

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144656.5 + phase: 0 /pseudo
(2301 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
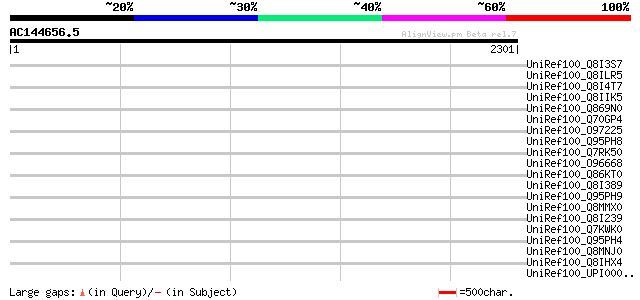
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8I3S7 Hypothetical protein PFE0905w [Plasmodium falci... 44 0.045
UniRef100_Q8ILR5 Hypothetical protein [Plasmodium falciparum] 42 0.22
UniRef100_Q8I4T7 Hypothetical protein [Plasmodium falciparum] 42 0.22
UniRef100_Q8IIK5 Hypothetical protein [Plasmodium falciparum] 41 0.38
UniRef100_Q869N0 Similar to Dictyostelium discoideum (Slime mold... 41 0.38
UniRef100_Q70GP4 Dd-STATb protein [Dictyostelium discoideum] 40 0.65
UniRef100_O97225 Hypothetical protein MAL3P2.2 [Plasmodium falci... 40 0.85
UniRef100_Q95PH8 Histidine kinase DhkH [Dictyostelium discoideum] 40 1.1
UniRef100_Q7RK50 Hypothetical protein [Plasmodium yoelii yoelii] 40 1.1
UniRef100_O96668 Cell surface protein DTFA [Dictyostelium discoi... 39 1.4
UniRef100_Q86KT0 Hypothetical protein [Dictyostelium discoideum] 39 1.9
UniRef100_Q8I389 Hypothetical protein PFI0295c [Plasmodium falci... 39 1.9
UniRef100_Q95PH9 Histidine kinase DhkG [Dictyostelium discoideum] 39 2.5
UniRef100_Q8MMX0 Similar to Dictyostelium discoideum (Slime mold... 39 2.5
UniRef100_Q8I239 Phosphatidylinositol-4-phosphate 5-kinase, puta... 39 2.5
UniRef100_Q7KWK0 Similar to Dictyostelium discoideum (Slime mold... 39 2.5
UniRef100_Q95PH4 Histidine kinase DhkM [Dictyostelium discoideum] 38 3.2
UniRef100_Q8MNJ0 Hypothetical protein [Dictyostelium discoideum] 38 3.2
UniRef100_Q8IHX4 Hypothetical protein [Plasmodium falciparum] 38 3.2
UniRef100_UPI000046C800 UPI000046C800 UniRef100 entry 38 4.2
>UniRef100_Q8I3S7 Hypothetical protein PFE0905w [Plasmodium falciparum]
Length = 1379
Score = 44.3 bits (103), Expect = 0.045
Identities = 22/68 (32%), Positives = 36/68 (52%), Gaps = 6/68 (8%)
Query: 384 NKLIQSTNISLPSHLLPMLFNHQIVSLDSHNILNKILNNNIPNQHTSNAHINNNPINNNS 443
N +I T + +H ++ +S NI+N + NN+ N H +N HINNN INNN
Sbjct: 1023 NNIIDDTMYNEKNH------DNNNMSCQDMNIVNYNIKNNLNNNHINNNHINNNDINNNH 1076
Query: 444 LDHKKCKL 451
+++ +
Sbjct: 1077 INNNNLNI 1084
>UniRef100_Q8ILR5 Hypothetical protein [Plasmodium falciparum]
Length = 3597
Score = 42.0 bits (97), Expect = 0.22
Identities = 24/64 (37%), Positives = 37/64 (57%), Gaps = 6/64 (9%)
Query: 399 LPMLFNHQIVSLDSHNILNK------ILNNNIPNQHTSNAHINNNPINNNSLDHKKCKLT 452
L +L N+ I ++ ++N+ N I NNNI N + +N +INNN INNN+ K K
Sbjct: 2391 LNILINNDINNMHNNNVNNNYVNNNNINNNNINNNNINNNNINNNNINNNNFSFGKKKFV 2450
Query: 453 QSLL 456
++L
Sbjct: 2451 ITVL 2454
>UniRef100_Q8I4T7 Hypothetical protein [Plasmodium falciparum]
Length = 788
Score = 42.0 bits (97), Expect = 0.22
Identities = 26/70 (37%), Positives = 40/70 (57%), Gaps = 13/70 (18%)
Query: 383 PNKLIQSTNISLPSHLLPMLFNHQIVSLDSHNILNKILNNN------IPNQHTSNAHINN 436
PN + NI +H+ N+ I LD+++I N IL+NN + N H +N H+NN
Sbjct: 161 PNNNFVNNNILDNNHI-----NNNI--LDNNHINNNILDNNHINNNHLNNNHLNNNHLNN 213
Query: 437 NPINNNSLDH 446
N +NNN L++
Sbjct: 214 NHLNNNHLNN 223
Score = 39.3 bits (90), Expect = 1.4
Identities = 17/39 (43%), Positives = 29/39 (73%), Gaps = 1/39 (2%)
Query: 410 LDSHNILNKILNNN-IPNQHTSNAHINNNPINNNSLDHK 447
L+++++ N LNNN + N H +N H+NNN +NN+ LD++
Sbjct: 206 LNNNHLNNNHLNNNHLNNNHFNNNHLNNNQLNNHLLDNR 244
>UniRef100_Q8IIK5 Hypothetical protein [Plasmodium falciparum]
Length = 2966
Score = 41.2 bits (95), Expect = 0.38
Identities = 32/90 (35%), Positives = 41/90 (45%), Gaps = 9/90 (10%)
Query: 411 DSHNILNKILNNNI-PNQHTSNAHINNNPINNNSLDHKKCKLTQSLLLTQSYSQDYSERI 469
D NI LN+N+ N H + HI N I N S DHKK K S ++ + S+ YSE
Sbjct: 476 DKENINENPLNDNVYKNYHHFDQHILNYNIYNTSNDHKKNKAVPSHIINRFRSERYSENS 535
Query: 470 *FKPSLLLQYQ--------KECQHGTDLIR 491
K YQ K C+ T L+R
Sbjct: 536 TMKYEGKNYYQSFMSQSALKRCRSYTILLR 565
>UniRef100_Q869N0 Similar to Dictyostelium discoideum (Slime mold). Nucleotide
exchange factor RasGEF N [Dictyostelium discoideum]
Length = 1672
Score = 41.2 bits (95), Expect = 0.38
Identities = 25/102 (24%), Positives = 54/102 (52%), Gaps = 4/102 (3%)
Query: 343 KESEKDV**KMELLLVEPRNLGTTSQERKNTMLMW*PMEEPNKLIQSTNISLPSHLLPML 402
K+++ + + + PRN+ ++ +T+ P++ P +I S+ + S + P+
Sbjct: 13 KDAKNKIEEPQQSTIASPRNICIST----STLSPPPPLQLPPTVIPSSTSTTNSMISPLY 68
Query: 403 FNHQIVSLDSHNILNKILNNNIPNQHTSNAHINNNPINNNSL 444
N +++ +++N N NNN N + +N + NNN NNN++
Sbjct: 69 TNSSLINNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNI 110
>UniRef100_Q70GP4 Dd-STATb protein [Dictyostelium discoideum]
Length = 1148
Score = 40.4 bits (93), Expect = 0.65
Identities = 20/64 (31%), Positives = 35/64 (54%)
Query: 380 MEEPNKLIQSTNISLPSHLLPMLFNHQIVSLDSHNILNKILNNNIPNQHTSNAHINNNPI 439
+E+ N +++ N+ L L M + + + + N N +NNN N + +N + NNN I
Sbjct: 233 VEDINAKLENENLQLKKELFEMSRKFKEIDIINLNNTNNNINNNNNNNNNNNNNNNNNSI 292
Query: 440 NNNS 443
NNN+
Sbjct: 293 NNNN 296
>UniRef100_O97225 Hypothetical protein MAL3P2.2 [Plasmodium falciparum]
Length = 2226
Score = 40.0 bits (92), Expect = 0.85
Identities = 30/82 (36%), Positives = 44/82 (53%), Gaps = 8/82 (9%)
Query: 385 KLIQSTNISLPSHLLPMLF--NHQIVSLDSH-NILNKILNNNIP--NQHTSNAHINNNPI 439
KLI ++I H+L F N QI ++ +H N+ + +LN N N H N + NNN
Sbjct: 1771 KLISISHI-FTRHILKFNFDKNEQIENITNHGNVKSNVLNPNTTHINNHPHNQYHNNNKY 1829
Query: 440 NNNSLDHKKCKLTQSLLLTQSY 461
NN + HK K+ Q LL ++Y
Sbjct: 1830 NNKNNYHK--KIIQELLRDKAY 1849
>UniRef100_Q95PH8 Histidine kinase DhkH [Dictyostelium discoideum]
Length = 1147
Score = 39.7 bits (91), Expect = 1.1
Identities = 31/102 (30%), Positives = 47/102 (45%), Gaps = 6/102 (5%)
Query: 385 KLIQSTNISLPSHLLPMLFNHQIVSLDSHNILNKILNNNIPNQHTSNAHINNNPINNNSL 444
K ++ +N H P L + + SLD +N N NNNI N + N + NNN NNN+
Sbjct: 891 KKLKKSNSDQSIHFSPSLTSSSLPSLDLNNNNNINNNNNINNNNNINNNNNNNNNNNNNN 950
Query: 445 DHKKCKL----TQSLLLTQSYSQDYSERI*FKPSLLLQYQKE 482
++ L + S+L + Q + LL YQK+
Sbjct: 951 NNNNNNLNHYNSDSILSSDLSPQQHQYH--HPNPLLANYQKK 990
>UniRef100_Q7RK50 Hypothetical protein [Plasmodium yoelii yoelii]
Length = 2522
Score = 39.7 bits (91), Expect = 1.1
Identities = 27/87 (31%), Positives = 49/87 (56%), Gaps = 13/87 (14%)
Query: 391 NISLPSHLLPMLFNHQIVSLDSH------NILNKILNNNIPNQHTSNAHINNNP---INN 441
NI++ ++ P ++QI+ L+++ N +N I NNNIP ++T N + N++P INN
Sbjct: 917 NINM-NNFSPYKKDNQIIQLNNYSKYEYNNFINNIKNNNIPYENTINNNFNDSPIDIINN 975
Query: 442 NSLDHKKCKLTQSL---LLTQSYSQDY 465
N L + Q++ +LT + +Y
Sbjct: 976 NHLYNISINSNQNVNNSILTDGHINNY 1002
>UniRef100_O96668 Cell surface protein DTFA [Dictyostelium discoideum]
Length = 1402
Score = 39.3 bits (90), Expect = 1.4
Identities = 28/104 (26%), Positives = 49/104 (46%), Gaps = 19/104 (18%)
Query: 384 NKLIQSTNISLPSHLLPMLFNHQIVSLDSHNILNKILNNNIPNQHTSNAHINNNP-INNN 442
NK +++ + + L FN+ I + +++NI N NNN N + +N + NNN ++NN
Sbjct: 687 NKNLKNQIFNYSNLLNSFTFNNNINNNNNNNINNNNNNNNNNNNNNNNNNNNNNSNLSNN 746
Query: 443 SLDHKK------------------CKLTQSLLLTQSYSQDYSER 468
++++ K CKL + L Y Q Y +R
Sbjct: 747 TIEYSKIKTKTVFNQFTIKQLINSCKLIVTFDLKNQYQQQYYKR 790
>UniRef100_Q86KT0 Hypothetical protein [Dictyostelium discoideum]
Length = 1173
Score = 38.9 bits (89), Expect = 1.9
Identities = 19/38 (50%), Positives = 29/38 (76%), Gaps = 1/38 (2%)
Query: 409 SLDSHNILNK-ILNNNIPNQHTSNAHINNNPINNNSLD 445
+++S+NI N I NNNI N + +N +INNN INNN+++
Sbjct: 453 NMNSNNINNNSINNNNINNNNINNNNINNNNINNNNIN 490
Score = 36.6 bits (83), Expect = 9.4
Identities = 17/39 (43%), Positives = 30/39 (76%), Gaps = 1/39 (2%)
Query: 409 SLDSHNI-LNKILNNNIPNQHTSNAHINNNPINNNSLDH 446
+++S+N+ N I NN+I N + +N +INNN INNN++++
Sbjct: 448 NMNSNNMNSNNINNNSINNNNINNNNINNNNINNNNINN 486
>UniRef100_Q8I389 Hypothetical protein PFI0295c [Plasmodium falciparum]
Length = 1807
Score = 38.9 bits (89), Expect = 1.9
Identities = 18/36 (50%), Positives = 25/36 (69%)
Query: 411 DSHNILNKILNNNIPNQHTSNAHINNNPINNNSLDH 446
D++N N I NN+I N +N INNN INNNS+++
Sbjct: 134 DNNNDNNSINNNSINNNSINNNSINNNSINNNSINN 169
Score = 38.9 bits (89), Expect = 1.9
Identities = 20/48 (41%), Positives = 31/48 (63%), Gaps = 1/48 (2%)
Query: 404 NHQIVSLDSHNILNKILNNN-IPNQHTSNAHINNNPINNNSLDHKKCK 450
N+ S+++++I N +NNN I N +N INNN INNNS+++ K
Sbjct: 136 NNDNNSINNNSINNNSINNNSINNNSINNNSINNNSINNNSINNNNIK 183
>UniRef100_Q95PH9 Histidine kinase DhkG [Dictyostelium discoideum]
Length = 3432
Score = 38.5 bits (88), Expect = 2.5
Identities = 24/65 (36%), Positives = 32/65 (48%), Gaps = 2/65 (3%)
Query: 379 PMEEPNKLIQSTNISLPSHLLPMLFNHQIVSLDSHNILNKILNNNIPNQHTSNAHINNNP 438
P PN Q+ +S +L L N + S H N I NNNI N + +N + NNN
Sbjct: 2924 PRPYPNN--QNNFLSSSPQMLSPLPNSPLSSSQQHQNNNNINNNNINNNNNNNNNNNNNN 2981
Query: 439 INNNS 443
NNN+
Sbjct: 2982 NNNNN 2986
>UniRef100_Q8MMX0 Similar to Dictyostelium discoideum (Slime mold). Protein tyrosine
kinase [Dictyostelium discoideum]
Length = 2165
Score = 38.5 bits (88), Expect = 2.5
Identities = 29/98 (29%), Positives = 49/98 (49%), Gaps = 9/98 (9%)
Query: 346 EKDV**KMELLLVEPRNLGTTSQERKNTMLMW*PMEEPNKLIQSTNISLPSHLLPMLFNH 405
+KDV K ++ E N+ +T+ NT P+ P+ + Q +NI+ + N+
Sbjct: 1135 QKDV--KSKINFYEHVNISSTNLNNVNTTSNT-PITNPSNVNQPSNITTATTATTTSTNN 1191
Query: 406 QIVSLDSHNILNKILNNNIPNQHTSNAHINNNPINNNS 443
++N+ N I NNN N + +N + NNN NNN+
Sbjct: 1192 ------NNNVNNSINNNNNNNNNNNNNNNNNNNNNNNN 1223
>UniRef100_Q8I239 Phosphatidylinositol-4-phosphate 5-kinase, putative [Plasmodium
falciparum]
Length = 1710
Score = 38.5 bits (88), Expect = 2.5
Identities = 23/77 (29%), Positives = 39/77 (49%), Gaps = 11/77 (14%)
Query: 404 NHQIVSLDS---HNILNKILNNNIPNQHTSNAHINNN--------PINNNSLDHKKCKLT 452
NH ++ D+ HN +N NNN N H +N+H NNN P+N +++ K
Sbjct: 199 NHNNINGDNNNNHNNINGDNNNNHNNSHNNNSHNNNNKAENSLGQPLNEKNINDPINKHR 258
Query: 453 QSLLLTQSYSQDYSERI 469
S + + + +Y+E+I
Sbjct: 259 NSQSIIYNINDEYNEKI 275
>UniRef100_Q7KWK0 Similar to Dictyostelium discoideum (Slime mold). LysA
[Dictyostelium discoideum]
Length = 1405
Score = 38.5 bits (88), Expect = 2.5
Identities = 33/101 (32%), Positives = 48/101 (46%), Gaps = 14/101 (13%)
Query: 383 PNKLIQSTNISLPSH----LLPMLFNHQIVSLDSHNI--LNKIL------NNNIPNQHTS 430
P KL+ TNI + + ML N S D +I LNK NNN N + +
Sbjct: 394 PPKLVLPTNIRISQESQDLMNKMLSNELEESTDEEDISNLNKKSPFITGNNNNNNNNNNN 453
Query: 431 NAHINNNPINNNSLDHKKCKLTQSL--LLTQSYSQDYSERI 469
N + NNN NNN+ +HK L Q L ++T+ + E++
Sbjct: 454 NNNNNNNNNNNNNNNHKPLDLHQELGNIITEKFEYVTFEKL 494
>UniRef100_Q95PH4 Histidine kinase DhkM [Dictyostelium discoideum]
Length = 1318
Score = 38.1 bits (87), Expect = 3.2
Identities = 25/73 (34%), Positives = 39/73 (53%), Gaps = 4/73 (5%)
Query: 411 DSHNILNKILNNNIPNQHTSNAHINNNPINNNSLDHKKCK----LTQSLLLTQSYSQDYS 466
+++N +N NNN N + N +INNN NNN++ KK K + S++ + S YS
Sbjct: 308 NNNNNINDSNNNNNSNNNNINNNINNNINNNNNIKKKKKKNEFTVYFSVIDSGSGIDPYS 367
Query: 467 ERI*FKPSLLLQY 479
+ F+P L Y
Sbjct: 368 TNLLFQPFSLSSY 380
>UniRef100_Q8MNJ0 Hypothetical protein [Dictyostelium discoideum]
Length = 1304
Score = 38.1 bits (87), Expect = 3.2
Identities = 44/168 (26%), Positives = 66/168 (39%), Gaps = 53/168 (31%)
Query: 377 W*PMEEPNKL----------IQSTNISLPSHLLPMLF----NHQIVSLDSHNILNKIL-- 420
W +EPNK S I + + LLP++ N+ + +D +I N L
Sbjct: 30 WVEYDEPNKYNVYNRCKFGNTHSLEIMIKNDLLPLIKDKIKNNHHIKIDHFSIKNLFLKL 89
Query: 421 ----------------NNNIPNQHTSNAHINNNPINNNSLDHKKCKLTQSLLLTQSYSQD 464
NNN N + +N NNN NNN +K +D
Sbjct: 90 SKKQPQQQNIQEEQTINNNNNNNNNNN---NNNNNNNNKNKYK--------------DED 132
Query: 465 YSERI*FKPSLLLQYQKECQHGTDLIRLVIFIKEPHVIILKVVMPLSM 512
Y E I L ++Y+++ TDLI++ + P VI L V P S+
Sbjct: 133 YLEII----ELFMKYRRDEFEITDLIKMAVKTYSPDVIRLLVNEPYSV 176
>UniRef100_Q8IHX4 Hypothetical protein [Plasmodium falciparum]
Length = 1791
Score = 38.1 bits (87), Expect = 3.2
Identities = 25/89 (28%), Positives = 43/89 (48%), Gaps = 3/89 (3%)
Query: 361 RNLGTTSQERKNTMLMW*PMEEPNKLIQSTNISLPSHLLPMLFNHQIVSLDSHNILNKIL 420
+N+G E N + E+ ++ + NI + ML N+ + +++ N +
Sbjct: 141 KNIGAIENELDNNLCRHNENEKKYDMVNNNNII---NNYDMLNNNNNILNNNNMHNNNMH 197
Query: 421 NNNIPNQHTSNAHINNNPINNNSLDHKKC 449
NNNI + +N INNN + NN+ H KC
Sbjct: 198 NNNIIINNNNNNMINNNNMLNNNSVHNKC 226
>UniRef100_UPI000046C800 UPI000046C800 UniRef100 entry
Length = 380
Score = 37.7 bits (86), Expect = 4.2
Identities = 16/40 (40%), Positives = 31/40 (77%), Gaps = 1/40 (2%)
Query: 410 LDSHNILNK-ILNNNIPNQHTSNAHINNNPINNNSLDHKK 448
++++N+ NK I NNN+ N+ +N ++NNN INNN++++ +
Sbjct: 331 INNNNVNNKGINNNNVNNKGINNNNVNNNGINNNNVNNNE 370
Score = 37.0 bits (84), Expect = 7.2
Identities = 16/39 (41%), Positives = 30/39 (76%), Gaps = 1/39 (2%)
Query: 410 LDSHNILNK-ILNNNIPNQHTSNAHINNNPINNNSLDHK 447
++++N+ NK I NNN+ N+ +N ++NN INNN++++K
Sbjct: 281 INNNNVNNKGINNNNVNNKGINNNNVNNKGINNNNVNNK 319
Score = 37.0 bits (84), Expect = 7.2
Identities = 16/39 (41%), Positives = 30/39 (76%), Gaps = 1/39 (2%)
Query: 410 LDSHNILNK-ILNNNIPNQHTSNAHINNNPINNNSLDHK 447
++++N+ NK I NNN+ N+ +N ++NN INNN++++K
Sbjct: 291 INNNNVNNKGINNNNVNNKGINNNNVNNKGINNNNVNNK 329
Score = 37.0 bits (84), Expect = 7.2
Identities = 16/39 (41%), Positives = 30/39 (76%), Gaps = 1/39 (2%)
Query: 410 LDSHNILNK-ILNNNIPNQHTSNAHINNNPINNNSLDHK 447
++++N+ NK I NNN+ N+ +N ++NN INNN++++K
Sbjct: 261 INNNNVNNKGINNNNVNNKGINNNNVNNKGINNNNVNNK 299
Score = 37.0 bits (84), Expect = 7.2
Identities = 16/39 (41%), Positives = 30/39 (76%), Gaps = 1/39 (2%)
Query: 410 LDSHNILNK-ILNNNIPNQHTSNAHINNNPINNNSLDHK 447
++++N+ NK I NNN+ N+ +N ++NN INNN++++K
Sbjct: 311 INNNNVNNKGINNNNVNNKGINNNNVNNKGINNNNVNNK 349
Score = 37.0 bits (84), Expect = 7.2
Identities = 16/39 (41%), Positives = 30/39 (76%), Gaps = 1/39 (2%)
Query: 410 LDSHNILNK-ILNNNIPNQHTSNAHINNNPINNNSLDHK 447
++++N+ NK I NNN+ N+ +N ++NN INNN++++K
Sbjct: 301 INNNNVNNKGINNNNVNNKGINNNNVNNKGINNNNVNNK 339
Score = 37.0 bits (84), Expect = 7.2
Identities = 16/39 (41%), Positives = 30/39 (76%), Gaps = 1/39 (2%)
Query: 410 LDSHNILNK-ILNNNIPNQHTSNAHINNNPINNNSLDHK 447
++++N+ NK I NNN+ N+ +N ++NN INNN++++K
Sbjct: 271 INNNNVNNKGINNNNVNNKGINNNNVNNKGINNNNVNNK 309
Score = 37.0 bits (84), Expect = 7.2
Identities = 16/39 (41%), Positives = 30/39 (76%), Gaps = 1/39 (2%)
Query: 410 LDSHNILNK-ILNNNIPNQHTSNAHINNNPINNNSLDHK 447
++++N+ NK I NNN+ N+ +N ++NN INNN++++K
Sbjct: 251 INNNNVNNKGINNNNVNNKGINNNNVNNKGINNNNVNNK 289
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.358 0.158 0.577
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,172,489,816
Number of Sequences: 2790947
Number of extensions: 113814890
Number of successful extensions: 524136
Number of sequences better than 10.0: 53
Number of HSP's better than 10.0 without gapping: 19
Number of HSP's successfully gapped in prelim test: 39
Number of HSP's that attempted gapping in prelim test: 520288
Number of HSP's gapped (non-prelim): 1475
length of query: 2301
length of database: 848,049,833
effective HSP length: 144
effective length of query: 2157
effective length of database: 446,153,465
effective search space: 962353024005
effective search space used: 962353024005
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 83 (36.6 bits)
Medicago: description of AC144656.5


