHKC 748
Query: 680 MHIGLLCVQELPKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSNS 739
+HIGLLCVQE +RPS+STV ML SEI +P P + AF+ N ESS+ S +S
Sbjct: 749 IHIGLLCVQEAANDRPSVSTVCSMLSSEIADIPEPKQPAFISRNNVPEAESSENSDLKDS 808
Query: 740 NNNVTLSDVIG 750
NNVT++DV G
Sbjct: 809 INNVTITDVTG 819
>UniRef100_Q9ZT06 Receptor-like protein kinase [Arabidopsis thaliana]
Length = 830
Score = 380 bits (976), Expect = e-104
Identities = 187/300 (62%), Positives = 238/300 (79%), Gaps = 1/300 (0%)
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+Q+G +IAVKRLS+ SGQG+EEF+NEV VISKLQHRNLVRLLG C+E E+MLVYEFMP
Sbjct: 531 LQEGLDIAVKRLSRTSGQGVEEFVNEVFVISKLQHRNLVRLLGFCIEGEERMLVYEFMPE 590
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
LDA+LFDP++++ LDW+ R NI++GI RG+MYLHRDSRLKIIHRDLKASNILLD+ +
Sbjct: 591 NCLDAYLFDPVKQRLLDWKTRFNIIDGICRGLMYLHRDSRLKIIHRDLKASNILLDENLN 650
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVS 630
PKISDFGLARI +G E DE +T RVVGTYGYM PEYAMGGLFSEKSDV+S GV+LLEIVS
Sbjct: 651 PKISDFGLARIFQGNE-DEVSTVRVVGTYGYMAPEYAMGGLFSEKSDVFSLGVILLEIVS 709
Query: 631 GRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQEL 690
GRRN+SFY + + +L +AWKLW I+L+D +++ FE+ + RC+H+GLLCVQ+
Sbjct: 710 GRRNSSFYNDGQNPNLSAYAWKLWNTGEDIALVDPVIFEECFENEIRRCVHVGLLCVQDH 769
Query: 691 PKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSNSNNNVTLSDVIG 750
+RPS++TV+ ML SE ++LP P + AF+ + + ESS QS S NNV+L+ + G
Sbjct: 770 ANDRPSVATVIWMLSSENSNLPEPKQPAFIPRRGTSEVESSGQSDPRASINNVSLTKITG 829
>UniRef100_Q9SXB5 T28P6.5 protein [Arabidopsis thaliana]
Length = 820
Score = 357 bits (917), Expect = 6e-97
Identities = 183/311 (58%), Positives = 226/311 (71%), Gaps = 12/311 (3%)
Query: 444 QGRFWPGIQ----DGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERG 499
QG F P + +GQEIAVKRLS+ASGQG+EE + EVVVISKLQHRNLV+L GCC+
Sbjct: 517 QGGFGPVYKGMLLEGQEIAVKRLSQASGQGLEELVTEVVVISKLQHRNLVKLFGCCIAGE 576
Query: 500 EKMLVYEFMPNKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLK 559
E+MLVYEFMP KSLD ++FDP + K LDW R I+ GI RG++YLHRDSRL+IIHRDLK
Sbjct: 577 ERMLVYEFMPKKSLDFYIFDPREAKLLDWNTRFEIINGICRGLLYLHRDSRLRIIHRDLK 636
Query: 560 ASNILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVY 619
ASNILLD+ +IPKISDFGLARI G E DEANT+RVVGTYGYM PEYAMGGLFSEKSDV+
Sbjct: 637 ASNILLDENLIPKISDFGLARIFPGNE-DEANTRRVVGTYGYMAPEYAMGGLFSEKSDVF 695
Query: 620 SFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRC 679
S GV+LLEI+SGRRN+ +L+ W +W E ++D E++D FE + +C
Sbjct: 696 SLGVILLEIISGRRNS-------HSTLLAHVWSIWNEGEINGMVDPEIFDQLFEKEIRKC 748
Query: 680 MHIGLLCVQELPKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSNS 739
+HI LLCVQ+ +RPS+STV +ML SE+ +P P + AF+ E S+ S
Sbjct: 749 VHIALLCVQDAANDRPSVSTVCMMLSSEVADIPEPKQPAFMPRNVGLEAEFSESIALKAS 808
Query: 740 NNNVTLSDVIG 750
NNVT++DV G
Sbjct: 809 INNVTITDVSG 819
>UniRef100_O49974 KI domain interacting kinase 1 [Zea mays]
Length = 848
Score = 351 bits (901), Expect = 4e-95
Identities = 171/297 (57%), Positives = 228/297 (76%), Gaps = 1/297 (0%)
Query: 454 GQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKSL 513
G+E+AVKRL + SGQG+EEF NEV++I+KLQHRNLVRLLGCC+ R EK+LVYE+MPNKSL
Sbjct: 552 GEEVAVKRLCRNSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIPREEKILVYEYMPNKSL 611
Query: 514 DAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKI 573
DAFLF+P +++ LDW+KR +I+EGIARG++YLHRDSRL+++HRDLKASNILLD +M PKI
Sbjct: 612 DAFLFNPEKQRLLDWKKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLDADMKPKI 671
Query: 574 SDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRR 633
SDFG+AR+ GG+ ++ NT RVVGT+GYM PEYAM G+FS KSDVY FGVL+LEI++G+R
Sbjct: 672 SDFGMARMF-GGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDVYGFGVLILEIITGKR 730
Query: 634 NNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPKE 693
SF+ +EDSL++ G+AW+ W E+N LID + + +LRC+HI LLCVQ+ E
Sbjct: 731 AVSFHCHEDSLNIAGYAWRQWNEDNAAELIDPVIRASCSVRQVLRCIHIALLCVQDHADE 790
Query: 694 RPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSNSNNNVTLSDVIG 750
RP I TV+LML ++ + LP P + + S + RS+S VT++ + G
Sbjct: 791 RPDIPTVILMLSNDSSSLPNPRPPTLMLRGREIESSKSSEKDRSHSIGTVTMTQLHG 847
>UniRef100_Q9SY89 T25B24.4 protein [Arabidopsis thaliana]
Length = 842
Score = 348 bits (894), Expect = 3e-94
Identities = 179/298 (60%), Positives = 223/298 (74%), Gaps = 2/298 (0%)
Query: 453 DGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKS 512
+G+EIAVKRLS S QG+EEF NE+++I+KLQHRNLVRLLGCC+E EKML+YE+MPNKS
Sbjct: 546 EGREIAVKRLSGKSKQGLEEFKNEILLIAKLQHRNLVRLLGCCIEDNEKMLLYEYMPNKS 605
Query: 513 LDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPK 572
LD FLFD ++ LDWRKR ++ GIARG++YLHRDSRLKIIHRDLKASNILLD EM PK
Sbjct: 606 LDRFLFDESKQGSLDWRKRWEVIGGIARGLLYLHRDSRLKIIHRDLKASNILLDTEMNPK 665
Query: 573 ISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGR 632
ISDFG+ARI + D ANT RVVGTYGYM PEYAM G+FSEKSDVYSFGVL+LEIVSGR
Sbjct: 666 ISDFGMARIFNYRQ-DHANTIRVVGTYGYMAPEYAMEGIFSEKSDVYSFGVLILEIVSGR 724
Query: 633 RNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPK 692
+N SF + D SL+G+AW LW + T +ID V D + +RC+H+G+LC Q+
Sbjct: 725 KNVSF-RGTDHGSLIGYAWHLWSQGKTKEMIDPIVKDTRDVTEAMRCIHVGMLCTQDSVI 783
Query: 693 ERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSNSNNNVTLSDVIG 750
RP++ +V+LML S+ + LPPP + F NS E + H S N+VT + ++G
Sbjct: 784 HRPNMGSVLLMLESQTSQLPPPRQPTFHSFLNSGDIELNFDGHDVASVNDVTFTTIVG 841
>UniRef100_Q7EZ32 Putative S-receptor kinase KIK1 [Oryza sativa]
Length = 853
Score = 348 bits (892), Expect = 4e-94
Identities = 170/297 (57%), Positives = 228/297 (76%), Gaps = 1/297 (0%)
Query: 454 GQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKSL 513
G+E+AVKRL + SGQG+EEF NEV++I+KLQHRNLVRLLGCC++ EK+LVYE+MPNKSL
Sbjct: 557 GEEVAVKRLCRKSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIQGEEKILVYEYMPNKSL 616
Query: 514 DAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKI 573
DAFLF+P ++ LDWRKR +I+EGIARG++YLHRDSRL+++HRDLKASNILLD +M PKI
Sbjct: 617 DAFLFNPEKQGLLDWRKRFDIIEGIARGLLYLHRDSRLRVVHRDLKASNILLDKDMNPKI 676
Query: 574 SDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRR 633
SDFG+AR+ GG+ ++ NT RVVGT+GYM PEYAM G+FS KSD+YSFGVL+LEI++G+R
Sbjct: 677 SDFGMARMF-GGDQNQFNTNRVVGTFGYMSPEYAMEGIFSVKSDIYSFGVLMLEIITGKR 735
Query: 634 NNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPKE 693
SF+ +DSL++ GFAW+ W E+ LID + + +LRC+HI LLCVQ+ +E
Sbjct: 736 ALSFHGQQDSLNIAGFAWRQWNEDKGEELIDPLIRASCSLRQVLRCIHIALLCVQDHAQE 795
Query: 694 RPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSNSNNNVTLSDVIG 750
RP I V+LML S+ + LP P + + S T S + +S+S V+++ + G
Sbjct: 796 RPDIPAVILMLSSDSSSLPMPRPPTLMLHGRSAETSKSSEKDQSHSIGTVSMTQLHG 852
>UniRef100_Q7EZ33 Putative S-receptor kinase KIK1 [Oryza sativa]
Length = 865
Score = 339 bits (869), Expect = 2e-91
Identities = 175/313 (55%), Positives = 231/313 (72%), Gaps = 8/313 (2%)
Query: 441 YAWQGRFWPGIQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGE 500
+ ++GR PG G+EIAVKRLS++SGQG+EEF NEV++I+KLQHRNLVRLLGCC++ E
Sbjct: 557 HVYKGRL-PG---GEEIAVKRLSRSSGQGLEEFKNEVILIAKLQHRNLVRLLGCCIQGEE 612
Query: 501 KMLVYEFMPNKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKA 560
K+LVYE+MPNKSLDAFLFDP ++ LDWR R I+EG+ARG++YLHRDSRL+++HRDLKA
Sbjct: 613 KILVYEYMPNKSLDAFLFDPERRGLLDWRTRFQIIEGVARGLLYLHRDSRLRVVHRDLKA 672
Query: 561 SNILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYS 620
SNILLD +M PKISDFG+ARI GG+ ++ NT RVVGT GYM PEYAM GLFS +SDVYS
Sbjct: 673 SNILLDRDMNPKISDFGMARIF-GGDQNQVNTNRVVGTLGYMSPEYAMEGLFSVRSDVYS 731
Query: 621 FGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCM 680
FG+L+LEI++G++N+SF+ E SL++VG+AW+LW + LID + LRC+
Sbjct: 732 FGILILEIITGQKNSSFHHMEGSLNIVGYAWQLWNGDRGQELIDPAIRGTCPAKEALRCV 791
Query: 681 HIGLLCVQELPKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESS---QQSHRS 737
H+ LLCVQ+ +RP I VVL L S+ + LP P F S S+ + S
Sbjct: 792 HMALLCVQDHAHDRPDIPYVVLTLGSDSSVLPTPRPPTFTLQCTSSSSGRDMYYRDKEES 851
Query: 738 NSNNNVTLSDVIG 750
S N++T++ + G
Sbjct: 852 YSANDLTVTMLQG 864
>UniRef100_Q9ZR08 Putative receptor kinase [Arabidopsis thaliana]
Length = 852
Score = 337 bits (865), Expect = 6e-91
Identities = 180/312 (57%), Positives = 231/312 (73%), Gaps = 9/312 (2%)
Query: 444 QGRFWP---GIQDG-QEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERG 499
QG F P G+ G QEIAVKRLS+ SGQG+EEF NEVV+I+KLQHRNLVRLLG CV
Sbjct: 540 QGGFGPVYKGMFPGDQEIAVKRLSRCSGQGLEEFKNEVVLIAKLQHRNLVRLLGYCVAGE 599
Query: 500 EKMLVYEFMPNKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLK 559
EK+L+YE+MP+KSLD F+FD ++LDW+ R NI+ GIARG++YLH+DSRL+IIHRDLK
Sbjct: 600 EKLLLYEYMPHKSLDFFIFDRKLCQRLDWKMRCNIILGIARGLLYLHQDSRLRIIHRDLK 659
Query: 560 ASNILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVY 619
SNILLD+EM PKISDFGLARI GG ANT RVVGTYGYM PEYA+ GLFS KSDV+
Sbjct: 660 TSNILLDEEMNPKISDFGLARIF-GGSETSANTNRVVGTYGYMSPEYALEGLFSFKSDVF 718
Query: 620 SFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRC 679
SFGV+++E +SG+RN F++ E SLSL+G AW LW E I L+D+ + ++ L+C
Sbjct: 719 SFGVVVIETISGKRNTGFHEPEKSLSLLGHAWDLWKAERGIELLDQALQESCETEGFLKC 778
Query: 680 MHIGLLCVQELPKERPSISTVVLML-ISEITHLPPPGKVAFVHNQ---NSRSTESSQQSH 735
+++GLLCVQE P +RP++S VV ML SE LP P + AFV + +S+++ S++
Sbjct: 779 LNVGLLCVQEDPNDRPTMSNVVFMLGSSEAATLPTPKQPAFVLRRCPSSSKASSSTKPET 838
Query: 736 RSNSNNNVTLSD 747
S + +TL D
Sbjct: 839 CSENELTITLED 850
>UniRef100_Q9SHX7 F1E22.15 [Arabidopsis thaliana]
Length = 1662
Score = 337 bits (864), Expect = 8e-91
Identities = 173/300 (57%), Positives = 222/300 (73%), Gaps = 7/300 (2%)
Query: 453 DGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKS 512
DG+EIAVKRLSK S QG +EFMNEV +I+KLQH NLVRLLGCCV++GEKML+YE++ N S
Sbjct: 1359 DGKEIAVKRLSKMSSQGTDEFMNEVRLIAKLQHINLVRLLGCCVDKGEKMLIYEYLENLS 1418
Query: 513 LDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPK 572
LD+ LFD + L+W+KR +I+ GIARG++YLH+DSR +IIHRDLKASN+LLD M PK
Sbjct: 1419 LDSHLFDQTRSSNLNWQKRFDIINGIARGLLYLHQDSRCRIIHRDLKASNVLLDKNMTPK 1478
Query: 573 ISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGR 632
ISDFG+ARI G E EANT+RVVGTYGYM PEYAM G+FS KSDV+SFGVLLLEI+SG+
Sbjct: 1479 ISDFGMARIF-GREETEANTRRVVGTYGYMSPEYAMDGIFSMKSDVFSFGVLLLEIISGK 1537
Query: 633 RNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFES----SMLRCMHIGLLCVQ 688
RN FY + L+L+GF W+ W E + ++D DA +LRC+ IGLLCVQ
Sbjct: 1538 RNKGFYNSNRDLNLLGFVWRHWKEGKELEIVDPINIDALSSEFPTHEILRCIQIGLLCVQ 1597
Query: 689 ELPKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSN--SNNNVTLS 746
E ++RP +S+V++ML SE T +P P + F ++S +SS + R + + N VTLS
Sbjct: 1598 ERAEDRPVMSSVMVMLGSETTAIPQPKRPGFCVGRSSLEVDSSSSTQRDDECTVNQVTLS 1657
Score = 336 bits (861), Expect = 2e-90
Identities = 171/300 (57%), Positives = 222/300 (74%), Gaps = 7/300 (2%)
Query: 453 DGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKS 512
DG+EIAVKRLSK S QG +EFMNEV +I+KLQH NLVRLLGCCV++GEKML+YE++ N S
Sbjct: 540 DGKEIAVKRLSKMSSQGTDEFMNEVRLIAKLQHINLVRLLGCCVDKGEKMLIYEYLENLS 599
Query: 513 LDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPK 572
LD+ LFD + L+W+KR +I+ GIARG++YLH+DSR +IIHRDLKASN+LLD M PK
Sbjct: 600 LDSHLFDQTRSSNLNWQKRFDIINGIARGLLYLHQDSRCRIIHRDLKASNVLLDKNMTPK 659
Query: 573 ISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGR 632
ISDFG+ARI G E EANT+RVVGTYGYM PEYAM G+FS KSDV+SFGVLLLEI+SG+
Sbjct: 660 ISDFGMARIF-GREETEANTRRVVGTYGYMSPEYAMDGIFSMKSDVFSFGVLLLEIISGK 718
Query: 633 RNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFES----SMLRCMHIGLLCVQ 688
RN FY + L+L+GF W+ W E N + ++D D+ +LRC+ IGLLCVQ
Sbjct: 719 RNKGFYNSNRDLNLLGFVWRHWKEGNELEIVDPINIDSLSSKFPTHEILRCIQIGLLCVQ 778
Query: 689 ELPKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSN--SNNNVTLS 746
E ++RP +S+V++ML SE T +P P + F ++ +SS + R + + N +TLS
Sbjct: 779 ERAEDRPVMSSVMVMLGSETTAIPQPKRPGFCIGRSPLEADSSSSTQRDDECTVNQITLS 838
>UniRef100_Q9S972 ARK2 product/receptor-like serine/threonine protein kinase ARK2
[Arabidopsis thaliana]
Length = 850
Score = 337 bits (864), Expect = 8e-91
Identities = 173/300 (57%), Positives = 222/300 (73%), Gaps = 7/300 (2%)
Query: 453 DGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKS 512
DG+EIAVKRLSK S QG +EFMNEV +I+KLQH NLVRLLGCCV++GEKML+YE++ N S
Sbjct: 547 DGKEIAVKRLSKMSSQGTDEFMNEVRLIAKLQHINLVRLLGCCVDKGEKMLIYEYLENLS 606
Query: 513 LDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPK 572
LD+ LFD + L+W+KR +I+ GIARG++YLH+DSR +IIHRDLKASN+LLD M PK
Sbjct: 607 LDSHLFDQTRSSNLNWQKRFDIINGIARGLLYLHQDSRCRIIHRDLKASNVLLDKNMTPK 666
Query: 573 ISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGR 632
ISDFG+ARI G E EANT+RVVGTYGYM PEYAM G+FS KSDV+SFGVLLLEI+SG+
Sbjct: 667 ISDFGMARIF-GREETEANTRRVVGTYGYMSPEYAMDGIFSMKSDVFSFGVLLLEIISGK 725
Query: 633 RNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFES----SMLRCMHIGLLCVQ 688
RN FY + L+L+GF W+ W E + ++D DA +LRC+ IGLLCVQ
Sbjct: 726 RNKGFYNSNRDLNLLGFVWRHWKEGKELEIVDPINIDALSSEFPTHEILRCIQIGLLCVQ 785
Query: 689 ELPKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSN--SNNNVTLS 746
E ++RP +S+V++ML SE T +P P + F ++S +SS + R + + N VTLS
Sbjct: 786 ERAEDRPVMSSVMVMLGSETTAIPQPKRPGFCVGRSSLEVDSSSSTQRDDECTVNQVTLS 845
>UniRef100_O04384 Serine/threonine kinase [Brassica oleracea]
Length = 850
Score = 337 bits (863), Expect = 1e-90
Identities = 176/306 (57%), Positives = 226/306 (73%), Gaps = 10/306 (3%)
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
++DGQEIAVKRLS SGQG++EF NE+++I+KLQHRNLVRLLGCC E EKMLVYE+MPN
Sbjct: 548 LEDGQEIAVKRLSGKSGQGVDEFKNEIILIAKLQHRNLVRLLGCCFEGEEKMLVYEYMPN 607
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
KSLD F+FD ++++ +DW+ R I+EGIARG++YLHRDSRL+IIHRDLK SN+LLD EM
Sbjct: 608 KSLDFFIFDEMKQELVDWKLRFAIIEGIARGLLYLHRDSRLRIIHRDLKVSNVLLDGEMN 667
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVS 630
PKISDFG+ARI GG +EANT RVVGTYGYM PEYAM GLFS KSDVYSFGVLLLEI+S
Sbjct: 668 PKISDFGMARIF-GGNQNEANTVRVVGTYGYMSPEYAMEGLFSVKSDVYSFGVLLLEIIS 726
Query: 631 GRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQEL 690
G+RN S +E SL+G+AW L+ + L+D ++ + LRC+H+ +LCVQ+
Sbjct: 727 GKRNTSLRASEHG-SLIGYAWFLYTHGRSEELVDPKIRATCNKREALRCIHVAMLCVQDS 785
Query: 691 PKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRS------TESSQQSHRSNSNNNVT 744
ERP+++ V+LML S+ LP P + F + S +SSQQ S+N +T
Sbjct: 786 AAERPNMAAVLLMLESDTATLPVPRQPTFTTSTRRNSMDVNFALDSSQQ--YIVSSNEIT 843
Query: 745 LSDVIG 750
+ V+G
Sbjct: 844 STVVLG 849
>UniRef100_Q8LLI2 S-locus receptor-like kinase RLK11 [Oryza sativa]
Length = 820
Score = 335 bits (860), Expect = 2e-90
Identities = 169/296 (57%), Positives = 219/296 (73%), Gaps = 2/296 (0%)
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
++ G E+AVKRL++ SGQGIEEF NEVV+I+KLQHRNLVRLLGCC+ EK+L+YE++PN
Sbjct: 523 LEGGTEVAVKRLNEGSGQGIEEFRNEVVLIAKLQHRNLVRLLGCCIHEDEKLLIYEYLPN 582
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
KSLDAFLFD +K LDW R I++GIA+G++YLH+DSRL IIHRDLKASNILLD EM
Sbjct: 583 KSLDAFLFDATRKYVLDWPTRFKIIKGIAKGLLYLHQDSRLTIIHRDLKASNILLDTEMN 642
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVS 630
PKISDFG+ARI G + +ANT RVVGTYGYM PEY +GG FS KSD YSFGVLLLEIVS
Sbjct: 643 PKISDFGIARIFHGNQ-QQANTTRVVGTYGYMSPEYVLGGAFSVKSDTYSFGVLLLEIVS 701
Query: 631 GRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQEL 690
G + +S + SL +AW+LW + N L+D+ D+ RC+H+GLLCVQ+
Sbjct: 702 GLKISSSKLTPNFFSLTAYAWRLWKDGNATELLDKFFVDSYPLHEAFRCIHVGLLCVQDH 761
Query: 691 PKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQS-HRSNSNNNVTL 745
P +RPS+S+VV ML +E T LP P + + +N + E++++S + N+ + TL
Sbjct: 762 PNDRPSMSSVVFMLENESTLLPAPKQPVYFEMKNHGTQEATEESVYSVNTMSTTTL 817
>UniRef100_Q40096 Receptor protein kinase [Ipomoea trifida]
Length = 853
Score = 335 bits (860), Expect = 2e-90
Identities = 170/304 (55%), Positives = 221/304 (71%), Gaps = 4/304 (1%)
Query: 445 GRFWPGIQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLV 504
G + G+ +G+EIAVKRLSK SGQG+EEF NE+ +I++LQHRNLVRLLGCCV+ EK+L+
Sbjct: 546 GCVYKGMVEGEEIAVKRLSKNSGQGVEEFKNELRLIARLQHRNLVRLLGCCVDMEEKILI 605
Query: 505 YEFMPNKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNIL 564
YE+M NKSLD+ LF+ + L+W+ R NI+ GIARG++YLH+DSR +IIHRDLKASNIL
Sbjct: 606 YEYMENKSLDSTLFNKQRSSLLNWQTRFNIICGIARGLLYLHQDSRFRIIHRDLKASNIL 665
Query: 565 LDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVL 624
LD EM PKISDFG+ARI G E D NTKRVVGTYGYM PEYAM GLFS KSDV+SFGVL
Sbjct: 666 LDKEMNPKISDFGMARIFGGDETDANNTKRVVGTYGYMSPEYAMDGLFSVKSDVFSFGVL 725
Query: 625 LLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGL 684
+LEIV+G++N FY + +L+G AW+LW E L+D + ++ ++RC+ +GL
Sbjct: 726 VLEIVTGKKNRGFYNQNNQQNLLGHAWRLWRERRGSELLDSAIGESYSLCEVMRCIQVGL 785
Query: 685 LCVQELPKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSNSNNNVT 744
LCVQE ++RP+++TVVLML SE LP P F +SS SN + + T
Sbjct: 786 LCVQEQAEDRPNMATVVLMLGSESATLPQPKHPGFCLGSRPADMDSS----TSNCDESCT 841
Query: 745 LSDV 748
++ V
Sbjct: 842 VNQV 845
>UniRef100_O81833 Putative receptor protein kinase [Arabidopsis thaliana]
Length = 815
Score = 335 bits (860), Expect = 2e-90
Identities = 172/315 (54%), Positives = 234/315 (73%), Gaps = 8/315 (2%)
Query: 436 FSF**YAWQGRFWP----GIQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRL 491
FS+ + +G F P ++DGQEIAVKRLS SGQG+EEF NEV +I+KLQHRNLVRL
Sbjct: 500 FSYVNFLGRGGFGPVYKGKLEDGQEIAVKRLSANSGQGVEEFKNEVKLIAKLQHRNLVRL 559
Query: 492 LGCCVERGEKMLVYEFMPNKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRL 551
LGCC++ E ML+YE+MPNKSLD F+FD + +LDW+KR NI+ G+ARGI+YLH+DSRL
Sbjct: 560 LGCCIQGEECMLIYEYMPNKSLDFFIFDERRSTELDWKKRMNIINGVARGILYLHQDSRL 619
Query: 552 KIIHRDLKASNILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGL 611
+IIHRDLKA N+LLD++M PKISDFGLA+ GG+ E++T RVVGTYGYMPPEYA+ G
Sbjct: 620 RIIHRDLKAGNVLLDNDMNPKISDFGLAKSF-GGDQSESSTNRVVGTYGYMPPEYAIDGH 678
Query: 612 FSEKSDVYSFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDRE-VWDA 670
FS KSDV+SFGVL+LEI++G+ N F + L+L+G WK+W+E+ I + + E + +
Sbjct: 679 FSVKSDVFSFGVLVLEIITGKTNRGFRHADHDLNLLGHVWKMWVEDREIEVPEEEWLEET 738
Query: 671 SFESSMLRCMHIGLLCVQELPKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTES 730
S +LRC+H+ LLCVQ+ P++RP++++VVLM S+ + LP P + F N+N S
Sbjct: 739 SVIPEVLRCIHVALLCVQQKPEDRPTMASVVLMFGSD-SSLPHPTQPGFFTNRNVPDI-S 796
Query: 731 SQQSHRSNSNNNVTL 745
S S RS + ++T+
Sbjct: 797 SSLSLRSQNEVSITM 811
>UniRef100_O81906 Serine/threonine kinase-like protein [Arabidopsis thaliana]
Length = 849
Score = 335 bits (858), Expect = 4e-90
Identities = 175/305 (57%), Positives = 226/305 (73%), Gaps = 9/305 (2%)
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
++DG+EIAVKRLS SGQG++EF NE+++I+KLQHRNLVRLLGCC E EKMLVYE+MPN
Sbjct: 548 LEDGREIAVKRLSGKSGQGVDEFKNEIILIAKLQHRNLVRLLGCCFEGEEKMLVYEYMPN 607
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
KSLD FLFD ++ +DW+ R +I+EGIARG++YLHRDSRL+IIHRDLK SN+LLD EM
Sbjct: 608 KSLDFFLFDETKQALIDWKLRFSIIEGIARGLLYLHRDSRLRIIHRDLKVSNVLLDAEMN 667
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVS 630
PKISDFG+ARI GG +EANT RVVGTYGYM PEYAM GLFS KSDVYSFGVLLLEIVS
Sbjct: 668 PKISDFGMARIF-GGNQNEANTVRVVGTYGYMSPEYAMEGLFSVKSDVYSFGVLLLEIVS 726
Query: 631 GRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQEL 690
G+RN S +E SL+G+AW L+ + L+D ++ + LRC+H+ +LCVQ+
Sbjct: 727 GKRNTSLRSSEHG-SLIGYAWYLYTHGRSEELVDPKIRVTCSKREALRCIHVAMLCVQDS 785
Query: 691 PKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSR-----STESSQQSHRSNSNNNVTL 745
ERP++++V+LML S+ L P + F + + + +SSQQ S+N +T
Sbjct: 786 AAERPNMASVLLMLESDTATLAAPRQPTFTSTRRNSIDVNFALDSSQQ--YIVSSNEITS 843
Query: 746 SDVIG 750
+ V+G
Sbjct: 844 TVVLG 848
>UniRef100_Q7XNW1 OSJNBb0015G09.5 protein [Oryza sativa]
Length = 846
Score = 334 bits (856), Expect = 7e-90
Identities = 164/301 (54%), Positives = 219/301 (72%), Gaps = 2/301 (0%)
Query: 445 GRFWPGIQDGQ-EIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKML 503
G+ + G+ +G E+AVKRLSK SGQG+EEF NEVV+I+KLQHRNLVRLLGCC+ EK+L
Sbjct: 540 GKVYKGVLEGGIEVAVKRLSKGSGQGVEEFRNEVVLIAKLQHRNLVRLLGCCIHEDEKLL 599
Query: 504 VYEFMPNKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNI 563
+YE++PN+SLDAFLFD +K LDW R I++G+ARG++YLH+DSRL IIHRDLK SNI
Sbjct: 600 IYEYLPNRSLDAFLFDANRKNTLDWPTRFKIIKGVARGLLYLHQDSRLTIIHRDLKTSNI 659
Query: 564 LLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGV 623
LLD EM PKISDFG+ARI GG +ANT RVVGTYGYM PEYA+ G FS KSD YSFGV
Sbjct: 660 LLDTEMSPKISDFGMARIF-GGNEQQANTTRVVGTYGYMSPEYALDGYFSVKSDTYSFGV 718
Query: 624 LLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIG 683
+LLE+VSG + +S + D +L+ +AW LW + N +D + ++ +LRC+H+G
Sbjct: 719 ILLEVVSGLKISSAHLKVDCSNLIAYAWSLWKDGNARDFVDSSIVESCPLHEVLRCIHLG 778
Query: 684 LLCVQELPKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSNSNNNV 743
LLC+Q+ P RP +S++V ML +E LP P + + + + E ++ S RS S N++
Sbjct: 779 LLCIQDQPSARPLMSSIVFMLENETAVLPAPKEPIYFTRREYGTDEDTRDSMRSRSLNHM 838
Query: 744 T 744
+
Sbjct: 839 S 839
>UniRef100_O48512 Receptor kinase 1 [Brassica campestris]
Length = 847
Score = 334 bits (856), Expect = 7e-90
Identities = 171/302 (56%), Positives = 222/302 (72%), Gaps = 7/302 (2%)
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ DG+EIAVKRLSK S QG +EF NEV +I++LQH NLVRLLGCCV++GEKML+YE++ N
Sbjct: 542 LPDGKEIAVKRLSKMSLQGTDEFKNEVRLIARLQHINLVRLLGCCVDKGEKMLIYEYLEN 601
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
SLD+ LFD I++ L W KR +I GIARG++YLH+DSR +IIHRDLKASN+LLD M
Sbjct: 602 LSLDSHLFDKIRRSNLSWPKRFDITNGIARGLLYLHQDSRFRIIHRDLKASNVLLDKNMT 661
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVS 630
PKISDFG+ARI G E EANT++VVGTYGYM PEYAM G+FS KSDV+SFGVLLLEI++
Sbjct: 662 PKISDFGMARIF-GREETEANTRKVVGTYGYMAPEYAMDGIFSMKSDVFSFGVLLLEIIT 720
Query: 631 GRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFES----SMLRCMHIGLLC 686
G+R+ FY + +L+GF W+ W E I ++D + D+S + +LRC+ IGLLC
Sbjct: 721 GKRSKGFYNSNRDNNLLGFVWRYWKEGKGIEIVDPIIMDSSLSALCTHEILRCIQIGLLC 780
Query: 687 VQELPKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSN--SNNNVT 744
VQE ++RP +STV++ML SE T +P P F ++ TESS + R + S N +T
Sbjct: 781 VQERAEDRPVMSTVMVMLGSETTAIPQPKPPGFCVGRSLFETESSSSTQRDDELSVNQIT 840
Query: 745 LS 746
LS
Sbjct: 841 LS 842
>UniRef100_Q8H785 Hypothetical protein [Arabidopsis thaliana]
Length = 658
Score = 332 bits (852), Expect = 2e-89
Identities = 163/264 (61%), Positives = 207/264 (77%), Gaps = 1/264 (0%)
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ DG E+AVKRLSK+SGQG EF NEVV+++KLQHRNLVRLLG C++ E++LVYE++PN
Sbjct: 356 LSDGTEVAVKRLSKSSGQGEVEFKNEVVLVAKLQHRNLVRLLGFCLDGEERVLVYEYVPN 415
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
KSLD FLFDP +K +LDW +R I+ G+ARGI+YLH+DSRL IIHRDLKASNILLD +M
Sbjct: 416 KSLDYFLFDPAKKGQLDWTRRYKIIGGVARGILYLHQDSRLTIIHRDLKASNILLDADMN 475
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVS 630
PKI+DFG+ARI G + E NT R+VGTYGYM PEYAM G +S KSDVYSFGVL+LEI+S
Sbjct: 476 PKIADFGMARIF-GLDQTEENTSRIVGTYGYMSPEYAMHGQYSMKSDVYSFGVLVLEIIS 534
Query: 631 GRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQEL 690
G++N+SFYQ + + LV +AW LW + L+D + + + ++RC+HIGLLCVQE
Sbjct: 535 GKKNSSFYQTDGAHDLVSYAWGLWSNGRPLELVDPAIVENCQRNEVVRCVHIGLLCVQED 594
Query: 691 PKERPSISTVVLMLISEITHLPPP 714
P ERP++ST+VLML S LP P
Sbjct: 595 PAERPTLSTIVLMLTSNTVTLPVP 618
>UniRef100_Q8GYA4 Putative receptor-like protein kinase 4 RLK4 [Arabidopsis thaliana]
Length = 669
Score = 332 bits (852), Expect = 2e-89
Identities = 163/264 (61%), Positives = 207/264 (77%), Gaps = 1/264 (0%)
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ DG E+AVKRLSK+SGQG EF NEVV+++KLQHRNLVRLLG C++ E++LVYE++PN
Sbjct: 367 LSDGTEVAVKRLSKSSGQGEVEFKNEVVLVAKLQHRNLVRLLGFCLDGEERVLVYEYVPN 426
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
KSLD FLFDP +K +LDW +R I+ G+ARGI+YLH+DSRL IIHRDLKASNILLD +M
Sbjct: 427 KSLDYFLFDPAKKGQLDWTRRYKIIGGVARGILYLHQDSRLTIIHRDLKASNILLDADMN 486
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVS 630
PKI+DFG+ARI G + E NT R+VGTYGYM PEYAM G +S KSDVYSFGVL+LEI+S
Sbjct: 487 PKIADFGMARIF-GLDQTEENTSRIVGTYGYMSPEYAMHGQYSMKSDVYSFGVLVLEIIS 545
Query: 631 GRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQEL 690
G++N+SFYQ + + LV +AW LW + L+D + + + ++RC+HIGLLCVQE
Sbjct: 546 GKKNSSFYQTDGAHDLVSYAWGLWSNGRPLELVDPAIVENCQRNEVVRCVHIGLLCVQED 605
Query: 691 PKERPSISTVVLMLISEITHLPPP 714
P ERP++ST+VLML S LP P
Sbjct: 606 PAERPTLSTIVLMLTSNTVTLPVP 629
>UniRef100_O65470 Serine/threonine kinase-like protein [Arabidopsis thaliana]
Length = 633
Score = 332 bits (852), Expect = 2e-89
Identities = 163/264 (61%), Positives = 207/264 (77%), Gaps = 1/264 (0%)
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ DG E+AVKRLSK+SGQG EF NEVV+++KLQHRNLVRLLG C++ E++LVYE++PN
Sbjct: 331 LSDGTEVAVKRLSKSSGQGEVEFKNEVVLVAKLQHRNLVRLLGFCLDGEERVLVYEYVPN 390
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
KSLD FLFDP +K +LDW +R I+ G+ARGI+YLH+DSRL IIHRDLKASNILLD +M
Sbjct: 391 KSLDYFLFDPAKKGQLDWTRRYKIIGGVARGILYLHQDSRLTIIHRDLKASNILLDADMN 450
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVS 630
PKI+DFG+ARI G + E NT R+VGTYGYM PEYAM G +S KSDVYSFGVL+LEI+S
Sbjct: 451 PKIADFGMARIF-GLDQTEENTSRIVGTYGYMSPEYAMHGQYSMKSDVYSFGVLVLEIIS 509
Query: 631 GRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQEL 690
G++N+SFYQ + + LV +AW LW + L+D + + + ++RC+HIGLLCVQE
Sbjct: 510 GKKNSSFYQTDGAHDLVSYAWGLWSNGRPLELVDPAIVENCQRNEVVRCVHIGLLCVQED 569
Query: 691 PKERPSISTVVLMLISEITHLPPP 714
P ERP++ST+VLML S LP P
Sbjct: 570 PAERPTLSTIVLMLTSNTVTLPVP 593
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.358 0.159 0.601
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,079,872,591
Number of Sequences: 2790947
Number of extensions: 39915942
Number of successful extensions: 324859
Number of sequences better than 10.0: 17610
Number of HSP's better than 10.0 without gapping: 9183
Number of HSP's successfully gapped in prelim test: 8428
Number of HSP's that attempted gapping in prelim test: 284924
Number of HSP's gapped (non-prelim): 22755
length of query: 750
length of database: 848,049,833
effective HSP length: 135
effective length of query: 615
effective length of database: 471,271,988
effective search space: 289832272620
effective search space used: 289832272620
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 79 (35.0 bits)
Medicago: description of AC144721.3


