

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144541.14 - phase: 0
(63 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
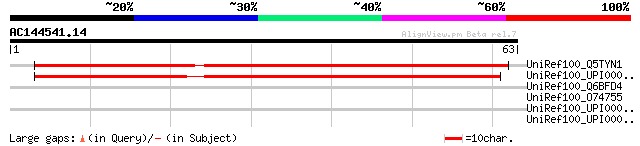
UniRef100_Q5TYN1 Novel protein [Brachydanio rerio] 58 6e-08
UniRef100_UPI000034D89F UPI000034D89F UniRef100 entry 54 1e-06
UniRef100_Q6BFD4 Hypothetical protein [Paramecium tetraurelia] 32 3.3
UniRef100_O74755 SIN3 family co-repressor; involved in transcrip... 32 4.3
UniRef100_UPI00002B84DF UPI00002B84DF UniRef100 entry 31 9.5
UniRef100_UPI000028A506 UPI000028A506 UniRef100 entry 31 9.5
>UniRef100_Q5TYN1 Novel protein [Brachydanio rerio]
Length = 175
Score = 58.2 bits (139), Expect = 6e-08
Identities = 28/59 (47%), Positives = 43/59 (72%), Gaps = 1/59 (1%)
Query: 4 ERSSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELLNFLNASF 62
+ SS PVQ+SK+L K G + S +SGKKI +K+ K+K+DK+ + R ELL+FLN+++
Sbjct: 118 DESSGPVQISKYLKEKKKSG-KYSMISGKKIKMKVKKSKKDKQRDKNRAELLDFLNSTY 175
>UniRef100_UPI000034D89F UPI000034D89F UniRef100 entry
Length = 160
Score = 53.9 bits (128), Expect = 1e-06
Identities = 28/58 (48%), Positives = 40/58 (68%), Gaps = 2/58 (3%)
Query: 4 ERSSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELLNFLNAS 61
E +S PVQ+SK+L K + S +SGKKI +K+ K+KEDK+ + R ELL FLN++
Sbjct: 104 EENSGPVQISKYLKDRKKG--KYSMISGKKIKMKVKKSKEDKKRDKNRAELLEFLNST 159
>UniRef100_Q6BFD4 Hypothetical protein [Paramecium tetraurelia]
Length = 899
Score = 32.3 bits (72), Expect = 3.3
Identities = 18/49 (36%), Positives = 26/49 (52%), Gaps = 2/49 (4%)
Query: 14 KFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELLNFLNASF 62
K L NK D + S K +L +D+ +D++ S RN L+N N SF
Sbjct: 767 KILKTNKQDS--KILTSDNKRILMVDELDKDQDVSSLRNSLVNLSNISF 813
>UniRef100_O74755 SIN3 family co-repressor; involved in transcriptional regulation
(Predicted); paired amphipathic helix; histone
deacetylase pathway (Predicted); similar to S. pombe
pst2 (Paralog); similar to S. cerevisiae YOL004W
[Schizosaccharomyces pombe]
Length = 1154
Score = 32.0 bits (71), Expect = 4.3
Identities = 17/42 (40%), Positives = 23/42 (54%)
Query: 5 RSSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKE 46
+S SPV S G KD+GV V K+IL + T++D E
Sbjct: 35 QSQSPVNTSLHNGDGKDNGVATEPVENKQILSERSVTRDDYE 76
>UniRef100_UPI00002B84DF UPI00002B84DF UniRef100 entry
Length = 174
Score = 30.8 bits (68), Expect = 9.5
Identities = 19/58 (32%), Positives = 27/58 (45%)
Query: 5 RSSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELLNFLNASF 62
RSSS S + R +SGK I LK ++ D ES R + L LN+++
Sbjct: 117 RSSSSSSSSSSSSESSSSERRYDHISGKVIKLKRKSSRVDDRLESNRLKTLKLLNSAY 174
>UniRef100_UPI000028A506 UPI000028A506 UniRef100 entry
Length = 156
Score = 30.8 bits (68), Expect = 9.5
Identities = 17/45 (37%), Positives = 27/45 (59%), Gaps = 1/45 (2%)
Query: 2 NGERSSSPVQL-SKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDK 45
NGE+S V + +KFLG D +RR+ + G+K+ K+ K + K
Sbjct: 25 NGEKSLVVVDVKNKFLGTISDGDIRRALLKGQKLSSKITKYFQKK 69
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.307 0.129 0.336
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 97,189,843
Number of Sequences: 2790947
Number of extensions: 3067054
Number of successful extensions: 9244
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 9238
Number of HSP's gapped (non-prelim): 6
length of query: 63
length of database: 848,049,833
effective HSP length: 39
effective length of query: 24
effective length of database: 739,202,900
effective search space: 17740869600
effective search space used: 17740869600
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC144541.14


