

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144617.10 + phase: 0
(337 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
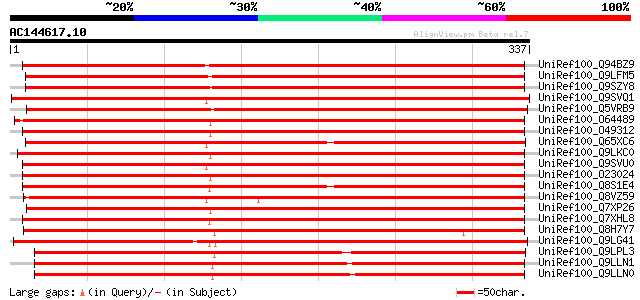
Sequences producing significant alignments: (bits) Value
UniRef100_Q94BZ9 Flavin monoxygenase-like protein floozy [Petuni... 493 e-138
UniRef100_Q9LFM5 Hypothetical protein F2I11_210 [Arabidopsis tha... 492 e-138
UniRef100_Q9SZY8 Dimethylaniline monooxygenase-like protein [Ara... 456 e-127
UniRef100_Q9SVQ1 Hypothetical protein F17N18.150 [Arabidopsis th... 431 e-119
UniRef100_Q5VRB9 Flavin-containing monooxygenase-like [Oryza sat... 426 e-118
UniRef100_O64489 F20D22.5 protein [Arabidopsis thaliana] 425 e-117
UniRef100_O49312 Putative flavin-containing monooxygenase [Arabi... 416 e-115
UniRef100_Q65XC6 Putative dimethylaniline monooxygenase [Oryza s... 415 e-114
UniRef100_Q9LKC0 Dimethylaniline monooxygenase-like [Arabidopsis... 414 e-114
UniRef100_Q9SVU0 Hypothetical protein F16A16.170 [Arabidopsis th... 412 e-114
UniRef100_O23024 T1G11.14 protein [Arabidopsis thaliana] 410 e-113
UniRef100_Q8S1E4 Putative dimethylaniline monooxygenase [Oryza s... 407 e-112
UniRef100_Q8VZ59 Hypothetical protein At5g25620 [Arabidopsis tha... 407 e-112
UniRef100_Q7XP26 OSJNBa0027H09.13 protein [Oryza sativa] 403 e-111
UniRef100_Q7XHL8 Putative dimethylaniline monooxygenase [Oryza s... 403 e-111
UniRef100_Q8H7Y7 Putative flavin-containing monooxygenase [Oryza... 385 e-106
UniRef100_Q9LG41 Dimethylaniline monooxygenase-like protein [Ory... 374 e-102
UniRef100_Q9LPL3 F24J8.6 protein [Arabidopsis thaliana] 288 2e-76
UniRef100_Q9LLN1 Hypothetical protein DUPR11.26 [Oryza sativa] 286 4e-76
UniRef100_Q9LLN0 Hypothetical protein DUPR11.28 [Oryza sativa] 286 6e-76
>UniRef100_Q94BZ9 Flavin monoxygenase-like protein floozy [Petunia hybrida]
Length = 412
Score = 493 bits (1270), Expect = e-138
Identities = 233/326 (71%), Positives = 278/326 (84%), Gaps = 2/326 (0%)
Query: 9 KCVWIHGPIIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPKHF 68
K +W++GPII+GAGPSG+AV+ACL E GVPSLILERSDCIASLWQ++TYDRLKLHLPK F
Sbjct: 12 KWLWVNGPIIIGAGPSGLAVSACLKENGVPSLILERSDCIASLWQHKTYDRLKLHLPKQF 71
Query: 69 CELPMMSFPQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDSTSQIWMVRTKE 128
C+LP+ FP FPKYPTK QFISY+ESYA HF I+P++ Q V AEFD S W V+T+
Sbjct: 72 CQLPLFGFPDNFPKYPTKRQFISYLESYAKHFSINPKYKQAVQVAEFDHVSGFWKVQTQ- 130
Query: 129 GDFQYFSPWLIVATGENAEPVFPTIHGMEHFHGPVVHTSDYKSGSEYKNKKVLVIGCGNS 188
+FQYFS WLIVATGENAEPV P I GM+ F GPV+HTS YKSG+E+ N++VLVIGCGN
Sbjct: 131 -NFQYFSKWLIVATGENAEPVIPNIQGMDKFKGPVMHTSLYKSGTEFNNQRVLVIGCGNF 189
Query: 189 GMEVSLDLCRHNAMPHLVARNSVHILPRDMFGFSTYGIAMGLYKWLPLKLVDKFLLLVSS 248
GMEVSLDLCRHNA+PH+VARNSVHILPR+M G ST+ +AM L K LPL++VDKFLLLV++
Sbjct: 190 GMEVSLDLCRHNAIPHMVARNSVHILPREMLGISTFSMAMALLKCLPLRIVDKFLLLVAN 249
Query: 249 FFLGNTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVMEGVKEITRNGAKF 308
LGNT+ G++RPKTGPIELK ATGKTPVLDVG ++QIK+G I+++ VKEIT+ GAKF
Sbjct: 250 LTLGNTDKLGLRRPKTGPIELKNATGKTPVLDVGALSQIKAGKIQIVHAVKEITKIGAKF 309
Query: 309 LDGQEKEFDAIILATGYKSNVPSWLK 334
+DG+E EFD+IILATGYKSNVPSWLK
Sbjct: 310 VDGKEGEFDSIILATGYKSNVPSWLK 335
>UniRef100_Q9LFM5 Hypothetical protein F2I11_210 [Arabidopsis thaliana]
Length = 411
Score = 492 bits (1266), Expect = e-138
Identities = 236/324 (72%), Positives = 276/324 (84%), Gaps = 2/324 (0%)
Query: 11 VWIHGPIIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPKHFCE 70
+++ GPIIVGAGPSG+AVAACLS +GVPS+ILER+DC+ASLWQ RTYDRLKLHLPKHFCE
Sbjct: 12 IFVPGPIIVGAGPSGLAVAACLSNRGVPSVILERTDCLASLWQKRTYDRLKLHLPKHFCE 71
Query: 71 LPMMSFPQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDSTSQIWMVRTKEGD 130
LP+M FP+ FPKYP+K FISY+ESYA F+I P FNQTV AEFD S +W V+T++G
Sbjct: 72 LPLMPFPKNFPKYPSKQLFISYVESYAARFNIKPVFNQTVEKAEFDDASGLWNVKTQDG- 130
Query: 131 FQYFSPWLIVATGENAEPVFPTIHGMEHFHGPVVHTSDYKSGSEYKNKKVLVIGCGNSGM 190
Y S WL+VATGENAEPVFP I G++ F GPVVHTS YKSGS + N+KVLV+GCGNSGM
Sbjct: 131 -VYTSTWLVVATGENAEPVFPNIPGLKKFTGPVVHTSAYKSGSAFANRKVLVVGCGNSGM 189
Query: 191 EVSLDLCRHNAMPHLVARNSVHILPRDMFGFSTYGIAMGLYKWLPLKLVDKFLLLVSSFF 250
EVSLDLCR+NA+PH+V RNSVH+LPRD FG ST+GIAM L KW PLKLVDKFLLL+++
Sbjct: 190 EVSLDLCRYNALPHMVVRNSVHVLPRDFFGLSTFGIAMTLLKWFPLKLVDKFLLLLANST 249
Query: 251 LGNTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVMEGVKEITRNGAKFLD 310
LGNT+ G++RPKTGPIELK TGKTPVLDVG I+ I+SG IKV + VKEITRNGAKFL+
Sbjct: 250 LGNTDLLGLRRPKTGPIELKNVTGKTPVLDVGAISLIRSGQIKVTQAVKEITRNGAKFLN 309
Query: 311 GQEKEFDAIILATGYKSNVPSWLK 334
G+E EFD+IILATGYKSNVP WLK
Sbjct: 310 GKEIEFDSIILATGYKSNVPDWLK 333
>UniRef100_Q9SZY8 Dimethylaniline monooxygenase-like protein [Arabidopsis thaliana]
Length = 414
Score = 456 bits (1172), Expect = e-127
Identities = 209/325 (64%), Positives = 271/325 (83%), Gaps = 2/325 (0%)
Query: 11 VWIHGPIIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPKHFCE 70
+ +HGPII+GAGPSG+A +ACLS +GVPSLILERSD IASLW+++TYDRL+LHLPKHFC
Sbjct: 16 ILVHGPIIIGAGPSGLATSACLSSRGVPSLILERSDSIASLWKSKTYDRLRLHLPKHFCR 75
Query: 71 LPMMSFPQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDSTSQIWMVRTKEGD 130
LP++ FP+ +PKYP+K++F++Y+ESYA HF I PRFN+ V +A +DS+S W V+T +
Sbjct: 76 LPLLDFPEYYPKYPSKNEFLAYLESYASHFRIAPRFNKNVQNAAYDSSSGFWRVKTHDNT 135
Query: 131 FQYFSPWLIVATGENAEPVFPTIHGMEHFHG-PVVHTSDYKSGSEYKNKKVLVIGCGNSG 189
+Y S WLIVATGENA+P FP I G + F G +VH S+YKSG E++ +KVLV+GCGNSG
Sbjct: 136 -EYLSKWLIVATGENADPYFPEIPGRKKFSGGKIVHASEYKSGEEFRRQKVLVVGCGNSG 194
Query: 190 MEVSLDLCRHNAMPHLVARNSVHILPRDMFGFSTYGIAMGLYKWLPLKLVDKFLLLVSSF 249
ME+SLDL RHNA PHLV RN+VH+LPR++ G ST+G+ M L K LPL+LVDKFLLL+++
Sbjct: 195 MEISLDLVRHNASPHLVVRNTVHVLPREILGVSTFGVGMTLLKCLPLRLVDKFLLLMANL 254
Query: 250 FLGNTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVMEGVKEITRNGAKFL 309
GNT+ G++RPKTGP+ELK TGK+PVLDVG ++ I+SG I++MEGVKEIT+ GAKF+
Sbjct: 255 SFGNTDRLGLRRPKTGPLELKNVTGKSPVLDVGAMSLIRSGMIQIMEGVKEITKKGAKFM 314
Query: 310 DGQEKEFDAIILATGYKSNVPSWLK 334
DGQEK+FD+II ATGYKSNVP+WL+
Sbjct: 315 DGQEKDFDSIIFATGYKSNVPTWLQ 339
>UniRef100_Q9SVQ1 Hypothetical protein F17N18.150 [Arabidopsis thaliana]
Length = 415
Score = 431 bits (1107), Expect = e-119
Identities = 202/340 (59%), Positives = 256/340 (74%), Gaps = 4/340 (1%)
Query: 2 DPFKEKVKCVWIHGPIIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLK 61
DP+ E+ +C+ I GPIIVG+GPSG+A AACL + +PSLILERS CIASLWQ++TYDRL+
Sbjct: 14 DPYVEETRCLMIPGPIIVGSGPSGLATAACLKSRDIPSLILERSTCIASLWQHKTYDRLR 73
Query: 62 LHLPKHFCELPMMSFPQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDSTSQI 121
LHLPK FCELP+M FP ++P YPTK QF+ Y+ESYA+HF + P FNQTV A+FD +
Sbjct: 74 LHLPKDFCELPLMPFPSSYPTYPTKQQFVQYLESYAEHFDLKPVFNQTVEEAKFDRRCGL 133
Query: 122 WMVRT----KEGDFQYFSPWLIVATGENAEPVFPTIHGMEHFHGPVVHTSDYKSGSEYKN 177
W VRT K+ +Y S WL+VATGENAE V P I G+ F GP++HTS YKSG +
Sbjct: 134 WRVRTTGGKKDETMEYVSRWLVVATGENAEEVMPEIDGIPDFGGPILHTSSYKSGEIFSE 193
Query: 178 KKVLVIGCGNSGMEVSLDLCRHNAMPHLVARNSVHILPRDMFGFSTYGIAMGLYKWLPLK 237
KK+LV+GCGNSGMEV LDLC NA+P LV R+SVH+LP++M G ST+GI+ L KW P+
Sbjct: 194 KKILVVGCGNSGMEVCLDLCNFNALPSLVVRDSVHVLPQEMLGISTFGISTSLLKWFPVH 253
Query: 238 LVDKFLLLVSSFFLGNTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVMEG 297
+VD+FLL +S LG+T+ G+ RPK GP+E K+ GKTPVLDVG +A+I+SG+IKV
Sbjct: 254 VVDRFLLRMSRLVLGDTDRLGLVRPKLGPLERKIKCGKTPVLDVGTLAKIRSGHIKVYPE 313
Query: 298 VKEITRNGAKFLDGQEKEFDAIILATGYKSNVPSWLKVKN 337
+K + A+F+DG+ FDAIILATGYKSNVP WLK N
Sbjct: 314 LKRVMHYSAEFVDGRVDNFDAIILATGYKSNVPMWLKGVN 353
>UniRef100_Q5VRB9 Flavin-containing monooxygenase-like [Oryza sativa]
Length = 364
Score = 426 bits (1096), Expect = e-118
Identities = 199/325 (61%), Positives = 250/325 (76%), Gaps = 1/325 (0%)
Query: 12 WIHGPIIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPKHFCEL 71
W+ G +IVGAGPSG+A AACL+ +GVP+ +LERSD +AS W++R YDRL LHLPK FCEL
Sbjct: 13 WVPGAVIVGAGPSGLAAAACLAARGVPATVLERSDSLASTWRHRMYDRLALHLPKRFCEL 72
Query: 72 PMMSFPQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDSTSQIWMVRTKEGDF 131
P++ FP+ +P YP+K QF++YME+YA + PRF TV A FD+ W VR G+
Sbjct: 73 PLLPFPEEYPTYPSKDQFVAYMEAYAAAAGVAPRFGATVEEAAFDAAVGAWRVRLDGGEV 132
Query: 132 QYFSPWLIVATGENAEPVFPTIHGMEHFHGPVVHTSDYKSGSEYKNKKVLVIGCGNSGME 191
+ WL+VATGENAEP P GM+ F G +HTS+YKSG ++ KKVLV+GCGNSGME
Sbjct: 133 -LMARWLVVATGENAEPRVPDFPGMQKFAGCAMHTSEYKSGEQFAGKKVLVVGCGNSGME 191
Query: 192 VSLDLCRHNAMPHLVARNSVHILPRDMFGFSTYGIAMGLYKWLPLKLVDKFLLLVSSFFL 251
VSLDLCRH A P +V RN+VH+LPR+MFG ST+GIAM L +WLP++LVD+FLL + L
Sbjct: 192 VSLDLCRHGAKPSMVVRNTVHVLPREMFGLSTFGIAMALLRWLPVQLVDRFLLTAAHLIL 251
Query: 252 GNTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVMEGVKEITRNGAKFLDG 311
GNT +G++RPKTGPIELK TG+TPVLDVG + IKSG IKV+ VKE+TR G +F DG
Sbjct: 252 GNTGQFGLRRPKTGPIELKNLTGRTPVLDVGTLDHIKSGKIKVVGAVKEMTRQGVRFTDG 311
Query: 312 QEKEFDAIILATGYKSNVPSWLKVK 336
+E++FD IILATGY+SNVPSWLKVK
Sbjct: 312 KEEQFDTIILATGYRSNVPSWLKVK 336
>UniRef100_O64489 F20D22.5 protein [Arabidopsis thaliana]
Length = 421
Score = 425 bits (1092), Expect = e-117
Identities = 196/334 (58%), Positives = 262/334 (77%), Gaps = 4/334 (1%)
Query: 4 FKEKVKCVWIHGPIIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLH 63
F E+ +CVW++GP+IVGAGPSG+A AACL +QGVP +++ERSDCIASLWQ RTYDRLKLH
Sbjct: 14 FSER-RCVWVNGPVIVGAGPSGLATAACLHDQGVPFVVVERSDCIASLWQKRTYDRLKLH 72
Query: 64 LPKHFCELPMMSFPQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDSTSQIWM 123
LPK FC+LP M FP +P+YPTK QFI Y+ESYA+ F I P FN++V SA FD TS +W
Sbjct: 73 LPKKFCQLPKMPFPDHYPEYPTKRQFIDYLESYANRFDIKPEFNKSVESARFDETSGLWR 132
Query: 124 VRTKEG--DFQYFSPWLIVATGENAEPVFPTIHG-MEHFHGPVVHTSDYKSGSEYKNKKV 180
VRT + +Y WL+VATGENAE V P I+G M F G V+H +YKSG +++ K+V
Sbjct: 133 VRTTSDGEEMEYICRWLVVATGENAERVVPEINGLMTEFDGEVIHACEYKSGEKFRGKRV 192
Query: 181 LVIGCGNSGMEVSLDLCRHNAMPHLVARNSVHILPRDMFGFSTYGIAMGLYKWLPLKLVD 240
LV+GCGNSGMEVSLDL HNA+ +V R+SVH+LPR++ G ST+GI++ + KWLPL LVD
Sbjct: 193 LVVGCGNSGMEVSLDLANHNAITSMVVRSSVHVLPREIMGKSTFGISVMMMKWLPLWLVD 252
Query: 241 KFLLLVSSFFLGNTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVMEGVKE 300
K LL++S LG+ ++YG+KRP GP+ELK TGKTPVLD+G + +IKSG+++++ +K+
Sbjct: 253 KLLLILSWLVLGSLSNYGLKRPDIGPMELKSMTGKTPVLDIGALEKIKSGDVEIVPAIKQ 312
Query: 301 ITRNGAKFLDGQEKEFDAIILATGYKSNVPSWLK 334
+R+ + +DGQ+ + DA++LATGY+SNVPSWL+
Sbjct: 313 FSRHHVELVDGQKLDIDAVVLATGYRSNVPSWLQ 346
>UniRef100_O49312 Putative flavin-containing monooxygenase [Arabidopsis thaliana]
Length = 431
Score = 416 bits (1070), Expect = e-115
Identities = 190/333 (57%), Positives = 253/333 (75%), Gaps = 7/333 (2%)
Query: 9 KCVWIHGPIIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPKHF 68
+C+W++GP+IVGAGPSG+AVAA L Q VP +ILER++CIASLWQNRTYDRLKLHLPK F
Sbjct: 25 RCIWVNGPVIVGAGPSGLAVAADLKRQEVPFVILERANCIASLWQNRTYDRLKLHLPKQF 84
Query: 69 CELPMMSFPQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDSTSQIWMVRTKE 128
C+LP + FP+ P+YPTK+QFI Y+ESYA HF + P+FN+TV SA++D +W V+T
Sbjct: 85 CQLPNLPFPEDIPEYPTKYQFIEYLESYATHFDLRPKFNETVQSAKYDKRFGLWRVQTVL 144
Query: 129 G-------DFQYFSPWLIVATGENAEPVFPTIHGMEHFHGPVVHTSDYKSGSEYKNKKVL 181
+F+Y WL+VATGENAE V P G+E F G V+H DYKSG Y+ K+VL
Sbjct: 145 RSELLGYCEFEYICRWLVVATGENAEKVVPEFEGLEDFGGDVLHAGDYKSGERYRGKRVL 204
Query: 182 VIGCGNSGMEVSLDLCRHNAMPHLVARNSVHILPRDMFGFSTYGIAMGLYKWLPLKLVDK 241
V+GCGNSGMEVSLDLC H+A P +V R+SVH+LPR++ G ST+ +++ + KW+P+ LVDK
Sbjct: 205 VVGCGNSGMEVSLDLCNHDASPSMVVRSSVHVLPREVLGKSTFELSVTMMKWMPVWLVDK 264
Query: 242 FLLLVSSFFLGNTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVMEGVKEI 301
LL+++ LGNT+ YG+KRP+ GP+ELK GKTPVLD+G I+ IKSG IK++ G+ +
Sbjct: 265 TLLVLTRLLLGNTDKYGLKRPEIGPLELKNTAGKTPVLDIGAISMIKSGKIKIVAGIAKF 324
Query: 302 TRNGAKFLDGQEKEFDAIILATGYKSNVPSWLK 334
+ +DG+ + D++ILATGY+SNVPSWLK
Sbjct: 325 GPGKVELVDGRVLQIDSVILATGYRSNVPSWLK 357
>UniRef100_Q65XC6 Putative dimethylaniline monooxygenase [Oryza sativa]
Length = 348
Score = 415 bits (1066), Expect = e-114
Identities = 193/331 (58%), Positives = 246/331 (74%), Gaps = 10/331 (3%)
Query: 11 VWIHGPIIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPKHFCE 70
VW+ GPI+VGAGPSG+A AACL E+G+ SL+LERS C+A LWQ + YDRL LHLP+ FCE
Sbjct: 3 VWVQGPIVVGAGPSGLAAAACLKEKGIDSLVLERSSCLAPLWQLKMYDRLSLHLPRQFCE 62
Query: 71 LPMMSFPQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDSTSQIWMVRT---- 126
LP+ FP ++P YPTK QF++Y+ESYA F I+P +N TV+ AEFD +W VRT
Sbjct: 63 LPLFPFPASYPDYPTKQQFVAYLESYAAKFGINPMYNHTVVCAEFDERLMLWRVRTTQAT 122
Query: 127 --KEGDFQYFSPWLIVATGENAEPVFPTIHGMEHFHGPVVHTSDYKSGSEYKNKKVLVIG 184
E D +Y S WL+VATGEN+E V P I G+E F G V+HTS YKSGS++ K VLV+G
Sbjct: 123 GMMEDDVEYVSQWLVVATGENSEAVLPVIDGLEEFRGSVIHTSAYKSGSKFAGKTVLVVG 182
Query: 185 CGNSGMEVSLDLCRHNAMPHLVARNSVHILPRDMFGFSTYGIAMGLYKWLPLKLVDKFLL 244
CGNSGMEV LDLC HN P +V VHILPR+M G T+ +AM L KWLP+ +VD+ LL
Sbjct: 183 CGNSGMEVCLDLCNHNGYPRIV----VHILPREMLGQPTFRLAMWLLKWLPIHIVDRILL 238
Query: 245 LVSSFFLGNTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVMEGVKEITRN 304
LV+ LG+T+ +G+KRP GP+ELK +GKTP+LD+G +A+IKSG+IKV ++ I
Sbjct: 239 LVARAILGDTSQFGLKRPSLGPLELKSLSGKTPILDIGTLAKIKSGDIKVRPAIRRIAGQ 298
Query: 305 GAKFLDGQEKEFDAIILATGYKSNVPSWLKV 335
KF+DG+ ++FDAI+LATGYKSNVP WLKV
Sbjct: 299 QVKFVDGRSEQFDAIVLATGYKSNVPCWLKV 329
>UniRef100_Q9LKC0 Dimethylaniline monooxygenase-like [Arabidopsis thaliana]
Length = 424
Score = 414 bits (1065), Expect = e-114
Identities = 187/335 (55%), Positives = 254/335 (75%), Gaps = 6/335 (1%)
Query: 6 EKVKCVWIHGPIIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLP 65
++ +C+W++GP+IVGAGPSG+A AACL E+GVP ++LER+DCIASLWQ RTYDR+KLHLP
Sbjct: 15 DRRRCIWVNGPVIVGAGPSGLATAACLREEGVPFVVLERADCIASLWQKRTYDRIKLHLP 74
Query: 66 KHFCELPMMSFPQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDSTSQIWMVR 125
K C+LP M FP+ +P+YPTK QFI Y+ESYA+ F I P+FN+ V SA +D TS +W ++
Sbjct: 75 KKVCQLPKMPFPEDYPEYPTKRQFIEYLESYANKFEITPQFNECVQSARYDETSGLWRIK 134
Query: 126 TKEG-----DFQYFSPWLIVATGENAEPVFPTIHGME-HFHGPVVHTSDYKSGSEYKNKK 179
T + +Y WL+VATGENAE V P I G+ F G V+H+ +YKSG +Y+ K
Sbjct: 135 TTSSSSSGSEMEYICRWLVVATGENAEKVVPEIDGLTTEFEGEVIHSCEYKSGEKYRGKS 194
Query: 180 VLVIGCGNSGMEVSLDLCRHNAMPHLVARNSVHILPRDMFGFSTYGIAMGLYKWLPLKLV 239
VLV+GCGNSGMEVSLDL HNA +V R+SVH+LPR++ G S++ I+M L KW PL LV
Sbjct: 195 VLVVGCGNSGMEVSLDLANHNANASMVVRSSVHVLPREILGKSSFEISMMLMKWFPLWLV 254
Query: 240 DKFLLLVSSFFLGNTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVMEGVK 299
DK LL+++ LGN YG+KRP GP+ELK+ +GKTPVLD+G + +IKSG ++++ G+K
Sbjct: 255 DKILLILAWLILGNLTKYGLKRPTMGPMELKIVSGKTPVLDIGAMEKIKSGEVEIVPGIK 314
Query: 300 EITRNGAKFLDGQEKEFDAIILATGYKSNVPSWLK 334
+R+ + +DGQ + DA++LATGY+SNVPSWL+
Sbjct: 315 RFSRSHVELVDGQRLDLDAVVLATGYRSNVPSWLQ 349
>UniRef100_Q9SVU0 Hypothetical protein F16A16.170 [Arabidopsis thaliana]
Length = 426
Score = 412 bits (1058), Expect = e-114
Identities = 187/332 (56%), Positives = 247/332 (74%), Gaps = 6/332 (1%)
Query: 9 KCVWIHGPIIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPKHF 68
+C+W++GP+IVGAGPSG+A AACL EQ VP ++LER+DCIASLWQ RTYDRLKLHLPK F
Sbjct: 18 RCIWVNGPVIVGAGPSGLATAACLHEQNVPFVVLERADCIASLWQKRTYDRLKLHLPKQF 77
Query: 69 CELPMMSFPQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDSTSQIWMVRT-- 126
C+LP M FP+ FP+YPTK QFI Y+ESYA F I+P+FN+ V +A FD TS +W V+T
Sbjct: 78 CQLPKMPFPEDFPEYPTKRQFIDYLESYATRFEINPKFNECVQTARFDETSGLWRVKTVS 137
Query: 127 ----KEGDFQYFSPWLIVATGENAEPVFPTIHGMEHFHGPVVHTSDYKSGSEYKNKKVLV 182
+ + +Y WL+VATGENAE V P I G+ F G V+H DYKSG ++ KKVLV
Sbjct: 138 KSESTQTEVEYICRWLVVATGENAERVMPEIDGLSEFSGEVIHACDYKSGEKFAGKKVLV 197
Query: 183 IGCGNSGMEVSLDLCRHNAMPHLVARNSVHILPRDMFGFSTYGIAMGLYKWLPLKLVDKF 242
+GCGNSGMEVSLDL H A P +V R+S+H++PR++ G ST+ +AM + +W PL LVDK
Sbjct: 198 VGCGNSGMEVSLDLANHFAKPSMVVRSSLHVMPREVMGKSTFELAMKMLRWFPLWLVDKI 257
Query: 243 LLLVSSFFLGNTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVMEGVKEIT 302
LL++S LGN YG+KRP+ GP+ELK GKTPVLD+G I +I+ G I V+ G+K
Sbjct: 258 LLVLSWMVLGNIEKYGLKRPEMGPMELKSVKGKTPVLDIGAIEKIRLGKINVVPGIKRFN 317
Query: 303 RNGAKFLDGQEKEFDAIILATGYKSNVPSWLK 334
N + ++G++ + D+++LATGY+SNVP WL+
Sbjct: 318 GNKVELVNGEQLDVDSVVLATGYRSNVPYWLQ 349
>UniRef100_O23024 T1G11.14 protein [Arabidopsis thaliana]
Length = 437
Score = 410 bits (1055), Expect = e-113
Identities = 188/333 (56%), Positives = 251/333 (74%), Gaps = 7/333 (2%)
Query: 9 KCVWIHGPIIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPKHF 68
+C+W++GP+IVGAGPSG+AVAA L +GVP +ILER++CIASLWQNRTYDRLKLHLPK F
Sbjct: 30 RCIWVNGPVIVGAGPSGLAVAAGLKREGVPFIILERANCIASLWQNRTYDRLKLHLPKQF 89
Query: 69 CELPMMSFPQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDSTSQIWMVRTKE 128
C+LP FP FP+YPTK QFI Y+ESYA +F I+P+FN+TV SA++D T +W V+T
Sbjct: 90 CQLPNYPFPDEFPEYPTKFQFIQYLESYAANFDINPKFNETVQSAKYDETFGLWRVKTIS 149
Query: 129 G-------DFQYFSPWLIVATGENAEPVFPTIHGMEHFHGPVVHTSDYKSGSEYKNKKVL 181
+F+Y W++VATGENAE V P G+E F G V+H DYKSG Y+ KKVL
Sbjct: 150 NMGQLGSCEFEYICRWIVVATGENAEKVVPDFEGLEDFGGDVLHAGDYKSGGRYQGKKVL 209
Query: 182 VIGCGNSGMEVSLDLCRHNAMPHLVARNSVHILPRDMFGFSTYGIAMGLYKWLPLKLVDK 241
V+GCGNSGMEVSLDL H A P +V R++VH+LPR++FG ST+ + + + K++P+ L DK
Sbjct: 210 VVGCGNSGMEVSLDLYNHGANPSMVVRSAVHVLPREIFGKSTFELGVTMMKYMPVWLADK 269
Query: 242 FLLLVSSFFLGNTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVMEGVKEI 301
+L ++ LGNT+ YG+KRPK GP+ELK GKTPVLD+G + +I+SG IK++ G+ +
Sbjct: 270 TILFLARIILGNTDKYGLKRPKIGPLELKNKEGKTPVLDIGALPKIRSGKIKIVPGIIKF 329
Query: 302 TRNGAKFLDGQEKEFDAIILATGYKSNVPSWLK 334
+ + +DG+ E D++ILATGY+SNVPSWLK
Sbjct: 330 GKGKVELIDGRVLEIDSVILATGYRSNVPSWLK 362
>UniRef100_Q8S1E4 Putative dimethylaniline monooxygenase [Oryza sativa]
Length = 430
Score = 407 bits (1047), Expect = e-112
Identities = 196/331 (59%), Positives = 250/331 (75%), Gaps = 9/331 (2%)
Query: 9 KCVWIHGPIIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPKHF 68
+C+ + GPIIVGAGPSG+AVAACL E+GV SL+LERS+CIASLWQ +TYDRL LHLP+ F
Sbjct: 47 RCIRVLGPIIVGAGPSGLAVAACLKEKGVDSLVLERSNCIASLWQLKTYDRLSLHLPRQF 106
Query: 69 CELPMMSFPQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDSTSQIWMVRTKE 128
CELP+M FP +P YP+K QF++Y+ESYA F I P +N+TV+ AE+D Q+W VRT+
Sbjct: 107 CELPLMPFPAYYPIYPSKQQFVAYLESYAARFGICPTYNRTVVCAEYDEQLQLWRVRTRA 166
Query: 129 -----GDFQYFSPWLIVATGENAEPVFPTIHGMEHFHGPVVHTSDYKSGSEYKNKKVLVI 183
+ +Y S WL+VATGENAE V P I G++ F G V+HTS YKSG + K+VLV+
Sbjct: 167 TGIMGEEVEYVSRWLVVATGENAEVVLPEIDGLDDFKGTVMHTSSYKSGGAFAGKRVLVV 226
Query: 184 GCGNSGMEVSLDLCRHNAMPHLVARNSVHILPRDMFGFSTYGIAMGLYKWLPLKLVDKFL 243
G GNSGMEV LDLC HNA PH+V VHILPR+M G ST+G++M L KWLP+ +VD+ L
Sbjct: 227 GSGNSGMEVCLDLCNHNANPHIV----VHILPREMLGQSTFGLSMWLLKWLPVHVVDRIL 282
Query: 244 LLVSSFFLGNTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVMEGVKEITR 303
LL++ LG+T G+KRP GP+ELK +GKTPVLDVG A+IKSG+IKV +K+I+
Sbjct: 283 LLIAQTMLGDTAQLGLKRPTIGPLELKSLSGKTPVLDVGTFAKIKSGDIKVRPAIKQISG 342
Query: 304 NGAKFLDGQEKEFDAIILATGYKSNVPSWLK 334
+F+D + +EFD I+LATGYKSNVP WLK
Sbjct: 343 RQVEFMDTRLEEFDVIVLATGYKSNVPFWLK 373
>UniRef100_Q8VZ59 Hypothetical protein At5g25620 [Arabidopsis thaliana]
Length = 417
Score = 407 bits (1047), Expect = e-112
Identities = 197/329 (59%), Positives = 248/329 (74%), Gaps = 5/329 (1%)
Query: 10 CVWIHGPIIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPKHFC 69
CV + GP+IVGAGPSG+A AACL E+G+ S++LERS+CIASLWQ +TYDRL LHLPK FC
Sbjct: 27 CV-VTGPVIVGAGPSGLATAACLKERGITSVLLERSNCIASLWQLKTYDRLHLHLPKQFC 85
Query: 70 ELPMMSFPQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDSTSQIWMVRT--K 127
ELP++ FP FP YPTK QFI Y+E YA F I P FNQTV SA FD +W V + +
Sbjct: 86 ELPIIPFPGDFPTYPTKQQFIEYLEDYARRFDIKPEFNQTVESAAFDENLGMWRVTSVGE 145
Query: 128 EGDFQYFSPWLIVATGENAEPVFPTIHGMEHFH--GPVVHTSDYKSGSEYKNKKVLVIGC 185
EG +Y WL+ ATGENAEPV P GM+ F G V HT YK+G ++ K+VLV+GC
Sbjct: 146 EGTTEYVCRWLVAATGENAEPVVPRFEGMDKFAAAGVVKHTCHYKTGGDFAGKRVLVVGC 205
Query: 186 GNSGMEVSLDLCRHNAMPHLVARNSVHILPRDMFGFSTYGIAMGLYKWLPLKLVDKFLLL 245
GNSGMEV LDLC A P LV R++VH+LPR+M G ST+G++M L KWLP++LVD+FLL+
Sbjct: 206 GNSGMEVCLDLCNFGAQPSLVVRDAVHVLPREMLGTSTFGLSMFLLKWLPIRLVDRFLLV 265
Query: 246 VSSFFLGNTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVMEGVKEITRNG 305
VS F LG+T G+ RP+ GP+ELK +GKTPVLDVG +A+IK+G+IKV G++ + R+
Sbjct: 266 VSRFILGDTTLLGLNRPRLGPLELKNISGKTPVLDVGTLAKIKTGDIKVCSGIRRLKRHE 325
Query: 306 AKFLDGQEKEFDAIILATGYKSNVPSWLK 334
+F +G+ + FDAIILATGYKSNVPSWLK
Sbjct: 326 VEFDNGKTERFDAIILATGYKSNVPSWLK 354
>UniRef100_Q7XP26 OSJNBa0027H09.13 protein [Oryza sativa]
Length = 419
Score = 403 bits (1035), Expect = e-111
Identities = 194/326 (59%), Positives = 240/326 (73%), Gaps = 3/326 (0%)
Query: 12 WIHGPIIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPKHFCEL 71
W+ GPIIVGAGPSG+AVAA L EQGVP +LER+DCIASLWQ RTYDRLKLHLPK FCEL
Sbjct: 19 WVSGPIIVGAGPSGLAVAASLREQGVPFTMLERADCIASLWQKRTYDRLKLHLPKQFCEL 78
Query: 72 PMMSFPQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDSTSQIWMVRTKEG-- 129
P M+FP +P+YPT+ QFI Y+E YA F I+P F TVLSA +D TS +W VR
Sbjct: 79 PRMAFPAHYPEYPTRRQFIDYLEDYAAAFDINPLFGHTVLSARYDETSGLWRVRASSSAG 138
Query: 130 -DFQYFSPWLIVATGENAEPVFPTIHGMEHFHGPVVHTSDYKSGSEYKNKKVLVIGCGNS 188
+ +Y WL+VATGENAE V P I G++ F G VVH +DYKSG Y+ K+VLV+GCGNS
Sbjct: 139 AEMEYIGSWLVVATGENAESVVPDIPGIDGFGGEVVHVADYKSGEAYRGKRVLVVGCGNS 198
Query: 189 GMEVSLDLCRHNAMPHLVARNSVHILPRDMFGFSTYGIAMGLYKWLPLKLVDKFLLLVSS 248
GMEVSLDLC H A P +V R++VH+LPR++ G ST+ +A+ L WLPL LVDK L+L++
Sbjct: 199 GMEVSLDLCDHGARPAMVVRDAVHVLPREVLGKSTFELAVLLMAWLPLWLVDKILVLLAW 258
Query: 249 FFLGNTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVMEGVKEITRNGAKF 308
LGN GI+RP TGP+ELK TG+TPVLD G +A+I+SG I V+ GV R A+
Sbjct: 259 LVLGNLAKLGIRRPATGPLELKNTTGRTPVLDYGALARIRSGEITVVPGVARFGRGFAEL 318
Query: 309 LDGQEKEFDAIILATGYKSNVPSWLK 334
DG+ DA++LATGY+SNVP WL+
Sbjct: 319 ADGRVIALDAVVLATGYRSNVPQWLQ 344
>UniRef100_Q7XHL8 Putative dimethylaniline monooxygenase [Oryza sativa]
Length = 398
Score = 403 bits (1035), Expect = e-111
Identities = 185/334 (55%), Positives = 255/334 (75%), Gaps = 8/334 (2%)
Query: 9 KCVWIHGPIIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPKHF 68
+ VW++GPI+VGAGP+G++VAACL E+GVPS++LER+DCIASLWQ RTYDRL+LHLPKHF
Sbjct: 4 RVVWVNGPIVVGAGPAGLSVAACLRERGVPSVLLERADCIASLWQRRTYDRLRLHLPKHF 63
Query: 69 CELPMMSFPQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDSTSQIWMVRTKE 128
CELP M FP +P+YP + QF+ Y+++YA + PRFNQ+V SA +D + +W VR ++
Sbjct: 64 CELPGMPFPDGYPEYPDRRQFVDYLQAYAARAGVEPRFNQSVTSARYDDAAGLWRVRAED 123
Query: 129 ------GDF-QYFSPWLIVATGENAEPVFPTIHGMEHFHGPVVHTSDYKSGSEYKNKKVL 181
GD +Y WL+VATGENAE V P I G + F GPV H ++YKSG+ Y+ K+VL
Sbjct: 124 VSVDAAGDVTEYIGRWLVVATGENAERVVPEIDGADDFEGPVSHVAEYKSGAAYRGKRVL 183
Query: 182 VIGCGNSGMEVSLDLCRHNAMPHLVARNS-VHILPRDMFGFSTYGIAMGLYKWLPLKLVD 240
V+GCGNSGMEV LDLC HNA+P +V R+S VH+LPR+M G +T+ +A+ L ++LPL +VD
Sbjct: 184 VVGCGNSGMEVCLDLCHHNALPAMVVRDSKVHVLPREMLGVATFSVAVFLLRFLPLWVVD 243
Query: 241 KFLLLVSSFFLGNTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVMEGVKE 300
+ L++++ FLG+ GI RP GP+ELK G+TPVLD+G +A+I+SG+I+V+ G++
Sbjct: 244 RILVVLAWLFLGDLAKIGITRPSRGPLELKNTRGRTPVLDIGALARIRSGDIEVVPGIRR 303
Query: 301 ITRNGAKFLDGQEKEFDAIILATGYKSNVPSWLK 334
+ R GA+ +DG+ DA+ILATGY+SNVP WLK
Sbjct: 304 LLRGGAELVDGRRVPADAVILATGYQSNVPQWLK 337
>UniRef100_Q8H7Y7 Putative flavin-containing monooxygenase [Oryza sativa]
Length = 444
Score = 385 bits (989), Expect = e-106
Identities = 187/355 (52%), Positives = 241/355 (67%), Gaps = 30/355 (8%)
Query: 10 CVWIHGPIIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPKHFC 69
C+ + GPIIVGAGPSG+AVAA L + G P ++ERS +A LW NRTYDRL+LHLPK FC
Sbjct: 20 CIVLDGPIIVGAGPSGLAVAATLRQHGAPFTVVERSGGVADLWTNRTYDRLRLHLPKVFC 79
Query: 70 ELPMMSFPQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDSTSQIWMVRTKEG 129
ELP ++FP FP YPTKH F+ Y+ SYA F I P +TV A +D + +W V T
Sbjct: 80 ELPHVAFPPDFPTYPTKHDFLRYLHSYAARFAIAPLLRRTVTRAWYDHPASLWRVTTTTT 139
Query: 130 DF-------QYFSPWLIVATGENAEPVFPTIHGMEHFHGPVVHTSDYKSGSEYKNKKVLV 182
+Y SPWL+VA+GENAE V P + G E F G +H+S+Y+SG ++ +VLV
Sbjct: 140 SSSATSVITEYASPWLVVASGENAEVVVPKVKGRERFAGEALHSSEYRSGERFRGMRVLV 199
Query: 183 IGCGNSGMEVSLDLCRHNAMPHLVARNSVHILPRDMFGFSTYGIAMGLYKWLPLKLVDKF 242
+GCGNSGME+ LDLC H AMP + R+ VH+LPR+MFG ST+GIAM L +WLP+K+VD+F
Sbjct: 200 VGCGNSGMEMCLDLCEHGAMPFMSVRSGVHVLPREMFGASTFGIAMKLLRWLPIKMVDRF 259
Query: 243 LLLVSSFFLGNTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIK--------- 293
LLLV+ LG+T YG+KRPK GP+E+K TGK+PVLDVG + IKSGNIK
Sbjct: 260 LLLVARMVLGDTEKYGLKRPKLGPLEIKNITGKSPVLDVGAWSLIKSGNIKKERMYDNSG 319
Query: 294 --------------VMEGVKEITRNGAKFLDGQEKEFDAIILATGYKSNVPSWLK 334
++ V+ + NGA+F+DG E FDA+I ATGY+SNVPSWL+
Sbjct: 320 YASGQRSFFLKWVEIVPEVESFSGNGARFVDGNEMAFDAVIFATGYRSNVPSWLQ 374
>UniRef100_Q9LG41 Dimethylaniline monooxygenase-like protein [Oryza sativa]
Length = 439
Score = 374 bits (959), Expect = e-102
Identities = 185/349 (53%), Positives = 243/349 (69%), Gaps = 17/349 (4%)
Query: 3 PFKEKVKCVWIHGPIIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKL 62
P + K V + G +IVGAGP+G+AV A L +GV ++LER CIASLW++RTYDRL L
Sbjct: 32 PVDDVDKVVDVPGAVIVGAGPAGVAVGALLGLRGVAYVVLERCGCIASLWRHRTYDRLCL 91
Query: 63 HLPKHFCELPMMSFPQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDSTSQIW 122
HLPK FCELP+ FP +FP+YPT+ QF+ Y+++YA F + P F + V+SAE+D S W
Sbjct: 92 HLPKRFCELPLRPFPASFPEYPTRDQFLGYLDAYAREFGVEPVFRRAVISAEYDGES--W 149
Query: 123 MVRTKE------GDFQ---------YFSPWLIVATGENAEPVFPTIHGMEHFHGPVVHTS 167
V T+E G Q Y S WL+VATGENAEPV P + G F G ++H+S
Sbjct: 150 WVYTREVVAAAAGGEQAVLGCTMTVYRSRWLVVATGENAEPVVPEMDGAGRFKGQMMHSS 209
Query: 168 DYKSGSEYKNKKVLVIGCGNSGMEVSLDLCRHNAMPHLVARNSVHILPRDMFGFSTYGIA 227
+Y++G Y KKVLV+GCGNSGMEVSLDLC HNA +V R++VH+LPR++ GFST+G++
Sbjct: 210 EYRNGDGYAGKKVLVVGCGNSGMEVSLDLCNHNARASMVVRDTVHVLPREILGFSTFGLS 269
Query: 228 MGLYKWLPLKLVDKFLLLVSSFFLGNTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQI 287
M L +WL ++ VD +LL+S G+T GI RP GP ELK +GKTPVLDVG +A+I
Sbjct: 270 MWLLRWLSVQTVDWLVLLLSFLVFGDTARLGIPRPSLGPFELKSVSGKTPVLDVGTLAKI 329
Query: 288 KSGNIKVMEGVKEITRNGAKFLDGQEKEFDAIILATGYKSNVPSWLKVK 336
KSG+IKV ++ +G +F+DG +EFD +ILATGYKSNVP WLK K
Sbjct: 330 KSGDIKVTPAIQCFQEHGVEFVDGSTEEFDVVILATGYKSNVPYWLKEK 378
>UniRef100_Q9LPL3 F24J8.6 protein [Arabidopsis thaliana]
Length = 391
Score = 288 bits (736), Expect = 2e-76
Identities = 143/322 (44%), Positives = 207/322 (63%), Gaps = 8/322 (2%)
Query: 17 IIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPKHFCELPMMSF 76
+I+GAGP+G+A +ACL+ +P++++ER C ASLW+ R+YDRLKLHL K FC+LP M F
Sbjct: 10 LIIGAGPAGLATSACLNRLNIPNIVVERDVCSASLWKRRSYDRLKLHLAKQFCQLPHMPF 69
Query: 77 PQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDSTSQIWMVRTKEGDF--QYF 134
P P + +K FI+Y++ YA F+++PR+N+ V SA F I V K Y
Sbjct: 70 PSNTPTFVSKLGFINYLDEYATRFNVNPRYNRNVKSAYFKDGQWIVKVVNKTTALIEVYS 129
Query: 135 SPWLIVATGENAEPVFPTIHGM-EHFHGPVVHTSDYKSGSEYKNKKVLVIGCGNSGMEVS 193
+ +++ ATGEN E V P I G+ E F G +H+S+YK+G ++ K VLV+GCGNSGME++
Sbjct: 130 AKFMVAATGENGEGVIPEIPGLVESFQGKYLHSSEYKNGEKFAGKDVLVVGCGNSGMEIA 189
Query: 194 LDLCRHNAMPHLVARNSVHILPRDMFGFSTYGIAMGLYKWLPLKLVDKFLLLVSSFFLGN 253
DL + NA +V R+ VH+L R I M L ++ P+KLVD+ LL++ N
Sbjct: 190 YDLSKCNANVSIVVRSQVHVLTR-----CIVRIGMSLLRFFPVKLVDRLCLLLAELRFRN 244
Query: 254 TNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVMEGVKEITRNGAKFLDGQE 313
T+ YG+ RP GP KL TG++ +DVG + +IKSG I+V+ +K I +F+DG
Sbjct: 245 TSRYGLVRPNNGPFLNKLITGRSATIDVGCVGEIKSGKIQVVTSIKRIEGKTVEFIDGNT 304
Query: 314 KEFDAIILATGYKSNVPSWLKV 335
K D+I+ ATGYKS+V WL+V
Sbjct: 305 KNVDSIVFATGYKSSVSKWLEV 326
>UniRef100_Q9LLN1 Hypothetical protein DUPR11.26 [Oryza sativa]
Length = 387
Score = 286 bits (733), Expect = 4e-76
Identities = 143/322 (44%), Positives = 204/322 (62%), Gaps = 7/322 (2%)
Query: 17 IIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPKHFCELPMMSF 76
+IVGAGP+G+A AACL+++ VP +I+ER ASLW++R YDRLKLHL K FCELP M++
Sbjct: 10 LIVGAGPAGLATAACLAQRHVPYIIVERESSTASLWRHRAYDRLKLHLAKEFCELPHMAY 69
Query: 77 PQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDSTSQIWMVRTKEGD----FQ 132
P P Y + F+ Y++SYA+ F I PR++ V SA D W+V ++ D +
Sbjct: 70 PAGTPTYVPRDMFVEYLDSYANQFGIRPRYHTAVESAIHDKGKNQWVVLVRDMDTSVVAR 129
Query: 133 YFSPWLIVATGENAEPVFPTIHGMEHFHGPVVHTSDYKSGSEYKNKKVLVIGCGNSGMEV 192
+ +L+VA GEN+ P I G+ F G +H+S YKSG Y K VLV+G GNSGME+
Sbjct: 130 LATQFLVVAAGENSAANIPPIPGLSRFEGEAIHSSAYKSGRAYTGKSVLVVGAGNSGMEI 189
Query: 193 SLDLCRHNAMPHLVARNSVHILPRDMFGFSTYGIAMGLYKWLPLKLVDKFLLLVSSFFLG 252
+ DL H A +V R+ VHI+ +++ YG+ M L + VD L++ ++F+ G
Sbjct: 190 AYDLATHGAHTSIVVRSPVHIMTKELI---WYGMTMVQNLGLNVTAVDSLLVMAANFYFG 246
Query: 253 NTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVMEGVKEITRNGAKFLDGQ 312
+ + +GI RPK GP+ LK TG++ V+DVG IK G IKV +G+ +I N +F G+
Sbjct: 247 DLSKHGIMRPKMGPLLLKSQTGRSAVIDVGTARLIKGGVIKVFQGISKINTNSVEFHGGR 306
Query: 313 EKEFDAIILATGYKSNVPSWLK 334
+ FDAI+ ATGYKS V +WLK
Sbjct: 307 QNSFDAIVFATGYKSTVNAWLK 328
>UniRef100_Q9LLN0 Hypothetical protein DUPR11.28 [Oryza sativa]
Length = 387
Score = 286 bits (732), Expect = 6e-76
Identities = 140/322 (43%), Positives = 204/322 (62%), Gaps = 7/322 (2%)
Query: 17 IIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPKHFCELPMMSF 76
+IVGAGP+G+A AACL+++ VP +I+ER C ASLW++R YDRLKLHL K FCELP M++
Sbjct: 10 LIVGAGPAGLATAACLAQRHVPYVIVERESCTASLWRHRAYDRLKLHLAKEFCELPHMAY 69
Query: 77 PQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDSTSQIWMVRTKEGD----FQ 132
P P Y + F+ Y++SY D F I PR++ + SA +D W V ++ D +
Sbjct: 70 PVGTPTYIPRDMFVEYLDSYTDQFGIRPRYHTAIESAIYDGGKNRWAVLARDTDTSVVTR 129
Query: 133 YFSPWLIVATGENAEPVFPTIHGMEHFHGPVVHTSDYKSGSEYKNKKVLVIGCGNSGMEV 192
+ +L+VATGEN+ P + G+ F G +H+S YKSG Y K VLV+G GNSGME+
Sbjct: 130 LTAQFLVVATGENSAASIPPVPGLTRFEGEAIHSSAYKSGRAYTGKNVLVVGAGNSGMEI 189
Query: 193 SLDLCRHNAMPHLVARNSVHILPRDMFGFSTYGIAMGLYKWLPLKLVDKFLLLVSSFFLG 252
+ DL H A +V R+ +HI+ +++ F G+ + L + D L++ ++F+ G
Sbjct: 190 AYDLATHGAHTSIVVRSPIHIMTKELIRF---GMTVVQNLGLTVTTADSLLVMAANFYFG 246
Query: 253 NTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVMEGVKEITRNGAKFLDGQ 312
+ + +GI RPK GP+ LK TG++ V+DVG IK G IKV +G+ +I N +F G+
Sbjct: 247 DLSKHGITRPKIGPLLLKSQTGRSAVIDVGTARLIKGGVIKVFQGISKIKTNSIEFHGGK 306
Query: 313 EKEFDAIILATGYKSNVPSWLK 334
+ FDAI+ ATGYKS V +WLK
Sbjct: 307 QIPFDAIVFATGYKSTVNTWLK 328
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.322 0.139 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 617,275,397
Number of Sequences: 2790947
Number of extensions: 26279420
Number of successful extensions: 76142
Number of sequences better than 10.0: 1518
Number of HSP's better than 10.0 without gapping: 764
Number of HSP's successfully gapped in prelim test: 755
Number of HSP's that attempted gapping in prelim test: 73226
Number of HSP's gapped (non-prelim): 2384
length of query: 337
length of database: 848,049,833
effective HSP length: 128
effective length of query: 209
effective length of database: 490,808,617
effective search space: 102579000953
effective search space used: 102579000953
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 75 (33.5 bits)
Medicago: description of AC144617.10


