

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
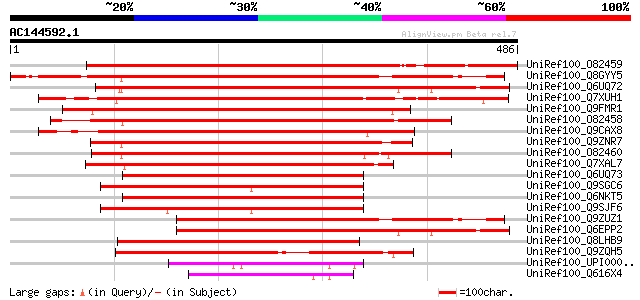
Query= AC144592.1 - phase: 0 /pseudo
(486 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O82459 Rac GTPase activating protein 2 [Lotus japonicus] 592 e-167
UniRef100_Q8GYY5 Putative rac GTPase activating protein [Arabido... 451 e-125
UniRef100_Q6UQ72 Rho GTPase activating protein 2 [Oryza sativa] 399 e-109
UniRef100_Q7XUH1 OSJNBa0020J04.11 protein [Oryza sativa] 397 e-109
UniRef100_Q9FMR1 Rac GTPase activating protein [Arabidopsis thal... 386 e-106
UniRef100_O82458 Rac GTPase activating protein 1 [Lotus japonicus] 382 e-104
UniRef100_Q9CAX8 Putative rac GTPase activating protein; 62102-6... 380 e-104
UniRef100_Q9ZNR7 Putative rac GTPase activating protein [Arabido... 378 e-103
UniRef100_O82460 Rac GTPase activating protein 3 [Lotus japonicus] 377 e-103
UniRef100_Q7XAL7 Rac GTPase activating protein 3-like protein [O... 363 8e-99
UniRef100_Q6UQ73 Rho GTPase activating protein 1 [Oryza sativa] 360 7e-98
UniRef100_Q9SGC6 T23G18.20 [Arabidopsis thaliana] 338 2e-91
UniRef100_Q6NKT5 At1g08340 [Arabidopsis thaliana] 337 6e-91
UniRef100_Q9SJF6 T27G7.4 [Arabidopsis thaliana] 330 7e-89
UniRef100_Q9ZUZ1 Putative rac GTPase activating protein [Arabido... 324 4e-87
UniRef100_Q6EPP2 Putative Rho GTPase activating protein 2 [Oryza... 319 1e-85
UniRef100_Q8LHB9 Putative rac GTPase activating protein [Oryza s... 305 1e-81
UniRef100_Q9ZQH5 Putative rac GTPase activating protein [Arabido... 277 4e-73
UniRef100_UPI00004996C7 UPI00004996C7 UniRef100 entry 99 2e-19
UniRef100_Q616X4 Hypothetical protein CBG15100 [Caenorhabditis b... 81 8e-14
>UniRef100_O82459 Rac GTPase activating protein 2 [Lotus japonicus]
Length = 424
Score = 592 bits (1525), Expect = e-167
Identities = 308/414 (74%), Positives = 343/414 (82%), Gaps = 11/414 (2%)
Query: 74 SNNNHQNQFAILDIVMAALKKSIVTCSVEREDVSSLDISWPTEVRHVSHVTFDRFNGFLG 133
S+N +QNQFAILDI++AALKKS+VTCSV+REDVSSLDISWPTEVRHVSHVTFDRFNGFLG
Sbjct: 15 SHNQNQNQFAILDILVAALKKSLVTCSVDREDVSSLDISWPTEVRHVSHVTFDRFNGFLG 74
Query: 134 LPSEFQPEVPTRVPSASVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEG 193
LP+E QPEVP +VP+AS KVFGVSAKSMQCSYD+RGNSVPTILL MQ +LYSEGGLKAEG
Sbjct: 75 LPTELQPEVPQKVPTASAKVFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEG 134
Query: 194 IFRITAENSQEAFVRDQLNKGVVPHGIDVHCLSGLIKAWFRELPTGVLDSLTPEQVMQCN 253
IFRI A+NSQE FVR QLN+G+VP G++VHCLSGLIKAWFRELPTGVLDSLTPEQVM CN
Sbjct: 135 IFRINADNSQEEFVRCQLNRGLVPRGVEVHCLSGLIKAWFRELPTGVLDSLTPEQVMHCN 194
Query: 254 TEDDCTNLVKLLPSTEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNMTQMADPLT 313
+E+DCTNLVKLLPSTEAALLDWAINLMADVVE+EQFNKMNARN+AMVFAPNMTQM DPLT
Sbjct: 195 SEEDCTNLVKLLPSTEAALLDWAINLMADVVEHEQFNKMNARNIAMVFAPNMTQMVDPLT 254
Query: 314 ALIHAVQVMNFLKTLILKMLREREESIDNARLLSHSMDFASCNDEFPPFSFNKEESSCEQ 373
ALIHAVQVMNFLKTLILK LRER+ES+ AR LS ++ SC + PF N+EESS +
Sbjct: 255 ALIHAVQVMNFLKTLILKTLRERDESMAKARQLSSLLNSPSCKGDSHPFKDNREESSAQP 314
Query: 374 IEDACDKNSSTTKRKFSRTSTLGRIEWSV-EKLRSSEEKRNREEVFRSFSSDSVTPRYES 432
+ D C K +FS R+EW V EK+ SSEEK S S S RYES
Sbjct: 315 V-DTC-ATMPPDKSEFS------RMEWCVDEKVWSSEEKGTGGGALESVSGGSSPSRYES 366
Query: 433 GPLENSYSYRRRYDSEHWSRLRNGVRKLCRHPVFQLSKPSKKPASLGIVNTREG 486
GPLE+ YR YDSEHW RLR GVR+LC+HPVFQLSK +KK A LGIVNTREG
Sbjct: 367 GPLES--RYRGIYDSEHWLRLRKGVRRLCQHPVFQLSKSTKKRADLGIVNTREG 418
>UniRef100_Q8GYY5 Putative rac GTPase activating protein [Arabidopsis thaliana]
Length = 455
Score = 451 bits (1160), Expect = e-125
Identities = 263/478 (55%), Positives = 313/478 (65%), Gaps = 52/478 (10%)
Query: 1 MTRLFRSKSCNIAGFSVSEFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDEI 60
MT RSKS GF EF P T + + ++DE EE S + + D
Sbjct: 1 MTNFSRSKSTGTIGFP--EFKP----TRPGPDKYENIHNDDDEYEEGHSTTSTDYYDAST 54
Query: 61 YSSNNASRQFTKGSNNNHQNQFAILDIVMAALKKSIV-TCSVERED---VSSLDISWPTE 116
S++ASR N + Q ++D++ A L+KS+V +C++ER + V+S+DI WPTE
Sbjct: 55 PLSSHASRS----GNGSGSGQLTVVDLLAAVLRKSLVMSCAMERGEDDVVASMDIGWPTE 110
Query: 117 VRHVSHVTFDRFNGFLGLPSEFQPEVPTRVPSASVKVFGVSAKSMQCSYDDRGNSVPTIL 176
V+HVSHVTFDRFNGFLGLPSE +PEVP R PSASV VFGVSAKSMQCSYDDRGNSVPTIL
Sbjct: 111 VKHVSHVTFDRFNGFLGLPSELEPEVPPRAPSASVSVFGVSAKSMQCSYDDRGNSVPTIL 170
Query: 177 LRMQKQLYSEGGLKAEGIFRITAENSQEAFVRDQLNKGVVPHGIDVHCLSGLIKAWFREL 236
LRMQK+LY+EGGLKAEGIFRI +N +E VR QLN GVVP GIDVHCL+GLIKAWFREL
Sbjct: 171 LRMQKRLYTEGGLKAEGIFRINPDNGKEEHVRRQLNCGVVPRGIDVHCLAGLIKAWFREL 230
Query: 237 PTGVLDSLTPEQVMQCNTEDDCTNLVKLLPSTEAALLDWAINLMADVVENEQFNKMNARN 296
PTGVLD LTPEQVM+CNTE+DC+ LV LLP E+A+LDWAI LMADVVE+EQFNKMNARN
Sbjct: 231 PTGVLDVLTPEQVMRCNTEEDCSRLVILLPPVESAILDWAIGLMADVVEHEQFNKMNARN 290
Query: 297 VAMVFAPNMTQMADPLTALIHAVQVMNFLKTLILKMLREREESIDNARLLSHSMDFASCN 356
VAMVFAPNMTQMADPLTALIHAVQVMNFLKTLIL L+ERE + AR L S
Sbjct: 291 VAMVFAPNMTQMADPLTALIHAVQVMNFLKTLILMNLKERENADAKARWLKKQTSDPS-- 348
Query: 357 DEFPPFSFNKEESSCEQIEDACDKNSSTTKRKFSRTSTLGRIEWSVEKLRSSEEKRNREE 416
EE + E + + KF R +TL R+E E+ + +KRN E
Sbjct: 349 ----------EEWESQHSEILSPEKPNNNNPKFLRVATLCRLEADNEEEFWNIKKRNDHE 398
Query: 417 VFRSFSSDSVTPRYESGPLENSYSYRRRYDSEHWSRLRNGVRKLCRHPVFQLSKPSKK 474
SS + GP V++LC+HP+FQLSK +KK
Sbjct: 399 GVLDTSSGN----GNIGP----------------------VQRLCKHPLFQLSKSTKK 430
>UniRef100_Q6UQ72 Rho GTPase activating protein 2 [Oryza sativa]
Length = 439
Score = 399 bits (1025), Expect = e-109
Identities = 223/414 (53%), Positives = 273/414 (65%), Gaps = 20/414 (4%)
Query: 83 AILDIVMAALKKSIVTCSVER--ED---------VSSLDISWPTEVRHVSHVTFDRFNGF 131
++++ V AAL++S++ CS R ED + + I PT+VRHVSHVTFDRF GF
Sbjct: 25 SVVETVAAALRRSLLLCSSVRAAEDEGAAAAAAAAAGMQIGRPTDVRHVSHVTFDRFVGF 84
Query: 132 LGLPSEFQPEVPTRVPSASVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKA 191
LGLP++ +P+VP PSASV VFGVS SMQCSYD+RGNSVPTILL MQK+LY GGL+A
Sbjct: 85 LGLPADLEPDVPRPAPSASVSVFGVSPTSMQCSYDNRGNSVPTILLTMQKKLYQLGGLQA 144
Query: 192 EGIFRITAENSQEAFVRDQLNKGVVPHGIDVHCLSGLIKAWFRELPTGVLDSLTPEQVMQ 251
EGIFRI A+NSQE VR+QLN GVVP G+D+HCL+GLIKAWFRELP+GVLDSLTPEQVM
Sbjct: 145 EGIFRINADNSQELHVREQLNMGVVPDGVDMHCLTGLIKAWFRELPSGVLDSLTPEQVMH 204
Query: 252 CNTEDDCTNLVKLLPSTEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNMTQMADP 311
CNTE++C L LP EAALLDWAINLMADVVE+E +NKMNARN+AMVFAPNMTQMADP
Sbjct: 205 CNTEEECALLASTLPPVEAALLDWAINLMADVVEHENYNKMNARNIAMVFAPNMTQMADP 264
Query: 312 LTALIHAVQVMNFLKTLILKMLREREESIDNARLLSHSMDFASCNDEFPPFSFNKEESSC 371
LTALIHAVQVMNFLKTLILK ++ REE+ + S S DE + + C
Sbjct: 265 LTALIHAVQVMNFLKTLILKTVKGREETAMPSSAFPSSSGSPSDKDEPQALEHLDKPTIC 324
Query: 372 --EQIEDACDKNSSTTKRKFSRTSTLGRIEWSVE----KLRSSEEKRNREEVFRSFSSDS 425
+Q D + +T R L + K R +++ N F SDS
Sbjct: 325 STQQNNDFPMISGATLDHFLFRAEPLRHNDAQGSAGRPKKRDNKDHDNSSREFSPIDSDS 384
Query: 426 VTPRYESGPLENSYSYRRRYDSEHWSRLRNGVRKLCRHPVFQLSKPSKKPASLG 479
+ S ++ + +D + R GV +LCRHPVFQLS+ KK G
Sbjct: 385 SSQASNSASKFSNDNVEGLFDR---FKFRKGVGRLCRHPVFQLSRSMKKSGEAG 435
>UniRef100_Q7XUH1 OSJNBa0020J04.11 protein [Oryza sativa]
Length = 479
Score = 397 bits (1020), Expect = e-109
Identities = 232/471 (49%), Positives = 285/471 (60%), Gaps = 44/471 (9%)
Query: 28 NNKKKEEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQFTKGSNNNHQNQFAILDI 87
++ ++EE E EE E EE+ SDD + S Q KG ++
Sbjct: 17 SSDEEEEDDEDEEGFEYEEILSDDGTD-------SPPPLMMQAEKGGGG-------LVGA 62
Query: 88 VMAALKKSIVTCSV---------EREDVSSLDISWPTEVRHVSHVTFDRFNGFLGLPSEF 138
V+ AL++S+V CS E E+ ++I PT+VRHVSHVTFDRF GFLGLP++
Sbjct: 63 VVGALRRSLVMCSAGKVGEEEDSEDEEEEGMEIGRPTDVRHVSHVTFDRFGGFLGLPADL 122
Query: 139 QPEVPTRVPSASVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRIT 198
+PEVP+ PSASV VFGVS S+QCS+D +GNSVPTILL MQ++LY GLK EGIFRI
Sbjct: 123 EPEVPSPTPSASVNVFGVSPTSLQCSFDHKGNSVPTILLMMQRKLYEREGLKIEGIFRIN 182
Query: 199 AENSQEAFVRDQLNKGVVPHGIDVHCLSGLIKAWFRELPTGVLDSLTPEQVMQCNTEDDC 258
AENSQE VR QLN GVVP +D+HCL+GLIKAWFRELPTGVLDSLTPEQVM CNTE+DC
Sbjct: 183 AENSQEICVRKQLNSGVVPDEVDLHCLAGLIKAWFRELPTGVLDSLTPEQVMHCNTEEDC 242
Query: 259 TNLVKLLPSTEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNMTQMADPLTALIHA 318
L +LP EAALLDWAINLMADVVE+E +NKMNARN+AMVFAPNMTQMADPLTALIHA
Sbjct: 243 ALLASMLPPVEAALLDWAINLMADVVEHENYNKMNARNIAMVFAPNMTQMADPLTALIHA 302
Query: 319 VQVMNFLKTLILKMLREREESIDNARLLSHSMDFASCNDEFPPFSFNKEESSCEQIEDAC 378
VQVMNFLKTLILK L+ERE + + + + PP N E C
Sbjct: 303 VQVMNFLKTLILKTLKEREAA--GTPKTTEPCSGSPNGQDKPPTPENLERPI------IC 354
Query: 379 DKNSSTTKRKFSRTSTLGRIEWSVEKLRSSEEKRNREEVFRSFSSDSVTPRYESGPLENS 438
K F +T ++ + ++ E N+ E E+ PL N
Sbjct: 355 SDQKGIDKPMFD-MATCDQLLFGPKQFLDHRE-NNKFEGPEKHDIGQPKRHSEASPLGND 412
Query: 439 YSYRRRYDSEHWSR-----------LRNGVRKLCRHPVFQLSKPSKKPASL 478
+ + + + R GV +LCRHPVFQLS+ KK A +
Sbjct: 413 SNNQVSSPGKEFGNRNVEGLFDKFSFRKGVERLCRHPVFQLSRSMKKSADV 463
>UniRef100_Q9FMR1 Rac GTPase activating protein [Arabidopsis thaliana]
Length = 466
Score = 386 bits (992), Expect = e-106
Identities = 201/342 (58%), Positives = 252/342 (72%), Gaps = 8/342 (2%)
Query: 51 DNDNDDDDEIYSSNNASRQFTKGSNNNH--QNQFAILDIVMAALKKSIVTCSVEREDVSS 108
D D D I + R T G + ++Q ++L +++A ++S+++C R ++ S
Sbjct: 55 DQDVDFHASIEEQDLRRRSSTDGGEEDDGGEDQISLLALLVAIFRRSLISCKSNRRELCS 114
Query: 109 LDISWPTEVRHVSHVTFDRFNGFLGLPSEFQPEVPTRVPSASVKVFGVSAKSMQCSYDDR 168
++I WPT VRHV+HVTFDRFNGFLGLP EF+PEVP R PSAS VFGVS +SMQ SYD R
Sbjct: 115 MEIGWPTNVRHVAHVTFDRFNGFLGLPVEFEPEVPRRAPSASATVFGVSTESMQLSYDSR 174
Query: 169 GNSVPTILLRMQKQLYSEGGLKAEGIFRITAENSQEAFVRDQLNKGVVPHGIDVHCLSGL 228
GN VPTILL MQ LYS+GGL+AEGIFR+TAENS+E VR+QLN+G +P IDVHCL+GL
Sbjct: 175 GNCVPTILLLMQNCLYSQGGLQAEGIFRLTAENSEEEAVREQLNRGFIPERIDVHCLAGL 234
Query: 229 IKAWFRELPTGVLDSLTPEQVMQCNTEDDCTNLVKLLPSTEAALLDWAINLMADVVENEQ 288
IKAWFRELPT VLDSL+PEQVMQC TE++ LV+LLP TEAALLDWAINLMADVV+ E
Sbjct: 235 IKAWFRELPTSVLDSLSPEQVMQCQTEEENVELVRLLPPTEAALLDWAINLMADVVQYEH 294
Query: 289 FNKMNARNVAMVFAPNMTQMADPLTALIHAVQVMNFLKTLILKMLREREES-IDNARLLS 347
NKMN+RN+AMVFAPNMTQM DPLTAL++AVQVMNFLKTLI K LRER++S ++ A
Sbjct: 295 LNKMNSRNIAMVFAPNMTQMDDPLTALMYAVQVMNFLKTLIEKTLRERQDSVVEQAHAFP 354
Query: 348 -HSMDFASCNDEFPPFSFN----KEESSCEQIEDACDKNSST 384
D + +FN EE+ + IE+A +++SS+
Sbjct: 355 LEPSDESGHQSPSQSLAFNTSEQSEETQSDNIENAENQSSSS 396
>UniRef100_O82458 Rac GTPase activating protein 1 [Lotus japonicus]
Length = 493
Score = 382 bits (981), Expect = e-104
Identities = 211/390 (54%), Positives = 260/390 (66%), Gaps = 34/390 (8%)
Query: 40 EEDEEEELESDDNDNDDDDEIYSSNNASRQFTKGSNNNHQNQFAILDIVMAALKKSIV-T 98
EEDEEEE E D +Q ++L +++A L+KS++ +
Sbjct: 46 EEDEEEEKEKD--------------------------REGDQLSLLTLLIATLRKSLIGS 79
Query: 99 CSVEREDV-----SSLDISWPTEVRHVSHVTFDRFNGFLGLPSEFQPEVPTRVPSASVKV 153
CS D SS++I WP+ VRHV+HVTFDRF+GFLGLP EF+PEVP R PSAS V
Sbjct: 80 CSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRFHGFLGLPVEFEPEVPRRPPSASTSV 139
Query: 154 FGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRITAENSQEAFVRDQLNK 213
FGVS +SMQ S+D RGNSVPTILL MQ+ LY+ GGL+AEGIFRI AENSQE VR+QLN+
Sbjct: 140 FGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVREQLNR 199
Query: 214 GVVPHGIDVHCLSGLIKAWFRELPTGVLDSLTPEQVMQCNTEDDCTNLVKLLPSTEAALL 273
GVVP+G+DVHCL+GLIKAWFRELPTG+LD L+PE+VMQ +E++C LV+LLP TEAALL
Sbjct: 200 GVVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQSQSEEECDQLVRLLPPTEAALL 259
Query: 274 DWAINLMADVVENEQFNKMNARNVAMVFAPNMTQMADPLTALIHAVQVMNFLKTLILKML 333
DWAINLMADV + E FNKMNARN+AMVFAPNMT MADPLTAL++AVQVMNFLKTL++K L
Sbjct: 260 DWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHMADPLTALMYAVQVMNFLKTLVVKTL 319
Query: 334 REREESIDNARLLSHSMDFASCNDEFPPFSFNKEESSCEQIEDACDKNSSTTKRKFSRTS 393
R REESI + + + F + K+ S E D D+++ + S+ S
Sbjct: 320 RVREESIVKSNPVPNLNSFDDDGHQSDSQVLPKDGS--ENGNDCSDEDTVFVSAEPSQPS 377
Query: 394 TLGRIEWSVEKLRSSEEKRNREEVFRSFSS 423
E E SE E F S S
Sbjct: 378 PTHHTEDGCETESGSETSPTPAENFLSSGS 407
>UniRef100_Q9CAX8 Putative rac GTPase activating protein; 62102-60058 [Arabidopsis
thaliana]
Length = 435
Score = 380 bits (976), Expect = e-104
Identities = 201/367 (54%), Positives = 255/367 (68%), Gaps = 36/367 (9%)
Query: 28 NNKKKEEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQFTKGSNNNHQNQFAILDI 87
N ++EE++E+EE E+E E + ++A L+I
Sbjct: 40 NISRREEEEEEEERSEKER------------ERFELSSA------------------LEI 69
Query: 88 VMAALKKSIVTCSVEREDVSSLDISWPTEVRHVSHVTFDRFNGFLGLPSEFQPEVPTRVP 147
+++A+++S++ V ED+ S++I PT+VRHV+HVTFDRF+GFLGLP EF+PEVP R P
Sbjct: 70 LVSAIRRSVIGGCVGEEDLCSMEIGVPTDVRHVAHVTFDRFHGFLGLPVEFEPEVPRRAP 129
Query: 148 SASVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRITAENSQEAFV 207
SAS VFGVS +SMQ SYD RGN VPTILL MQ LYS GGL+ EGIFRI EN QE ++
Sbjct: 130 SASATVFGVSTESMQLSYDTRGNIVPTILLMMQSHLYSRGGLRVEGIFRINGENGQEEYI 189
Query: 208 RDQLNKGVVPHGIDVHCLSGLIKAWFRELPTGVLDSLTPEQVMQCNTEDDCTNLVKLLPS 267
R++LNKG++P IDVHCL+ LIKAWFRELP+GVLDSL+PEQVM+ +ED+C LV+LLPS
Sbjct: 190 REELNKGIIPDNIDVHCLASLIKAWFRELPSGVLDSLSPEQVMESESEDECVELVRLLPS 249
Query: 268 TEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNMTQMADPLTALIHAVQVMNFLKT 327
TEA+LLDWAINLMADVVE EQ NKMNARN+AMVFAPNMTQM DPLTAL++AVQVMNFLKT
Sbjct: 250 TEASLLDWAINLMADVVEMEQLNKMNARNIAMVFAPNMTQMLDPLTALMYAVQVMNFLKT 309
Query: 328 LILKMLREREESID------NARLLSHSMDFASCNDEFPPFSFNKEESSCEQIEDACDKN 381
LI+K L++R+ES D N H+ D +S NKEE+ + DK
Sbjct: 310 LIVKTLKDRKESRDKLVPASNPSPRDHNGDQSSSRQLLHLMKANKEETLDNFEAEMKDKE 369
Query: 382 SSTTKRK 388
S + +
Sbjct: 370 ESADEEE 376
>UniRef100_Q9ZNR7 Putative rac GTPase activating protein [Arabidopsis thaliana]
Length = 424
Score = 378 bits (970), Expect = e-103
Identities = 190/321 (59%), Positives = 242/321 (75%), Gaps = 20/321 (6%)
Query: 78 HQNQFAILDIVMAALKKSIVTCSVE-RED-----------VSSLDISWPTEVRHVSHVTF 125
+Q Q ++++ ++ AL+KS+V+C V+ R+D V ++I WPT VRH++HVTF
Sbjct: 29 NQQQLSLVEFLLTALRKSVVSCRVDNRQDDGGVGGGISSAVHHMEIGWPTNVRHITHVTF 88
Query: 126 DRFNGFLGLPSEFQPEVPTRVPSASVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYS 185
DRF+GFLGLP E Q E+P RVPSASV VFGVSA+SMQCSYD++GNSVPTILL MQ++LYS
Sbjct: 89 DRFHGFLGLPHELQVEIPCRVPSASVSVFGVSAESMQCSYDEKGNSVPTILLLMQERLYS 148
Query: 186 EGGLKAEGIFRITAENSQEAFVRDQLNKGVVPHGIDVHCLSGLIKAWFRELPTGVLDSLT 245
+ GLKAEGIFRI ENSQE VRDQLN+G+VP IDVHCL+GLIKAWFRELP+GVLD L+
Sbjct: 149 QQGLKAEGIFRINPENSQEEHVRDQLNRGIVPENIDVHCLAGLIKAWFRELPSGVLDGLS 208
Query: 246 PEQVMQCNTEDDCTNLVKLLPSTEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNM 305
PE+V+ CNTED+ L+K L TE+ALL+WA++LMADVVE E+ NKMNARN+AMVFAPNM
Sbjct: 209 PEEVLNCNTEDESVELIKQLKPTESALLNWAVDLMADVVEEEESNKMNARNIAMVFAPNM 268
Query: 306 TQMADPLTALIHAVQVMNFLKTLILKMLREREESIDNARLLSHSMDFASCNDEFPPFSFN 365
TQM DPLTAL+HAVQVMN LKTLI K L EREE+ + S S S D
Sbjct: 269 TQMTDPLTALMHAVQVMNLLKTLITKTLAEREENATGSEGYSPSHSSNSQTD-------- 320
Query: 366 KEESSCEQIEDACDKNSSTTK 386
+ + + +E +C+ ++ ++
Sbjct: 321 SDSDNAQDMEVSCESQATDSE 341
>UniRef100_O82460 Rac GTPase activating protein 3 [Lotus japonicus]
Length = 432
Score = 377 bits (967), Expect = e-103
Identities = 204/356 (57%), Positives = 254/356 (71%), Gaps = 12/356 (3%)
Query: 79 QNQFAILDIVMAALKKSIVTCSVERED-----VSSLDISWPTEVRHVSHVTFDRFNGFLG 133
QNQ + ++AALKKS+V CSV+ D V ++I WPT V+HV+HVTFDRFNGFLG
Sbjct: 5 QNQGSPAAFLLAALKKSMVACSVDSPDDVISAVHPMEIGWPTNVKHVTHVTFDRFNGFLG 64
Query: 134 LPSEFQPEVPTRVPSASVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEG 193
LP E + VP VPSASV VFGVSA+SM CSYD +GNSVPTILL MQ++LYS+GGL AEG
Sbjct: 65 LPLELEVHVPAPVPSASVSVFGVSAESMHCSYDSKGNSVPTILLLMQERLYSQGGLMAEG 124
Query: 194 IFRITAENSQEAFVRDQLNKGVVPHGIDVHCLSGLIKAWFRELPTGVLDSLTPEQVMQCN 253
IFRI EN QE +RDQLN+GVVP IDVHCL+GLIKAWFRELP+GVLD L+PEQV++CN
Sbjct: 125 IFRINPENGQEEHLRDQLNRGVVPDNIDVHCLAGLIKAWFRELPSGVLDGLSPEQVLECN 184
Query: 254 TEDDCTNLVKLLPSTEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNMTQMADPLT 313
TE++ LVK L TE ALL+WA++LMADVVE E+ NKM+ARN+AMVFAPNMTQM+DPLT
Sbjct: 185 TEEEFVQLVKQLKPTELALLNWALDLMADVVEEEEHNKMDARNIAMVFAPNMTQMSDPLT 244
Query: 314 ALIHAVQVMNFLKTLILKMLREREE--SIDNARLLSHSMDFASCNDEFPP----FSFNKE 367
AL+HAVQVMN LKTLILK L EREE + + + SHS D S DE+ ++ +
Sbjct: 245 ALMHAVQVMNLLKTLILKTLSEREEATTAGYSSMSSHSSDRQS-EDEYDSQLEMYTSAEL 303
Query: 368 ESSCEQIEDACDKNSSTTKRKFSRTSTLGRIEWSVEKLRSSEEKRNREEVFRSFSS 423
S +D + NS ++ + S E +++L +++ EE RS S
Sbjct: 304 RGSQSDCDDHVNNNSLNSEEEVDSASVSEIEECFLKQLNENKQGGFAEEPARSTCS 359
>UniRef100_Q7XAL7 Rac GTPase activating protein 3-like protein [Oryza sativa]
Length = 487
Score = 363 bits (931), Expect = 8e-99
Identities = 185/302 (61%), Positives = 227/302 (74%), Gaps = 9/302 (2%)
Query: 73 GSNNNHQNQFAILDIVMAALKKSIVT-CSVEREDVSS-----LDISWPTEVRHVSHVTFD 126
G Q +L +++AAL++S+V C + D + ++I WPT+VRHV+HVTFD
Sbjct: 32 GEGKAEGQQGQVLALLLAALRRSVVLPCQMADADDPAAVAWGMEIGWPTDVRHVAHVTFD 91
Query: 127 RFNGFLGLPSEFQPEVPTRVPSASVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSE 186
R NGFLGLP+EF+ E+P VPSAS VFGVS +SMQC +DD GNSVP ILL MQ++LY++
Sbjct: 92 RLNGFLGLPAEFELEIPGHVPSASASVFGVSPESMQCCFDDNGNSVPKILLLMQERLYAQ 151
Query: 187 GGLKAEGIFRITAENSQEAFVRDQLNKGVVPHGIDVHCLSGLIKAWFRELPTGVLDSLTP 246
GLKAEGIFRIT ENSQE VR+QLN+G+VP IDVHCL+ LIKAWFRELP GVLDSL+P
Sbjct: 152 DGLKAEGIFRITPENSQEENVREQLNRGLVPDDIDVHCLASLIKAWFRELPEGVLDSLSP 211
Query: 247 EQVMQCNTEDDCTNLVKLLPSTEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNMT 306
EQV+ CNTE++C LV+LLP T+AALL+W + MADVVE E+ NKMNARNVAMVFAPNMT
Sbjct: 212 EQVLHCNTEEECVELVRLLPPTQAALLNWVVEFMADVVEEEESNKMNARNVAMVFAPNMT 271
Query: 307 QMADPLTALIHAVQVMNFLKTLILKMLREREESIDNARLLSHSMDFASCNDEFPPFSFNK 366
QM+DPLTAL+HAVQVMN LKTLILK LRERE +S +S +DE +
Sbjct: 272 QMSDPLTALMHAVQVMNLLKTLILKTLREREHDESEYSAISSQ---SSSSDELDEMHHHV 328
Query: 367 EE 368
E+
Sbjct: 329 EQ 330
>UniRef100_Q6UQ73 Rho GTPase activating protein 1 [Oryza sativa]
Length = 364
Score = 360 bits (923), Expect = 7e-98
Identities = 173/231 (74%), Positives = 203/231 (86%)
Query: 109 LDISWPTEVRHVSHVTFDRFNGFLGLPSEFQPEVPTRVPSASVKVFGVSAKSMQCSYDDR 168
++I WPT+VRHV+HVTFDRF+GFLGLP EF+ E+P RVPSAS VFGVSA+SMQC+YD +
Sbjct: 1 MEIGWPTDVRHVAHVTFDRFHGFLGLPVEFEVEMPCRVPSASASVFGVSAESMQCTYDGK 60
Query: 169 GNSVPTILLRMQKQLYSEGGLKAEGIFRITAENSQEAFVRDQLNKGVVPHGIDVHCLSGL 228
GNSVPTILL MQ++LY++GGLKAEGIFRI EN QE VRDQLNKGVVP IDVHCL+ L
Sbjct: 61 GNSVPTILLHMQERLYAQGGLKAEGIFRINPENDQEEHVRDQLNKGVVPEDIDVHCLASL 120
Query: 229 IKAWFRELPTGVLDSLTPEQVMQCNTEDDCTNLVKLLPSTEAALLDWAINLMADVVENEQ 288
IKAWFRELP GVLDSL+PEQV+QCN+E + LV LL T+AALL+WA+ LMADVVE E+
Sbjct: 121 IKAWFRELPEGVLDSLSPEQVLQCNSEGEFLELVTLLRPTQAALLNWAVELMADVVEEEE 180
Query: 289 FNKMNARNVAMVFAPNMTQMADPLTALIHAVQVMNFLKTLILKMLREREES 339
NKMNARN+AMVFAPNMTQM+DPLTAL+HAVQVMNFLKTLIL+ LRER+++
Sbjct: 181 LNKMNARNIAMVFAPNMTQMSDPLTALMHAVQVMNFLKTLILRTLRERDDA 231
>UniRef100_Q9SGC6 T23G18.20 [Arabidopsis thaliana]
Length = 409
Score = 338 bits (867), Expect = 2e-91
Identities = 166/257 (64%), Positives = 204/257 (78%), Gaps = 5/257 (1%)
Query: 88 VMAALKKSIVTCSVEREDVSSLDISWPTEVRHVSHVTFDRFNGFLGLPSEFQPEVPTRVP 147
V+ L + T VE D ++DI PT +RHV+HVTFDRF+GFLGLPSEF+P+VP + P
Sbjct: 53 VVLELARCFSTAEVEDNDRPAMDIGGPTNIRHVAHVTFDRFDGFLGLPSEFEPDVPRKAP 112
Query: 148 SASVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRITAENSQEAFV 207
SAS VFGVS +SMQ SYD RGN VP ILL +Q +LY +GGL+AEG+FRIT ENS+E FV
Sbjct: 113 SASATVFGVSTESMQLSYDSRGNCVPVILLLLQSRLYDQGGLQAEGVFRITGENSEEEFV 172
Query: 208 RDQLNKGVVPHGIDVHCLSGLIK-----AWFRELPTGVLDSLTPEQVMQCNTEDDCTNLV 262
R+QLNKG++P GIDVHCL+GLIK AWFRELP GVLD L EQVMQC +++D +V
Sbjct: 173 REQLNKGIIPDGIDVHCLAGLIKVLVVIAWFRELPRGVLDPLPSEQVMQCESDEDFVKVV 232
Query: 263 KLLPSTEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNMTQMADPLTALIHAVQVM 322
+LLP TEA+LL+WAINLMADV++ E NKMN+RN+A+VFAPNM+QMADPLTAL++AVQVM
Sbjct: 233 RLLPQTEASLLNWAINLMADVIQFEHVNKMNSRNLALVFAPNMSQMADPLTALMYAVQVM 292
Query: 323 NFLKTLILKMLREREES 339
LK+L K +RERE S
Sbjct: 293 KLLKSLTEKTVREREAS 309
Score = 40.4 bits (93), Expect = 0.12
Identities = 16/29 (55%), Positives = 23/29 (79%)
Query: 31 KKEEQQEQEEEDEEEELESDDNDNDDDDE 59
+KE+ E+EEEDEEEE E +D D D+++E
Sbjct: 325 EKEKDNEEEEEDEEEEEEEEDEDEDEEEE 353
Score = 36.6 bits (83), Expect = 1.7
Identities = 13/38 (34%), Positives = 27/38 (70%)
Query: 24 AQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDEIY 61
A+ +K E++E++EE+EEEE + D+++ ++ D +Y
Sbjct: 321 AEDGEKEKDNEEEEEDEEEEEEEEDEDEDEEEEGDGVY 358
Score = 35.4 bits (80), Expect = 3.7
Identities = 16/51 (31%), Positives = 27/51 (52%)
Query: 30 KKKEEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQFTKGSNNNHQN 80
K EE++E EEE+EEEE E +D + + D + + K + H++
Sbjct: 328 KDNEEEEEDEEEEEEEEDEDEDEEEEGDGVYIIKEEEASEIIKVVADEHKS 378
>UniRef100_Q6NKT5 At1g08340 [Arabidopsis thaliana]
Length = 331
Score = 337 bits (863), Expect = 6e-91
Identities = 160/231 (69%), Positives = 195/231 (84%)
Query: 109 LDISWPTEVRHVSHVTFDRFNGFLGLPSEFQPEVPTRVPSASVKVFGVSAKSMQCSYDDR 168
+DI PT +RHV+HVTFDRF+GFLGLPSEF+P+VP + PSAS VFGVS +SMQ SYD R
Sbjct: 1 MDIGGPTNIRHVAHVTFDRFDGFLGLPSEFEPDVPRKAPSASATVFGVSTESMQLSYDSR 60
Query: 169 GNSVPTILLRMQKQLYSEGGLKAEGIFRITAENSQEAFVRDQLNKGVVPHGIDVHCLSGL 228
GN VP ILL +Q +LY +GGL+AEG+FRIT ENS+E FVR+QLNKG++P GIDVHCL+GL
Sbjct: 61 GNCVPVILLLLQSRLYDQGGLQAEGVFRITGENSEEEFVREQLNKGIIPDGIDVHCLAGL 120
Query: 229 IKAWFRELPTGVLDSLTPEQVMQCNTEDDCTNLVKLLPSTEAALLDWAINLMADVVENEQ 288
IKAWFRELP GVLD L EQVMQC +++D +V+LLP TEA+LL+WAINLMADV++ E
Sbjct: 121 IKAWFRELPRGVLDPLPSEQVMQCESDEDFVKVVRLLPQTEASLLNWAINLMADVIQFEH 180
Query: 289 FNKMNARNVAMVFAPNMTQMADPLTALIHAVQVMNFLKTLILKMLREREES 339
NKMN+RN+A+VFAPNM+QMADPLTAL++AVQVM LK+L K +RERE S
Sbjct: 181 VNKMNSRNLALVFAPNMSQMADPLTALMYAVQVMKLLKSLTEKTVREREAS 231
Score = 40.4 bits (93), Expect = 0.12
Identities = 16/29 (55%), Positives = 23/29 (79%)
Query: 31 KKEEQQEQEEEDEEEELESDDNDNDDDDE 59
+KE+ E+EEEDEEEE E +D D D+++E
Sbjct: 247 EKEKDNEEEEEDEEEEEEEEDEDEDEEEE 275
Score = 36.6 bits (83), Expect = 1.7
Identities = 13/38 (34%), Positives = 27/38 (70%)
Query: 24 AQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDEIY 61
A+ +K E++E++EE+EEEE + D+++ ++ D +Y
Sbjct: 243 AEDGEKEKDNEEEEEDEEEEEEEEDEDEDEEEEGDGVY 280
Score = 35.4 bits (80), Expect = 3.7
Identities = 16/51 (31%), Positives = 27/51 (52%)
Query: 30 KKKEEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQFTKGSNNNHQN 80
K EE++E EEE+EEEE E +D + + D + + K + H++
Sbjct: 250 KDNEEEEEDEEEEEEEEDEDEDEEEEGDGVYIIKEEEASEIIKVVADEHKS 300
>UniRef100_Q9SJF6 T27G7.4 [Arabidopsis thaliana]
Length = 420
Score = 330 bits (845), Expect = 7e-89
Identities = 166/268 (61%), Positives = 204/268 (75%), Gaps = 16/268 (5%)
Query: 88 VMAALKKSIVTCSVEREDVSSLDISWPTEVRHVSHVTFDRFNGFLGLPSEFQPEVPTRVP 147
V+ L + T VE D ++DI PT +RHV+HVTFDRF+GFLGLPSEF+P+VP + P
Sbjct: 53 VVLELARCFSTAEVEDNDRPAMDIGGPTNIRHVAHVTFDRFDGFLGLPSEFEPDVPRKAP 112
Query: 148 SA-----------SVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFR 196
SA S VFGVS +SMQ SYD RGN VP ILL +Q +LY +GGL+AEG+FR
Sbjct: 113 SARFHIIILFVFGSATVFGVSTESMQLSYDSRGNCVPVILLLLQSRLYDQGGLQAEGVFR 172
Query: 197 ITAENSQEAFVRDQLNKGVVPHGIDVHCLSGLIK-----AWFRELPTGVLDSLTPEQVMQ 251
IT ENS+E FVR+QLNKG++P GIDVHCL+GLIK AWFRELP GVLD L EQVMQ
Sbjct: 173 ITGENSEEEFVREQLNKGIIPDGIDVHCLAGLIKVLVVIAWFRELPRGVLDPLPSEQVMQ 232
Query: 252 CNTEDDCTNLVKLLPSTEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNMTQMADP 311
C +++D +V+LLP TEA+LL+WAINLMADV++ E NKMN+RN+A+VFAPNM+QMADP
Sbjct: 233 CESDEDFVKVVRLLPQTEASLLNWAINLMADVIQFEHVNKMNSRNLALVFAPNMSQMADP 292
Query: 312 LTALIHAVQVMNFLKTLILKMLREREES 339
LTAL++AVQVM LK+L K +RERE S
Sbjct: 293 LTALMYAVQVMKLLKSLTEKTVREREAS 320
Score = 40.4 bits (93), Expect = 0.12
Identities = 16/29 (55%), Positives = 23/29 (79%)
Query: 31 KKEEQQEQEEEDEEEELESDDNDNDDDDE 59
+KE+ E+EEEDEEEE E +D D D+++E
Sbjct: 336 EKEKDNEEEEEDEEEEEEEEDEDEDEEEE 364
Score = 36.6 bits (83), Expect = 1.7
Identities = 13/38 (34%), Positives = 27/38 (70%)
Query: 24 AQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDEIY 61
A+ +K E++E++EE+EEEE + D+++ ++ D +Y
Sbjct: 332 AEDGEKEKDNEEEEEDEEEEEEEEDEDEDEEEEGDGVY 369
Score = 35.4 bits (80), Expect = 3.7
Identities = 16/51 (31%), Positives = 27/51 (52%)
Query: 30 KKKEEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQFTKGSNNNHQN 80
K EE++E EEE+EEEE E +D + + D + + K + H++
Sbjct: 339 KDNEEEEEDEEEEEEEEDEDEDEEEEGDGVYIIKEEEASEIIKVVADEHKS 389
>UniRef100_Q9ZUZ1 Putative rac GTPase activating protein [Arabidopsis thaliana]
Length = 301
Score = 324 bits (830), Expect = 4e-87
Identities = 184/314 (58%), Positives = 211/314 (66%), Gaps = 38/314 (12%)
Query: 161 MQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRITAENSQEAFVRDQLNKGVVPHGI 220
MQCSYDDRGNSVPTILLRMQK+LY+EGGLKAEGIFRI +N +E VR QLN GVVP GI
Sbjct: 1 MQCSYDDRGNSVPTILLRMQKRLYTEGGLKAEGIFRINPDNGKEEHVRRQLNCGVVPRGI 60
Query: 221 DVHCLSGLIKAWFRELPTGVLDSLTPEQVMQCNTEDDCTNLVKLLPSTEAALLDWAINLM 280
DVHCL+GLIKAWFRELPTGVLD LTPEQVM+CNTE+DC+ LV LLP E+A+LDWAI LM
Sbjct: 61 DVHCLAGLIKAWFRELPTGVLDVLTPEQVMRCNTEEDCSRLVILLPPVESAILDWAIGLM 120
Query: 281 ADVVENEQFNKMNARNVAMVFAPNMTQMADPLTALIHAVQVMNFLKTLILKMLREREESI 340
ADVVE+EQFNKMNARNVAMVFAPNMTQMADPLTALIHAVQVMNFLKTLIL L+ERE +
Sbjct: 121 ADVVEHEQFNKMNARNVAMVFAPNMTQMADPLTALIHAVQVMNFLKTLILMNLKERENAD 180
Query: 341 DNARLLSHSMDFASCNDEFPPFSFNKEESSCEQIEDACDKNSSTTKRKFSRTSTLGRIEW 400
AR L S EE + E + + KF R +TL R+E
Sbjct: 181 AKARWLKKQTSDPS------------EEWESQHSEILSPEKPNNNNPKFLRVATLCRLEA 228
Query: 401 SVEKLRSSEEKRNREEVFRSFSSDSVTPRYESGPLENSYSYRRRYDSEHWSRLRNGVRKL 460
E+ + +KRN E SS + GP V++L
Sbjct: 229 DNEEEFWNIKKRNDHEGVLDTSSGN----GNIGP----------------------VQRL 262
Query: 461 CRHPVFQLSKPSKK 474
C+HP+FQLSK +KK
Sbjct: 263 CKHPLFQLSKSTKK 276
>UniRef100_Q6EPP2 Putative Rho GTPase activating protein 2 [Oryza sativa]
Length = 326
Score = 319 bits (818), Expect = 1e-85
Identities = 178/325 (54%), Positives = 213/325 (64%), Gaps = 9/325 (2%)
Query: 161 MQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRITAENSQEAFVRDQLNKGVVPHGI 220
MQCSYD+RGNSVPTILL MQK+LY GGL+AEGIFRI A+NSQE VR+QLN GVVP G+
Sbjct: 1 MQCSYDNRGNSVPTILLTMQKKLYQLGGLQAEGIFRINADNSQELHVREQLNMGVVPDGV 60
Query: 221 DVHCLSGLIKAWFRELPTGVLDSLTPEQVMQCNTEDDCTNLVKLLPSTEAALLDWAINLM 280
D+HCL+GLIKAWFRELP+GVLDSLTPEQVM CNTE++C L LP EAALLDWAINLM
Sbjct: 61 DMHCLTGLIKAWFRELPSGVLDSLTPEQVMHCNTEEECALLASTLPPVEAALLDWAINLM 120
Query: 281 ADVVENEQFNKMNARNVAMVFAPNMTQMADPLTALIHAVQVMNFLKTLILKMLREREESI 340
ADVVE+E +NKMNARN+AMVFAPNMTQMADPLTALIHAVQVMNFLKTLILK ++ REE+
Sbjct: 121 ADVVEHENYNKMNARNIAMVFAPNMTQMADPLTALIHAVQVMNFLKTLILKTVKGREETA 180
Query: 341 DNARLLSHSMDFASCNDEFPPFSFNKEESSC--EQIEDACDKNSSTTKRKFSRTSTLGRI 398
+ S S DE + + C +Q D + +T R L
Sbjct: 181 MPSSAFPSSSGSPSDKDEPQALEHLDKPTICSTQQNNDFPMISGATLDHFLFRAEPLRHN 240
Query: 399 EWSVE----KLRSSEEKRNREEVFRSFSSDSVTPRYESGPLENSYSYRRRYDSEHWSRLR 454
+ K R +++ N F SDS + S ++ + +D + R
Sbjct: 241 DAQGSAGRPKKRDNKDHDNSSREFSPIDSDSSSQASNSASKFSNDNVEGLFDR---FKFR 297
Query: 455 NGVRKLCRHPVFQLSKPSKKPASLG 479
GV +LCRHPVFQLS+ KK G
Sbjct: 298 KGVGRLCRHPVFQLSRSMKKSGEAG 322
>UniRef100_Q8LHB9 Putative rac GTPase activating protein [Oryza sativa]
Length = 258
Score = 305 bits (782), Expect = 1e-81
Identities = 151/233 (64%), Positives = 179/233 (76%), Gaps = 1/233 (0%)
Query: 104 EDVSSLDISWPTEVRHVSHVTFDRFNGFLGLPSEFQPEVPTRVPSASVKVFGVSAKSMQC 163
E ++I WPT+VRHV+HVTFDRF+GF GLP E QPEV PSAS VFGVS +SMQC
Sbjct: 20 EQQPPMEIGWPTDVRHVAHVTFDRFHGFQGLPVELQPEVAGNAPSASKTVFGVSTESMQC 79
Query: 164 SYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRITAENSQEAFVRDQLNKGVVPH-GIDV 222
SYD RGNSVP+ILL MQ++LY +GGLKAEGIFRI A+++QE VR+QLN GV+P G+DV
Sbjct: 80 SYDARGNSVPSILLLMQRRLYEQGGLKAEGIFRIAADDAQEQAVREQLNSGVLPEGGVDV 139
Query: 223 HCLSGLIKAWFRELPTGVLDSLTPEQVMQCNTEDDCTNLVKLLPSTEAALLDWAINLMAD 282
HCL+GLIKAWFRELP G+LDSL +V +C + DDC L LP+ +AALLDWA+ LMAD
Sbjct: 140 HCLAGLIKAWFRELPGGMLDSLPAAEVTRCQSADDCARLCARLPAAKAALLDWAVQLMAD 199
Query: 283 VVENEQFNKMNARNVAMVFAPNMTQMADPLTALIHAVQVMNFLKTLILKMLRE 335
V E+ NKM +RNVAMVFAPNMT DP TAL HAV VMNFL LI + L +
Sbjct: 200 VAREERSNKMGSRNVAMVFAPNMTHAMDPFTALKHAVHVMNFLTMLIDRALND 252
>UniRef100_Q9ZQH5 Putative rac GTPase activating protein [Arabidopsis thaliana]
Length = 368
Score = 277 bits (709), Expect = 4e-73
Identities = 150/293 (51%), Positives = 195/293 (66%), Gaps = 35/293 (11%)
Query: 102 EREDVSSLDISWPTEVRHVSHVTFDRFNGFLGLPSEFQPEVPTRVPSASVKVFGVSAKSM 161
E + ++DIS PT + HV+HVT+DRF+GFLGLPSEF+P+VP + PSAS VFGVS +SM
Sbjct: 80 EENERHAMDISRPTNISHVAHVTYDRFDGFLGLPSEFEPDVPKKPPSASATVFGVSTESM 139
Query: 162 QCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRITAENSQEAFVRDQLNKGVVPHGID 221
Q SYD RGN VPTIL +Q +LY +GGL+ EGIFRIT +NS+E F+R++LNKGV+P GID
Sbjct: 140 QLSYDSRGNCVPTILTLLQSRLYDQGGLQVEGIFRITGDNSEEEFIREELNKGVLPEGID 199
Query: 222 VHCLSGLIKAWFRELPTGVLDSLTPEQVMQCNTEDDCTNLVKLLPSTEAALLDWAINLMA 281
+HCL+GLIKAWFRELP GVLDSL +QVMQC + +D VK+
Sbjct: 200 IHCLAGLIKAWFRELPKGVLDSLPSQQVMQCESGED---FVKVF---------------- 240
Query: 282 DVVENEQFNKMNARNVAMVFAPNMTQMADPLTALIHAVQVMNFLKTLILKMLREREESID 341
E NKM +RN+A+VFAPNM+QMADPLTAL++AVQVMN L+ L K LRER+ +
Sbjct: 241 -----EVVNKMTSRNLALVFAPNMSQMADPLTALMYAVQVMNLLRNLTDKTLRERKIATS 295
Query: 342 NARLLSHSMDFASCNDEFPPFSFNK-------EESSCEQIEDACDKNSSTTKR 387
N + + N E +N+ EE E+ ED D ++ ++R
Sbjct: 296 NVDPCDNRSEAEDGNVE----EYNQEVEIYVLEEVEEEEGEDVDDLDNEESER 344
>UniRef100_UPI00004996C7 UPI00004996C7 UniRef100 entry
Length = 324
Score = 99.4 bits (246), Expect = 2e-19
Identities = 59/197 (29%), Positives = 107/197 (53%), Gaps = 10/197 (5%)
Query: 153 VFGVSAKSMQC-SYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRITAENSQEAFVRDQL 211
VFGV +S++ + R +P I++ + +GG +EG+FR+ E ++++L
Sbjct: 125 VFGVDPESLEWFKHPTRDLILPQIIITLDIAFREKGGFTSEGVFRLAGEQGMVKSLKERL 184
Query: 212 NK--GVVPHGI---DVHCLSGLIKAWFRELPTGVLDSLTPEQVMQCNTEDDCTNLVKLLP 266
NK G++ + V +S LIK WFRELP +L+ L+ +Q+ + + L
Sbjct: 185 NKSNGIITEDMMDATVDDISNLIKLWFRELPRPILNVLSCDQIFYSTEPAESYKAFESLN 244
Query: 267 STEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNM--TQMADPLTALIHAVQVMNF 324
ALL W +LM +V +N + NKM +N+A+V APN+ + DP+ L+ + + + F
Sbjct: 245 EKSRALLTWLFDLMIEVAKNRETNKMTIQNLAIVIAPNLYEPESTDPMEGLVMSQKAVQF 304
Query: 325 LKTLI--LKMLREREES 339
++ ++ L+ L+E+ S
Sbjct: 305 VQNILNYLEALKEQNGS 321
>UniRef100_Q616X4 Hypothetical protein CBG15100 [Caenorhabditis briggsae]
Length = 646
Score = 80.9 bits (198), Expect = 8e-14
Identities = 54/163 (33%), Positives = 83/163 (50%), Gaps = 5/163 (3%)
Query: 172 VPTILLRMQKQLYSEGGLKAEGIFRITAENSQEAFVRDQLNKGVVPHGIDVHCLSGLIKA 231
+P +L + + LY GG + EGIFR+ + Q A R QL+ + P D + + L+K
Sbjct: 472 LPWLLTTLIELLYQSGGRRTEGIFRVAGDPEQLATARGQLDGWLAPKMHDANVPACLLKL 531
Query: 232 WFRELPTG-VLDSLTPEQVMQCNTEDDCTNLVKLLPSTEAALLDWAINLMADVVENEQF- 289
W R+LP +L +L ++ +T + LV LLP +L I L+ D+ E
Sbjct: 532 WLRQLPVPLILPNLYQRALVASDTPAEAIRLVDLLPDINRLVLVRVIALLQDLSREEVVA 591
Query: 290 -NKMNARNVAMVFAPNM--TQMADPLTALIHAVQVMNFLKTLI 329
KM+ N+AMV APN+ + DP + + M+FLK LI
Sbjct: 592 KTKMDTSNLAMVIAPNILRCESEDPRVIFENTRREMSFLKVLI 634
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.314 0.129 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 813,048,302
Number of Sequences: 2790947
Number of extensions: 35728853
Number of successful extensions: 530571
Number of sequences better than 10.0: 4156
Number of HSP's better than 10.0 without gapping: 2562
Number of HSP's successfully gapped in prelim test: 1662
Number of HSP's that attempted gapping in prelim test: 424464
Number of HSP's gapped (non-prelim): 41876
length of query: 486
length of database: 848,049,833
effective HSP length: 131
effective length of query: 355
effective length of database: 482,435,776
effective search space: 171264700480
effective search space used: 171264700480
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 77 (34.3 bits)
Medicago: description of AC144592.1


