

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144539.1 + phase: 0
(163 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
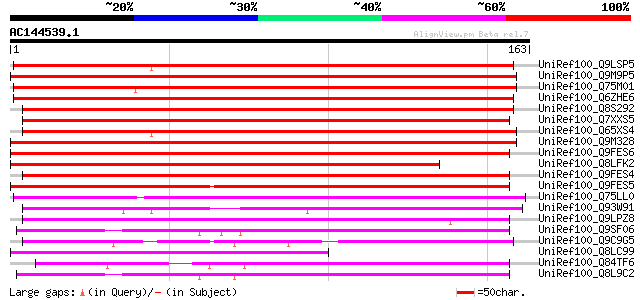
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LSP5 Gb|AAF26101.1 [Arabidopsis thaliana] 196 2e-49
UniRef100_Q9M9P5 T17B22.4 protein [Arabidopsis thaliana] 174 7e-43
UniRef100_Q75M01 Hypothetical protein P0676G05.13 [Oryza sativa] 173 1e-42
UniRef100_Q6ZHE6 Universal stress protein / early nodulin ENOD18... 171 8e-42
UniRef100_Q8S292 P0414E03.3 protein [Oryza sativa] 165 4e-40
UniRef100_Q7XXS5 Hypothetical protein [Oryza sativa] 158 5e-38
UniRef100_Q65XS4 Hypothetical protein P0685E10.5 [Oryza sativa] 152 2e-36
UniRef100_Q9M328 Hypothetical protein F5K20_290 [Arabidopsis tha... 151 7e-36
UniRef100_Q9FES6 Early nodulin ENOD18 [Vicia faba] 141 7e-33
UniRef100_Q8LFK2 Hypothetical protein At3g03270 [Arabidopsis tha... 140 1e-32
UniRef100_Q9FES4 ENOD18 protein [Vicia faba] 139 2e-32
UniRef100_Q9FES5 Early nodulin ENOD18 [Vicia faba] 137 1e-31
UniRef100_Q75LL0 Putative stress-related protein [Oryza sativa] 86 3e-16
UniRef100_Q93W91 Hypothetical protein At3g62550 [Arabidopsis tha... 84 1e-15
UniRef100_Q9LPZ8 T23J18.3 [Arabidopsis thaliana] 79 3e-14
UniRef100_Q9SF06 F26K24.22 protein [Arabidopsis thaliana] 79 4e-14
UniRef100_Q9C9G5 Hypothetical protein T22E19.7 [Arabidopsis thal... 77 2e-13
UniRef100_Q8LC99 Hypothetical protein [Arabidopsis thaliana] 76 3e-13
UniRef100_Q84TF6 At1g09740 [Arabidopsis thaliana] 76 3e-13
UniRef100_Q8L9C2 Ethylene-responsive protein, putative [Arabidop... 75 6e-13
>UniRef100_Q9LSP5 Gb|AAF26101.1 [Arabidopsis thaliana]
Length = 163
Score = 196 bits (498), Expect = 2e-49
Identities = 91/158 (57%), Positives = 117/158 (73%), Gaps = 1/158 (0%)
Query: 2 ASARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPE-EYEHGEMQLWEVTGSPL 60
+ RR+G+A+DFS CS KA W +DN+V++GD+LILI I + YE GEMQLWE GSP
Sbjct: 4 SGGRRIGVAVDFSDCSKKALSWAIDNVVRDGDHLILITIAHDMNYEEGEMQLWETVGSPF 63
Query: 61 TPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPL 120
P+ EF ++ + KKY +K D E L I TA +K + V++K+YWGD REK+C A EQ+PL
Sbjct: 64 IPMSEFSDAAVMKKYALKPDAETLDIVNTAARKKTITVVMKIYWGDPREKICAAAEQIPL 123
Query: 121 DGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKS 158
L MGNRGLG L+R IMGSVSN+VVNN +CPVTVVK+
Sbjct: 124 SSLVMGNRGLGGLKRMIMGSVSNHVVNNVACPVTVVKA 161
>UniRef100_Q9M9P5 T17B22.4 protein [Arabidopsis thaliana]
Length = 159
Score = 174 bits (441), Expect = 7e-43
Identities = 81/159 (50%), Positives = 115/159 (71%)
Query: 1 MASARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPL 60
M AR +G+ MD+SP S A +W +N++++GD +ILI ++P+ +H L+E TGSPL
Sbjct: 1 MGKARTVGVGMDYSPTSKLALRWAAENLLEDGDTVILIHVQPQNADHTRKILFEETGSPL 60
Query: 61 TPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPL 120
PL EF +L K+Y + DPEVL + T KKV V+ KVYWGD REKLC+A+E + L
Sbjct: 61 IPLEEFREVNLSKQYGLAYDPEVLDVLDTLSRAKKVKVVAKVYWGDPREKLCDAVENLKL 120
Query: 121 DGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKSS 159
D + +G+RGLG+L+R ++GSVSN+VV NA+CPVTVVK++
Sbjct: 121 DSIVLGSRGLGSLKRILLGSVSNHVVTNATCPVTVVKAN 159
>UniRef100_Q75M01 Hypothetical protein P0676G05.13 [Oryza sativa]
Length = 167
Score = 173 bits (439), Expect = 1e-42
Identities = 83/159 (52%), Positives = 114/159 (71%), Gaps = 1/159 (0%)
Query: 2 ASARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILI-IIRPEEYEHGEMQLWEVTGSPL 60
A+ R +G A+DFS S A +W DN+++ GD+LIL+ +++ +YE GE LWE TGSPL
Sbjct: 7 AAERWVGAAVDFSEGSRAALRWAADNLLRAGDHLILLHVLKDPDYEQGETLLWEATGSPL 66
Query: 61 TPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPL 120
PL +F + KKY K D E L + T QK+VVV+ KV WGD REKLC+AI ++P+
Sbjct: 67 IPLSDFSEPTIAKKYGAKPDAETLDMLNTVARQKEVVVVFKVLWGDPREKLCQAINEIPM 126
Query: 121 DGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKSS 159
L +G+RGLG L+R ++GSVS+YVVNNA+CPVTVVK++
Sbjct: 127 SCLVIGSRGLGKLKRVLLGSVSDYVVNNATCPVTVVKTA 165
>UniRef100_Q6ZHE6 Universal stress protein / early nodulin ENOD18-like [Oryza sativa]
Length = 165
Score = 171 bits (432), Expect = 8e-42
Identities = 81/157 (51%), Positives = 112/157 (70%)
Query: 2 ASARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPLT 61
A R +G+ MD+SP S A +W VDN+VK GD +IL+ + P+ + +LW+ TGSPL
Sbjct: 3 AEKRTIGLGMDYSPSSKAAAKWAVDNLVKAGDRIILVHVLPKGADASHKELWKSTGSPLI 62
Query: 62 PLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPLD 121
PL EF+ ++ +Y I D EVL+I + K+V VL KVYWGDAREKLCEA++ + ++
Sbjct: 63 PLLEFMEMNVQARYGINPDKEVLEILQAESKSKQVEVLAKVYWGDAREKLCEAVDDLKVN 122
Query: 122 GLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKS 158
+G RGLG L+RA++GSVSNYVVNNA+CPVTVV++
Sbjct: 123 TFVLGCRGLGPLKRALLGSVSNYVVNNATCPVTVVRA 159
>UniRef100_Q8S292 P0414E03.3 protein [Oryza sativa]
Length = 162
Score = 165 bits (417), Expect = 4e-40
Identities = 74/154 (48%), Positives = 113/154 (73%)
Query: 5 RRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPLTPLG 64
RR+G+AMDFSP S KA QW DN++++GD L+L+ IR + + LW TGSPL PL
Sbjct: 8 RRIGVAMDFSPSSKKALQWAADNLLRKGDTLVLLHIRHHGRDEAKNVLWSHTGSPLIPLE 67
Query: 65 EFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPLDGLT 124
E + + + ++Y+I +D EV + +K++ V++K+YWG+ REK+CEA+ ++ L+ L
Sbjct: 68 ELMETAVRQRYDIPSDEEVFDMLNAVSREKELSVVLKMYWGEPREKVCEAVGELNLESLV 127
Query: 125 MGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKS 158
MG+RGLG ++R ++GSV+NYV++NASCPVTVVK+
Sbjct: 128 MGSRGLGQIQRILLGSVTNYVLSNASCPVTVVKA 161
>UniRef100_Q7XXS5 Hypothetical protein [Oryza sativa]
Length = 165
Score = 158 bits (399), Expect = 5e-38
Identities = 72/153 (47%), Positives = 109/153 (71%)
Query: 5 RRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPLTPLG 64
RR+G+AMD+S S +A W + N+++ GD+L+++ + E + LW +GSPL PL
Sbjct: 11 RRIGVAMDYSASSKRALDWAIANLLRRGDHLVVLHVLHHGGEEAKHALWGKSGSPLIPLS 70
Query: 65 EFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPLDGLT 124
EF + ++Y + D EVL + TA Q ++ V+ K+YWGDAREKLC+A+E+ +D L
Sbjct: 71 EFRDPTAMQQYGVHCDAEVLDMLDTAARQLELTVVAKLYWGDAREKLCDAVEEQKIDTLV 130
Query: 125 MGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVK 157
MG+RGLG+++R ++GSV+NYV++NASCPVTVVK
Sbjct: 131 MGSRGLGSIQRILLGSVTNYVLSNASCPVTVVK 163
>UniRef100_Q65XS4 Hypothetical protein P0685E10.5 [Oryza sativa]
Length = 203
Score = 152 bits (385), Expect = 2e-36
Identities = 72/156 (46%), Positives = 103/156 (65%), Gaps = 1/156 (0%)
Query: 5 RRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPE-EYEHGEMQLWEVTGSPLTPL 63
R +G+AMDFS CS A +W ++ + GD L+L+ ++P +YE G LWE GSP+ PL
Sbjct: 27 RNIGVAMDFSACSKAALRWAAASLARPGDRLVLVHVKPSFQYEQGVAHLWEQQGSPMIPL 86
Query: 64 GEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPLDGL 123
E + + + Y + D E + I T+A QK V V+ KVYWG+ +KL EA + +PL L
Sbjct: 87 VELADPRVSRIYGVAPDAETIGILTSAANQKGVEVVAKVYWGEPAKKLTEAAQGIPLHWL 146
Query: 124 TMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKSS 159
+GNRGLG ++R +MGSVS YV N+A+CPVTVV+ +
Sbjct: 147 VVGNRGLGAVKRVLMGSVSTYVANHATCPVTVVREN 182
>UniRef100_Q9M328 Hypothetical protein F5K20_290 [Arabidopsis thaliana]
Length = 160
Score = 151 bits (381), Expect = 7e-36
Identities = 73/159 (45%), Positives = 103/159 (63%)
Query: 1 MASARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPL 60
M R +GIAMDFS S A +W ++N+ +GD + +I P + LW +GSPL
Sbjct: 1 MPKDRNIGIAMDFSESSKNALKWAIENLADKGDTIYIIHTLPLSGDESRNSLWFKSGSPL 60
Query: 61 TPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPL 120
PL EF ++ +KY +KTD L + T QK+V V+ K+YWGDAREKL +A++ + L
Sbjct: 61 IPLAEFREPEIMEKYGVKTDIACLDMLDTGSRQKEVHVVTKLYWGDAREKLVDAVKDLKL 120
Query: 121 DGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKSS 159
D + MG+RGL L+R IMGSVS++V+ +A CPVTVVK +
Sbjct: 121 DSIVMGSRGLSALQRIIMGSVSSFVIQHAPCPVTVVKDN 159
>UniRef100_Q9FES6 Early nodulin ENOD18 [Vicia faba]
Length = 165
Score = 141 bits (355), Expect = 7e-33
Identities = 67/157 (42%), Positives = 103/157 (64%)
Query: 1 MASARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPL 60
M R++G+ +DFS S A +W + N+ +GD LI I + +L+ TGSPL
Sbjct: 1 MVEDRKVGVGIDFSKNSKNALKWAIVNMADKGDTFYLIHINSNSSDESRSKLFAKTGSPL 60
Query: 61 TPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPL 120
PL E + + K+Y ++TD EV+ + A QK+V V+ K+YWGDAR+KL ++IE + L
Sbjct: 61 IPLEELKEAGVMKQYGVQTDVEVIDLLEIAATQKEVSVVAKLYWGDARQKLMDSIEDLKL 120
Query: 121 DGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVK 157
D L +G+RGL T++R ++GSVSN+V+ ++ CPVT+VK
Sbjct: 121 DALVLGSRGLSTIKRILLGSVSNFVMVHSPCPVTIVK 157
>UniRef100_Q8LFK2 Hypothetical protein At3g03270 [Arabidopsis thaliana]
Length = 201
Score = 140 bits (353), Expect = 1e-32
Identities = 65/135 (48%), Positives = 93/135 (68%)
Query: 1 MASARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPL 60
M AR +G+ MD+SP S A +W +N++++GD +ILI ++P+ +H L+E TGSPL
Sbjct: 1 MGKARTVGVGMDYSPTSKLALRWAAENLLEDGDTVILIHVQPQNADHTRKILFEETGSPL 60
Query: 61 TPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPL 120
PL EF +L K+Y + DPEVL + T KKV V+ KVYWGD REKLC+A+E + L
Sbjct: 61 IPLEEFREVNLSKQYGLAYDPEVLDVLDTLSRAKKVKVVAKVYWGDPREKLCDAVENLKL 120
Query: 121 DGLTMGNRGLGTLRR 135
D + +G+RGLG+L+R
Sbjct: 121 DSIVLGSRGLGSLKR 135
>UniRef100_Q9FES4 ENOD18 protein [Vicia faba]
Length = 164
Score = 139 bits (351), Expect = 2e-32
Identities = 66/153 (43%), Positives = 102/153 (66%)
Query: 5 RRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPLTPLG 64
R++G+ +DFS S A +W + N+ +GD LI I + +L+ TGSPL PL
Sbjct: 4 RKVGVGIDFSKNSKNALKWAIVNMADKGDTFYLIHINSNSSDESRNKLFAKTGSPLIPLE 63
Query: 65 EFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPLDGLT 124
E + + K+Y ++TD EV+ + A QK+V V+ K+YWGDAR+KL ++IE + LD L
Sbjct: 64 ELKEAGVMKQYGVQTDVEVIDLLEIAATQKEVSVVAKLYWGDARQKLMDSIEDLKLDALV 123
Query: 125 MGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVK 157
+G+RGL T++R ++GSVSN+V+ ++ CPVT+VK
Sbjct: 124 LGSRGLSTIKRILLGSVSNFVMVHSPCPVTIVK 156
>UniRef100_Q9FES5 Early nodulin ENOD18 [Vicia faba]
Length = 164
Score = 137 bits (345), Expect = 1e-31
Identities = 67/157 (42%), Positives = 103/157 (64%), Gaps = 1/157 (0%)
Query: 1 MASARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPL 60
M R++G+ +DFS S A +W + N+ +GD LI I + +L+ TGSPL
Sbjct: 1 MVEDRKVGVGIDFSKNSKNALKWAIVNMADKGDTFYLIHINSNSSDESRSKLFAKTGSPL 60
Query: 61 TPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPL 120
PL E + + K+Y ++TD EV+ + A QK+V V+ K+YWGDAR+KL ++IE + L
Sbjct: 61 IPL-ELKEAGVMKQYGVQTDVEVIDLLEIAATQKEVSVVAKLYWGDARQKLMDSIEDLKL 119
Query: 121 DGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVK 157
D L +G+RGL T++R ++GSVSN+V+ ++ CPVT+VK
Sbjct: 120 DALVLGSRGLSTIKRILLGSVSNFVMVHSPCPVTIVK 156
>UniRef100_Q75LL0 Putative stress-related protein [Oryza sativa]
Length = 182
Score = 85.9 bits (211), Expect = 3e-16
Identities = 46/161 (28%), Positives = 86/161 (52%), Gaps = 2/161 (1%)
Query: 2 ASARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPLT 61
A ++ +A+D S S+ A W +DN++ + +++ +HG +
Sbjct: 22 AGTMKVVVAVDASEESLNALSWALDNVIGRRAGAVSVVV--VHAQHGPDHFVYPVAAHAI 79
Query: 62 PLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPLD 121
+ +K + + +V+ A +Q++V + GDA+E +C+A+E++ D
Sbjct: 80 AYAPASAIESMRKAQEEISRKVVSRALDVCKQREVSATGAIVEGDAKEAICQAVEEMHAD 139
Query: 122 GLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKSSGQH 162
L +G+RGLG ++RA +GSVS+Y+V++A CPV VVK + H
Sbjct: 140 MLVLGSRGLGKIKRAFLGSVSDYLVHHACCPVLVVKPTKAH 180
>UniRef100_Q93W91 Hypothetical protein At3g62550 [Arabidopsis thaliana]
Length = 162
Score = 84.3 bits (207), Expect = 1e-15
Identities = 52/163 (31%), Positives = 86/163 (51%), Gaps = 15/163 (9%)
Query: 5 RRLGIAMDFSPCSIKAFQWTVDNIVKEGDN--LILIIIRPE--EYEHGEMQLWEVTGSPL 60
R++ +A+D S S++A W++DN+ G N LIL+ ++P Y + + VTG P+
Sbjct: 7 RKIVVAVDESEESMEALSWSLDNLFPYGSNNTLILLYVKPPLPVYSSLDAAGFIVTGDPV 66
Query: 61 TPLGEFINSDLPKKYEIKTDPEVLKIATTAIE--QKKVVVLVKVYWGDAREKLCEAIEQV 118
L KKYE + V+ + T + + + + +V GDA+E +C A++++
Sbjct: 67 AAL---------KKYEYELVESVMARSRTVYQDYESDINIERRVGRGDAKEVICNAVQKL 117
Query: 119 PLDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKSSGQ 161
+D L MG G +RA++GSVS Y CPV +VK Q
Sbjct: 118 RVDMLVMGTHDYGFFKRALLGSVSEYCAKRVKCPVVIVKKQAQ 160
>UniRef100_Q9LPZ8 T23J18.3 [Arabidopsis thaliana]
Length = 875
Score = 79.3 bits (194), Expect = 3e-14
Identities = 49/156 (31%), Positives = 81/156 (51%), Gaps = 3/156 (1%)
Query: 5 RRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPLTPLG 64
R++GIA+D S S A QW V N ++ GD ++L+ ++P +G P
Sbjct: 671 RKIGIAVDLSDESAYAVQWAVQNYLRSGDAVVLLHVQPTSVLYGADWGAMDLSPQWDPNN 730
Query: 65 EFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPLDGLT 124
E L ++I T+ + +A +E + V D +E+LC +E++ L L
Sbjct: 731 EESQRKLEDDFDIVTNKKASDVAQPLVEADIPFKIHIVKDHDMKERLCLEVERLGLSTLI 790
Query: 125 MGNRGLGTLRRAI---MGSVSNYVVNNASCPVTVVK 157
MG+RG G +R+ +GSVS+Y V++ +CPV VV+
Sbjct: 791 MGSRGFGATKRSSKGRLGSVSDYSVHHCACPVVVVR 826
>UniRef100_Q9SF06 F26K24.22 protein [Arabidopsis thaliana]
Length = 200
Score = 79.0 bits (193), Expect = 4e-14
Identities = 51/168 (30%), Positives = 88/168 (52%), Gaps = 18/168 (10%)
Query: 3 SARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGS---- 58
+ +R+ +A+D S S A QW +D+ +L+ E E G + + V
Sbjct: 31 TTKRMVVAIDESDSSFYALQWVIDHFSN-----LLLTTAAAEAESGMLTVIHVQSPFNHF 85
Query: 59 ---PLTPLGE---FINSDL---PKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDARE 109
P P G + +S + KK + +T +L A K++ V G+A+E
Sbjct: 86 AAFPAGPGGATAVYASSSMIESVKKAQQETSAALLSRALQMCRAKQIRTETLVLEGEAKE 145
Query: 110 KLCEAIEQVPLDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVK 157
+CEA+E++ +D L +G+RGLG ++RA +GSVS+Y ++A+CP+ +VK
Sbjct: 146 MICEAVEKMHVDLLVVGSRGLGKIKRAFLGSVSDYCAHHANCPILIVK 193
>UniRef100_Q9C9G5 Hypothetical protein T22E19.7 [Arabidopsis thaliana]
Length = 160
Score = 76.6 bits (187), Expect = 2e-13
Identities = 54/159 (33%), Positives = 87/159 (53%), Gaps = 15/159 (9%)
Query: 5 RRLGIAMDFSPCSIKAFQWTVDNIVKE--GDNLILIIIRPEEYEHGEMQLWEVTGSPLTP 62
+++ +A+D S CS +A QWT+ + ++IL +P H ++ + P
Sbjct: 10 KQVMVAIDESECSKRALQWTLVYLKDSLADSDIILFTAQP----HLDLSCVYASSYGAAP 65
Query: 63 LGEFINS--DLPKKYEIKTDPEVLKI-ATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVP 119
+ E INS + K + E KI A T + +KV+ +G+ +E +CEA E++
Sbjct: 66 I-ELINSLQESHKNAGLNRLDEGTKICAETGVTPRKVLE-----FGNPKEAICEAAEKLG 119
Query: 120 LDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKS 158
+D L +G+ G G L+R +GSVSNY VNNA CPV VV++
Sbjct: 120 VDMLVVGSHGKGALQRTFLGSVSNYCVNNAKCPVLVVRT 158
>UniRef100_Q8LC99 Hypothetical protein [Arabidopsis thaliana]
Length = 126
Score = 75.9 bits (185), Expect = 3e-13
Identities = 37/100 (37%), Positives = 55/100 (55%)
Query: 1 MASARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPL 60
M R +GIAMDFS S A +W ++N+ +GD + +I P + LW +GSPL
Sbjct: 1 MPKDRNIGIAMDFSESSKNALKWAIENLADKGDTIYIIHTLPLSGDESRNSLWFKSGSPL 60
Query: 61 TPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLV 100
PL EF ++ +KY +KTD L + T QK+V + +
Sbjct: 61 IPLAEFREPEIMEKYGVKTDIACLDMLDTGSRQKEVFIRI 100
>UniRef100_Q84TF6 At1g09740 [Arabidopsis thaliana]
Length = 171
Score = 75.9 bits (185), Expect = 3e-13
Identities = 47/160 (29%), Positives = 84/160 (52%), Gaps = 18/160 (11%)
Query: 9 IAMDFSPCSIKAFQWTVDNIV----KEGDNLILIIIRPEEYEHGEMQLWEVTGSPLT-PL 63
+A+D S S++A +W +DN+ + +++ ++P + SP T P
Sbjct: 12 VAVDGSEVSMEALRWALDNLKLSSSSSDSSFVVLHVQPSPSVAAGV-------SPGTIPF 64
Query: 64 GEFINSDLP------KKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQ 117
G ++P ++++ + +L+ A+ +K V V +V GD + K+CEA+E
Sbjct: 65 GGPSGLEVPAFTAAIEQHQKRITDTILEHASQICAEKSVNVKTQVVIGDPKYKICEAVEN 124
Query: 118 VPLDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVK 157
+ D L MG+R G ++R +GSVSNY N+A CPV ++K
Sbjct: 125 LHADLLVMGSRAYGRIKRMFLGSVSNYCTNHAHCPVVIIK 164
>UniRef100_Q8L9C2 Ethylene-responsive protein, putative [Arabidopsis thaliana]
Length = 199
Score = 75.1 bits (183), Expect = 6e-13
Identities = 49/167 (29%), Positives = 86/167 (51%), Gaps = 17/167 (10%)
Query: 3 SARRLGIAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGS---- 58
+ +R+ +A+D S S A Q +D+ +L+ E E G + + V
Sbjct: 31 TTKRMVVAIDESDSSFYALQLVIDHFSN-----LLLTTAAAEAESGMLTVIHVQSPFNHF 85
Query: 59 ---PLTPLGEFINS-----DLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREK 110
P P G + + + KK + +T +L A K++ V G+A+E
Sbjct: 86 AAFPAGPGGATVYASSSMIESVKKAQQETSAALLSRALQMCRAKQIRTETLVLEGEAKEM 145
Query: 111 LCEAIEQVPLDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVK 157
+CEA+E++ +D L +G+RGLG ++RA +GSVS+Y ++A+CP+ +VK
Sbjct: 146 ICEAVEKMHVDLLVVGSRGLGKIKRAFLGSVSDYCAHHANCPILIVK 192
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.318 0.135 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 266,613,628
Number of Sequences: 2790947
Number of extensions: 10646936
Number of successful extensions: 24504
Number of sequences better than 10.0: 326
Number of HSP's better than 10.0 without gapping: 279
Number of HSP's successfully gapped in prelim test: 47
Number of HSP's that attempted gapping in prelim test: 24104
Number of HSP's gapped (non-prelim): 386
length of query: 163
length of database: 848,049,833
effective HSP length: 117
effective length of query: 46
effective length of database: 521,509,034
effective search space: 23989415564
effective search space used: 23989415564
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 69 (31.2 bits)
Medicago: description of AC144539.1


