

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144538.7 + phase: 1 /pseudo
(632 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
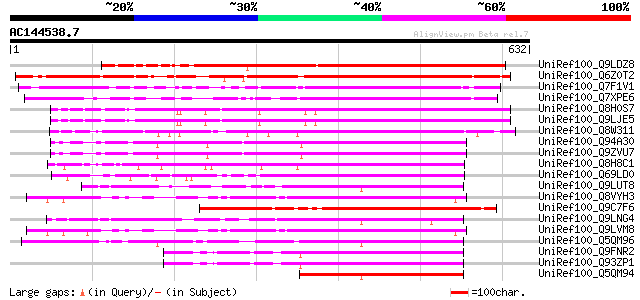
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LDZ8 Gb|AAF35955.1 [Arabidopsis thaliana] 556 e-157
UniRef100_Q6Z0T2 Putative rubisco subunit binding-protein beta s... 521 e-146
UniRef100_Q7F1V1 Putative Rubisco subunit binding-protein beta s... 461 e-128
UniRef100_Q7XPE6 OSJNBa0060N03.15 protein [Oryza sativa] 382 e-104
UniRef100_Q8H0S7 Hypothetical protein At3g13460 [Arabidopsis tha... 374 e-102
UniRef100_Q9LJE5 Gb|AAD10646.1 [Arabidopsis thaliana] 374 e-102
UniRef100_Q8W311 Putative RNA-binding protein [Oryza sativa] 350 7e-95
UniRef100_Q94A30 At1g55500/T5A14_10 [Arabidopsis thaliana] 348 2e-94
UniRef100_Q9ZVU7 T5A14.10 protein [Arabidopsis thaliana] 348 2e-94
UniRef100_Q8H8C1 Putative RNA-binding protein [Oryza sativa] 342 2e-92
UniRef100_Q69LD0 High-glucose-regulated protein 8-like [Oryza sa... 324 6e-87
UniRef100_Q9LUT8 Arabidopsis thaliana genomic DNA, chromosome 3,... 308 2e-82
UniRef100_Q8VYH3 Hypothetical protein [Arabidopsis thaliana] 306 2e-81
UniRef100_Q9C7F6 Hypothetical protein F13K9.7 [Arabidopsis thali... 305 3e-81
UniRef100_Q9LNG4 F21D18.17 [Arabidopsis thaliana] 298 2e-79
UniRef100_Q9LVM8 Similarity to unknown protein [Arabidopsis thal... 296 1e-78
UniRef100_Q5QM96 Rubisco subunit binding-protein beta subunit-li... 286 2e-75
UniRef100_Q9FNR2 Similarity to unknown protein [Arabidopsis thal... 284 5e-75
UniRef100_Q93ZP1 AT5g61020/maf19_20 [Arabidopsis thaliana] 284 5e-75
UniRef100_Q5QM94 Rubisco subunit binding-protein beta subunit-li... 278 3e-73
>UniRef100_Q9LDZ8 Gb|AAF35955.1 [Arabidopsis thaliana]
Length = 503
Score = 556 bits (1434), Expect = e-157
Identities = 295/505 (58%), Positives = 373/505 (73%), Gaps = 32/505 (6%)
Query: 112 MPYGPYSPVTSPLPSVGGDAQLYSPQQFPYT--PPYYNQLVPPPNLPYLNSPTPVSQPEL 169
MPYGPYSP SPLPS G QLYSPQQFP++ PYY Q+VPP ++ Y+ SPT QPEL
Sbjct: 1 MPYGPYSPAASPLPSEG---QLYSPQQFPFSGASPYYQQVVPP-SMQYITSPT---QPEL 53
Query: 170 TNLLGIDQQVESNFFGPRASYPS--VGSFARGSFPVAPGSFSFHESQQGFDGSRSGGLWS 227
T+L+G+DQQ ++ GPR SY +G F G+ P + F E QQGFDG G+WS
Sbjct: 54 TSLVGVDQQGDN--IGPRQSYHPHPIGPF-NGNQP----NLGFPEWQQGFDG----GIWS 102
Query: 228 DSSKPSERQRSFMPLSPSVPQQPIGSVRSFGPSAGMASHQQRSLYGFGSSSNPYGRGYLS 287
D SKPS+ R +SP++ QP+GS S+G + M S +QRS YGFGS SN Y RGY+
Sbjct: 103 DWSKPSDMHRHSSSISPALSPQPLGSYGSYGQNIPMGSQRQRSFYGFGSGSNSYNRGYMH 162
Query: 288 N----PGSSFGGSAISGLSANDRSFLSLENSRRHGR--ETASFCRCNGTLDILSEQNRGP 341
+ GS++G IS + ++ ++ ++NSR GR + + NGT DIL+EQNRGP
Sbjct: 163 SGGRGQGSNYGSRLISNVGMGNQGWIGVDNSRGRGRVSDPSLGGAYNGTFDILNEQNRGP 222
Query: 342 RASKLKNHISSE-NNAIDGSKNNASTAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVH 400
RASK K + E ++A D KNN +AK +ES N DF TD+ +AK F+IKSYSEDNVH
Sbjct: 223 RASKPKTQVLEELDSAADSKKNNKGSAKEHEES-NNADFVTDYTNAKLFIIKSYSEDNVH 281
Query: 401 KSIKYGVWASTPNGNRKLDAAYCQAKEKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDK 460
KSIKY VWASTPNGN+KLDAAY +AK++++ C +FL FSVNAS+QFCGVAEMVGPV+F+K
Sbjct: 282 KSIKYNVWASTPNGNKKLDAAYREAKDEKEPCPLFLLFSVNASSQFCGVAEMVGPVDFEK 341
Query: 461 SVDFWQQDKWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEML 520
SVD+WQQDKWSGQFPVKWHIIKDVPNSQFRHI+LENNDNKPVTNSRDTQEVKL+QGIEML
Sbjct: 342 SVDYWQQDKWSGQFPVKWHIIKDVPNSQFRHIILENNDNKPVTNSRDTQEVKLEQGIEML 401
Query: 521 TIFKNYETDTSILDDFDFYEDRQKAMQERKARQQSSIMPTGLVGG-SEHRSSSDS-TGDF 578
IFKNY+ DTSILDDF FYE+R+K +Q+RKAR+Q S+ G+V G +EH+ +S + DF
Sbjct: 402 KIFKNYDADTSILDDFGFYEEREKIIQDRKARRQPSLPSAGVVAGENEHKPASAALPTDF 461
Query: 579 IKQVSKNFSLVVRLDDNNNEVIAAN 603
+K +SK+F+ VVRLD+ + E + A+
Sbjct: 462 MKNMSKSFAQVVRLDEGSKEAVKAS 486
>UniRef100_Q6Z0T2 Putative rubisco subunit binding-protein beta subunit [Oryza
sativa]
Length = 624
Score = 521 bits (1343), Expect = e-146
Identities = 298/618 (48%), Positives = 383/618 (61%), Gaps = 73/618 (11%)
Query: 8 LHPSIPESSKFSN-SSQGTTTIGHSRDITSQSGSFGSGADQPLYPPNVYAPQAQAFYYGG 66
L P+I S N S+G G TS G G YP +Y+ QAQ FYY G
Sbjct: 46 LEPTISHDSNGINLPSEGQAQAG-----TSNIG----GGHNAAYPQTMYSSQAQPFYYQG 96
Query: 67 --FDNGNGEWDEYPSYVNNGGIEIGSPGVYNENQSLVFHSGYGFNPQMPYGPYSPVTSPL 124
+DN + EWD YP YV+ G+E G VYN++ L++H GYG++P Y YSP+++P+
Sbjct: 97 PGYDNPSNEWDGYPPYVSVEGLEAGPAVVYNDDPQLMYHGGYGYDP---YAHYSPISTPV 153
Query: 125 PS-VGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPELTNLLGIDQQVESNF 183
P+ V GD QLYSPQQF ++ PYY Q VPP +PYL+SPTP+SQ E ++ ID
Sbjct: 154 PAAVSGDGQLYSPQQFSFSAPYYQQSVPP-GMPYLSSPTPISQGEA--MVPID------- 203
Query: 184 FGPRASYPSVGSFARGSFPVAPGSFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFMPLS 243
P+ G+F + ++P SF F + F S G + S
Sbjct: 204 -------PTQGAFIAET--LSPNSFLFGPRPEWFRSSEGNGSFP---------------S 239
Query: 244 PSVPQQPIGSVRS-FGPS-----AGMASHQQRSLYGFGSSSNPYGR-----GYLSNPGSS 292
P+ QP G V FG S +GM S Q R YGFG+ S+ YGR GY P ++
Sbjct: 240 PAASPQPAGGVSGPFGQSNFPMASGMQSPQHRPFYGFGTPSDSYGRVFSHGGYF--PQAT 297
Query: 293 FGGSAISGLSANDRSFLSLENSRRHGRETASFCRCNGTLDILSEQNRGPRASKLKNHISS 352
G N S + +E RR GR A C CNG+LD L+EQ+RGPRA++ K
Sbjct: 298 NYGGPFPSFGLNGTSSIPMEKGRRRGRGNALLCSCNGSLDFLNEQSRGPRATRPKKQPE- 356
Query: 353 ENNAIDGSKNNASTAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTP 412
DG K+ +A E NRPDF T++K+A+FF+IKSYSEDNVHKSIKYGVWAST
Sbjct: 357 -----DGGKDEKPSAGVDCELYNRPDFVTEYKNARFFIIKSYSEDNVHKSIKYGVWASTT 411
Query: 413 NGNRKLDAAYCQAKEKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSG 472
NGN+KLD+AY +AKEK++ C IFL FSVNASAQFCGVAEM+GPV+F+KSVD+WQQDKW+G
Sbjct: 412 NGNKKLDSAYREAKEKEEHCPIFLLFSVNASAQFCGVAEMIGPVDFEKSVDYWQQDKWTG 471
Query: 473 QFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSI 532
QFPVKWHI+KDVPN+ FRHI+LENNDNKPVTNSRDTQEVKL+QG+EML IFK++E D SI
Sbjct: 472 QFPVKWHIVKDVPNNLFRHIILENNDNKPVTNSRDTQEVKLEQGMEMLKIFKDHEEDASI 531
Query: 533 LDDFDFYEDRQKAMQERKAR-QQSSIMPTGLVGGSEHRSSSDSTGDFIKQVSKNFSLVVR 591
LDDFDFYE+R++A+ + KAR Q +P+ V E + D + ++K+F+ VR
Sbjct: 532 LDDFDFYEERERALLDNKARLHQQHQLPSSTV--VEPKKPLTVATDLVGHITKSFAQAVR 589
Query: 592 LDDNNNEVIAANRDKLAS 609
L + N V + DK AS
Sbjct: 590 LGEAKN-VSPNSADKGAS 606
>UniRef100_Q7F1V1 Putative Rubisco subunit binding-protein beta subunit [Oryza
sativa]
Length = 577
Score = 461 bits (1187), Expect = e-128
Identities = 268/589 (45%), Positives = 344/589 (57%), Gaps = 83/589 (14%)
Query: 11 SIPESSKFSNSSQGTTTIGHSRDITSQSGSFGSGADQPLYPPNVYAPQAQAFYYGGFDNG 70
S+P+ + S S +G ++ G G +QP Y PNVYAPQ Q Y GG+ N
Sbjct: 34 SVPQGASSSKSPKGAQ---------EKASFLGKGGEQPFYQPNVYAPQPQTIYSGGYLNH 84
Query: 71 NGEWDEYPSYVNNGGIEIGSPGVYNENQSLVFHSGYGFNPQMPYGPYSPVTSPLPSVGGD 130
G+W+EYP YVN G+ SPG+YN+NQS++ GY NPQM YG YSP GD
Sbjct: 85 LGQWEEYPHYVNMEGLHSVSPGIYNDNQSIMLSPGYANNPQMMYGAYSPGV-------GD 137
Query: 131 AQLYSPQQFPYTPPYYNQLVPP--PNLPYLNSPTPVSQPELTNLLGIDQQVESNFFGPRA 188
Q Y P FP++ PYY PP P++ Y NS T +SQ D ++ +F P
Sbjct: 138 GQPYLPLHFPFSSPYYQ---PPASPSMGYSNSATGMSQG--------DPMLQQEYFLPDG 186
Query: 189 SYPSVGSFARGSFPVAPGSFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFMPLSPSVPQ 248
+ PG +H+ FD + + SS P + PL+
Sbjct: 187 LL----------YSPTPG---YHQPFGSFDRAST----QPSSTPGLFGQGNTPLA----- 224
Query: 249 QPIGSVRSFGPSAGMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGLSANDRSF 308
FG G S+Y GS G FGG+ S S R F
Sbjct: 225 --------FGMHHG-------SMYAPGSYKP-------RQQGGKFGGTTPSWSSG--RRF 260
Query: 309 LSLENSRRHGRETASFCRCNGTLDILSEQNRGPRASKLKNHISSENNAIDGSKNNASTAK 368
+ + S + + F NG L+ L+EQNRGPRA+K K +EN++ID KN +
Sbjct: 261 GTFDLSANQQKGSMPFGIQNGALEFLNEQNRGPRATKPKKQ-DTENSSID-DKNEKNVPL 318
Query: 369 FQDESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKLDAAYCQAKEK 428
E NRPDF T++KDAKFFVIKSY+ED+VH+SIKY VWAST +GNRKLD+AY AKEK
Sbjct: 319 VDSELYNRPDFVTEYKDAKFFVIKSYTEDHVHRSIKYNVWASTASGNRKLDSAYRLAKEK 378
Query: 429 QDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKWHIIKDVPNSQ 488
+D C IFLFFSVN S QFCGVAEM+GPV+FDKSVD+WQQDKWSGQFPVKWHIIKDVPN+
Sbjct: 379 EDYCPIFLFFSVNGSGQFCGVAEMIGPVDFDKSVDYWQQDKWSGQFPVKWHIIKDVPNNL 438
Query: 489 FRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDFYEDRQKAMQE 548
RHI+LENNDNKPVTNSRDTQEVKL+ G++MLTIFKN+E++T+IL+DFDFYE R+KA+QE
Sbjct: 439 LRHIILENNDNKPVTNSRDTQEVKLEHGLQMLTIFKNHESETNILEDFDFYEQREKALQE 498
Query: 549 RKARQQSSIMPTGLVGGSEHRSSSDSTGDFIKQVSKNFSLVVRLDDNNN 597
+ RQQ P + + + + G+ + +S F V+L + N
Sbjct: 499 NR-RQQQPASPE-----LQKPAENKALGELMAHISDTFGQTVQLKETEN 541
>UniRef100_Q7XPE6 OSJNBa0060N03.15 protein [Oryza sativa]
Length = 574
Score = 382 bits (982), Expect = e-104
Identities = 234/579 (40%), Positives = 310/579 (53%), Gaps = 102/579 (17%)
Query: 19 SNSSQGTTTIGHSRDITSQSGSFGSGADQP-LYPPNVYAPQAQAFYYGGFDNGNGEWDEY 77
S + G +++ ++ + G G +Q +Y PNVY PQ + GG+ N G+W+EY
Sbjct: 47 STALHGASSLKSTKSAQEKGSFLGKGGEQHFIYQPNVYTPQPHTVFSGGYLNHLGQWEEY 106
Query: 78 PSYVNNGGIEIGSPGVYNENQSLVFHSGYGFNPQMPYGPYSPVTSPLPSVGGDAQLYSPQ 137
P + G + SP V +S Y SP+P++G D+Q Y P
Sbjct: 107 PHVASADGTDAASP---------VMYSSY---------------SPVPTMG-DSQPYYPL 141
Query: 138 QFPYTPPYYNQLVPP--PNLPYLNSPTPVSQPELTNLLGIDQQVESNFFGPRASYPSVGS 195
P + PYY PP P++ Y NS T +SQ F P Y
Sbjct: 142 HCPLSSPYYQ---PPASPSMGYSNSATGMSQ-----------------FDPMQEYYLPDG 181
Query: 196 FARGSFPVAPGSFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFMPLSPSVPQQPIGSVR 255
P FH+ FDG++ QQ + +
Sbjct: 182 LLYSPTP------GFHQHFGSFDGTQM-------------------------QQSVTGIF 210
Query: 256 SFGPSAGMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGLSANDRSFLSLENSR 315
G + Q S+Y GS R + N FGGS SA R F +
Sbjct: 211 GQGNIPLASGMHQGSMYSSGSYK---ARQQVGN----FGGST-PNWSAASRRFSPFDRGF 262
Query: 316 RHGRETASFCRCNGTLDILSEQNRGPRASKLKNHISSENNAIDGSKNNASTAKFQDESLN 375
+H + G+L+ ++EQNRGPRA+K K + NN+ KN S N
Sbjct: 263 KHDK---------GSLEFMNEQNRGPRATKPKKEV---NNSSTEDKNRKSALINDSNLYN 310
Query: 376 RPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKLDAAYCQAKEKQDACRIF 435
+ DF +++DAKFFVIKSY+ED+VHKSIKYGVWAST +GNRKLDAAY +AKEK+ C IF
Sbjct: 311 QHDFVIEYEDAKFFVIKSYTEDHVHKSIKYGVWASTASGNRKLDAAYREAKEKEATCPIF 370
Query: 436 LFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKWHIIKDVPNSQFRHIVLE 495
LFFSVN S QFCGVAEM+GPV+FDKSVD+WQQDKWSGQFPVKWHIIKDVPNS RHI+LE
Sbjct: 371 LFFSVNGSGQFCGVAEMIGPVDFDKSVDYWQQDKWSGQFPVKWHIIKDVPNSLLRHIILE 430
Query: 496 NNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDFYEDRQKAMQERKARQQS 555
NN+NKPVTNSRDTQEV+L G++MLTIFKN+E +T+IL+DFDFYE R+KAM + + RQ+
Sbjct: 431 NNENKPVTNSRDTQEVRLDHGLQMLTIFKNHEVETTILEDFDFYEQREKAMLDIRQRQKQ 490
Query: 556 SIMPTGLVGGSEHRSSSDSTGDFIKQVSKNFSLVVRLDD 594
+ + + + D + Q+S F+ V+L +
Sbjct: 491 QHTDSEV---QKPMVEAKEPVDLMNQISATFARAVQLGE 526
>UniRef100_Q8H0S7 Hypothetical protein At3g13460 [Arabidopsis thaliana]
Length = 667
Score = 374 bits (961), Expect = e-102
Identities = 231/611 (37%), Positives = 331/611 (53%), Gaps = 68/611 (11%)
Query: 50 YPPNVYAPQAQAFYYGGFDNGNGEWDEYPSYVNNGGIEIGSPGVYNENQSLVFHSGYGFN 109
Y PN Y P YY + +G EW +YP+Y N G+++ S G+Y EN ++V+ GYG+
Sbjct: 72 YVPNPYNPYQ---YYNVYGSGQ-EWTDYPAYTNPEGVDMNS-GIYGENGTVVYPQGYGYA 126
Query: 110 PQMPYGPYSPVTSPLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPEL 169
PYSP TSP P +GG+ QLY QQ+ Y P Y+ P + PY +S +QP+L
Sbjct: 127 AY----PYSPATSPAPQLGGEGQLYGAQQYQY-PNYF-----PNSGPYASSVATPTQPDL 176
Query: 170 TNLLGIDQQVESNFFGPRASYPSVGSFARGSFPV------------------APG---SF 208
+ + AS ++ + GS PV APG +
Sbjct: 177 SANKPAGVKTLPADSNNVASAAAITKGSNGSAPVKPTNQATLNTSSNLYGMGAPGGGLAA 236
Query: 209 SFHESQQGFDGSRSGGLWSDSSKPSERQR--------SFMPLSPSVPQQPIGSVRSFGPS 260
+ + + ++G + W D SK S+ QR S S +VP + RS S
Sbjct: 237 GYQDPRYAYEGYYAPVPWHDGSKYSDVQRPVSGSGVASSYSKSSTVPSSRNQNYRS--NS 294
Query: 261 AGMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGLS---------ANDRSFLSL 311
+ HQ S+ G+G++ Y R Y + +G + S L N R + +
Sbjct: 295 HYTSVHQPSSVTGYGTAQGYYNRMYQNKLYGQYGSTGRSALGYGSSGYDSRTNGRGWAAT 354
Query: 312 ENSRRH-GRETASFCRCNGTLDILSEQNRGPRASKLKNHISSENNAID-----GSKNNAS 365
+N R GR + + +D L+E NRGPRA KN + +++++ G N
Sbjct: 355 DNKYRSWGRGNSYYYGNENNVDGLNELNRGPRAKGTKNQKGNLDDSLEVKEQTGESNVTE 414
Query: 366 TAKFQD-------ESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKL 418
+ + E N+ DF D+ +A FF+IKSYSED+VHKSIKY VWASTPNGN+KL
Sbjct: 415 VGEADNTCVVPDREQYNKEDFPVDYANAMFFIIKSYSEDDVHKSIKYNVWASTPNGNKKL 474
Query: 419 DAAYCQAKEKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKW 478
AAY +A++K C IFLFFSVNAS QF G+AEM GPV+F+ +V++WQQDKW+G FP+KW
Sbjct: 475 AAAYQEAQQKAGGCPIFLFFSVNASGQFVGLAEMTGPVDFNTNVEYWQQDKWTGSFPLKW 534
Query: 479 HIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDF 538
HI+KDVPNS +HI LENN+NKPVTNSRDTQEVKL+QG++++ IFK + + T ILDDF F
Sbjct: 535 HIVKDVPNSLLKHITLENNENKPVTNSRDTQEVKLEQGLKIVKIFKEHSSKTCILDDFSF 594
Query: 539 YEDRQKAMQERKARQQSSIMPTGLVGGSEHRSSSDSTGDFIKQVSKNFSLVVRLDDNNNE 598
YE RQK + E+KA+Q + V + S++ + + S V++ +N +
Sbjct: 595 YEVRQKTILEKKAKQTQKQVSEEKVTDEKKESATAGSASKESPAAVQTSSDVKVAENGSV 654
Query: 599 VIAANRDKLAS 609
D +A+
Sbjct: 655 AKPVTGDVVAN 665
>UniRef100_Q9LJE5 Gb|AAD10646.1 [Arabidopsis thaliana]
Length = 667
Score = 374 bits (960), Expect = e-102
Identities = 231/611 (37%), Positives = 330/611 (53%), Gaps = 68/611 (11%)
Query: 50 YPPNVYAPQAQAFYYGGFDNGNGEWDEYPSYVNNGGIEIGSPGVYNENQSLVFHSGYGFN 109
Y PN Y P YY + +G EW +YP+Y N G+++ S G+Y EN ++V+ GYG+
Sbjct: 72 YVPNPYNPYQ---YYNVYGSGQ-EWTDYPAYTNPEGVDMNS-GIYGENGTVVYPQGYGYA 126
Query: 110 PQMPYGPYSPVTSPLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPEL 169
PYSP TSP P +GG+ QLY QQ+ Y P Y+ P + PY +S +QP+L
Sbjct: 127 AY----PYSPATSPAPQLGGEGQLYGAQQYQY-PNYF-----PNSGPYASSVATPTQPDL 176
Query: 170 TNLLGIDQQVESNFFGPRASYPSVGSFARGSFPV------------------APG---SF 208
+ + AS + + GS PV APG +
Sbjct: 177 SANKPAGVKTLPADSNNVASAAGITKGSNGSAPVKPTNQATLNTSSNLYGMGAPGGGLAA 236
Query: 209 SFHESQQGFDGSRSGGLWSDSSKPSERQR--------SFMPLSPSVPQQPIGSVRSFGPS 260
+ + + ++G + W D SK S+ QR S S +VP + RS S
Sbjct: 237 GYQDPRYAYEGYYAPVPWHDGSKYSDVQRPVSGSGVASSYSKSSTVPSSRNQNYRS--NS 294
Query: 261 AGMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGLS---------ANDRSFLSL 311
+ HQ S+ G+G++ Y R Y + +G + S L N R + +
Sbjct: 295 HYTSVHQPSSVTGYGTAQGYYNRMYQNKLYGQYGSTGRSALGYGSSGYDSRTNGRGWAAT 354
Query: 312 ENSRRH-GRETASFCRCNGTLDILSEQNRGPRASKLKNHISSENNAID-----GSKNNAS 365
+N R GR + + +D L+E NRGPRA KN + +++++ G N
Sbjct: 355 DNKYRSWGRGNSYYYGNENNVDGLNELNRGPRAKGTKNQKGNLDDSLEVKEQTGESNVTE 414
Query: 366 TAKFQD-------ESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKL 418
+ + E N+ DF D+ +A FF+IKSYSED+VHKSIKY VWASTPNGN+KL
Sbjct: 415 VGEADNTCVVPDREQYNKEDFPVDYANAMFFIIKSYSEDDVHKSIKYNVWASTPNGNKKL 474
Query: 419 DAAYCQAKEKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKW 478
AAY +A++K C IFLFFSVNAS QF G+AEM GPV+F+ +V++WQQDKW+G FP+KW
Sbjct: 475 AAAYQEAQQKAGGCPIFLFFSVNASGQFVGLAEMTGPVDFNTNVEYWQQDKWTGSFPLKW 534
Query: 479 HIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDF 538
HI+KDVPNS +HI LENN+NKPVTNSRDTQEVKL+QG++++ IFK + + T ILDDF F
Sbjct: 535 HIVKDVPNSLLKHITLENNENKPVTNSRDTQEVKLEQGLKIVKIFKEHSSKTCILDDFSF 594
Query: 539 YEDRQKAMQERKARQQSSIMPTGLVGGSEHRSSSDSTGDFIKQVSKNFSLVVRLDDNNNE 598
YE RQK + E+KA+Q + V + S++ + + S V++ +N +
Sbjct: 595 YEVRQKTILEKKAKQTQKQVSEEKVTDEKKESATAESASKESPAAVQTSSDVKVAENGSV 654
Query: 599 VIAANRDKLAS 609
D +A+
Sbjct: 655 AKPVTGDVVAN 665
>UniRef100_Q8W311 Putative RNA-binding protein [Oryza sativa]
Length = 688
Score = 350 bits (898), Expect = 7e-95
Identities = 237/622 (38%), Positives = 325/622 (52%), Gaps = 87/622 (13%)
Query: 49 LYPPNVYAPQAQAFYYGGFDNGNGEWDEYPSYVNNGGIEIGSPGVYNENQSLVFHSGYGF 108
LY N Y P A +Y G+D EWD Y + G E+ VY + GYG+
Sbjct: 99 LYQTNGYGPSAY-YYPTGYDGSANEWDS--RYAAHDGTEMPPQSVYGDMY------GYGY 149
Query: 109 NPQMPYGPYSPVTSPLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPE 168
PYGPY P SP+P+VG D Q Y Q + Y YY Q P +N+ SQ E
Sbjct: 150 ---APYGPY-PSGSPVPTVGHDGQSYGAQHYQYPGQYYQQPAPTNASHGVNAVN--SQSE 203
Query: 169 LTNLLGIDQQV-------------------ESNFFGPRASYPSV-----GSFARGSFPVA 204
+ ++ +V S+ + ++ +V GS+ G+
Sbjct: 204 MPSVAAHQARVPVESAKASANGTANGMANTNSSSLARKQTHQNVSVLNNGSYGGGTLQGG 263
Query: 205 P-----GSFSFHESQQGFDGS--RSGGLWSDSSKPSERQRSFMPLSPSVPQQPIGSVRSF 257
P G H Q +DG +G S+++ S S+ + S Q+P ++
Sbjct: 264 PSANNYGHSGLHSPVQWYDGPVYSNGHQRSNTNSTSYGSNSYSAKNQS--QRPTANL--- 318
Query: 258 GPSAGMASHQQRSLYGFGSSSNPY-GRGYLSN----PGSSFGGSAISGLSANDRSFLSLE 312
M H Q G G +S Y R Y N +G + +GL + S
Sbjct: 319 -----MGMHAQIPSSGMGLTSPSYHTRMYPDNRLYGQYGQYGNALKTGLGFGSNMYNSRN 373
Query: 313 N-------SRRHGRETASFCRCNGTLDILSEQNRGPRASKLKNH--------ISSENNAI 357
N S+ R ASF + D +E NRGPR+ K+ I+ + A+
Sbjct: 374 NGRWGIVDSKYKPRGRASFGFGSENQDGFTELNRGPRSGGFKHQKQFGPSVTIAVKGQAL 433
Query: 358 DGSKNNASTAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRK 417
++A N+ F +KDAKFFVIKSYSED+VHKSIKY VWASTPNGN+K
Sbjct: 434 PSVGKQENSAIPDKGQFNQEGFPVTYKDAKFFVIKSYSEDDVHKSIKYNVWASTPNGNKK 493
Query: 418 LDAAYCQAKEKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVK 477
LDA Y +A+EK C +FLFFSVN S QF GVAEMVGPV+F+K+VD+WQQDKW+G FP+K
Sbjct: 494 LDAGYREAQEKSSECPVFLFFSVNTSGQFVGVAEMVGPVDFEKTVDYWQQDKWNGCFPIK 553
Query: 478 WHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFD 537
WH++KDVPN+ +HI L+NNDNKPVTNSRDTQEVKL+QG+EML IFK++ + TSILDDF
Sbjct: 554 WHVVKDVPNNILKHITLDNNDNKPVTNSRDTQEVKLEQGLEMLKIFKDHISKTSILDDFG 613
Query: 538 FYEDRQKAMQERKARQQSSIMPTGLVGGSEH---RSSSDSTGDFIKQ-VSKNFSLVVRLD 593
FYE+RQK MQE++A+QQ + G + + H +++ D KQ +SK + +V
Sbjct: 614 FYENRQKLMQEKRAKQQ-LLQGQGSLDNASHEKEKNAIDGKSTAQKQALSKEGTPIV--- 669
Query: 594 DNNNEVIAANRDKLASDVPTGN 615
E++ A++ + S V GN
Sbjct: 670 ---GEMLNASKSAVESSVTNGN 688
>UniRef100_Q94A30 At1g55500/T5A14_10 [Arabidopsis thaliana]
Length = 549
Score = 348 bits (894), Expect = 2e-94
Identities = 216/534 (40%), Positives = 292/534 (54%), Gaps = 61/534 (11%)
Query: 50 YPPNVYAPQAQAFYYGGFDNGNGEWDEYPSYVNNGGIEIGSPGVYNENQSLVFHSGYGFN 109
Y PNVY Q +YYG +Y Y N+ +++ S GYG+
Sbjct: 29 YVPNVYQ---QPYYYG-------YGSDYTGYTNSESVDMTS--------------GYGYA 64
Query: 110 PQMPYGPYSPVTSPLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPEL 169
PYSP TSP P +GGD QLY QQ+ Y P + + P+ +S +Q +L
Sbjct: 65 AF----PYSPATSPAPQLGGDGQLYGAQQYQYPFP-----LTASSGPFASSVPASTQSKL 115
Query: 170 TNLLGIDQQ---VESNFFGPRASYPSVGSFARGSFPVAPG-SFSFHESQQGFDGSRSGGL 225
+ + + G P S G+ + G + + + + +DG +
Sbjct: 116 STNKAANSASAGIPKGMNGSAPVKPLNQSALYGNSALGGGLAAGYQDPRYSYDGFYTPVS 175
Query: 226 WSDSSKPSERQRSF---------------MPLSPSVPQQPIGSVRSFGPSAGMASHQQRS 270
W D S S+ QRS +P + + S A M + +
Sbjct: 176 WHDGSNFSDVQRSVSGSGVASSYSKANNNVPATRNQNSSSNSHYTSMYQPASMTGYAAQG 235
Query: 271 LYGFGSSSNPYGRGYLSNPGSSFG-GSAISGLSANDRSFLSLENSRRHGRETASFCRCNG 329
Y S + YG+ Y S S G GS+ G N+R +L+ +N R S+ N
Sbjct: 236 YYDRVSPNKSYGQ-YGSTVRSGMGYGSSGYGSRTNERGWLNTDNKYRSRGRGNSYFYGNE 294
Query: 330 TLDILSEQNRGPRA--SKLKNHISSEN---NAIDGSKNNAS-TAKFQD-ESLNRPDFATD 382
+D L+E NRGPRA +K +SSE D S + T D E NR DF +
Sbjct: 295 NIDGLNELNRGPRAKGTKATEEVSSEEVKKQTFDESNTEETVTCVLPDREECNRDDFPVE 354
Query: 383 FKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKLDAAYCQAKEKQDACRIFLFFSVNA 442
+KDAKFF+IKSYSED+VHKSIKY VWASTPNGN+KLDAAY +A++K C +FLFFSVNA
Sbjct: 355 YKDAKFFIIKSYSEDDVHKSIKYNVWASTPNGNKKLDAAYQEAQQKSSGCPVFLFFSVNA 414
Query: 443 SAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPV 502
S QF G+AEM GPV+F+K++++WQQDKW+G FP+KWHI+KDVPNS +HI LE N+NKPV
Sbjct: 415 SGQFIGLAEMKGPVDFNKNIEYWQQDKWTGSFPLKWHILKDVPNSLLKHITLEYNENKPV 474
Query: 503 TNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDFYEDRQKAMQERKARQQSS 556
TNSRDTQEVKL+QG++++ IFK + + T ILDDF FYE RQK + E+KA+QQ S
Sbjct: 475 TNSRDTQEVKLEQGLKVVKIFKEHNSKTCILDDFSFYEARQKTILEKKAKQQQS 528
>UniRef100_Q9ZVU7 T5A14.10 protein [Arabidopsis thaliana]
Length = 580
Score = 348 bits (894), Expect = 2e-94
Identities = 216/534 (40%), Positives = 292/534 (54%), Gaps = 61/534 (11%)
Query: 50 YPPNVYAPQAQAFYYGGFDNGNGEWDEYPSYVNNGGIEIGSPGVYNENQSLVFHSGYGFN 109
Y PNVY Q +YYG +Y Y N+ +++ S GYG+
Sbjct: 72 YVPNVYQ---QPYYYG-------YGSDYTGYTNSESVDMTS--------------GYGYA 107
Query: 110 PQMPYGPYSPVTSPLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPEL 169
PYSP TSP P +GGD QLY QQ+ Y P + + P+ +S +Q +L
Sbjct: 108 AF----PYSPATSPAPQLGGDGQLYGAQQYQYPFP-----LTASSGPFASSVPASTQSKL 158
Query: 170 TNLLGIDQQ---VESNFFGPRASYPSVGSFARGSFPVAPG-SFSFHESQQGFDGSRSGGL 225
+ + + G P S G+ + G + + + + +DG +
Sbjct: 159 STNKAANSASAGIPKGMNGSAPVKPLNQSALYGNSALGGGLAAGYQDPRYSYDGFYTPVS 218
Query: 226 WSDSSKPSERQRSF---------------MPLSPSVPQQPIGSVRSFGPSAGMASHQQRS 270
W D S S+ QRS +P + + S A M + +
Sbjct: 219 WHDGSNFSDVQRSVSGSGVASSYSKANNNVPATRNQNSSSNSHYTSMYQPASMTGYAAQG 278
Query: 271 LYGFGSSSNPYGRGYLSNPGSSFG-GSAISGLSANDRSFLSLENSRRHGRETASFCRCNG 329
Y S + YG+ Y S S G GS+ G N+R +L+ +N R S+ N
Sbjct: 279 YYDRVSPNKSYGQ-YGSTVRSGMGYGSSGYGSRTNERGWLNTDNKYRSRGRGNSYFYGNE 337
Query: 330 TLDILSEQNRGPRA--SKLKNHISSEN---NAIDGSKNNAS-TAKFQD-ESLNRPDFATD 382
+D L+E NRGPRA +K +SSE D S + T D E NR DF +
Sbjct: 338 NIDGLNELNRGPRAKGTKATEEVSSEEVKKQTFDESNTEETVTCVLPDREECNRDDFPVE 397
Query: 383 FKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKLDAAYCQAKEKQDACRIFLFFSVNA 442
+KDAKFF+IKSYSED+VHKSIKY VWASTPNGN+KLDAAY +A++K C +FLFFSVNA
Sbjct: 398 YKDAKFFIIKSYSEDDVHKSIKYNVWASTPNGNKKLDAAYQEAQQKSSGCPVFLFFSVNA 457
Query: 443 SAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPV 502
S QF G+AEM GPV+F+K++++WQQDKW+G FP+KWHI+KDVPNS +HI LE N+NKPV
Sbjct: 458 SGQFIGLAEMKGPVDFNKNIEYWQQDKWTGSFPLKWHILKDVPNSLLKHITLEYNENKPV 517
Query: 503 TNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDFYEDRQKAMQERKARQQSS 556
TNSRDTQEVKL+QG++++ IFK + + T ILDDF FYE RQK + E+KA+QQ S
Sbjct: 518 TNSRDTQEVKLEQGLKVVKIFKEHNSKTCILDDFSFYEARQKTILEKKAKQQQS 571
>UniRef100_Q8H8C1 Putative RNA-binding protein [Oryza sativa]
Length = 716
Score = 342 bits (877), Expect = 2e-92
Identities = 224/567 (39%), Positives = 298/567 (52%), Gaps = 74/567 (13%)
Query: 47 QPLYPPNVYAPQAQAFYYG---------GFDNGNGEWDEYPSYVNNGGIEIGSPGVYNEN 97
Q PN++ A+YYG G+D EWD+YP YVN G+EI +P VY +
Sbjct: 90 QDFMDPNMF--YLPAYYYGELCWFAKLPGYDGSVSEWDDYPRYVNPDGVEI-TPAVYGDI 146
Query: 98 QSLVFHSGYGFNPQMPYGPYSPVTSPLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNL-- 155
GYG+ PYG YSP +SP+P+V D Q++ Q + Y YY P P+
Sbjct: 147 Y------GYGY---APYGAYSPASSPVPTV--DGQMFG-QHYQYPTSYYQPPTPVPSTTQ 194
Query: 156 ----PYLN--SPTPVSQPELTNLLGIDQ-QVESNF----FGPRASYPSVGSFARGSFPVA 204
P N PT + P T G V SN G S+ P+
Sbjct: 195 GDLQPSANPDKPTAKADPAKTTTNGAPNGTVHSNSGTVPLGSSQQNSSLTPDGTYRAPLL 254
Query: 205 PG--SFSFHESQQGFDGSRSGGLWSDSSKPSERQR-----SFMPLSP------------- 244
G S + +S G+D + + W D S + Q+ + M S
Sbjct: 255 GGVPSAGYLDSTYGYDSTGAHFAWYDGSAYTNGQQRTTTTNHMSSSTFSNGSSARTQNKG 314
Query: 245 SVPQQP-IGSVRSFGPSAGMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGLSA 303
S PQQ + + R + A +Y S + YG Y + S G +G +
Sbjct: 315 STPQQMGMNNRRPTTTTGSAAPTYPNRMYPSTRSYSQYGNSYKTGLSYSTNGYGSNGYGS 374
Query: 304 ND-------RSFLSLENSRR-HGRETASFCRCNGTLDILSEQNRGPRASKLKNH------ 349
N R LS++N + GR + N + D E NRGPR+ + KN
Sbjct: 375 NGYDSRLYGRWGLSMDNRYKPRGRGNGYYGFGNESQDGTIELNRGPRSGRFKNQKLFGHT 434
Query: 350 --ISSENNAIDGSKNNASTAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGV 407
I+ + ++ S + +T NR DF + DAKFFVIKSYSED++HKSIKY V
Sbjct: 435 VTIAVKGQSLPTSDSMNATDVPDRTQFNRDDFPVQYDDAKFFVIKSYSEDDIHKSIKYNV 494
Query: 408 WASTPNGNRKLDAAYCQAKEKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQ 467
WAST NGN+KLDAAY +A+ K C IFLFFSVN S QF GVAEM G V+F+K++++WQQ
Sbjct: 495 WASTTNGNKKLDAAYQEAQAKSSKCPIFLFFSVNTSGQFVGVAEMTGAVDFEKTLEYWQQ 554
Query: 468 DKWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYE 527
DKW+G +KWHI+KDVPN+ +HI+LENN+NKPVTNSRDTQEV L QGI+ML IFK +
Sbjct: 555 DKWNGSLSLKWHIVKDVPNNILKHIILENNENKPVTNSRDTQEVNLDQGIQMLKIFKEHV 614
Query: 528 TDTSILDDFDFYEDRQKAMQERKARQQ 554
+ TSILDDF FYE+RQK MQE++ +QQ
Sbjct: 615 SKTSILDDFAFYENRQKLMQEKRVKQQ 641
>UniRef100_Q69LD0 High-glucose-regulated protein 8-like [Oryza sativa]
Length = 602
Score = 324 bits (830), Expect = 6e-87
Identities = 214/538 (39%), Positives = 289/538 (52%), Gaps = 74/538 (13%)
Query: 52 PNVYAPQAQAFYYGGFD----NGNGEWDEYPSYVNNGGIEIGSPGVYNENQSLVFHSGYG 107
P V + A YYG + G W EY +Y+++ G E + G Y + YG
Sbjct: 22 PRVEDYKDAAMYYGTYPAYLYGAYGGWGEYSTYLSHDGAETPTAGAYGDMY-------YG 74
Query: 108 FNPQMPYGPYSPVTSPLPSVGGDAQLYSPQQFPYTPPY---YNQLVPPPNLPYLNSPTPV 164
Y PY TS G D+Q+Y Q + Y P Y N P N +P +
Sbjct: 75 ------YSPYGYSTS-----GHDSQMYGSQHYQYQPTYNKQQNTTGKPSNNGKTENPAAL 123
Query: 165 SQPELTNLLGIDQ---QVESNFFGPRASYPSVGSFARGSFPVAPGSFSFHESQ------- 214
Q +++ G+D Q ++N P+AS + GS GS+ G F +++Q
Sbjct: 124 PQGDVS-ANGVDSLKGQKKTNLL-PKASQNTPGS--NGSYGRPSGRFGNYQNQTNRTTYP 179
Query: 215 ----QGFDG--------SRSGGLWSDSSKPSERQRSFMP--LSPSVPQQPIGSVRSFGPS 260
Q F+G +RS + SK R ++ P + P P+G PS
Sbjct: 180 CYSSQIFNGKQQKLPTGNRSLTTSNSKSKGQSRNQNTYPHLMGLQTPTSPLGP-----PS 234
Query: 261 AGMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGLSANDRSFLSLENSRRHGRE 320
AS +YG+ SS G Y GS GS + G + LS GR
Sbjct: 235 IYSAS----GMYGYNGSSYGSGLWY----GSHLYGSGLYG----GWNALSDGKYNPRGRG 282
Query: 321 TASFCRCNGTLDILSEQNRGPRASKLKNH--ISSENNAIDGSKNNASTAKFQ--DESLNR 376
S+ +G D +E RGPR+ N + + + G + +AS + + NR
Sbjct: 283 NGSYGYIHGNQDGFNELRRGPRSGLFNNQQGVGATVAPVKGQELSASDSSLSVMKDQYNR 342
Query: 377 PDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKLDAAYCQAKEKQDACRIFL 436
DF + DAKFF+IKSYSED+VHKSIKY VWAST NGN+KLDAAY +AKEK +FL
Sbjct: 343 ADFVETYSDAKFFIIKSYSEDDVHKSIKYNVWASTSNGNKKLDAAYQEAKEKSSDSSVFL 402
Query: 437 FFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKWHIIKDVPNSQFRHIVLEN 496
FSVNAS QF G+AEMVG V+F+K+++ WQQDKW+G FPVKWHI+KDVPNS +HI+LEN
Sbjct: 403 LFSVNASGQFVGLAEMVGRVDFNKTLEHWQQDKWTGCFPVKWHIVKDVPNSLLKHIILEN 462
Query: 497 NDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDFYEDRQKAMQERKARQQ 554
N+NKPVTN RDT EVKL+ G+++L IFK++ TS+LDDFDFY++R+K MQERKA+ Q
Sbjct: 463 NENKPVTNCRDTHEVKLEPGLQVLKIFKDHVCKTSLLDDFDFYDNREKMMQERKAKHQ 520
>UniRef100_Q9LUT8 Arabidopsis thaliana genomic DNA, chromosome 3, P1 clone: MGD8
[Arabidopsis thaliana]
Length = 1455
Score = 308 bits (790), Expect = 2e-82
Identities = 189/471 (40%), Positives = 254/471 (53%), Gaps = 65/471 (13%)
Query: 88 IGSPGVYNENQSLVFHS-GYGFNPQMPYGPYSPVTSPLPSVGGDAQLYSPQQFPYTPPYY 146
+G G NEN + ++ YG+ Q PY PY+P P S+G D+ QQ+ PPY
Sbjct: 881 VGLQGGQNENAPYICYTPSYGY-AQSPYNPYNPYI-PGASIGVDSSFVGFQQYYSDPPYE 938
Query: 147 NQLVPPPNLPYLNSPTPVSQPELTNLLGIDQQVESNFFGPRASYPSVGSFARGSFPVAPG 206
+ P +PY+ P VS +L+ Q + GS R
Sbjct: 939 SAASSPTYVPYVIQPDMVSNSSTDSLVANGGQSDGR-----------GSMQR-------- 979
Query: 207 SFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFMPLSPSVPQQPIGSVRSFGPSAGMASH 266
+GS GL D+ K + + P P S ++H
Sbjct: 980 -----------NGSAIAGLPKDAPKSTTTGQYKQPGIPK------------NVSTTASAH 1016
Query: 267 QQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGLSANDRSFLSLENSRRHGRETASFCR 326
+ + ++ PYG+ ++N G S+I+ S S + ++R T S
Sbjct: 1017 SLQGKTAYANTLLPYGKSDIAN-----GVSSIASYSYKPCS--KIYDARGDNNTTGS--- 1066
Query: 327 CNGTLDILSEQNRGPRASKLKNH-ISSENNAIDGSKNNASTAKFQDESLNRPDFATDFKD 385
SEQNRG R + +N I G+ + + N+ DF+ ++ D
Sbjct: 1067 -----TYTSEQNRGSRTRRSRNQLIVKAYTTKAGNADAEGNIVINPDRYNKEDFSIEYSD 1121
Query: 386 AKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKLDAAYCQAK----EKQDACRIFLFFSVN 441
A+FFVIKSYSED+VHKSIKYGVW+ST NGN+KL + Y A+ EK C IFLFFSVN
Sbjct: 1122 ARFFVIKSYSEDDVHKSIKYGVWSSTLNGNKKLQSVYEDAQRIATEKSRECPIFLFFSVN 1181
Query: 442 ASAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKP 501
+S FCGVAEM GPV+FD+ +DFWQQDKWSG FPVKWHIIKDVPNS FRHI+L NN+NKP
Sbjct: 1182 SSGLFCGVAEMTGPVSFDRDMDFWQQDKWSGSFPVKWHIIKDVPNSYFRHIILHNNENKP 1241
Query: 502 VTNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDFYEDRQKAMQERKAR 552
VTNSRDTQE+ L+QG+E+L +FK++ TS+LDDF +YEDRQ+ MQE +AR
Sbjct: 1242 VTNSRDTQEIILKQGLEVLKLFKHHAEKTSLLDDFMYYEDRQRLMQEERAR 1292
>UniRef100_Q8VYH3 Hypothetical protein [Arabidopsis thaliana]
Length = 527
Score = 306 bits (783), Expect = 2e-81
Identities = 208/560 (37%), Positives = 284/560 (50%), Gaps = 103/560 (18%)
Query: 21 SSQGTTTIGHSRDITSQSGSFGSG---ADQPLYPPNVYAPQAQAFYY------GGFDNGN 71
+ G + H S S+ S + P NV ++Y G+ G
Sbjct: 25 ADNGVQQVSHDDGRNIVSSSYSSSVVPSPDPAPYYNVLQTPTNLYHYDWDIANSGYAQGI 84
Query: 72 GEWDEYPSYVNNGGIEIGSPGV-YNENQSLVF-HSGYGFNPQMPYGPYSPVTSPLPSVGG 129
WD YP Y + P V YN+N SL++ + GYGFNP PSV
Sbjct: 85 NTWDGYPRYATTTPEAMHVPPVVYNDNSSLMYQYPGYGFNPY-------------PSVML 131
Query: 130 DAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPELTNLLGIDQQVESNFFGPRAS 189
+ Q+ P +P YY P P P + L D
Sbjct: 132 EGQI------PVSPAYY--------------PQPFGAPSAMHYLPSD------------- 158
Query: 190 YPSVGSFARGSFPVAPGSFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFMPLSPSVPQQ 249
+ P S ++ + G G D S L+ +P
Sbjct: 159 -------------IDPTSAAYMIPYGQYGGGNYSGNQGDIS-----------LTSHIPYP 194
Query: 250 PIGSVRSFGPSAGMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGLSANDRSFL 309
+ GP AS Q +L+G G +S+ GY S S ++R L
Sbjct: 195 QTMGI--LGPYDHNAS--QVALHGSGVASSSSLGGYYHVGSYQNPSSTPSYYGVDNRVRL 250
Query: 310 SLENSRRHGRETASFCRCNGTLDILSEQNRGPRAS-KLKNHISSEN-NAIDGSKNNASTA 367
+ + +R ++ S + T D+ NRGPRAS ++K+ SS+ + I S +++STA
Sbjct: 251 TPDIGKRREKDQGSI---SSTSDLYG--NRGPRASSRVKSKNSSKPCSTIGDSASDSSTA 305
Query: 368 KFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKLDAAYCQAKE 427
N P+F TD+K+AKFF++KS+SEDNVH+SIKY VWASTP+GN+KLD AY A++
Sbjct: 306 GPNPSLYNHPEFVTDYKNAKFFIVKSFSEDNVHRSIKYNVWASTPHGNKKLDTAYRDAEK 365
Query: 428 KQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKWHIIKDVPNS 487
C IFLFFSVNAS QFCGV+EMVGPV+F+K +WQQD+WSGQFPVKWHI+KD+PN+
Sbjct: 366 MGGKCPIFLFFSVNASGQFCGVSEMVGPVDFEKDAGYWQQDRWSGQFPVKWHIVKDIPNN 425
Query: 488 QFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDFYED------ 541
+F HI+L+NNDNKPVT+SRD+QEVKL+QGIEML IFK YE TSILDDF +Y++
Sbjct: 426 RFCHILLQNNDNKPVTHSRDSQEVKLRQGIEMLRIFKEYEAHTSILDDFGYYDELEGQKV 485
Query: 542 -----RQKAMQERKARQQSS 556
R+KA +E + +Q S
Sbjct: 486 GEDGTRKKAGEEETSVEQLS 505
>UniRef100_Q9C7F6 Hypothetical protein F13K9.7 [Arabidopsis thaliana]
Length = 542
Score = 305 bits (780), Expect = 3e-81
Identities = 176/364 (48%), Positives = 233/364 (63%), Gaps = 20/364 (5%)
Query: 232 PSERQRSFMPLSPSVP-QQPIGSVRSFGPSAGMASHQQRSLYGFGSSSNPYGR--GYLSN 288
P + F PL+ + P + +GS M F S++N R Y
Sbjct: 191 PFSGENMFPPLASTRPCMRHLGSAELLANDFNMGPAHGGLHDAFESAANLSYREQAYAQC 250
Query: 289 PGSSFGGSAISGLSANDRSFLSLENSRRHGRETASFCRCNGTLDILSEQNRGPRASKLKN 348
S F S+ S ND + L+ +R F C LD+L+E NRGPRAS+L
Sbjct: 251 KRSPFSASSSSPTWENDYNLPPLDEARSESYN--DFSHCPAMLDMLTESNRGPRASRL-- 306
Query: 349 HISSENNAIDGSKNNASTAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVW 408
+S++ I + + +F + L + F+DAKFFVIKSYSEDNVHKSIK+ VW
Sbjct: 307 --NSKSKMISYDRVD----RFCQQEL-----LSQFRDAKFFVIKSYSEDNVHKSIKHCVW 355
Query: 409 ASTPNGNRKLDAAYCQAKEKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQD 468
AST NGN+KLDAAY +AK+K AC +FL FSVNAS+QFCGVAEMVGPV+F+ SV++WQQD
Sbjct: 356 ASTKNGNKKLDAAYREAKKKDVACPVFLLFSVNASSQFCGVAEMVGPVDFNTSVEYWQQD 415
Query: 469 KWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYET 528
+WSG FPV+W I+KDVPNS FRHI++E+NDNKPVTNSRDTQEV L++GIEML IF + E
Sbjct: 416 RWSGHFPVQWLIVKDVPNSLFRHIIIESNDNKPVTNSRDTQEVGLEKGIEMLDIFISCEM 475
Query: 529 DTSILDDFDFYEDRQKAMQERKARQQSSIMPTGLVGGSEHRSSSDSTGDFIKQVSKNFSL 588
+SILDDF+FYE+RQ A+Q+RKARQ++ + L S S DF++++SK+F+
Sbjct: 476 RSSILDDFNFYEERQIAIQDRKARQRAVLESLALSATSAPTYSLHD--DFVREMSKHFAE 533
Query: 589 VVRL 592
+ L
Sbjct: 534 ALAL 537
>UniRef100_Q9LNG4 F21D18.17 [Arabidopsis thaliana]
Length = 664
Score = 298 bits (764), Expect = 2e-79
Identities = 200/540 (37%), Positives = 269/540 (49%), Gaps = 69/540 (12%)
Query: 46 DQPLYPPNVYAPQAQAFYYGGFDNGNGEWDEYPSYVNNGGIEIGSPGVYNENQSLVFHS- 104
DQ +Y P + +Y G+++ G+W+ + + G E+ G NEN + ++
Sbjct: 13 DQGMYYP---VDASYGYYCTGYESP-GDWENHQMFFGVDGSEVQYTGGQNENSPYICYTP 68
Query: 105 GYGFNPQMPYGPYSPVTSPLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPV 164
YG+ Q PY P++P P S+G D+ +PQQF PPY + P +PY P V
Sbjct: 69 SYGY-AQSPYNPFNPYI-PGASIGVDSAFVAPQQFYSIPPYQSVATSPTFVPYAIQPEIV 126
Query: 165 SQPELTNLL--GIDQQVESNFFGPRASYPSVGSFARGSFPVAPGSFSFHESQQGFDGSRS 222
S +L+ G + S+ G R + + + + P P S
Sbjct: 127 SNSSTNSLVETGSANRGRSDGRGSRQRSGTATAGLQRNDPKLPAGNSL------------ 174
Query: 223 GGLWSDSSKPSERQRSFMPLSPSVPQQPIGSVRSFGPSAGMASHQQRSLYGFGSSSNPYG 282
G S+ +P+ Q + S G R G + SSS
Sbjct: 175 -GKISEKPRPNSGQSRQSEMDKSDSTSSSGQARQ-----GRVTSVSAQPVDVVSSSRV-- 226
Query: 283 RGYLSNPGSSFGGSAISGLSANDRSFLSLENSRRHGRETASFCRCNGTLDILSEQNRGPR 342
SSF I+ ND S ++ N+ + N D + EQNRG R
Sbjct: 227 --------SSFRQLDIAPPQLNDFSKIATNNNNIRPKLYGG--HANIIPDTVREQNRGRR 276
Query: 343 ASKLKNH-ISSENNAIDGSKNNASTAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHK 401
+ L N I G+ + N+ D D+ +AKFFVIKSYSED+VHK
Sbjct: 277 SRALGNQLIVKAYTTKAGNADAEGNIVINPSQYNKEDLRIDYSNAKFFVIKSYSEDDVHK 336
Query: 402 SIKYGVWASTPNGNRKLDAAYCQAK----EKQDACRIFLFFSVNASAQFCGVAEMVGPVN 457
SIKY VW+ST +GN+KL +AY A+ EK C IFLFFSVNAS FCG+AEM GPV+
Sbjct: 337 SIKYNVWSSTLHGNKKLQSAYEDAQRIATEKSCECPIFLFFSVNASGLFCGMAEMTGPVS 396
Query: 458 FDKSVDFWQQDKWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVK----- 512
FDK +DFWQQDKWSG FPVKWHIIKDVPNS FRHI+L+NN+NKPVTNSRDTQEV
Sbjct: 397 FDKDMDFWQQDKWSGSFPVKWHIIKDVPNSYFRHIILQNNENKPVTNSRDTQEVNFPTRP 456
Query: 513 --------------------LQQGIEMLTIFKNYETDTSILDDFDFYEDRQKAMQERKAR 552
L+QG+E+L IFK++ TS+LDDF +YE RQ+ MQ+ + R
Sbjct: 457 RFEDFHIQNLLIYHFLLQIMLKQGLEVLKIFKDHMERTSLLDDFVYYESRQRVMQDERTR 516
>UniRef100_Q9LVM8 Similarity to unknown protein [Arabidopsis thaliana]
Length = 552
Score = 296 bits (758), Expect = 1e-78
Identities = 208/585 (35%), Positives = 284/585 (47%), Gaps = 128/585 (21%)
Query: 21 SSQGTTTIGHSRDITSQSGSFGSG---ADQPLYPPNVYAPQAQAFYY------GGFDNGN 71
+ G + H S S+ S + P NV ++Y G+ G
Sbjct: 25 ADNGVQQVSHDDGRNIVSSSYSSSVVPSPDPAPYYNVLQTPTNLYHYDWDIANSGYAQGI 84
Query: 72 GEWDEYPSYVNNGGIEIGSPGV--------------------------YNENQSLVF-HS 104
WD YP Y + P V YN+N SL++ +
Sbjct: 85 NTWDGYPRYATTTPEAMHVPPVSMTCPIYVLSSSCYLINLVILYPQVVYNDNSSLMYQYP 144
Query: 105 GYGFNPQMPYGPYSPVTSPLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPV 164
GYGFNP PSV + Q+ P +P YY P P
Sbjct: 145 GYGFNPY-------------PSVMLEGQI------PVSPAYY--------------PQPF 171
Query: 165 SQPELTNLLGIDQQVESNFFGPRASYPSVGSFARGSFPVAPGSFSFHESQQGFDGSRSGG 224
P + L D + P S ++ + G G
Sbjct: 172 GAPSAMHYLPSD--------------------------IDPTSAAYMIPYGQYGGGNYSG 205
Query: 225 LWSDSSKPSERQRSFMPLSPSVPQQPIGSVRSFGPSAGMASHQQRSLYGFGSSSNPYGRG 284
D S L+ +P + GP AS Q +L+G G +S+ G
Sbjct: 206 NQGDIS-----------LTSHIPYPQTMGI--LGPYDHNAS--QVALHGSGVASSSSLGG 250
Query: 285 YLSNPGSSFGGSAISGLSANDRSFLSLENSRRHGRETASFCRCNGTLDILSEQNRGPRAS 344
Y S S ++R L+ + +R ++ S + T D+ NRGPRAS
Sbjct: 251 YYHVGSYQNPSSTPSYYGVDNRVRLTPDIGKRREKDQGSI---SSTSDLYG--NRGPRAS 305
Query: 345 -KLKNHISSEN-NAIDGSKNNASTAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHKS 402
++K+ SS+ + I S +++STA N P+F TD+K+AKFF++KS+SEDNVH+S
Sbjct: 306 SRVKSKNSSKPCSTIGDSASDSSTAGPNPSLYNHPEFVTDYKNAKFFIVKSFSEDNVHRS 365
Query: 403 IKYGVWASTPNGNRKLDAAYCQAKEKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSV 462
IKY VWASTP+GN+KLD AY A++ C IFLFFSVNAS QFCGV+EMVGPV+F+K
Sbjct: 366 IKYNVWASTPHGNKKLDTAYRDAEKMGGKCPIFLFFSVNASGQFCGVSEMVGPVDFEKDA 425
Query: 463 DFWQQDKWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTI 522
+WQQD+WSGQFPVKWHI+KD+PN++F HI+L+NNDNKPVT+SRD+QEVKL+QGIEML I
Sbjct: 426 GYWQQDRWSGQFPVKWHIVKDIPNNRFCHILLQNNDNKPVTHSRDSQEVKLRQGIEMLRI 485
Query: 523 FKNYETDTSILDDFDFYED-----------RQKAMQERKARQQSS 556
FK YE TSILDDF +Y++ R+KA +E + +Q S
Sbjct: 486 FKEYEAHTSILDDFGYYDELEGQKVGEDGTRKKAGEEETSVEQLS 530
>UniRef100_Q5QM96 Rubisco subunit binding-protein beta subunit-like [Oryza sativa]
Length = 609
Score = 286 bits (731), Expect = 2e-75
Identities = 205/558 (36%), Positives = 278/558 (49%), Gaps = 64/558 (11%)
Query: 15 SSKFSNSSQGTTTIGHSRDITS--QSGSFGSGADQPLYPPNVYAPQAQAFYYGGFDNGN- 71
S+K SN + T S D S SG S + +YYG + +
Sbjct: 24 STKTSNVNLPATKGASSSDAISCISSGDAASTVKESEMNQEASVGDQGMYYYGYYYPASF 83
Query: 72 GEWDEYPSYVNNGGIEIGSPGVYNENQS-LVFHSGY--GFNPQMPYGPYSPVTSPLPSVG 128
G +DE +V G+E+ V +N S L + GY G+ YSP+ P G
Sbjct: 84 GGYDENGYFVGYNGLEVHPTVVQGDNGSYLCYLPGYENGYT-------YSPIV-PGVIAG 135
Query: 129 GDAQLYSPQQFPYTPPYYNQL-VPPPNLPYLNSPTPVSQPELTNLLGIDQQ------VES 181
D Q S + PYY+ + + P+ P + + PEL D V+
Sbjct: 136 VDGQYISKE------PYYSTISMQDPSTPGIFAQPVAYGPELVPAYTWDPSFALLDGVQG 189
Query: 182 NFFGP-RASYPSVGSFARGSFPVAPGSFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFM 240
G + +YP+ ++ P + S + + GS S D+ S
Sbjct: 190 RPVGVHQTNYPARPKYSSNKLPSSKASRNTKSASDTIKGSSSA---LDTMSTSANG---Y 243
Query: 241 PLSPSVPQQPIGSVRSFGP-SAGMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAIS 299
P S + + S+ P S+ A H + S +
Sbjct: 244 PSSKTANKASGASISKGYPLSSKFAVHTNQGKDNLYQSKD-------------------I 284
Query: 300 GLSANDRSFLSLENSRRHGRETA-SFCRCNGTLDILSEQNRGPRASKLKNHISSENNAID 358
G+ + RS+ S E + + C + L S+ + P+ ++A
Sbjct: 285 GMKESGRSWNSTEKLKARSKLNGYGDCDISDNLTDNSKNSLSPQGGHY-----GLSSAGG 339
Query: 359 GSKNNASTAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKL 418
G+ S ++ N PDF T + A FFVIKSYSED++HKSIKY VWASTPNGN++L
Sbjct: 340 GNDVTPSPVAMSRDAYNLPDFVTKYDQALFFVIKSYSEDDIHKSIKYNVWASTPNGNKRL 399
Query: 419 DAAYCQAKE----KQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQF 474
D A+ A+E K C +FLFFSVNAS QFCGVAEMVGPV+F+++++FWQQDKW+G F
Sbjct: 400 DNAFKLAQERVAEKGTKCPMFLFFSVNASGQFCGVAEMVGPVDFNRNMNFWQQDKWNGFF 459
Query: 475 PVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSILD 534
PVKWHIIKDVPN QFRHI+LENN+NKPVTNSRDTQEVK QG EML IFKN+ TSILD
Sbjct: 460 PVKWHIIKDVPNPQFRHIILENNENKPVTNSRDTQEVKFPQGSEMLNIFKNFSCKTSILD 519
Query: 535 DFDFYEDRQKAMQERKAR 552
DFDFYE+RQK MQ+R+ +
Sbjct: 520 DFDFYENRQKVMQDRRGK 537
>UniRef100_Q9FNR2 Similarity to unknown protein [Arabidopsis thaliana]
Length = 493
Score = 284 bits (727), Expect = 5e-75
Identities = 158/374 (42%), Positives = 221/374 (58%), Gaps = 37/374 (9%)
Query: 188 ASYPSVGSFARGSFPVAPGSFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFMPLSPSVP 247
A+Y + GS+ +G++ ++ + G+ GS + G + Q ++
Sbjct: 80 AAYNAKGSYGKGAYAYGYYPPAYQYPRHGYTGSYASG-------KTNLQYQYLT------ 126
Query: 248 QQPIGSVRSFGPSAGMASHQQRSLYGF-GSSSNPYGRGYLSNPGSSFGGSAISGLSANDR 306
QQ G G S G S YG G +N YG G + + A +
Sbjct: 127 QQ--GRSAGNGQSYGGYMDNIYSNYGMCGPYTNGYGYGSYGYDSWKY----MPNWYAVNN 180
Query: 307 SFLSLENSRRHGRETASFCRCNGTLDILSEQNRGPRASKLKNHISS--------ENNAID 358
++ +G+E ++ L+E NRGPRA + S E +
Sbjct: 181 TYKPRNGYHGYGKEN---------IEGLNEMNRGPRAKGFNSQDGSKVMAVSLKEQRVTE 231
Query: 359 GSKNNASTAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKL 418
K + + + N+ DF + +AKF+VIKSYSED++HKSIKY VW+STPNGN+KL
Sbjct: 232 TEKLSEDVSLLDPKDYNKIDFPETYTEAKFYVIKSYSEDDIHKSIKYSVWSSTPNGNKKL 291
Query: 419 DAAYCQAKEKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKW 478
DA+Y +AK+K D C +FL FSVN S QF G+AEMVGPV+F+K+V++WQQDKW G FPVKW
Sbjct: 292 DASYNEAKQKSDGCPVFLLFSVNTSGQFVGLAEMVGPVDFNKTVEYWQQDKWIGCFPVKW 351
Query: 479 HIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDF 538
H +KD+PNS RHI LENN+NKPVTNSRDTQEVKL+QGI+++ IFK++ + T ILDDF+F
Sbjct: 352 HFVKDIPNSSLRHITLENNENKPVTNSRDTQEVKLEQGIKVIKIFKDHASKTCILDDFEF 411
Query: 539 YEDRQKAMQERKAR 552
YE+RQK +QERK++
Sbjct: 412 YENRQKIIQERKSK 425
>UniRef100_Q93ZP1 AT5g61020/maf19_20 [Arabidopsis thaliana]
Length = 495
Score = 284 bits (727), Expect = 5e-75
Identities = 158/374 (42%), Positives = 221/374 (58%), Gaps = 37/374 (9%)
Query: 188 ASYPSVGSFARGSFPVAPGSFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFMPLSPSVP 247
A+Y + GS+ +G++ ++ + G+ GS + G + Q ++
Sbjct: 82 AAYNAKGSYGKGAYAYGYYPPAYQYPRHGYTGSYASG-------KTNLQYQYLT------ 128
Query: 248 QQPIGSVRSFGPSAGMASHQQRSLYGF-GSSSNPYGRGYLSNPGSSFGGSAISGLSANDR 306
QQ G G S G S YG G +N YG G + + A +
Sbjct: 129 QQ--GRSAGNGQSYGGYMDNIYSNYGMCGPYTNGYGYGSYGYDSWKY----MPNWYAVNN 182
Query: 307 SFLSLENSRRHGRETASFCRCNGTLDILSEQNRGPRASKLKNHISS--------ENNAID 358
++ +G+E ++ L+E NRGPRA + S E +
Sbjct: 183 TYKPRNGYHGYGKEN---------IEGLNEMNRGPRAKGFNSQDGSKVMAVSLKEQRVTE 233
Query: 359 GSKNNASTAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKL 418
K + + + N+ DF + +AKF+VIKSYSED++HKSIKY VW+STPNGN+KL
Sbjct: 234 TEKLSEDVSLLDPKDYNKIDFPETYTEAKFYVIKSYSEDDIHKSIKYSVWSSTPNGNKKL 293
Query: 419 DAAYCQAKEKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKW 478
DA+Y +AK+K D C +FL FSVN S QF G+AEMVGPV+F+K+V++WQQDKW G FPVKW
Sbjct: 294 DASYNEAKQKSDGCPVFLLFSVNTSGQFVGLAEMVGPVDFNKTVEYWQQDKWIGCFPVKW 353
Query: 479 HIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDF 538
H +KD+PNS RHI LENN+NKPVTNSRDTQEVKL+QGI+++ IFK++ + T ILDDF+F
Sbjct: 354 HFVKDIPNSSLRHITLENNENKPVTNSRDTQEVKLEQGIKVIKIFKDHASKTCILDDFEF 413
Query: 539 YEDRQKAMQERKAR 552
YE+RQK +QERK++
Sbjct: 414 YENRQKIIQERKSK 427
>UniRef100_Q5QM94 Rubisco subunit binding-protein beta subunit-like [Oryza sativa]
Length = 458
Score = 278 bits (712), Expect = 3e-73
Identities = 133/203 (65%), Positives = 160/203 (78%), Gaps = 4/203 (1%)
Query: 354 NNAIDGSKNNASTAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPN 413
++A G+ S ++ N PDF T + A FFVIKSYSED++HKSIKY VWASTPN
Sbjct: 184 SSAGGGNDVTPSPVAMSRDAYNLPDFVTKYDQALFFVIKSYSEDDIHKSIKYNVWASTPN 243
Query: 414 GNRKLDAAYCQAKE----KQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDK 469
GN++LD A+ A+E K C +FLFFSVNAS QFCGVAEMVGPV+F+++++FWQQDK
Sbjct: 244 GNKRLDNAFKLAQERVAEKGTKCPMFLFFSVNASGQFCGVAEMVGPVDFNRNMNFWQQDK 303
Query: 470 WSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETD 529
W+G FPVKWHIIKDVPN QFRHI+LENN+NKPVTNSRDTQEVK QG EML IFKN+
Sbjct: 304 WNGFFPVKWHIIKDVPNPQFRHIILENNENKPVTNSRDTQEVKFPQGSEMLNIFKNFSCK 363
Query: 530 TSILDDFDFYEDRQKAMQERKAR 552
TSILDDFDFYE+RQK MQ+R+ +
Sbjct: 364 TSILDDFDFYENRQKVMQDRRGK 386
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.313 0.132 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,188,942,536
Number of Sequences: 2790947
Number of extensions: 57592348
Number of successful extensions: 131156
Number of sequences better than 10.0: 634
Number of HSP's better than 10.0 without gapping: 163
Number of HSP's successfully gapped in prelim test: 495
Number of HSP's that attempted gapping in prelim test: 129253
Number of HSP's gapped (non-prelim): 1659
length of query: 632
length of database: 848,049,833
effective HSP length: 134
effective length of query: 498
effective length of database: 474,062,935
effective search space: 236083341630
effective search space used: 236083341630
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 78 (34.7 bits)
Medicago: description of AC144538.7


