

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144539.4 - phase: 0
(587 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
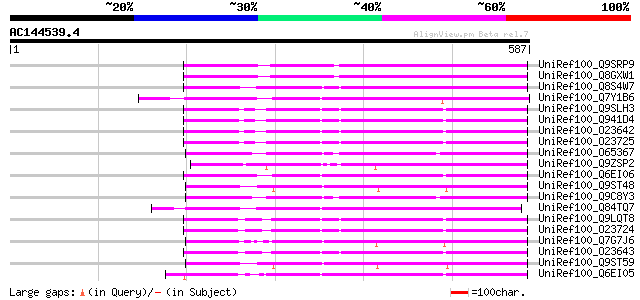
UniRef100_Q9SRP9 RGA1-like protein [Arabidopsis thaliana] 190 1e-46
UniRef100_Q8GXW1 Putative RGA1 [Arabidopsis thaliana] 190 1e-46
UniRef100_Q8S4W7 GAI-like protein 1 [Vitis vinifera] 187 1e-45
UniRef100_Q7Y1B6 GAI-like protein [Lycopersicon esculentum] 186 2e-45
UniRef100_Q9SLH3 Putative RGA1, giberellin repsonse modulation p... 185 4e-45
UniRef100_Q941D4 At2g01570/F2I9.19 [Arabidopsis thaliana] 185 4e-45
UniRef100_O23642 RGA1 protein [Arabidopsis thaliana] 185 4e-45
UniRef100_O23725 GRS protein [Arabidopsis thaliana] 184 5e-45
UniRef100_O65367 RGA-like protein [Arabidopsis thaliana] 183 1e-44
UniRef100_Q9ZSP2 Lateral suppressor protein [Lycopersicon escule... 182 2e-44
UniRef100_Q6EI06 Gibberellic acid insensitive phloem [Cucurbita ... 181 4e-44
UniRef100_Q9ST48 Gibberellin response modulator [Zea mays] 181 5e-44
UniRef100_Q9C8Y3 Gibberellin regulatory protein, putative; 49974... 181 7e-44
UniRef100_Q84TQ7 GIA/RGA-like gibberellin response modulator [Go... 178 3e-43
UniRef100_Q9LQT8 F10B6.34 [Arabidopsis thaliana] 178 4e-43
UniRef100_O23724 GAI protein [Arabidopsis thaliana] 178 4e-43
UniRef100_Q7G7J6 Gibberellin-insensitive protein OsGAI [Oryza sa... 177 6e-43
UniRef100_O23643 RGA2 protein [Arabidopsis thaliana] 177 6e-43
UniRef100_Q9ST59 Gibberellin response modulator [Triticum aestivum] 176 1e-42
UniRef100_Q6EI05 Gibberellic acid insensitive phloem B [Cucurbit... 176 1e-42
>UniRef100_Q9SRP9 RGA1-like protein [Arabidopsis thaliana]
Length = 547
Score = 190 bits (482), Expect = 1e-46
Identities = 119/393 (30%), Positives = 195/393 (49%), Gaps = 21/393 (5%)
Query: 197 LSDEEKENVSLAESLLACAEKVGYQQYERARKLLSQIESLSSKTGNPVKRVVHYFAEALC 256
L D ++ V L +L+ACAE + + A L+ ++ +L+ + +V YFA+AL
Sbjct: 170 LVDSQETGVRLVHALVACAEAIHQENLNLADALVKRVGTLAGSQAGAMGKVATYFAQALA 229
Query: 257 QRIDKE-TGRFSVSSNNMQKMESLFDPQEVSKDLNPAMIALYEDLPFSQVSIFTCVQALL 315
+RI ++ T V + E + + + YE P+ + + FT QA+L
Sbjct: 230 RRIYRDYTAETDVCAAVNPSFEEVLE------------MHFYESCPYLKFAHFTANQAIL 277
Query: 316 ENVNDAKKIHVIDLEIRKGCQWTILMQALQSRNECP--LELLKITAIESGNSDTSKHIVE 373
E V A+++HVIDL + +G QW LMQAL R P L I ++ NSD+ ++
Sbjct: 278 EAVTTARRVHVIDLGLNQGMQWPALMQALALRPGGPPSFRLTGIGPPQTENSDS----LQ 333
Query: 374 DTGKRLKDFAQSLNIPFSFDIVVVSDLLHIREELFKIDSE-ETVAVYSQFALRSKIQQPD 432
G +L FAQ++ + F F + L + E+F+ E ET+ V S F L + +
Sbjct: 334 QLGWKLAQFAQNMGVEFEFKGLAAESLSDLEPEMFETRPESETLVVNSVFELHRLLARSG 393
Query: 433 KLETIMRVIRTINPIVMVVAEIEANHNSKSFVNRFIEALFYFSAYFDCFETCMKGDEKNR 492
+E ++ ++ I P ++ V E EANHN F++RF EAL Y+S+ FD E ++R
Sbjct: 394 SIEKLLNTVKAIKPSIVTVVEQEANHNGIVFLDRFNEALHYYSSLFDSLEDSYSLPSQDR 453
Query: 493 FILESMYFSHGIRNIVAEEGAERKSRNVKIDVWRAFFTRFGMVETELSMKSLYQAELVAK 552
++ +Y I N+VA EG++R R+ WR G L + QA ++
Sbjct: 454 -VMSEVYLGRQILNVVAAEGSDRVERHETAAQWRIRMKSAGFDPIHLGSSAFKQASMLLS 512
Query: 553 RFACGYACTFDMNGHCLLVGWKGTPINSVSVWK 585
+A G + N CL++GW+ P+ + S WK
Sbjct: 513 LYATGDGYRVEENDGCLMIGWQTRPLITTSAWK 545
>UniRef100_Q8GXW1 Putative RGA1 [Arabidopsis thaliana]
Length = 547
Score = 190 bits (482), Expect = 1e-46
Identities = 119/393 (30%), Positives = 195/393 (49%), Gaps = 21/393 (5%)
Query: 197 LSDEEKENVSLAESLLACAEKVGYQQYERARKLLSQIESLSSKTGNPVKRVVHYFAEALC 256
L D ++ V L +L+ACAE + + A L+ ++ +L+ + +V YFA+AL
Sbjct: 170 LVDSQETGVRLVHALVACAEAIHQENLNLADALVKRVGTLTGSQAGAMGKVATYFAQALA 229
Query: 257 QRIDKE-TGRFSVSSNNMQKMESLFDPQEVSKDLNPAMIALYEDLPFSQVSIFTCVQALL 315
+RI ++ T V + E + + + YE P+ + + FT QA+L
Sbjct: 230 RRIYRDYTAETDVCAAVNPSFEEVLE------------MHFYESCPYLKFAHFTANQAIL 277
Query: 316 ENVNDAKKIHVIDLEIRKGCQWTILMQALQSRNECP--LELLKITAIESGNSDTSKHIVE 373
E V A+++HVIDL + +G QW LMQAL R P L I ++ NSD+ ++
Sbjct: 278 EAVTTARRVHVIDLGLNQGMQWPALMQALALRPGGPPSFRLTGIGPPQTENSDS----LQ 333
Query: 374 DTGKRLKDFAQSLNIPFSFDIVVVSDLLHIREELFKIDSE-ETVAVYSQFALRSKIQQPD 432
G +L FAQ++ + F F + L + E+F+ E ET+ V S F L + +
Sbjct: 334 QLGWKLAQFAQNMGVEFEFKGLAAESLSDLEPEMFETRPESETLVVNSVFELHRLLARSG 393
Query: 433 KLETIMRVIRTINPIVMVVAEIEANHNSKSFVNRFIEALFYFSAYFDCFETCMKGDEKNR 492
+E ++ ++ I P ++ V E EANHN F++RF EAL Y+S+ FD E ++R
Sbjct: 394 SIEKLLNTVKAIKPSIVTVVEQEANHNGIVFLDRFNEALHYYSSLFDSLEDSYSLPSQDR 453
Query: 493 FILESMYFSHGIRNIVAEEGAERKSRNVKIDVWRAFFTRFGMVETELSMKSLYQAELVAK 552
++ +Y I N+VA EG++R R+ WR G L + QA ++
Sbjct: 454 -VMSEVYLGRQILNVVAAEGSDRVERHETAAQWRIRMKSAGFDPIHLGSSAFKQASMLLS 512
Query: 553 RFACGYACTFDMNGHCLLVGWKGTPINSVSVWK 585
+A G + N CL++GW+ P+ + S WK
Sbjct: 513 LYATGDGYRVEENDGCLMIGWQTRPLITTSAWK 545
>UniRef100_Q8S4W7 GAI-like protein 1 [Vitis vinifera]
Length = 590
Score = 187 bits (474), Expect = 1e-45
Identities = 114/391 (29%), Positives = 193/391 (49%), Gaps = 21/391 (5%)
Query: 197 LSDEEKENVSLAESLLACAEKVGYQQYERARKLLSQIESLSSKTGNPVKRVVHYFAEALC 256
L D ++ + L +L+ACAE V + + A L+ QI L+ +++V YFAE L
Sbjct: 204 LVDSQETGIRLVHTLMACAEAVQQENLKLAEALVKQIGFLAVSQAGAMRKVATYFAEGLA 263
Query: 257 QRIDKETGRFSVSSNNMQKMESLFDPQEVSKDLNPAM-IALYEDLPFSQVSIFTCVQALL 315
+RI + L+ + + + + + YE P+ + + FT QA+L
Sbjct: 264 RRIYR-----------------LYPDKPLDSSFSDILQMHFYETCPYLKFAHFTANQAIL 306
Query: 316 ENVNDAKKIHVIDLEIRKGCQWTILMQALQSRNECPLELLKITAIESGNSDTSKHIVEDT 375
E K++HVID +++G QW LMQAL R P ++T I ++D + H+ E
Sbjct: 307 EAFEGKKRVHVIDFSMKQGMQWPALMQALALRPGGPPSF-RLTGIGPPSTDNTDHLHE-V 364
Query: 376 GKRLKDFAQSLNIPFSFDIVVVSDLLHIREELFKIDSEETVAVYSQFALRSKIQQPDKLE 435
G +L A+++++ F + V + L + + ++ E+VAV S F L S + +P +E
Sbjct: 365 GWKLAQLAETIHVEFEYRGFVANSLADLDASMLELRDGESVAVNSVFELHSLLARPGGIE 424
Query: 436 TIMRVIRTINPIVMVVAEIEANHNSKSFVNRFIEALFYFSAYFDCFETC-MKGDEKNRFI 494
++ ++ + P ++ + E EANHN F++RF E+L Y+S FD E C + +
Sbjct: 425 RVLSAVKDMKPDIVTIVEQEANHNGPVFLDRFTESLHYYSTLFDSLEGCGVSPVNTQDKL 484
Query: 495 LESMYFSHGIRNIVAEEGAERKSRNVKIDVWRAFFTRFGMVETELSMKSLYQAELVAKRF 554
+ +Y I N+VA EG ER R+ + WRA G L + QA ++ F
Sbjct: 485 MSEVYLGQQICNVVACEGPERVERHETLAQWRARLGSAGFDPVNLGSNAFKQASMLLALF 544
Query: 555 ACGYACTFDMNGHCLLVGWKGTPINSVSVWK 585
A G + N CL++GW P+ + S W+
Sbjct: 545 AGGDGYRVEENNGCLMLGWHTRPLIATSAWQ 575
>UniRef100_Q7Y1B6 GAI-like protein [Lycopersicon esculentum]
Length = 588
Score = 186 bits (472), Expect = 2e-45
Identities = 127/457 (27%), Positives = 207/457 (44%), Gaps = 47/457 (10%)
Query: 146 GLVTNNENEKKLLSTIEIMKMAGTRFIQSSSSSKSASGLILNHPFGFSFSGLSDEEKENV 205
G V N+++ K+ ST + + SS+++ L D ++ V
Sbjct: 152 GAVFNSDSNKRHRSTTSSFSTTSSSMVTDSSATRPVV--------------LVDSQETGV 197
Query: 206 SLAESLLACAEKVGYQQYERARKLLSQIESLSSKTGNPVKRVVHYFAEALCQRIDKETGR 265
L +L+ACAE V + A +L+ I L+ +++V YFAEAL +RI K +
Sbjct: 198 RLVHTLMACAEAVQQENLTLADQLVRHIGILAVSQSGAMRKVATYFAEALARRIYKIYPQ 257
Query: 266 FSVSSNNMQKMESLFDPQEVSKDLNPAMIALYEDLPFSQVSIFTCVQALLENVNDAKKIH 325
S+ S+ ++ F YE P+ + + FT QA+LE K+H
Sbjct: 258 DSMESSYTDVLQMHF----------------YETCPYLKFAHFTANQAILEAFTGCNKVH 301
Query: 326 VIDLEIRKGCQWTILMQALQSRNECPLELLKITAIESGNSDTSKHIVEDTGKRLKDFAQS 385
VID +++G QW LMQAL R P ++T I D + + + G +L A++
Sbjct: 302 VIDFSLKQGMQWPALMQALALRPGGP-PAFRLTGIGPPQPDNTDAL-QQVGWKLAQLAET 359
Query: 386 LNIPFSFDIVVVSDLLHIREELFKIDSEET--VAVYSQFALRSKIQQPDKLETIMRVIRT 443
+ + F F V + L + + I ET VA+ S F L + +P +E ++ I+
Sbjct: 360 IGVEFEFRGFVANSLADLDATILDIRPSETEAVAINSVFELHRLLSRPGAIEKVLNSIKQ 419
Query: 444 INPIVMVVAEIEANHNSKSFVNRFIEALFYFSAYFDCFETCMKGD-------------EK 490
INP ++ + E EANHN+ F++RF EAL Y+S FD E+
Sbjct: 420 INPKIVTLVEQEANHNAGVFIDRFNEALHYYSTMFDSLESSGSSSSASPTGILPQPPVNN 479
Query: 491 NRFILESMYFSHGIRNIVAEEGAERKSRNVKIDVWRAFFTRFGMVETELSMKSLYQAELV 550
++ +Y I N+VA EG++R R+ ++ WR G L + QA ++
Sbjct: 480 QDLVMSEVYLGRQICNVVACEGSDRVERHETLNQWRVRMNSSGFDPVHLGSNAFKQASML 539
Query: 551 AKRFACGYACTFDMNGHCLLVGWKGTPINSVSVWKFI 587
FA G + N CL++GW P+ + S WK +
Sbjct: 540 LALFAGGDGYRVEENDGCLMLGWHTRPLIATSAWKLL 576
>UniRef100_Q9SLH3 Putative RGA1, giberellin repsonse modulation protein [Arabidopsis
thaliana]
Length = 587
Score = 185 bits (469), Expect = 4e-45
Identities = 117/391 (29%), Positives = 187/391 (46%), Gaps = 22/391 (5%)
Query: 197 LSDEEKENVSLAESLLACAEKVGYQQYERARKLLSQIESLSSKTGNPVKRVVHYFAEALC 256
L D ++ V L +L+ACAE + A L+ QI L+ +++V YFAEAL
Sbjct: 211 LVDSQENGVRLVHALMACAEAIQQNNLTLAEALVKQIGCLAVSQAGAMRKVATYFAEALA 270
Query: 257 QRIDKETGRFSVSSNNMQKMESLFDPQEVSKDLNPAMIALYEDLPFSQVSIFTCVQALLE 316
+RI R S N + S + + YE P+ + + FT QA+LE
Sbjct: 271 RRIY----RLSPPQNQIDHCLS-----------DTLQMHFYETCPYLKFAHFTANQAILE 315
Query: 317 NVNDAKKIHVIDLEIRKGCQWTILMQALQSRNECPLELLKITAIESGNSDTSKHIVEDTG 376
K++HVID + +G QW LMQAL R P ++T I D S H+ E G
Sbjct: 316 AFEGKKRVHVIDFSMNQGLQWPALMQALALREGGP-PTFRLTGIGPPAPDNSDHLHE-VG 373
Query: 377 KRLKDFAQSLNIPFSFDIVVVSDLLHIREELFKIDSEET--VAVYSQFALRSKIQQPDKL 434
+L A+++++ F + V + L + + ++ +T VAV S F L + +P +
Sbjct: 374 CKLAQLAEAIHVEFEYRGFVANSLADLDASMLELRPSDTEAVAVNSVFELHKLLGRPGGI 433
Query: 435 ETIMRVIRTINPIVMVVAEIEANHNSKSFVNRFIEALFYFSAYFDCFETCMKGDEKNRFI 494
E ++ V++ I P++ V E E+NHN F++RF E+L Y+S FD E +K +
Sbjct: 434 EKVLGVVKQIKPVIFTVVEQESNHNGPVFLDRFTESLHYYSTLFDSLEGVPNSQDK---V 490
Query: 495 LESMYFSHGIRNIVAEEGAERKSRNVKIDVWRAFFTRFGMVETELSMKSLYQAELVAKRF 554
+ +Y I N+VA EG +R R+ + W F G+ L + QA ++ F
Sbjct: 491 MSEVYLGKQICNLVACEGPDRVERHETLSQWGNRFGSSGLAPAHLGSNAFKQASMLLSVF 550
Query: 555 ACGYACTFDMNGHCLLVGWKGTPINSVSVWK 585
G + + CL++GW P+ + S WK
Sbjct: 551 NSGQGYRVEESNGCLMLGWHTRPLITTSAWK 581
>UniRef100_Q941D4 At2g01570/F2I9.19 [Arabidopsis thaliana]
Length = 587
Score = 185 bits (469), Expect = 4e-45
Identities = 117/391 (29%), Positives = 187/391 (46%), Gaps = 22/391 (5%)
Query: 197 LSDEEKENVSLAESLLACAEKVGYQQYERARKLLSQIESLSSKTGNPVKRVVHYFAEALC 256
L D ++ V L +L+ACAE + A L+ QI L+ +++V YFAEAL
Sbjct: 211 LVDSQENGVRLVHALMACAEAIQQNNLTLAEALVKQIGCLAVSQAGAMRKVATYFAEALA 270
Query: 257 QRIDKETGRFSVSSNNMQKMESLFDPQEVSKDLNPAMIALYEDLPFSQVSIFTCVQALLE 316
+RI R S N + S + + YE P+ + + FT QA+LE
Sbjct: 271 RRIY----RLSPPQNQIDHCLS-----------DTLQMHFYETCPYLKFAHFTANQAILE 315
Query: 317 NVNDAKKIHVIDLEIRKGCQWTILMQALQSRNECPLELLKITAIESGNSDTSKHIVEDTG 376
K++HVID + +G QW LMQAL R P ++T I D S H+ E G
Sbjct: 316 AFEGKKRVHVIDFSMNQGLQWPALMQALALREGGP-PTFRLTGIGPPAPDNSDHLHE-VG 373
Query: 377 KRLKDFAQSLNIPFSFDIVVVSDLLHIREELFKIDSEET--VAVYSQFALRSKIQQPDKL 434
+L A+++++ F + V + L + + ++ +T VAV S F L + +P +
Sbjct: 374 CKLAQLAEAIHVEFEYRGFVANSLADLDASMLELRPSDTEAVAVNSVFELHKLLGRPGGI 433
Query: 435 ETIMRVIRTINPIVMVVAEIEANHNSKSFVNRFIEALFYFSAYFDCFETCMKGDEKNRFI 494
E ++ V++ I P++ V E E+NHN F++RF E+L Y+S FD E +K +
Sbjct: 434 EKVLGVVKQIKPVIFTVVEQESNHNGPVFLDRFTESLHYYSTLFDSLEGVPNSQDK---V 490
Query: 495 LESMYFSHGIRNIVAEEGAERKSRNVKIDVWRAFFTRFGMVETELSMKSLYQAELVAKRF 554
+ +Y I N+VA EG +R R+ + W F G+ L + QA ++ F
Sbjct: 491 MSEVYLGKQICNLVACEGPDRVERHETLSQWGNRFGSSGLAPAHLGSNAFKQASMLLSVF 550
Query: 555 ACGYACTFDMNGHCLLVGWKGTPINSVSVWK 585
G + + CL++GW P+ + S WK
Sbjct: 551 NSGQGYRVEESNGCLMLGWHTRPLITTSAWK 581
>UniRef100_O23642 RGA1 protein [Arabidopsis thaliana]
Length = 587
Score = 185 bits (469), Expect = 4e-45
Identities = 117/391 (29%), Positives = 187/391 (46%), Gaps = 22/391 (5%)
Query: 197 LSDEEKENVSLAESLLACAEKVGYQQYERARKLLSQIESLSSKTGNPVKRVVHYFAEALC 256
L D ++ V L +L+ACAE + A L+ QI L+ +++V YFAEAL
Sbjct: 211 LVDSQENGVRLVHALMACAEAIQQNNLTLAEALVKQIGCLAVSQAGAMRKVATYFAEALA 270
Query: 257 QRIDKETGRFSVSSNNMQKMESLFDPQEVSKDLNPAMIALYEDLPFSQVSIFTCVQALLE 316
+RI R S N + S + + YE P+ + + FT QA+LE
Sbjct: 271 RRIY----RLSPPQNQIDHCLS-----------DTLQMHFYETCPYLKFAHFTANQAILE 315
Query: 317 NVNDAKKIHVIDLEIRKGCQWTILMQALQSRNECPLELLKITAIESGNSDTSKHIVEDTG 376
K++HVID + +G QW LMQAL R P ++T I D S H+ E G
Sbjct: 316 AFEGKKRVHVIDFSMNQGLQWPALMQALALREGGP-PTFRLTGIGPPAPDNSDHLHE-VG 373
Query: 377 KRLKDFAQSLNIPFSFDIVVVSDLLHIREELFKIDSEET--VAVYSQFALRSKIQQPDKL 434
+L A+++++ F + V + L + + ++ +T VAV S F L + +P +
Sbjct: 374 CKLAQLAEAIHVEFEYRGFVANSLADLDASMLELRPSDTEAVAVNSVFELHKLLGRPGGI 433
Query: 435 ETIMRVIRTINPIVMVVAEIEANHNSKSFVNRFIEALFYFSAYFDCFETCMKGDEKNRFI 494
E ++ V++ I P++ V E E+NHN F++RF E+L Y+S FD E +K +
Sbjct: 434 EKVLGVVKQIKPVIFTVVEQESNHNGPVFLDRFTESLHYYSTLFDSLEGVPNSQDK---V 490
Query: 495 LESMYFSHGIRNIVAEEGAERKSRNVKIDVWRAFFTRFGMVETELSMKSLYQAELVAKRF 554
+ +Y I N+VA EG +R R+ + W F G+ L + QA ++ F
Sbjct: 491 MSEVYLGKQICNLVACEGPDRVERHETLSQWGNRFGSSGLAPAHLGSNAFKQASMLLSVF 550
Query: 555 ACGYACTFDMNGHCLLVGWKGTPINSVSVWK 585
G + + CL++GW P+ + S WK
Sbjct: 551 NSGQGYRVEESNGCLMLGWHTRPLITTSAWK 581
>UniRef100_O23725 GRS protein [Arabidopsis thaliana]
Length = 587
Score = 184 bits (468), Expect = 5e-45
Identities = 117/391 (29%), Positives = 187/391 (46%), Gaps = 22/391 (5%)
Query: 197 LSDEEKENVSLAESLLACAEKVGYQQYERARKLLSQIESLSSKTGNPVKRVVHYFAEALC 256
L D ++ V L +L+ACAE + A L+ QI L+ +++V YFAEAL
Sbjct: 211 LVDSQENGVRLVHALMACAEAIQQNNLTLAEALVKQIGCLAVSQAGAMRKVATYFAEALA 270
Query: 257 QRIDKETGRFSVSSNNMQKMESLFDPQEVSKDLNPAMIALYEDLPFSQVSIFTCVQALLE 316
+RI R S N + S + + YE P+ + + FT QA+LE
Sbjct: 271 RRIY----RLSPPQNQIDHCLS-----------DTLQMHFYETCPYLKFAHFTANQAILE 315
Query: 317 NVNDAKKIHVIDLEIRKGCQWTILMQALQSRNECPLELLKITAIESGNSDTSKHIVEDTG 376
K++HVID + +G QW LMQAL R P ++T I D S H+ E G
Sbjct: 316 AFEGKKRVHVIDFSMNQGLQWPALMQALALREGGP-PTFRLTGIGPPAPDNSDHLHE-VG 373
Query: 377 KRLKDFAQSLNIPFSFDIVVVSDLLHIREELFKIDSEET--VAVYSQFALRSKIQQPDKL 434
+L A+++++ F + V + L + + ++ +T VAV S F L + +P +
Sbjct: 374 CKLAQLAEAVHVEFEYRGFVANSLADLDASMLELRPSDTEAVAVNSVFELHKLLGRPGGI 433
Query: 435 ETIMRVIRTINPIVMVVAEIEANHNSKSFVNRFIEALFYFSAYFDCFETCMKGDEKNRFI 494
E ++ V++ I P++ V E E+NHN F++RF E+L Y+S FD E +K +
Sbjct: 434 EKVLGVVKQIKPVIFTVVEQESNHNGPVFLDRFTESLHYYSTLFDSLEGVPNSQDK---V 490
Query: 495 LESMYFSHGIRNIVAEEGAERKSRNVKIDVWRAFFTRFGMVETELSMKSLYQAELVAKRF 554
+ +Y I N+VA EG +R R+ + W F G+ L + QA ++ F
Sbjct: 491 MSEVYLGKQICNLVACEGPDRVERHETLSQWGNRFGSSGLAPAHLGSNAFKQASMLLSVF 550
Query: 555 ACGYACTFDMNGHCLLVGWKGTPINSVSVWK 585
G + + CL++GW P+ + S WK
Sbjct: 551 NSGQGYRVEESNGCLMLGWHTRPLITTSAWK 581
>UniRef100_O65367 RGA-like protein [Arabidopsis thaliana]
Length = 662
Score = 183 bits (464), Expect = 1e-44
Identities = 116/388 (29%), Positives = 191/388 (48%), Gaps = 26/388 (6%)
Query: 199 DEEKENVSLAESLLACAEKVGYQQYERARKLLSQIESLSSKTGNPVKRVVHYFAEALCQR 258
D ++ V L +LLACAE V + A L+ + L+S +++V YFAE L +R
Sbjct: 295 DSQETGVRLVHALLACAEAVQQNNLKLADALVKHVGLLASSQAGAMRKVATYFAEGLARR 354
Query: 259 IDKETGRFSVSSNNMQKMESLFDPQEVSKDLNPAMIALYEDLPFSQVSIFTCVQALLENV 318
I + R V+S++ + I YE P+ + + FT QA+LE
Sbjct: 355 IYRIYPRDDVASSSFS---------------DTLQIHFYESCPYLKFAHFTANQAILEVF 399
Query: 319 NDAKKIHVIDLEIRKGCQWTILMQALQSRNECPLELLKITAIESGNSDTSKHIVEDTGKR 378
A+K+HVIDL + G QW L+QAL R P + ++T I +D +++ G +
Sbjct: 400 ATAEKVHVIDLGLNHGLQWPALIQALALRPNGPPDF-RLTGIGYSLTD-----IQEVGWK 453
Query: 379 LKDFAQSLNIPFSFDIVVVSDLLHIREELFKI-DSEETVAVYSQFALRSKIQQPDKLETI 437
L A ++ + F F + +++L ++ E+ I E+VAV S F L + P ++
Sbjct: 454 LGQLASTIGVNFEFKSIALNNLSDLKPEMLDIRPGLESVAVNSVFELHRLLAHPGSIDKF 513
Query: 438 MRVIRTINPIVMVVAEIEANHNSKSFVNRFIEALFYFSAYFDCFETCMKGDEKNRFILES 497
+ I++I P +M V E EANHN F++RF E+L Y+S+ FD E G ++
Sbjct: 514 LSTIKSIRPDIMTVVEQEANHNGTVFLDRFTESLHYYSSLFDSLE----GPPSQDRVMSE 569
Query: 498 MYFSHGIRNIVAEEGAERKSRNVKIDVWRAFFTRFGMVETELSMKSLYQAELVAKRFACG 557
++ I N+VA EG +R R+ ++ WR F G + + QA ++ +A
Sbjct: 570 LFLGRQILNLVACEGEDRVERHETLNQWRNRFGLGGFKPVSIGSNAYKQASMLLALYAGA 629
Query: 558 YACTFDMNGHCLLVGWKGTPINSVSVWK 585
+ N CLL+GW+ P+ + S W+
Sbjct: 630 DGYNVEENEGCLLLGWQTRPLIATSAWR 657
>UniRef100_Q9ZSP2 Lateral suppressor protein [Lycopersicon esculentum]
Length = 428
Score = 182 bits (463), Expect = 2e-44
Identities = 120/392 (30%), Positives = 203/392 (51%), Gaps = 21/392 (5%)
Query: 205 VSLAESLLACAEKVGYQQYERARKLLSQIESLSSKTGNPVKRVVHYFAEALCQRIDKETG 264
+ + + L++CAE + + A++LL+ + + SS G+ +R+VH F AL R+++
Sbjct: 47 IQIRQLLISCAELISQSDFSAAKRLLTILSTNSSPFGDSTERLVHQFTRALSLRLNRYIS 106
Query: 265 RFSVSSNNMQKMESLFDPQEVSKDL---NPAMIALYEDLPFSQVSIFTCVQALLENVN-D 320
S +++ M +E+ S L + ++L + PF + + T QA+LE +N +
Sbjct: 107 --STTNHFMTPVETTPTDSSSSSSLALIQSSYLSLNQVTPFIRFTQLTANQAILEAINGN 164
Query: 321 AKKIHVIDLEIRKGCQWTILMQALQSRNECPLELLKITAIESGNS-DTSKHIVEDTGKRL 379
+ IH++D +I G QW LMQAL R P L+IT +GN DT + TG RL
Sbjct: 165 HQAIHIVDFDINHGVQWPPLMQALADRYPAPT--LRITG--TGNDLDTLRR----TGDRL 216
Query: 380 KDFAQSLNIPFSFDIVVVSDLLHIREELFKIDS------EETVAVYSQFALRSKIQQPDK 433
FA SL + F F + +++ H +E I S +ET+A+ F L ++ +K
Sbjct: 217 AKFAHSLGLRFQFHPLYIANNNHDHDEDPSIISSIVLLPDETLAINCVFYLHRLLKDREK 276
Query: 434 LETIMRVIRTINPIVMVVAEIEANHNSKSFVNRFIEALFYFSAYFDCFETCMKGDEKNRF 493
L + ++++NP ++ +AE EANHN F+ RFIEAL Y++A FD E + + R
Sbjct: 277 LRIFLHRVKSMNPKIVTIAEKEANHNHPLFLQRFIEALDYYTAVFDSLEATLPPGSRERM 336
Query: 494 ILESMYFSHGIRNIVAEEGAERKSRNVKIDVWRAFFTRFGMVETELSMKSLYQAELVAKR 553
+E ++F I +IVA EG +RK R+ + W G LS +L QA+L+ +
Sbjct: 337 TVEQVWFGREIVDIVAMEGDKRKERHERFRSWEVMLRSCGFSNVALSPFALSQAKLLLRL 396
Query: 554 FACGYACTFDMNGHCLLVGWKGTPINSVSVWK 585
++ + +GW+ P+ S+S W+
Sbjct: 397 HYPSEGYQLGVSSNSFFLGWQNQPLFSISSWR 428
>UniRef100_Q6EI06 Gibberellic acid insensitive phloem [Cucurbita maxima]
Length = 579
Score = 181 bits (460), Expect = 4e-44
Identities = 113/391 (28%), Positives = 187/391 (46%), Gaps = 23/391 (5%)
Query: 197 LSDEEKENVSLAESLLACAEKVGYQQYERARKLLSQIESLSSKTGNPVKRVVHYFAEALC 256
L D ++ + L +L+ CAE V A L+ +I+ L+ +++V +FAEAL
Sbjct: 201 LVDSQENGIQLVHALMVCAEAVQQNNLNLAEALVKRIDYLAVSQAGAMRKVATFFAEALA 260
Query: 257 QRIDKETGRFSVSSNNMQKMESLFDPQEVSKDLNPAMIALYEDLPFSQVSIFTCVQALLE 316
+RI + + + + ++ F YE P+ + + FT QA+LE
Sbjct: 261 RRIYRLCPENPLDRSVLDMLQMHF----------------YESCPYLKFAHFTANQAILE 304
Query: 317 NVNDAKKIHVIDLEIRKGCQWTILMQALQSRNECPLELLKITAIESGNSDTSKHIVEDTG 376
K++HVID + +G QW L+QAL R P ++T I D S ++ +D G
Sbjct: 305 AFEGKKRVHVIDFSMNQGIQWPALIQALALRPSGP-PTFRLTGIGPPAPDNSDYL-QDVG 362
Query: 377 KRLKDFAQSLNIPFSFDIVVVSDLLHIREELFKI--DSEETVAVYSQFALRSKIQQPDKL 434
+L FA++L++ F + V + L + + ++ E+V V S F L + +P +
Sbjct: 363 WKLVKFAETLHVEFEYRGFVANSLADLDASMLELRPSEVESVVVNSVFELHQLLARPGAI 422
Query: 435 ETIMRVIRTINPIVMVVAEIEANHNSKSFVNRFIEALFYFSAYFDCFETCMKGDEKNRFI 494
E ++ V++ + P ++ V E EANHN FV RF E+L Y+S FD E +K +
Sbjct: 423 EKVLSVVKQMKPEIVTVVEQEANHNGPVFVERFTESLHYYSTLFDSLECSPNSQDK---M 479
Query: 495 LESMYFSHGIRNIVAEEGAERKSRNVKIDVWRAFFTRFGMVETELSMKSLYQAELVAKRF 554
+ MY I N+VA EGA+R R+ + WR + G L + QA ++ F
Sbjct: 480 MSEMYLGKQICNVVACEGADRVERHETLTQWRTRLSSAGFDPIHLGSNAFKQASILLALF 539
Query: 555 ACGYACTFDMNGHCLLVGWKGTPINSVSVWK 585
G + N L++GW P+ + S WK
Sbjct: 540 GSGEGYRVEENEGSLMLGWHTRPLIATSAWK 570
>UniRef100_Q9ST48 Gibberellin response modulator [Zea mays]
Length = 630
Score = 181 bits (459), Expect = 5e-44
Identities = 118/409 (28%), Positives = 192/409 (46%), Gaps = 42/409 (10%)
Query: 199 DEEKENVSLAESLLACAEKVGYQQYERARKLLSQIESLSSKTGNPVKRVVHYFAEALCQR 258
D ++ + L +LLACAE V + + A L+ QI L+S G +++V YF EAL +R
Sbjct: 235 DTQEAGIRLVHALLACAEAVQQENFSAAEALVKQIPMLASSQGGAMRKVAAYFGEALARR 294
Query: 259 IDKETGRFSVSSNNMQKMESLFDPQEVSKDLNPAMIAL-----YEDLPFSQVSIFTCVQA 313
+ + F P S L+ A L YE P+ + + FT QA
Sbjct: 295 VYR------------------FRPPPDSSLLDAAFADLLHAHFYESCPYLKFAHFTANQA 336
Query: 314 LLENVNDAKKIHVIDLEIRKGCQWTILMQALQSRNECPLELLKITAIESGNSDTSKHIVE 373
+LE +++HV+D I++G QW L+QAL R P ++T + D + + +
Sbjct: 337 ILEAFAGCRRVHVVDFGIKQGMQWPALLQALALRPGGPPSF-RLTGVGPPQPDETDAL-Q 394
Query: 374 DTGKRLKDFAQSLNIPFSFDIVVVSDLLHIREELFKIDSEET------VAVYSQFALRSK 427
G +L FA ++ + F + +V + L + + + + ++T +AV S F L
Sbjct: 395 QVGWKLAQFAHTIRVDFQYRGLVAATLADLEPFMLQPEGDDTDDEPEVIAVNSVFELHRL 454
Query: 428 IQQPDKLETIMRVIRTINPIVMVVAEIEANHNSKSFVNRFIEALFYFSAYFDCFETCMKG 487
+ QP LE ++ +R + P ++ V E EANHNS +F++RF E+L Y+S FD E G
Sbjct: 455 LAQPGALEKVLGTVRAVRPRIVTVVEQEANHNSGTFLDRFTESLHYYSTMFDSLEGAGAG 514
Query: 488 DEKNR-----------FILESMYFSHGIRNIVAEEGAERKSRNVKIDVWRAFFTRFGMVE 536
++ ++ +Y I N+VA EGAER R+ + WR+ G
Sbjct: 515 SGQSTDASPAAAGGTDQVMSEVYLGRQICNVVACEGAERTERHETLGQWRSRLGGSGFAP 574
Query: 537 TELSMKSLYQAELVAKRFACGYACTFDMNGHCLLVGWKGTPINSVSVWK 585
L + QA + FA G + CL +GW P+ + S W+
Sbjct: 575 VHLGSNAYKQASTLLALFAGGDGYRVEEKDGCLTLGWHTRPLIATSAWR 623
>UniRef100_Q9C8Y3 Gibberellin regulatory protein, putative; 49974-51509 [Arabidopsis
thaliana]
Length = 511
Score = 181 bits (458), Expect = 7e-44
Identities = 115/388 (29%), Positives = 190/388 (48%), Gaps = 26/388 (6%)
Query: 199 DEEKENVSLAESLLACAEKVGYQQYERARKLLSQIESLSSKTGNPVKRVVHYFAEALCQR 258
D ++ V L +LLACAE V + A L+ + L+S +++V YFAE L +R
Sbjct: 144 DSQETGVRLVHALLACAEAVQQNNLKLADALVKHVGLLASSQAGAMRKVATYFAEGLARR 203
Query: 259 IDKETGRFSVSSNNMQKMESLFDPQEVSKDLNPAMIALYEDLPFSQVSIFTCVQALLENV 318
I + R V+ ++ + I YE P+ + + FT QA+LE
Sbjct: 204 IYRIYPRDDVALSSFS---------------DTLQIHFYESCPYLKFAHFTANQAILEVF 248
Query: 319 NDAKKIHVIDLEIRKGCQWTILMQALQSRNECPLELLKITAIESGNSDTSKHIVEDTGKR 378
A+K+HVIDL + G QW L+QAL R P + ++T I +D +++ G +
Sbjct: 249 ATAEKVHVIDLGLNHGLQWPALIQALALRPNGPPDF-RLTGIGYSLTD-----IQEVGWK 302
Query: 379 LKDFAQSLNIPFSFDIVVVSDLLHIREELFKI-DSEETVAVYSQFALRSKIQQPDKLETI 437
L A ++ + F F + +++L ++ E+ I E+VAV S F L + P ++
Sbjct: 303 LGQLASTIGVNFEFKSIALNNLSDLKPEMLDIRPGLESVAVNSVFELHRLLAHPGSIDKF 362
Query: 438 MRVIRTINPIVMVVAEIEANHNSKSFVNRFIEALFYFSAYFDCFETCMKGDEKNRFILES 497
+ I++I P +M V E EANHN F++RF E+L Y+S+ FD E G ++
Sbjct: 363 LSTIKSIRPDIMTVVEQEANHNGTVFLDRFTESLHYYSSLFDSLE----GPPSQDRVMSE 418
Query: 498 MYFSHGIRNIVAEEGAERKSRNVKIDVWRAFFTRFGMVETELSMKSLYQAELVAKRFACG 557
++ I N+VA EG +R R+ ++ WR F G + + QA ++ +A
Sbjct: 419 LFLGRQILNLVACEGEDRVERHETLNQWRNRFGLGGFKPVSIGSNAYKQASMLLALYAGA 478
Query: 558 YACTFDMNGHCLLVGWKGTPINSVSVWK 585
+ N CLL+GW+ P+ + S W+
Sbjct: 479 DGYNVEENEGCLLLGWQTRPLIATSAWR 506
>UniRef100_Q84TQ7 GIA/RGA-like gibberellin response modulator [Gossypium hirsutum]
Length = 537
Score = 178 bits (452), Expect = 3e-43
Identities = 124/421 (29%), Positives = 192/421 (45%), Gaps = 33/421 (7%)
Query: 161 IEIMKMAGTRFIQSSSSSKSASGLILNHPFGFSFSGLSDEEKENVSLAESLLACAEKVGY 220
+EI + SSSSS + ++L D ++ V L +L+ACAE V
Sbjct: 136 LEITRKRAKTESSSSSSSTTTRPVVL-----------IDSQEAGVRLVHTLMACAEAVQQ 184
Query: 221 QQYERARKLLSQIESLSSKTGNPVKRVVHYFAEALCQRIDKETGRFSVSSNNMQKMESLF 280
+ A L+ I L+S +++V YFAEAL +RI + +F
Sbjct: 185 DNLKLADALVKHIGLLASSQTGAMRKVATYFAEALARRIYR-----------------IF 227
Query: 281 DPQEVSKDLNPAM-IALYEDLPFSQVSIFTCVQALLENVNDAKKIHVIDLEIRKGCQWTI 339
P + N + I YE P+ + + FT QA+LE + A ++HVID +++G QW
Sbjct: 228 PPDSLDPSYNDKLQIPFYETCPYLKFAHFTANQAILEAFSMASRVHVIDFGLKQGMQWPA 287
Query: 340 LMQALQSRNECPLELLKITAIESGNSDTSKHIVEDTGKRLKDFAQSLNIPFSFDIVVVSD 399
LMQAL R P ++T I D + + + G +L A+ + I F F V +
Sbjct: 288 LMQALALRPGGP-PAFRLTGIGPPQPDNTDAL-QQVGWKLAQLAERIGIEFEFRGFVANS 345
Query: 400 LLHIREELFKIDSEE--TVAVYSQFALRSKIQQPDKLETIMRVIRTINPIVMVVAEIEAN 457
L + E+ I E VAV + F L + +P +E ++ I+ + P ++ V E EAN
Sbjct: 346 LADLEPEMLDIRPPEIEVVAVNAVFELHPLLARPGGIEKVVSSIKAMKPKIVTVVEQEAN 405
Query: 458 HNSKSFVNRFIEALFYFSAYFDCFETCMKGDEKNRFILESMYFSHGIRNIVAEEGAERKS 517
HN F++RF EAL Y+S FD E + +Y I N+VA EG +R
Sbjct: 406 HNGPVFLDRFTEALHYYSTLFDSLEGSGVAPASQDLAMSELYLGRQICNVVACEGMDRVE 465
Query: 518 RNVKIDVWRAFFTRFGMVETELSMKSLYQAELVAKRFACGYACTFDMNGHCLLVGWKGTP 577
R+ + WR G+ L + QA ++ FA G + N CL++GW P
Sbjct: 466 RHEPLTQWRTRMETAGVSPVHLGSNAYKQASMLLALFASGDGYRVEENNGCLMLGWHTRP 525
Query: 578 I 578
+
Sbjct: 526 L 526
>UniRef100_Q9LQT8 F10B6.34 [Arabidopsis thaliana]
Length = 533
Score = 178 bits (451), Expect = 4e-43
Identities = 117/391 (29%), Positives = 190/391 (47%), Gaps = 22/391 (5%)
Query: 197 LSDEEKENVSLAESLLACAEKVGYQQYERARKLLSQIESLSSKTGNPVKRVVHYFAEALC 256
L D ++ V L +LLACAE V + A L+ QI L+ +++V YFAEAL
Sbjct: 159 LVDSQENGVRLVHALLACAEAVQKENLTVAEALVKQIGFLAVSQIGAMRKVATYFAEALA 218
Query: 257 QRIDKETGRFSVSSNNMQKMESLFDPQEVSKDLNPAMIALYEDLPFSQVSIFTCVQALLE 316
+RI + +S + SL D ++ YE P+ + + FT QA+LE
Sbjct: 219 RRI------YRLSPSQSPIDHSLSDTLQMH---------FYETCPYLKFAHFTANQAILE 263
Query: 317 NVNDAKKIHVIDLEIRKGCQWTILMQALQSRNECPLELLKITAIESGNSDTSKHIVEDTG 376
K++HVID + +G QW LMQAL R P + ++T I D ++ E G
Sbjct: 264 AFQGKKRVHVIDFSMSQGLQWPALMQALALRPGGP-PVFRLTGIGPPAPDNFDYLHE-VG 321
Query: 377 KRLKDFAQSLNIPFSFDIVVVSDLLHIREELFKIDSEE--TVAVYSQFALRSKIQQPDKL 434
+L A+++++ F + V + L + + ++ E +VAV S F L + +P +
Sbjct: 322 CKLAHLAEAIHVEFEYRGFVANTLADLDASMLELRPSEIESVAVNSVFELHKLLGRPGAI 381
Query: 435 ETIMRVIRTINPIVMVVAEIEANHNSKSFVNRFIEALFYFSAYFDCFETCMKGDEKNRFI 494
+ ++ V+ I P + V E E+NHNS F++RF E+L Y+S FD E G +K +
Sbjct: 382 DKVLGVVNQIKPEIFTVVEQESNHNSPIFLDRFTESLHYYSTLFDSLEGVPSGQDK---V 438
Query: 495 LESMYFSHGIRNIVAEEGAERKSRNVKIDVWRAFFTRFGMVETELSMKSLYQAELVAKRF 554
+ +Y I N+VA +G +R R+ + WR F G + + QA ++ F
Sbjct: 439 MSEVYLGKQICNVVACDGPDRVERHETLSQWRNRFGSAGFAAAHIGSNAFKQASMLLALF 498
Query: 555 ACGYACTFDMNGHCLLVGWKGTPINSVSVWK 585
G + + CL++GW P+ + S WK
Sbjct: 499 NGGEGYRVEESDGCLMLGWHTRPLIATSAWK 529
>UniRef100_O23724 GAI protein [Arabidopsis thaliana]
Length = 532
Score = 178 bits (451), Expect = 4e-43
Identities = 117/391 (29%), Positives = 190/391 (47%), Gaps = 22/391 (5%)
Query: 197 LSDEEKENVSLAESLLACAEKVGYQQYERARKLLSQIESLSSKTGNPVKRVVHYFAEALC 256
L D ++ V L +LLACAE V + A L+ QI L+ +++V YFAEAL
Sbjct: 158 LVDSQENGVRLVHALLACAEAVQKENLTVAEALVKQIGFLAVSQIGAMRKVATYFAEALA 217
Query: 257 QRIDKETGRFSVSSNNMQKMESLFDPQEVSKDLNPAMIALYEDLPFSQVSIFTCVQALLE 316
+RI + +S + SL D ++ YE P+ + + FT QA+LE
Sbjct: 218 RRI------YRLSPSQSPIDHSLSDTLQMH---------FYETCPYLKFAHFTANQAILE 262
Query: 317 NVNDAKKIHVIDLEIRKGCQWTILMQALQSRNECPLELLKITAIESGNSDTSKHIVEDTG 376
K++HVID + +G QW LMQAL R P + ++T I D ++ E G
Sbjct: 263 AFQGKKRVHVIDFSMSQGLQWPALMQALALRPGGP-PVFRLTGIGPPAPDNFDYLHE-VG 320
Query: 377 KRLKDFAQSLNIPFSFDIVVVSDLLHIREELFKIDSEE--TVAVYSQFALRSKIQQPDKL 434
+L A+++++ F + V + L + + ++ E +VAV S F L + +P +
Sbjct: 321 CKLAHLAEAIHVEFEYRGFVANTLADLDASMLELRPSEIESVAVNSVFELHKLLGRPGAI 380
Query: 435 ETIMRVIRTINPIVMVVAEIEANHNSKSFVNRFIEALFYFSAYFDCFETCMKGDEKNRFI 494
+ ++ V+ I P + V E E+NHNS F++RF E+L Y+S FD E G +K +
Sbjct: 381 DKVLGVVNQIKPEIFTVVEQESNHNSPIFLDRFTESLHYYSTLFDSLEGVPSGQDK---V 437
Query: 495 LESMYFSHGIRNIVAEEGAERKSRNVKIDVWRAFFTRFGMVETELSMKSLYQAELVAKRF 554
+ +Y I N+VA +G +R R+ + WR F G + + QA ++ F
Sbjct: 438 MSEVYLGKQICNVVACDGPDRVERHETLSQWRNRFGSAGFAAAHIGSNAFKQASMLLALF 497
Query: 555 ACGYACTFDMNGHCLLVGWKGTPINSVSVWK 585
G + + CL++GW P+ + S WK
Sbjct: 498 NGGEGYRVEESDGCLMLGWHTRPLIATSAWK 528
>UniRef100_Q7G7J6 Gibberellin-insensitive protein OsGAI [Oryza sativa]
Length = 625
Score = 177 bits (450), Expect = 6e-43
Identities = 119/405 (29%), Positives = 193/405 (47%), Gaps = 34/405 (8%)
Query: 199 DEEKENVSLAESLLACAEKVGYQQYERARKLLSQIESLSSKTGNPVKRVVHYFAEALCQR 258
D ++ + L +LLACAE V + + A L+ QI +L++ G +++V YF EAL +R
Sbjct: 233 DTQEAGIRLVHALLACAEAVQQENFAAAEALVKQIPTLAASQGGAMRKVAAYFGEALARR 292
Query: 259 IDKETGRFSVSSNNMQKMESLFDPQEVSKDLNPAMIALYEDLPFSQVSIFTCVQALLENV 318
+ RF + + + +++ F DL A YE P+ + + FT QA+LE
Sbjct: 293 VY----RFRPADSTL--LDAAF------ADLLHAHF--YESCPYLKFAHFTANQAILEAF 338
Query: 319 NDAKKIHVIDLEIRKGCQWTILMQALQSRNECPLELLKITAIESGNSDTSKHIVEDTGKR 378
++HV+D I++G QW L+QAL R P ++T + D + + + G +
Sbjct: 339 AGCHRVHVVDFGIKQGMQWPALLQALALRPGGPPSF-RLTGVGPPQPDETDAL-QQVGWK 396
Query: 379 LKDFAQSLNIPFSFDIVVVSDLLHIREELFKIDSE-------ETVAVYSQFALRSKIQQP 431
L FA ++ + F + +V + L + + + + E E +AV S F L + QP
Sbjct: 397 LAQFAHTIRVDFQYRGLVAATLADLEPFMLQPEGEADANEEPEVIAVNSVFELHRLLAQP 456
Query: 432 DKLETIMRVIRTINPIVMVVAEIEANHNSKSFVNRFIEALFYFSAYFDCFETCMKGDEK- 490
LE ++ + + P ++ V E EANHNS SF++RF E+L Y+S FD E G +
Sbjct: 457 GALEKVLGTVHAVRPRIVTVVEQEANHNSGSFLDRFTESLHYYSTMFDSLEGGSSGQAEL 516
Query: 491 ----------NRFILESMYFSHGIRNIVAEEGAERKSRNVKIDVWRAFFTRFGMVETELS 540
++ +Y I N+VA EGAER R+ + WR R G L
Sbjct: 517 SPPAAGGGGGTDQVMSEVYLGRQICNVVACEGAERTERHETLGQWRNRLGRAGFEPVHLG 576
Query: 541 MKSLYQAELVAKRFACGYACTFDMNGHCLLVGWKGTPINSVSVWK 585
+ QA + FA G + CL +GW P+ + S W+
Sbjct: 577 SNAYKQASTLLALFAGGDGYRVEEKEGCLTLGWHTRPLIATSAWR 621
>UniRef100_O23643 RGA2 protein [Arabidopsis thaliana]
Length = 532
Score = 177 bits (450), Expect = 6e-43
Identities = 117/391 (29%), Positives = 190/391 (47%), Gaps = 22/391 (5%)
Query: 197 LSDEEKENVSLAESLLACAEKVGYQQYERARKLLSQIESLSSKTGNPVKRVVHYFAEALC 256
L D ++ V L +LLACAE V + A L+ QI L+ +++V YFAEAL
Sbjct: 158 LVDSQENGVRLVHALLACAEAVQKENLTVAEALVKQIGFLAVSQIGAMRQVATYFAEALA 217
Query: 257 QRIDKETGRFSVSSNNMQKMESLFDPQEVSKDLNPAMIALYEDLPFSQVSIFTCVQALLE 316
+RI + +S + SL D ++ YE P+ + + FT QA+LE
Sbjct: 218 RRI------YRLSPSQSPIDHSLSDTLQMH---------FYETCPYLKFAHFTANQAILE 262
Query: 317 NVNDAKKIHVIDLEIRKGCQWTILMQALQSRNECPLELLKITAIESGNSDTSKHIVEDTG 376
K++HVID + +G QW LMQAL R P + ++T I D ++ E G
Sbjct: 263 AFQGKKRVHVIDFSMSQGLQWPALMQALALRPGGP-PVFRLTGIGPPAPDNFDYLHE-VG 320
Query: 377 KRLKDFAQSLNIPFSFDIVVVSDLLHIREELFKIDSEE--TVAVYSQFALRSKIQQPDKL 434
+L A+++++ F + V + L + + ++ E +VAV S F L + +P +
Sbjct: 321 CKLAHLAEAIHVEFEYRGFVANTLADLDASMLELRPSEIESVAVNSVFELHKLLGRPGAI 380
Query: 435 ETIMRVIRTINPIVMVVAEIEANHNSKSFVNRFIEALFYFSAYFDCFETCMKGDEKNRFI 494
+ ++ V+ I P + V E E+NHNS F++RF E+L Y+S FD E G +K +
Sbjct: 381 DKVLGVVNQIKPEIFTVVEQESNHNSPIFLDRFTESLHYYSTLFDSLEGVPSGQDK---V 437
Query: 495 LESMYFSHGIRNIVAEEGAERKSRNVKIDVWRAFFTRFGMVETELSMKSLYQAELVAKRF 554
+ +Y I N+VA +G +R R+ + WR F G + + QA ++ F
Sbjct: 438 MSEVYLGKQICNVVACDGPDRVERHETLSQWRNRFGSAGFAAAHIGSNAFKQASMLLALF 497
Query: 555 ACGYACTFDMNGHCLLVGWKGTPINSVSVWK 585
G + + CL++GW P+ + S WK
Sbjct: 498 NGGEGYRVEESDGCLMLGWHTRPLIATSAWK 528
>UniRef100_Q9ST59 Gibberellin response modulator [Triticum aestivum]
Length = 623
Score = 176 bits (447), Expect = 1e-42
Identities = 118/414 (28%), Positives = 189/414 (45%), Gaps = 47/414 (11%)
Query: 199 DEEKENVSLAESLLACAEKVGYQQYERARKLLSQIESLSSKTGNPVKRVVHYFAEALCQR 258
D ++ + L +LLACAE V + A L+ QI L++ G +++V YF EAL +R
Sbjct: 226 DTQEAGIRLVHALLACAEAVQQENLSAAEALVKQIPLLAASQGGAMRKVAAYFGEALARR 285
Query: 259 IDKETGRFSVSSNNMQKMESLFDPQEVSKDLNPAMIAL-----YEDLPFSQVSIFTCVQA 313
+ + F PQ S L+ A L YE P+ + + FT QA
Sbjct: 286 VFR------------------FRPQPDSSLLDAAFADLLHAHFYESCPYLKFAHFTANQA 327
Query: 314 LLENVNDAKKIHVIDLEIRKGCQWTILMQALQSRNECPLELLKITAIESGNSDTSKHIVE 373
+LE +++HV+D I++G QW L+QAL R P ++T + D + + +
Sbjct: 328 ILEAFAGCRRVHVVDFGIKQGMQWPALLQALALRPGGPPSF-RLTGVGPPQPDETDAL-Q 385
Query: 374 DTGKRLKDFAQSLNIPFSFDIVVVSDLLHIREELFKIDSEE-------TVAVYSQFALRS 426
G +L FA ++ + F + +V + L + + + + EE +AV S F +
Sbjct: 386 QVGWKLAQFAHTIRVDFQYRGLVAATLADLEPFMLQPEGEEDPNEEPEVIAVNSVFEMHR 445
Query: 427 KIQQPDKLETIMRVIRTINPIVMVVAEIEANHNSKSFVNRFIEALFYFSAYFDCFETCMK 486
+ QP LE ++ +R + P ++ V E EANHNS +F++RF E+L Y+S FD E
Sbjct: 446 LLAQPGALEKVLGTVRAVRPRIVTVVEQEANHNSGTFLDRFTESLHYYSTMFDSLEGGSS 505
Query: 487 GDEKNRF---------------ILESMYFSHGIRNIVAEEGAERKSRNVKIDVWRAFFTR 531
G + ++ +Y I N+VA EGAER R+ + WR
Sbjct: 506 GGGPSEVSSGAAAAPAAAGTDQVMSEVYLGRQICNVVACEGAERTERHETLGQWRNRLGN 565
Query: 532 FGMVETELSMKSLYQAELVAKRFACGYACTFDMNGHCLLVGWKGTPINSVSVWK 585
G L + QA + FA G + CL +GW P+ + S W+
Sbjct: 566 AGFETVHLGSNAYKQASTLLALFAGGDGYKVEEKEGCLTLGWHTRPLIATSAWR 619
>UniRef100_Q6EI05 Gibberellic acid insensitive phloem B [Cucurbita maxima]
Length = 587
Score = 176 bits (447), Expect = 1e-42
Identities = 122/416 (29%), Positives = 192/416 (45%), Gaps = 28/416 (6%)
Query: 177 SSKSASGLILNHPFGFSFSG-----LSDEEKENVSLAESLLACAEKVGYQQYERARKLLS 231
SS+S S L G S S L D ++ + L +L+ACAE V A L
Sbjct: 183 SSESDSDLFSTSAIGASNSATRPIVLVDSQENGIQLVHALMACAEAVQQNNLNLAEALEK 242
Query: 232 QIESLSSKTGNPVKRVVHYFAEALCQRIDKETGRFSVSSNNMQKMESLFDPQEVSKDLNP 291
+I L+ +++V +FAEAL +RI + V N P + S +
Sbjct: 243 RIGYLAVSQAGAMRKVATFFAEALARRI------YRVCPEN---------PLDHSMS-DM 286
Query: 292 AMIALYEDLPFSQVSIFTCVQALLENVNDAKKIHVIDLEIRKGCQWTILMQALQSRNECP 351
+ YE P+ + + FT QA+LE K++HVID + +G QW L+QAL R P
Sbjct: 287 LQLHFYESSPYLKFAHFTANQAILEAFEGKKRVHVIDFSMNQGMQWPALLQALALRPSGP 346
Query: 352 LELLKITAIESGNSDTSKHIVEDTGKRLKDFAQSLNIPFSFDIVVVSDLLHIREELFKI- 410
++T I D S ++ +D G +L +++N+ F + V + L + + ++
Sbjct: 347 -PAFRLTGIGPPAPDNSDYL-QDVGWKLAKLVETINVEFEYRGFVANSLADLDASMLELR 404
Query: 411 -DSEETVAVYSQFALRSKIQQPDKLETIMRVIRTINPIVMVVAEIEANHNSKSFVNRFIE 469
E+V V S F L + +P +E +M V++ + P +M V E EANHN F++RF E
Sbjct: 405 PSEVESVVVNSVFELHKLLARPGAIEKVMSVVKQMKPEIMTVVEQEANHNGPVFMDRFTE 464
Query: 470 ALFYFSAYFDCFETCMKGDEKNRFILESMYFSHGIRNIVAEEGAERKSRNVKIDVWRAFF 529
+L Y+S FD E+ +K ++ MY I N+VA EG++R + + WR
Sbjct: 465 SLHYYSTLFDSLESSPNNQDK---MMSEMYLGKQICNVVACEGSDRVEWHETLTQWRTRL 521
Query: 530 TRFGMVETELSMKSLYQAELVAKRFACGYACTFDMNGHCLLVGWKGTPINSVSVWK 585
G L + QA ++ F G + N L +GW P+ S WK
Sbjct: 522 CSSGFEPIHLGSNAFKQASMLLALFGSGEGYRVEENNGSLTLGWHTRPLIVTSAWK 577
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.135 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 960,959,456
Number of Sequences: 2790947
Number of extensions: 39771238
Number of successful extensions: 105672
Number of sequences better than 10.0: 213
Number of HSP's better than 10.0 without gapping: 157
Number of HSP's successfully gapped in prelim test: 56
Number of HSP's that attempted gapping in prelim test: 105028
Number of HSP's gapped (non-prelim): 239
length of query: 587
length of database: 848,049,833
effective HSP length: 133
effective length of query: 454
effective length of database: 476,853,882
effective search space: 216491662428
effective search space used: 216491662428
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 78 (34.7 bits)
Medicago: description of AC144539.4


