

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144538.1 - phase: 0 /pseudo
(942 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
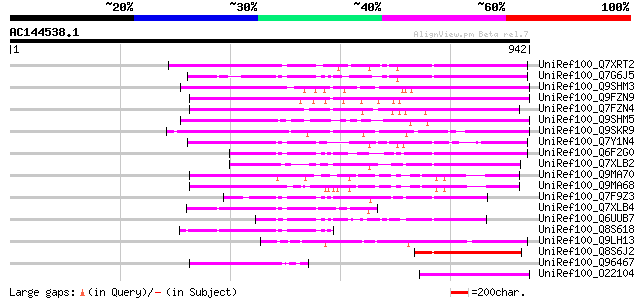
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q7XRT2 OSJNBa0042F21.11 protein [Oryza sativa] 360 8e-98
UniRef100_Q7G6J5 Putative polyprotein [Oryza sativa] 290 2e-76
UniRef100_Q9SHM3 F7F22.17 [Arabidopsis thaliana] 289 3e-76
UniRef100_Q9FZN9 Retroelement pol polyprotein-like [Arabidopsis ... 277 9e-73
UniRef100_Q7FZN4 T17A2.3 protein [Arabidopsis thaliana] 271 5e-71
UniRef100_Q9SHM5 F7F22.15 [Arabidopsis thaliana] 270 1e-70
UniRef100_Q9SKR9 Putative Athila retroelement ORF1 protein [Arab... 255 4e-66
UniRef100_Q7Y1N4 Putative retrotransposon gag protein [Oryza sat... 255 5e-66
UniRef100_Q6F2G0 Hypothetical protein OSJNBa0083K01.26 [Oryza sa... 238 8e-61
UniRef100_Q7XLB2 OSJNBa0011K22.17 protein [Oryza sativa] 237 1e-60
UniRef100_Q9MA70 F2J6.11 protein [Arabidopsis thaliana] 222 5e-56
UniRef100_Q9MA68 F2J6.13 protein [Arabidopsis thaliana] 220 1e-55
UniRef100_Q7F9Z3 OSJNBa0079C19.24 protein [Oryza sativa] 205 5e-51
UniRef100_Q7XLB4 OSJNBa0011K22.15 protein [Oryza sativa] 181 7e-44
UniRef100_Q6UUB7 Hypothetical protein [Oryza sativa] 173 2e-41
UniRef100_Q8S618 Hypothetical protein OSJNBb0048O22.17 [Oryza sa... 163 3e-38
UniRef100_Q9LH13 Similarity to Athila retroelement ORF1 protein ... 162 3e-38
UniRef100_Q8S6J2 Hypothetical protein OSJNBa0019N10.34 [Oryza sa... 155 5e-36
UniRef100_Q96467 Hypothetical protein [Hordeum vulgare] 155 7e-36
UniRef100_O22104 Vicia faba mRNA expressed from retrotransposon-... 150 1e-34
>UniRef100_Q7XRT2 OSJNBa0042F21.11 protein [Oryza sativa]
Length = 920
Score = 360 bits (925), Expect = 8e-98
Identities = 237/681 (34%), Positives = 355/681 (51%), Gaps = 58/681 (8%)
Query: 289 WIESWSKRKGLGADLVRKYEDSETKIRLPKVRSLARPFSGSPTEDPNLHISSFLRLSG-- 346
W E + + +++ D + KI+ K R A PF G P ED N H+ FL +
Sbjct: 231 WREKVYHNSKIYTERTKRWHDKQIKIKKFKPRDKASPFCGKPNEDANAHLLQFLEICSMY 290
Query: 347 TIKE-NQEAVRLHLFPFSLRDRASAWFHSLEVGSITSWDDMRRAFLARFSPPSKTAKLRD 405
TIK + +AVRL LFPFSL RA WF++ +I +WD +FL++F P KT LR
Sbjct: 291 TIKGVSPDAVRLRLFPFSLLRRAKQWFYA-NCAAINTWDKCSTSFLSKFFPIGKTNALRG 349
Query: 406 QITRFNQKDGESLYEA*ERFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSVDAAAGGA 465
+I+RF Q ES+ EA ER +E + CPHHG++ WLI+ FYNGL+ ++ +DAAAGGA
Sbjct: 350 RISRFQQTRDESIPEAWERLQEYVAACPHHGMDDWLILQNFYNGLTLMSRDHLDAAAGGA 409
Query: 466 LMNKNYTEAYALIEDMAQNHYQWTNERAVTTPTPSKKEAGIYEVSEYNHLAAKVEALTQK 525
+K A LIE M N W+ ER T ++ G++ V E LAAK++ L +
Sbjct: 410 FFSKTVQGAVELIEKMVSNT-GWSKERLQT------RQRGMHTVKETELLAAKLDLLMKC 462
Query: 526 I---EKLNVNAAQPSPASPTCEVCGITGHTGVDCQLGSAANIEQLNYAQYNQGMRPNQNF 582
+ EK + + TCEVCG TGH+G DC + + M +N
Sbjct: 463 LDDHEKRPQGTVKALDSHVTCEVCGGTGHSGNDC------------LETHEEAMYMGKNN 510
Query: 583 YKNPQGSYGQTAP----PGYTNNQRVAQKSSLEILLENCMMNQNKQLQELKNQTGSLNDS 638
PQG G P PG NN + + SL L+ Q K L+ + + +
Sbjct: 511 EYRPQGGEGWNQPRPYYPGGNNNGNFSNQPSLMDLV----FAQAKTTDALRKKLAANDKI 566
Query: 639 LSKLNTKVDSIAT-------HTKMLETQISQVAQQVAISSQTPGVFPGQTETNPKAHVNA 691
L +N K+D A+ KM+ETQ++Q+A V + G PGQ +++ + +V A
Sbjct: 567 LENINVKLDGFASAFQNQLSFNKMIETQLAQLASLVPANES--GRIPGQPDSSIE-NVKA 623
Query: 692 ISLGGNKLEET------------ITKAKSVKGEIVKLLGEKNAIKTPLDKNKTLNPLRLT 739
I+ G K + S + ++ +K + D P +
Sbjct: 624 ITTRGGKSTRDPPYPNPAGTNGMSKETPSTDSDDKEIQPDKTVPQEYCDTRLLPFPQQSR 683
Query: 740 KLNLEAQFAKFLNILKRICIKIPFAEALSRMPLYAKFLREIFSKKKAIDHKETIALTRE- 798
K +++ QFA F+ ++++I I +P +A+ ++P YA++L++I + K+ + E + LT
Sbjct: 684 KPSVDKQFACFVEVIQKIHINVPLLDAM-QVPTYARYLKDILNNKRPLPTTEVVKLTEHC 742
Query: 799 SSAIVKKQPQKLRDPGSFAIPCVIGKETVDKALCHLGASVSLLPLSLFKRMGIGELKPTE 858
S+ I+ K P+K +DPG I C IG + D+ALC LGASVS++P +F ++ L PT
Sbjct: 743 SNLILHKLPEKKKDPGCPTITCSIGAQQFDQALCDLGASVSVMPKDVFDKLNFTVLAPTP 802
Query: 859 MILKLADRSTIPLAGYIEDIPVKIEGIYIPTDFVVVDIEEDHDVPIILGRPFLATAGAII 918
M L+LAD S G ED+P KI +IP DFVV+D++ + P+ILGRPFL+TAGA I
Sbjct: 803 MRLQLADSSVHYPVGIAEDVPAKIRDFFIPVDFVVLDMDTGKETPLILGRPFLSTAGANI 862
Query: 919 DVQSGRIVFQASDAMIGFELE 939
DV +G I F + FE +
Sbjct: 863 DVGTGSIRFHTNGKEEKFEFQ 883
>UniRef100_Q7G6J5 Putative polyprotein [Oryza sativa]
Length = 689
Score = 290 bits (741), Expect = 2e-76
Identities = 207/636 (32%), Positives = 308/636 (47%), Gaps = 90/636 (14%)
Query: 323 ARPFSGSPTEDPNLHISSFLRLSGTIK---ENQEAVRLHLFPFSLRDRASAWFHSLEVGS 379
A PF G P ED N H+ FL + T + + V+L LFPFSL RA WF++ +
Sbjct: 42 ASPFCGKPNEDANAHLQQFLEICSTYTIKGVSPDVVKLRLFPFSLLGRAKQWFYANRT-A 100
Query: 380 ITSWDDMRRAFLARFSPPSKTAKLRDQITRFNQKDGESLYEA*ERFKEMLRLCPHHGLEK 439
+ +WD AFL++F P EA ER +E + CPH G++
Sbjct: 101 VNTWDKCSTAFLSKFFP----------------------MEAWERLQEYVAACPHLGMDD 138
Query: 440 WLIVHTFYNGLSYTTKMSVDAAAGGALMNKNYTEAYALIEDMAQNHYQWTNERAVTTPTP 499
WLI+ FYNGL+ ++ +DAAA GA +K A LIE M N W+ ER T
Sbjct: 139 WLILQNFYNGLTPMSRDHLDAAARGAFFSKTVQGAVELIEKMVSN-MGWSEERLQT---- 193
Query: 500 SKKEAGIYEVSEYNHLAAKVEALTQKI---EKLNVNAAQPSPASPTCEVCGITGHTGVDC 556
++ G++ V E LAAK++ L +++ EK + + CEVCG TGH+G DC
Sbjct: 194 --RQRGMHTVKETELLAAKLDLLMKRLDDHEKGPQRTVKALDSHVMCEVCGNTGHSGNDC 251
Query: 557 QLGSAANIEQLNYAQYNQGMRPNQNFYKNPQGSYGQTAPPGYTNNQRVAQKSSLEILLEN 616
+E A Y N N Y PQG G P Y N
Sbjct: 252 -------LETCEEAMY----MGNNNGYC-PQGGQGWNQPRPYYQGGN-----------NN 288
Query: 617 CMMNQNKQLQELKNQTGSLNDSLSKLNTKVDSIATHTKMLETQISQVAQQVAISSQTPGV 676
L++L D+LSK D +A+ ET G
Sbjct: 289 SNFPNQPSLKDLVFAQAKTTDALSKKLAANDKLASLVPANET----------------GR 332
Query: 677 FPGQTETNPKAHVNAISLGGNKLEET-----------ITKAKSVKGEIVKLLGEKNAIKT 725
PGQ + + + +V AI++ K I+K K + +N +
Sbjct: 333 IPGQPDPSIE-NVKAITMRRGKSTRDPPYPNPVGTNEISKGAPSNDSADKEVQPENTVPQ 391
Query: 726 PLDKNKTLN-PLRLTKLNLEAQFAKFLNILKRICIKIPFAEALSRMPLYAKFLREIFSKK 784
+ L+ P R+ K +++ QFA+F+ ++++I I +P +A+ ++P YA++L++I + K
Sbjct: 392 EYCDTRLLSFPQRMRKPSVDEQFARFVEVIQKIHINVPLLDAM-QVPTYARYLKDILNNK 450
Query: 785 KAIDHKETIALTRE-SSAIVKKQPQKLRDPGSFAIPCVIGKETVDKALCHLGASVSLLPL 843
+ + E + LT + S+ I+ K P+K +DPG I C IG + D+ C+LGASVS++
Sbjct: 451 RPLPTTEVVKLTEQCSNLILHKLPEKKKDPGCPTITCSIGAQQFDQVSCNLGASVSVVLK 510
Query: 844 SLFKRMGIGELKPTEMILKLADRSTIPLAGYIEDIPVKIEGIYIPTDFVVVDIEEDHDVP 903
+F ++ L PT M L+LAD S AG ED+PVKI +IP DFVV+D++ + P
Sbjct: 511 DVFDKLNFTVLAPTPMRLQLADSSVRYPAGIAEDVPVKIRDFFIPVDFVVLDMDTGKETP 570
Query: 904 IILGRPFLATAGAIIDVQSGRIVFQASDAMIGFELE 939
+ILGRPFL+TAGA IDV +G I F + FE +
Sbjct: 571 LILGRPFLSTAGANIDVGTGSIRFHINGKKEKFEFQ 606
>UniRef100_Q9SHM3 F7F22.17 [Arabidopsis thaliana]
Length = 1799
Score = 289 bits (739), Expect = 3e-76
Identities = 227/755 (30%), Positives = 359/755 (47%), Gaps = 147/755 (19%)
Query: 310 SETKIRLPKVRSLARP---FSGSPTEDPNLHISSFLRLSGTIKEN---QEAVRLHLFPFS 363
+ T + K R+ P F G P EDP H+ F RL K N ++ +L LFPFS
Sbjct: 18 NRTAREIRKRRTTVNPGNKFHGLPMEDPLDHLDEFDRLCNLTKINGVSEDGFKLRLFPFS 77
Query: 364 LRDRASAWFHSLEVGSITSWDDMRRAFLARFSPPSKTAKLRDQITRFNQKDGESLYEA*E 423
L D+A W +L SIT+WDD ++ FL++F ++TA+LR++I+ F+QK GES EA E
Sbjct: 78 LGDKAHIWEKNLPHDSITTWDDCKKVFLSKFFSNARTARLRNEISGFSQKTGESFCEAWE 137
Query: 424 RFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSVDAAAGGALMNKNYTEAYALIEDMAQ 483
RFK CPHHG K ++ T Y G+ +M +D A+ G NK+ E + L+E++AQ
Sbjct: 138 RFKGYTNQCPHHGFTKASLLSTLYRGVLPRIRMLLDTASNGNFQNKDVEEGWELVENLAQ 197
Query: 484 N--HYQWTNERAVTTPTPSKKEAGIYEVSEYNHLAAKVEALTQKIEKLNVNA-------- 533
+ +Y +R V S + +++AL K++++ ++
Sbjct: 198 SDGNYNEDCDRTVRGTADSDDKH-----------RKEIKALNDKLDRILLSQHKHVHFLV 246
Query: 534 ------AQPSPASPTCEVCGITGHTG--------------VDCQLGSAANIEQLNY---- 569
Q + EV I + G + + + AN + Y
Sbjct: 247 DDEQYEVQDGEGNQLEEVSYINNNQGGYKGYNYFKTNNPNLSYRSTNVANPQDQVYPPQQ 306
Query: 570 --------AQYNQGMRPNQNFYKNPQGSYGQTAPPGYTNNQRVA-------QKSSLEILL 614
YNQG P Q F QG+Y Q PPG+ Q K L+ LL
Sbjct: 307 QQSQNKPFVPYNQGFVPKQQF----QGNY-QQQPPGFAPPQHQGPAAPDADMKQMLQQLL 361
Query: 615 E---NCMMNQNKQLQELKNQTGSLNDSLSKLNTKVDSIATHTKMLETQISQVAQQVAISS 671
+C M K++ EL N+ L+ S + LN K++++ T + LE + + ++
Sbjct: 362 HGQASCSMEMAKKISELHNK---LDCSYNDLNVKMETLDTKVRYLEGHSTSSS-----AT 413
Query: 672 QTPGVFPGQTETNPKAHVNAISLGGNKLEETITKAKSVK-------GEIVKL-------- 716
+ PG+ NPK + +AI+L K T + K+V GE + L
Sbjct: 414 KQTSQLPGKAVQNPKEYAHAITLRSGKALPTREEPKTVTEDSEDQDGEDLSLEKDQADKP 473
Query: 717 ----------LGEKNAIKTPLD------------------------------------KN 730
L + ++ PLD K+
Sbjct: 474 LDLSLEQPLDLSLQQSLDPPLDSFTRPTTRPVIPAASPTAPKPVAVKNKEKVFVPPPYKS 533
Query: 731 KTLNPLRLTKLNLEAQFAKFLNILKRICIKIPFAEALSRMPLYAKFLREIFSKKKAIDHK 790
+ P R K + A F +K + ++IP +AL+ +P KFL+++ ++ + +
Sbjct: 534 QLPFPGRHKKALADKYRAMFAKNIKEVELRIPLVDALALIPDSHKFLKDLIVERIQ-EVQ 592
Query: 791 ETIALTRESSAIVKKQ--PQKLRDPGSFAIPCVIGKETVDKALCHLGASVSLLPLSLFKR 848
+ L+ E SAI++K+ P+KL DPGSF +PC +G ++ LC LGASVSL+PLS+ KR
Sbjct: 593 GMVVLSHECSAIIQKKIIPKKLSDPGSFTLPCSLGPLAFNRCLCDLGASVSLMPLSVAKR 652
Query: 849 MGIGELKPTEMILKLADRSTIPLAGYIEDIPVKIEGIYIPTDFVVVDIEEDHDVPIILGR 908
+G + K + L LADRS G +E++P++I + IPTDFVV++++E+ P+ILGR
Sbjct: 653 LGFTQYKSCNISLILADRSVRIPHGLLENLPIRIGAVEIPTDFVVLEMDEEPKDPLILGR 712
Query: 909 PFLATAGAIIDVQSGRIVFQ-ASDAMIGFELENVM 942
PFLATAGA+IDV+ G+I D + F++++ M
Sbjct: 713 PFLATAGAMIDVKKGKIDLNLGKDFRMTFDVKDAM 747
>UniRef100_Q9FZN9 Retroelement pol polyprotein-like [Arabidopsis thaliana]
Length = 1864
Score = 277 bits (709), Expect = 9e-73
Identities = 214/701 (30%), Positives = 336/701 (47%), Gaps = 90/701 (12%)
Query: 326 FSGSPTEDPNLHISSFLRLSGTIKEN---QEAVRLHLFPFSLRDRASAWFHSLEVGSITS 382
F G P EDP H+ F RL K N ++ +L LFPFSL D+A W SL GSITS
Sbjct: 86 FHGLPMEDPLDHLDEFDRLCSLTKINGVSEDGFKLRLFPFSLGDKAHQWEKSLLQGSITS 145
Query: 383 WDDMRRAFLARFSPPSKTAKLRDQITRFNQKDGESLYEA*ERFKEMLRLCPHHGLEKWLI 442
W+D ++AFLA+F S+TA+LR+ I+ F Q + E+ EA ERFK CPHHG K +
Sbjct: 146 WNDCKKAFLAKFFSNSRTARLRNDISGFTQTNNETFCEAWERFKGYQTQCPHHGFSKASL 205
Query: 443 VHTFYNGLSYTTKMSVDAAAGGALMNKNYTEAYALIEDMAQN--HYQWTNERAVTTPTPS 500
+ T Y G+ +M +D A+ G +NK+ + + L+E++AQ+ +Y +R+V T + S
Sbjct: 206 LSTLYRGVLPKIRMLLDTASNGNFLNKDVEDGWELVENLAQSDGNYNEDYDRSVRTSSDS 265
Query: 501 KKEAGIYEVSEYNHLAAKVEALTQKI-------EKLNVNAAQPSPASPTCEVCGITG--- 550
E E+ N K+ + QK E V + + V G
Sbjct: 266 D-EKHRREMKAMNDKLDKLLLVQQKHIHFLGDDETFQVQDGETMQSEEVSYVQNQGGYNK 324
Query: 551 --------HTGVDCQLGSAANIEQLNYAQ-----------YNQGMR--PNQNFYKNPQGS 589
H + + + AN + Y YNQG P Q + QG+
Sbjct: 325 GFNNFKQNHPNLSYRSTNVANPQDQVYPSQQQNQPRPFVPYNQGQGYVPKQQY----QGN 380
Query: 590 YG-QTAPPGYTNNQR----VAQKSSLEILLENCMMNQNKQLQELKNQTGSLND----SLS 640
Y Q PPG+T Q+ S L+ +L+ + Q +L + +++ S +
Sbjct: 381 YQPQLPPPGFTQQQQQPASTTPDSDLKNMLQQILQGQATGAMDLSKRMAEIHNKVDCSYN 440
Query: 641 KLNTKVDSIATHTKMLETQISQVAQQV-----------------AISSQTPGVFPGQTET 683
+N KV+++ + + +E Q A AI+ ++ P +
Sbjct: 441 DINIKVEALTSKIRYIEGQTGSTAAPKFTGPSRKSMSNSEEYAHAITLRSGKELPTKESP 500
Query: 684 NPKAHVNAISLG------GNKLEETITK----------AKSVKGEIVKLLGEK---NAIK 724
N + G GN E+ I + A + + K K N
Sbjct: 501 NQNTEDSVDQDGEDFCQNGNSAEKAIEEPILDQPTRLLAPAASPLVEKPAATKTKDNVFV 560
Query: 725 TPLDKNKTLNPLRLTKLNLEAQFAKFLNILKRICIKIPFAEALSRMPLYAKFLREIFSKK 784
P K P R K+ ++ A LK + + +P + L+ +P K+++++ +++
Sbjct: 561 PPPYKPPLPFPGRFKKVMIQKYKALLEKQLKNLEVTMPLVDCLALIPDSNKYVKDMITER 620
Query: 785 KAIDHKETIALTRESSAIVKKQ--PQKLRDPGSFAIPCVIGKETVDKALCHLGASVSLLP 842
+ + + L+ E SAI++++ P+KL DPGSF +PC +G +K LC LGASVSL+P
Sbjct: 621 IK-EVQGMVVLSHECSAIIQQKIIPKKLGDPGSFTLPCALGPLAFNKCLCDLGASVSLMP 679
Query: 843 LSLFKRMGIGELKPTEMILKLADRSTIPLAGYIEDIPVKIEGIYIPTDFVVVDIEEDHDV 902
LS+ K++G + KP + L LADRS G +ED+PV I + +PTDFVV++++E+
Sbjct: 680 LSVAKKLGFNKYKPCNISLILADRSVRIPHGLLEDLPVMIGMVEVPTDFVVLEMDEEPKD 739
Query: 903 PIILGRPFLATAGAIIDVQSGRIVFQ-ASDAMIGFELENVM 942
P+ILGRPFL T GAIIDV+ G+I D + F++ N M
Sbjct: 740 PLILGRPFLVTVGAIIDVKKGKIDLNLGRDLKMTFDITNTM 780
>UniRef100_Q7FZN4 T17A2.3 protein [Arabidopsis thaliana]
Length = 1428
Score = 271 bits (694), Expect = 5e-71
Identities = 205/653 (31%), Positives = 317/653 (48%), Gaps = 73/653 (11%)
Query: 326 FSGSPTEDPNLHISSFLRLSGTIKEN---QEAVRLHLFPFSLRDRASAWFHSLEVGSITS 382
F G P EDP H+ F RL K N ++ +L LFPFSL D+A W +L SIT+
Sbjct: 54 FHGLPMEDPLDHLDEFDRLCNLTKINGVSEDGFKLRLFPFSLGDKAHIWEKNLSHDSITT 113
Query: 383 WDDMRRAFLARFSPPSKTAKLRDQITRFNQKDGESLYEA*ERFKEMLRLCPHHGLEKWLI 442
WDD ++AFL++F ++TA+LR++I F+QK GES EA ERFK CPHH K +
Sbjct: 114 WDDYKKAFLSKFFSNARTARLRNEIYGFSQKTGESFCEAWERFKGYTNQCPHHSFTKASL 173
Query: 443 VHTFYNGLSYTTKMSVDAAAGGALMNKNYTEAYALIEDMAQN--HYQWTNERAVTTPTPS 500
+ T Y G+ +M +D A+ G NK+ E + L+E++AQ+ +Y +R V T
Sbjct: 174 LSTLYRGVLPRIRMLLDTASNGNFQNKDVEEGWELVENLAQSDGNYNEDCDRTV-RGTAD 232
Query: 501 KKEAGIYEVSEYNHLAAKVEALTQKIEKLNVNAAQPSPASPTCEVCGITGHTGVDCQLGS 560
+ E+ N ++ QK V+ Q +V G+ + L
Sbjct: 233 SDDKHRKEIKALNDKLDRILLSQQKHVHFLVDDEQ-------YQVQDGEGNQLEEVYLPQ 285
Query: 561 AANIEQLNYAQYNQGMRPNQNFYKNPQGSYGQTAPPGYTNNQ---RVAQKSSLEILLENC 617
+ + YNQG P Q F QG+Y PPG+ Q A + ++ +++
Sbjct: 286 QKQGQNKPFVLYNQGFVPKQQF----QGNYQPPPPPGFVTQQNQCHAAPDAKMKQMVKQL 341
Query: 618 MMNQNKQLQELKNQTGSLNDSL----SKLNTKVDSIATHTKMLETQISQVAQQVAISSQT 673
+ Q E+ + L+ L + LN KV+++ T + LE Q + S+ +
Sbjct: 342 LQGQASSSMEIAKKLSELHHKLDCSYNDLNAKVEALNTKVRYLEGQ--------SASTFS 393
Query: 674 PGV--FPGQTETNPKAHVNAIS--------LGGNKLEETITK---------AKSVKGEIV 714
P V PG++ NPK + A + L + + IT+ + V+ +V
Sbjct: 394 PKVTRLPGKSIQNPKEYATAHAITICHDRELPTRHVLDLITRDNDVQEGEASTQVEASVV 453
Query: 715 KL-------------LGEKNAIKTPLDKNKTLNPLRLTKLN---LEAQFAKFLNI----L 754
+ EK AI + K PL L L +A ++ ++ L
Sbjct: 454 EFNHSAGSRHLTQSTSEEKAAIIERMVKRFKPTPLPLRALPWTFRKAWMERYKSVAAKQL 513
Query: 755 KRICIKIPFAEALSRMPLYAKFLREIFSKKKAIDHKETIALTRESSAIVKKQ--PQKLRD 812
I +P E L+ + K +R + ++ + H S K+ +KL D
Sbjct: 514 DEIEAVMPLMEVLNLILDPHKVVRNLILERIKMYHDSDDESDATPSRAADKRIVQEKLED 573
Query: 813 PGSFAIPCVIGKETVDKALCHLGASVSLLPLSLFKRMGIGELKPTEMILKLADRSTIPLA 872
PGSF +PC IG+ LC LGASV+L+PLS+ +R+ + KP ++ L LADRS+
Sbjct: 574 PGSFTLPCSIGEFAFSDCLCDLGASVNLMPLSMARRLEFIQYKPCDLTLILADRSSRKHF 633
Query: 873 GYIEDIPVKIEGIYIPTDFVVVDIEEDHDVPIILGRPFLATAGAIIDVQSGRI 925
G ++D+PV I G+ +PTDFVV+D+E +H P+ILGRPFLA+ GA+IDV+ G+I
Sbjct: 634 GMLKDLPVMINGVEVPTDFVVLDMEVEHKDPLILGRPFLASVGAVIDVKEGKI 686
>UniRef100_Q9SHM5 F7F22.15 [Arabidopsis thaliana]
Length = 1862
Score = 270 bits (691), Expect = 1e-70
Identities = 206/669 (30%), Positives = 331/669 (48%), Gaps = 102/669 (15%)
Query: 310 SETKIRLPKVRSLARP---FSGSPTEDPNLHISSFLRLSGTIKEN---QEAVRLHLFPFS 363
+ T + K R+ P F G P EDP H+ F RL K N ++ +L LFPFS
Sbjct: 153 NRTAREIRKRRTTVNPGNKFHGLPMEDPLDHLDEFDRLCNLTKINGVSEDGFKLRLFPFS 212
Query: 364 LRDRASAWFHSLEVGSITSWDDMRRAFLARFSPPSKTAKLRDQITRFNQKDGESLYEA*E 423
L D+A W +L SIT+WDD ++AFL++F ++TA+LR++I+ F+QK GES EA E
Sbjct: 213 LGDKAHIWEKNLPHDSITTWDDCKKAFLSKFFSNARTARLRNEISGFSQKTGESFCEAWE 272
Query: 424 RFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSVDAAAGGALMNKNYTEAYALIEDMAQ 483
RFK C HHG K ++ T Y G+ +M +D A+ G NK+ E + L+E++AQ
Sbjct: 273 RFKGYTNQCSHHGFTKASLLSTLYRGVLPRIRMLLDTASNGNFQNKDVEEGWELVENLAQ 332
Query: 484 NHYQWTNERAVTTPTPSKKEAGIYEVSEYNHLAAKVEALTQKIEKLNVNAAQPSPASPTC 543
++ + NE T + S+ H +++AL K++++ ++ +
Sbjct: 333 SNGNY-NENCDRTVRGTAD-------SDDKH-RKEIKALNDKLDRILLSQHK-------- 375
Query: 544 EVCGITGHTGVDCQLGSAANIEQLNYAQYNQGMRPNQNFYKNPQGSYGQTAPPGYTNNQR 603
V + + Q G +E+++Y NQG N +K TNN
Sbjct: 376 HVHFLVDDEQYEVQDGEGNQLEEVSYINNNQGGYKGYNNFK--------------TNNPN 421
Query: 604 VAQKSSLEILLENCMMNQNKQLQELKNQTGSLNDSLSKLNTKVDSIATHTKMLETQISQV 663
++ +S+ + N Q+ + Q S +K + T L
Sbjct: 422 LSYRSTN-------VANPQDQVYPPQQQ-----QSQNKPFVPYNQATKQTSQL------- 462
Query: 664 AQQVAISSQTPGVFPGQTETNPKAHVNAISLGGNKLEETITKAKSVKGEIVKLLGEKNAI 723
PG+ NPK + +AI+L K T + K+V + GE ++
Sbjct: 463 --------------PGKAVQNPKEYAHAITLHSGKALPTREEPKTVTEDSEDQDGEDLSL 508
Query: 724 KT-----PLDKNKTLNPLRLT-KLNLEAQFAKFLNILKR--------------------- 756
K PLD + PL L+ + +L+ F R
Sbjct: 509 KKDQADKPLDLSLE-QPLDLSLQQSLDPPLDSFTRPTTRPVIPAASPTAPKPVAVKNKEK 567
Query: 757 ICIKIPFAEALSRMPLYAKFLREIFSKKKAIDHKETIALTRESSAIVKKQ--PQKLRDPG 814
+ ++IP +AL+ +P KFL+++ ++ + + + L+ E SAI++K+ P+KL DPG
Sbjct: 568 VELRIPLVDALALIPDSHKFLKDLIVERIQ-EVQGMVVLSHECSAIIQKKIIPKKLSDPG 626
Query: 815 SFAIPCVIGKETVDKALCHLGASVSLLPLSLFKRMGIGELKPTEMILKLADRSTIPLAGY 874
SF +PC +G ++ LC LGASVSL+PLS+ KR+G + K + L LADRS G
Sbjct: 627 SFTLPCSLGPLAFNRCLCDLGASVSLMPLSVAKRLGFTQYKSCNISLILADRSVRIPHGL 686
Query: 875 IEDIPVKIEGIYIPTDFVVVDIEEDHDVPIILGRPFLATAGAIIDVQSGRIVFQ-ASDAM 933
+E++P++I + IPTDFVV++++E+ P+ILGRPFLATAGA+IDV+ G+I D
Sbjct: 687 LENLPIRIGAVEIPTDFVVLEMDEEPKDPLILGRPFLATAGAMIDVKKGKIDLNLGKDFR 746
Query: 934 IGFELENVM 942
+ F++++ M
Sbjct: 747 MTFDVKDAM 755
>UniRef100_Q9SKR9 Putative Athila retroelement ORF1 protein [Arabidopsis thaliana]
Length = 929
Score = 255 bits (652), Expect = 4e-66
Identities = 204/695 (29%), Positives = 338/695 (48%), Gaps = 71/695 (10%)
Query: 285 TNIG---WIESWSKRKGLGADLVRKYEDSETKIRLPKVRSL-ARPFSGSPTEDPNLHISS 340
TNIG + + ++R G+ ++R +++ KI+ + + F G P EDP H+
Sbjct: 44 TNIGAGDFPHNHNQRHGI---VLRPVQNNNFKIKSGLIAMVQGNKFHGLPMEDPLDHLDE 100
Query: 341 FLRLSGTIKEN---QEAVRLHLFPFSLRDRASAWFHSLEVGSITSWDDMRRAFLARFSPP 397
RL G K N ++ +L LFPFSL D+A W +L SIT+WDD + AFLA+F
Sbjct: 101 LERLCGLTKINGVSEDGFKLRLFPFSLGDKAHLWEKTLPKNSITTWDDCKMAFLAKFFSN 160
Query: 398 SKTAKLRDQITRFNQKDGESLYEA*ERFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMS 457
S+TA+LR++I+ F K ES EA ERFK CPHHG + +++T Y G+ +M
Sbjct: 161 SRTARLRNEISGFTLKQNESFCEAWERFKGYQTKCPHHGFSQASLLNTLYRGVLPKIRML 220
Query: 458 VDAAAGGALMNKNYTEAYALIEDMAQNHYQWTNERAVTTPTPSKKEAGIYEVSEYNHLAA 517
+D A+ G + K+ E + L+E++AQ+ + + + T S + + E L
Sbjct: 221 LDTASNGNFLKKDIEEGWELVENLAQSGGNYNEDYGRSIYTSSNTDDKHHR--EMKALND 278
Query: 518 KVEALTQKIEKLNVNAAQPSP------ASPTCEVCGITGHTGVDCQLGSAANIEQLNYAQ 571
K++ + Q +K ++ P + C + G +AA +
Sbjct: 279 KLDKIIQMQQKHVHFISEDEPFQVQEGENDQCAEIRYVHNQGPSLPSAAAAAEPTKPFVP 338
Query: 572 YNQ--GMRPNQNFYKNPQGSY-GQTAPPGYTNNQRVA---QKSSLEILLENCMMNQNKQL 625
YNQ G P Q F QG Y Q PPG+T +Q+ A Q S + +L+ + Q
Sbjct: 339 YNQSLGFVPKQQF----QGGYQQQQPPPGFTPHQQQAHAPQNSDIMTVLQQLIQGQATGA 394
Query: 626 QELKNQTGSLNDSLSK----LNTKVDSIATHTKMLETQISQVAQQVAISSQTPGVFPGQT 681
E+ + +N+ + + LN+K +S++T + +E ++ + + P PG+
Sbjct: 395 MEIAKKLSKVNNKVDRQGVELNSKFESMSTRMRYVEGILASPS-----VNNNPCQLPGKA 449
Query: 682 ETNPKAH--VNAISLGGNKLEETITKAKSVKGEIVKLLGE---------KNAIKTPLDKN 730
NPK + +AI++ ++ T S + V L GE N I+ P+ +
Sbjct: 450 IQNPKQYSTAHAITITHDRELPTRYVPTSNTEDSVILEGEDFYQDDVLADNPIEEPILTS 509
Query: 731 KTLNPLRLTKLNLEAQFAKFLNILKRICIKIPFAEALSRMPLYAKFLREIFSKKKAI--D 788
+ P + FAK K I +P +F + + +K +A+
Sbjct: 510 QPTRP----QAPPATPFAKKTEAAKTKDIDFVPPPYKPLLPFPGRFKKVLVAKYRALLEK 565
Query: 789 HKETIALTRESSAIVKKQPQKLRDPGSFAIPCVIGKETVDKALCHLGASVSLLPLSLFKR 848
H + I L + A++ + ++L T + LC LGASVSL+PLS+ K+
Sbjct: 566 HIKDIPLV-DCLALIHDEHKQL---------------TFNNCLCDLGASVSLMPLSVVKK 609
Query: 849 MGIGELKPTEMILKLADRSTIPLAGYIEDIPVKIEGIYIPTDFVVVDIEEDHDVPIILGR 908
+G KP ++ L LADR++ G +ED+PV I G+ +PTDFVV++++ + P+ILGR
Sbjct: 610 LGFVHYKPCDLTLILADRTSKTPFGLLEDVPVMINGVEVPTDFVVLEMDGESKDPLILGR 669
Query: 909 PFLATAGAIIDVQSGRIVFQ-ASDAMIGFELENVM 942
PFLA+ GA+IDV+ G+I D + F++ + M
Sbjct: 670 PFLASVGAVIDVKQGKINLNLGEDVKMKFDIRDAM 704
>UniRef100_Q7Y1N4 Putative retrotransposon gag protein [Oryza sativa]
Length = 697
Score = 255 bits (651), Expect = 5e-66
Identities = 198/642 (30%), Positives = 297/642 (45%), Gaps = 115/642 (17%)
Query: 323 ARPFSGSPTEDPNLHISSFLRLSG--TIKE-NQEAVRLHLFPFSLRDRASAWFHSLEVGS 379
A PF G P ED N H+ FL + TIK + VRL LFPFSL R WF++ +
Sbjct: 42 ASPFCGKPNEDANAHLQQFLEICSMYTIKGVSPNTVRLRLFPFSLLRRVKQWFYA-NCAA 100
Query: 380 ITSWDDMRRAFLARFSPPSKTAKLRDQITRFNQKDGESLYEA*ERFKEMLRLCPHHGLEK 439
+WD AFL++F P KT LR +I+ F Q ES+ EA ER +E + CPHHG++
Sbjct: 101 ANTWDKCSMAFLSKFFPMGKTNALRGRISSFQQTMDESIPEAWERLQEYVAACPHHGMDD 160
Query: 440 WLIVHTFYNGLSYTTKMSVDAAAGGALMNKNYTEAYALIEDMAQNHYQWTNERAVTTPTP 499
WLI+ FYNGL+ ++ +DAAAGGA +K A IE M N W+ ER T
Sbjct: 161 WLILQNFYNGLTPMSRDHLDAAAGGAFFSKTVQGAVEFIEKMVSN-MGWSEERLQT---- 215
Query: 500 SKKEAGIYEVSEYNHLAAKVEALTQKI---EKLNVNAAQPSPASPTCEVCGITGHTGVDC 556
++ G++ V E L AK++ L +++ EK+ + + TCEVCG TGH G +C
Sbjct: 216 --RQRGMHTVKETELLPAKLDLLMKRLDDHEKMPQGTVKALDSHVTCEVCGDTGHPGNNC 273
Query: 557 QLGSAANIEQLNYAQYNQGMRPNQNFYKNPQGSYGQTAPPGYTNNQRVAQKSSLEILLEN 616
E+ Y N G R QG +
Sbjct: 274 ----PETREEAMYMGNNNGYR--------RQG---------------------------D 294
Query: 617 CMMNQNKQLQELKNQTGSLNDSLSKLNTKVDSIAT-------HTKMLETQISQVAQQVAI 669
+ Q K L + + + + N K+D A+ KM+ETQ++Q+A +
Sbjct: 295 LVFAQAKTTDSLSKKLAANDKIVENRNVKLDGFASAFQNQLCFNKMMETQLAQLAS--LV 352
Query: 670 SSQTPGVFPGQTETNPKAHVNAISLGGNKLEE-----TITKAKSVKGEI-------VKLL 717
+ G PGQ +++ + +V AI+ G K T+ + E ++
Sbjct: 353 PANETGRIPGQPDSSIE-NVKAITTRGGKSTRDPPYPNPTRTNGLSKEAPSNGLADKEVQ 411
Query: 718 GEKNAIKTPLDKNKTLNPLRLTKLNLEAQFAKFLNILKRICIKIPFAEALSRMPLYAKFL 777
EK + D P R+ K +++ QFA+F+ ++++I I +P +A+ ++P YA+ L
Sbjct: 412 PEKTMPQEYCDIRLLPFPQRMRKPSVDEQFARFVEVIQKIHINMPLLDAM-QVPTYARCL 470
Query: 778 REIFSKKKAIDHKETIALTRESSAIVKKQPQKLRDPGSFAIPCVIGKETVDKALCHLGAS 837
++I + K+ + E + LT +QP + PG
Sbjct: 471 KDILNNKRPLPTTEVVKLTG-----TMQQPDTPQAPGE---------------------- 503
Query: 838 VSLLPLSLFKRMGIGELKPTEMILKLADRSTIPLAGYIEDIPVKIEGIYIPTDFVVVDIE 897
+R G PT M L+L D S A +D+PVKI +IP DFVV+D++
Sbjct: 504 --------EERSG----APTPMRLQLVDSSIRYTARIAKDVPVKIRYFFIPVDFVVLDMD 551
Query: 898 EDHDVPIILGRPFLATAGAIIDVQSGRIVFQASDAMIGFELE 939
+ P ILGRPFL+TAGA IDV +G I F + FE +
Sbjct: 552 TGKETPFILGRPFLSTAGANIDVGTGSIYFHINGKEEKFEFQ 593
>UniRef100_Q6F2G0 Hypothetical protein OSJNBa0083K01.26 [Oryza sativa]
Length = 585
Score = 238 bits (606), Expect = 8e-61
Identities = 176/547 (32%), Positives = 263/547 (47%), Gaps = 69/547 (12%)
Query: 399 KTAKLRDQITRFNQKDGESLYEA*ERFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSV 458
KT L +I+ F Q ES+ EA E +E + CPHHG++ WLI+ FYN L+ ++ +
Sbjct: 3 KTDALCGRISSFQQTRDESIPEAWELLQEYVAACPHHGMDDWLILQNFYNRLTPMSRDHL 62
Query: 459 DAAAGGALMNKNYTEAYALIEDMAQNHYQWTNERAVTTPTPSKKEAGIYEVSEYNHLAAK 518
DAAAGGA +K LIE M N W+ ER T ++ G++ V E LAAK
Sbjct: 63 DAAAGGAFFSKTVQGTVELIEKMVSN-MGWSKERLQT------RQRGMHTVKETELLAAK 115
Query: 519 VEALTQKI---EKLNVNAAQPSPASPTCEVCGITGHTGVDCQLGSAANIEQLNYAQYNQG 575
++ L +++ EK + + TCEVCG T H+G DC E A Y
Sbjct: 116 LDLLMKRLDNHEKRPQGTVKALDSHVTCEVCGGTDHSGNDCP-------ETREEAMY--- 165
Query: 576 MRPNQNFYKNPQGSYGQTAPPGYTN--NQRVAQKSSLEILLENCMMNQNKQLQELKNQTG 633
M N N Y+ PQG G P Y N Q + L L+ N+TG
Sbjct: 166 MGNNNNGYR-PQGVQGWNQPRSYYQGGNNNETQLAQLAYLVP-------------ANETG 211
Query: 634 SLNDSLSKLNTKVDSIATHTKMLETQISQVAQQVAISSQTPGVFPGQTETNPKAHVNAIS 693
+ V +I T G T + N
Sbjct: 212 RIPGQPYSSIENVKAITTR--------------------------GGKSTRDPPYPNPAG 245
Query: 694 LGGNKLEETITKAKSVKGEIVKLLGEKNAIKTPLDKNKTLNPLRLTKLNLEAQFAKFLNI 753
G +ET + + K ++ EK + D P K +++ FA+F+ +
Sbjct: 246 TNGMS-KETPSNDSADK----EIQPEKTVPQEYCDTRLLPFPQWSRKTSVDELFARFVEV 300
Query: 754 LKRICIKIPFAEALSRMPLYAKFLREIFSKKKAIDHKETIALTRE-SSAIVKKQPQKLRD 812
++RI I + +A+ ++P+YA++L++I + K+ + E + LT S+ I+ K P+K +
Sbjct: 301 IQRIHINVLLLDAM-QVPIYARYLKDILNNKRPLPTTEVVKLTEHCSNVILHKLPEKKKY 359
Query: 813 PGSFAIPCVIGKETVDKALCHLGASVSLLPLSLFKRMGIGELKPTEMILKLADRSTIPLA 872
I C IG + D+ALC LGASVS++P +F R+ L PT M L++AD S A
Sbjct: 360 SECPTITCSIGAQQFDQALCDLGASVSVMPKDVFDRLNFTVLAPTPMRLQMADSSVRYQA 419
Query: 873 GYIEDIPVKIEGIYIPTDFVVVDIEEDHDVPIILGRPFLATAGAIIDVQSGRIVFQASDA 932
G ED+P+KI +IP DFVV+D++ + P+ILGRPFL+TAGA IDV++ I F ++
Sbjct: 420 GIAEDVPIKIWDFFIPVDFVVLDMDTRKETPLILGRPFLSTAGANIDVETSSICFHINEK 479
Query: 933 MIGFELE 939
FE +
Sbjct: 480 EEKFEFQ 486
>UniRef100_Q7XLB2 OSJNBa0011K22.17 protein [Oryza sativa]
Length = 551
Score = 237 bits (605), Expect = 1e-60
Identities = 173/540 (32%), Positives = 260/540 (48%), Gaps = 84/540 (15%)
Query: 399 KTAKLRDQITRFNQKDGESLYEA*ERFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSV 458
KT LR +I+ F Q ES+ EA ER +E + CPHHG++ WLI+ FYNGL+ + +
Sbjct: 3 KTNALRGRISSFQQTRDESIPEAWERLQEYMAACPHHGMDDWLILQNFYNGLTPMSHDHL 62
Query: 459 DAAAGGALMNKNYTEAYALIEDMAQNHYQWTNERAVTTPTPSKKEAGIYEVSEYNHLAAK 518
DAAA GA +K A LIE M + ++ KK G +V + +H+
Sbjct: 63 DAAARGAFFSKTVQGAVDLIEKMLDLLMKRLDDH-------DKKPQGTVKVLD-SHV--- 111
Query: 519 VEALTQKIEKLNVNAAQPSPASPTCEVCGITGHTGVDCQLGSAANIEQLNYAQYNQGMRP 578
TCEVCG TGH+G DC + + Y N G P
Sbjct: 112 -----------------------TCEVCGNTGHSGNDC-----LDTREAMYMGNNNGYCP 143
Query: 579 NQNFYKNPQGSYGQTAPPGYTNNQRVAQKSSLEIL---LENCMMNQNKQLQELKNQTGSL 635
Q G P Y + + A + L + L++ + Q K + L + +
Sbjct: 144 --------QRGQGWNQPCPYVDRKHEAWEICLTPVQPSLKDLVFAQAKTIDALSKKLATN 195
Query: 636 NDSLSKLNTKVDSIAT-------HTKMLETQISQVAQQVAISSQTPGVFPGQTETNPKAH 688
+ L +N K+D A+ KMLETQ++Q A V
Sbjct: 196 DKILENINVKLDGFASAFQNQLRFNKMLETQLAQFASLVP-------------------- 235
Query: 689 VNAISLGGNKLEETITKAKSVKGEIVKLLGEKNAIKTPLDKNKTLNPLRLTKLNLEAQFA 748
A G N + + + + S E+ EK + D R K +++ QFA
Sbjct: 236 --ANETGTNGIAKEVPSSDSADKEVQL---EKIVPQEYCDTWLLSFHQRSRKPSVDEQFA 290
Query: 749 KFLNILKRICIKIPFAEALSRMPLYAKFLREIFSKKKAIDHKETIALTRE-SSAIVKKQP 807
+F ++++I I +P +A+ ++P YA++L+++ + K+ + E + LT + SS I+ K P
Sbjct: 291 RFAEVIQKIHINVPLLDAM-QVPTYARYLKDMLNNKRLLPTMEVVKLTEQCSSVILHKLP 349
Query: 808 QKLRDPGSFAIPCVIGKETVDKALCHLGASVSLLPLSLFKRMGIGELKPTEMILKLADRS 867
+K +DPG I C IG + D+A C LGAS+S++P + ++ L PT M L+LAD S
Sbjct: 350 EKKKDPGCPTITCSIGAQQFDQAFCDLGASISVMPKDVPDKLNFTVLTPTLMCLQLADLS 409
Query: 868 TIPLAGYIEDIPVKIEGIYIPTDFVVVDIEEDHDVPIILGRPFLATAGAIIDVQSGRIVF 927
G ED+PVKI +IP DFVV+D++ + P+ILGRPFL+TAGA IDV G I F
Sbjct: 410 VHYPTGIAEDVPVKIRDFFIPVDFVVLDMDMGKETPLILGRPFLSTAGANIDVGMGSIRF 469
>UniRef100_Q9MA70 F2J6.11 protein [Arabidopsis thaliana]
Length = 823
Score = 222 bits (565), Expect = 5e-56
Identities = 187/639 (29%), Positives = 288/639 (44%), Gaps = 124/639 (19%)
Query: 326 FSGSPTEDPNLHISSFLRLSGTIKEN---QEAVRLHLFPFSLRDRASAWFHSLEVGSITS 382
F G P EDP H+ +F RL K N +++ +L LFPFSL D+A W +L V S+ +
Sbjct: 90 FHGLPMEDPLDHLDNFDRLCSLTKINGVSEDSFKLRLFPFSLGDKAHLWEKTLPVDSVDT 149
Query: 383 WDDMRRAFLARFSPPSKTAKLRDQITRFNQKDGESLYEA*ERFKEMLRLCPHHGLEKWLI 442
DD ++AFLA+F S+TA+LR++I+ FNQK+ ES EA ERFK CPHHG +K +
Sbjct: 150 LDDCKKAFLAKFFSNSRTARLRNEISGFNQKNSESFAEAWERFKGYSTQCPHHGFKKASL 209
Query: 443 VHTFYNGLSYTTKMSVDAAAGGALMNKNYTEAYALIEDMAQN------HYQWTNERAVTT 496
+ T Y G +M +D + G +NK+ E + L+E++AQ+ +Y TN + +
Sbjct: 210 LSTLYRGALPKIRMLLDTTSNGNFLNKDVAEGWELVENLAQSDGNYNENYDRTNRGSSDS 269
Query: 497 PTPSKKEAGIYEVSEYNHLAAKVEALTQKIEKLNVNAAQ---------PSPASPTCEVCG 547
K+ + + N E T +K N+ + + A+P +V
Sbjct: 270 EDRHKRRTKLLMIRLTNWCLLNREMSTTLQKKSLHNSKKGRILLLRRSNNVANPQDQVYP 329
Query: 548 ITGHTGVDCQLGSAANIEQLNYAQYNQGMRPNQNFYKNPQGSYGQTAPPGYTN--NQRVA 605
+ + YNQG QNF PPG+T Q A
Sbjct: 330 PQNQPP-----------QAKPFIPYNQGYNQKQNF-----------GPPGFTQQPQQTSA 367
Query: 606 QKSSLEILLE-------NCMMNQNKQLQELKNQTGSLNDSLSKLNTKVDSIATHTKMLET 658
Q S ++ LL +C M +K+L EL T ++ S + LN K+D++ T K +E
Sbjct: 368 QDSEMKTLLHQLVQGQASCSMTMDKKLAEL---TTRIDCSYNDLNIKIDALNTRVKTMEG 424
Query: 659 QISQVAQQVAISSQTPGVFPGQTETNPKAHVNAISLGGNKLEETITKAKSVKGEIVKLLG 718
I+ + + + PG P + NPK + +AI+ T A + G
Sbjct: 425 HIASTS-----APKHPGQLPRNSVQNPKEYAHAIT-------TVSTSATADSG------- 465
Query: 719 EKNAIKTPLDKNKTLNPLRLTKLNLEAQFAKFLNILKRICIKIPFAEALSRMPLY----A 774
+ + + L P ++ L+ FA + I A P + A
Sbjct: 466 --------IQEGEVLRPRSRQEIELDF-FAWLVERAYDPSNPIRIPPAYEPKPYFPERIA 516
Query: 775 KFLREIFSKKKAI------DHKETIALTRESSAIVKKQPQKLRDPGSFAIPC--VIGKET 826
+ IF K K + + +E I L ++ ++PQ+ + + C +I ++
Sbjct: 517 QVNARIFQKHKMMFIKCIKELEEKIPLVDTPKEVIMERPQEAQQIVELSFECSAIIQRKV 576
Query: 827 VDKALCHLGASVSLLPLSLFKRMGIGELKPTEMILKLADRSTIPLAGYIEDIPVKIEGIY 886
+ K KLADRS G +ED+PVKI I
Sbjct: 577 IPK--------------------------------KLADRSVRVPHGMLEDLPVKIGSIE 604
Query: 887 IPTDFVVVDIEEDHDVPIILGRPFLATAGAIIDVQSGRI 925
IPTDFVV++++E+ P+ILGRPFLATAGA+IDVQ G+I
Sbjct: 605 IPTDFVVLEMDEEPKDPLILGRPFLATAGALIDVQMGKI 643
>UniRef100_Q9MA68 F2J6.13 protein [Arabidopsis thaliana]
Length = 780
Score = 220 bits (561), Expect = 1e-55
Identities = 193/661 (29%), Positives = 296/661 (44%), Gaps = 141/661 (21%)
Query: 326 FSGSPTEDPNLHISSFLRLSGTIKEN---QEAVRLHLFPFSLRDRASAWFHSLEVGSITS 382
F G P EDP H+ +F R K N +++ +L LFPFSL D A W +L V S+ +
Sbjct: 54 FHGLPMEDPLDHLDNFDRFCSLTKINGVSEDSFKLRLFPFSLGDEAHLWEKTLLVDSVDT 113
Query: 383 WDDMRRAFLARFSPPSKTAKLRDQITRFNQKDGESLYEA*ERFKEMLRLCPHHGLEKWLI 442
WDD ++AFLA+F S+TA+LR++I+ FNQK+ ES EA ERFK CPHHG +K +
Sbjct: 114 WDDCKKAFLAKFFSNSRTARLRNEISGFNQKNSESFAEAWERFKRYSTQCPHHGFKKASL 173
Query: 443 VHTFYNGLSYTTKMSVDAAAGGALMNKNYTEAYALIEDMAQNHYQWTNERAVTTPTPSKK 502
+ T Y G +M +D + G +NK+ E + L+E++AQ+ + + T S
Sbjct: 174 LSTLYRGALPKIRMLLDTTSNGNFLNKDVAEGWELVENLAQSDGNYNEDYDRTNRGSSDS 233
Query: 503 EAGIYEVSEYNHLAAKVEALTQKIEKLNVNAAQPSPASPTCEVCGITGHTGVDCQLGSAA 562
E + +++AL KI+KL V A Q + V IT Q G
Sbjct: 234 E---------DRHKKEIKALNDKIDKL-VLAQQRN-------VHYITEEELTQLQEGENL 276
Query: 563 NIEQLNYAQ----YNQGM------RPNQNFYKN-----------PQGSYGQT-------- 593
+E+++Y Q YN+G PN ++ N PQ Q
Sbjct: 277 TVEEVSYLQNQGGYNKGFNNYKPPHPNLSYRSNNVANPQDQVYPPQNQPPQAKPFIPYNQ 336
Query: 594 --------APPGYTN--NQRVAQKSSLEILLE-------NCMMNQNKQLQELKNQTGSLN 636
PP +T Q AQ S ++ LL +C + +K+L EL T ++
Sbjct: 337 GYNQKQNFRPPSFTQQPQQTSAQDSEMKTLLHQLVQGQASCSITMDKKLAEL---TTRID 393
Query: 637 DSLSKLNTKVDSIATHTKMLETQISQVAQQVAISSQTPGVFPGQTETNPKAHVNAISLGG 696
S + LN K+D++ T K +E I+ + + + PG ++ NPK + +AI+
Sbjct: 394 CSYNDLNIKIDALNTRVKTMEGHIASTS-----APKHPGQLSRKSVQNPKEYAHAIT--- 445
Query: 697 NKLEETITKAKSVKGEIVKLLGEKNAIKTPLDKNKTLNPLRLTKLNLEAQFAKFLNILKR 756
T A + G + + + L P ++ L+ FA +
Sbjct: 446 ----TVSTSATADSG---------------IQEGEVLRPRSRQEIELDF-FALLVERAYD 485
Query: 757 ICIKIPFAEALSRMPLY----AKFLREIFSKKKAI------DHKETIALTRESSAIVKKQ 806
I A P + A+ IF K K + + +E I L ++ ++
Sbjct: 486 PSNPIRIPPAYEPKPYFPERIAQVNARIFQKHKMMFIKCIKELEEKIPLVDTPKEVIMER 545
Query: 807 PQKLRD--PGSFAIPCVIGKETVDKALCHLGASVSLLPLSLFKRMGIGELKPTEMILKLA 864
PQ+ + SF +I ++ + K L A
Sbjct: 546 PQEAQQIIELSFECSAIIQRKVIPKKL--------------------------------A 573
Query: 865 DRSTIPLAGYIEDIPVKIEGIYIPTDFVVVDIEEDHDVPIILGRPFLATAGAIIDVQSGR 924
DRS G +ED+PVKI I IPTDF+V++++E+ P+ILGRPFLATAGA+IDVQ G+
Sbjct: 574 DRSVRVPHGMLEDLPVKIGSIEIPTDFMVLEMDEEPKDPLILGRPFLATAGALIDVQMGK 633
Query: 925 I 925
I
Sbjct: 634 I 634
>UniRef100_Q7F9Z3 OSJNBa0079C19.24 protein [Oryza sativa]
Length = 461
Score = 205 bits (522), Expect = 5e-51
Identities = 156/490 (31%), Positives = 240/490 (48%), Gaps = 43/490 (8%)
Query: 389 AFLARFSPPSKTAKLRDQITRFNQKDGESLYEA*ERFKEMLRLCPHHGLEKWLIVHTFYN 448
AFL++F P KT LR +I+ F Q ES+ EA ER PHHG++ LI+ FYN
Sbjct: 2 AFLSKFFPMGKTNALRGRISNFQQTRDESIPEAWER--------PHHGMDNSLILQNFYN 53
Query: 449 GLSYTTKMSVDAAAGGALMNKNYTEAYALIEDMAQNHYQWTNERAVTTPTPSKKEAGIYE 508
GL+ + +DAAAGGA +K A LIE M N W+ ER T ++ GI+
Sbjct: 54 GLTTMSHDHLDAAAGGAFFSKTVRGAIDLIEKMVSN-MGWSKERLQT------RQQGIHT 106
Query: 509 VSEYNHLAAKVEALTQKIEKLNVNAAQPSPASP---TCEVCGITGHTGVDCQLGSAANIE 565
V E LAAK++ L ++++ + A TCEVCG TGH+G DC E
Sbjct: 107 VKETELLAAKLDLLMKRLDDHDNRPQDTVKALDLHVTCEVCGGTGHSGNDCP-------E 159
Query: 566 QLNYAQYNQGMRPNQNFYKNPQGSYGQTAPPGYTNNQRVAQKSSLEILLENCMMNQNKQL 625
A Y M N N Y+ PQG G P Y + S + L++ + Q K
Sbjct: 160 THEEAMY---MGKNNNEYR-PQGGQGWNQPRPYYQGENNNSNFSNQPSLKDLIFVQAKTT 215
Query: 626 QELKNQTGSLNDSLSKLNTKVDSIATH-------TKMLETQISQVAQQVAISSQTPGVFP 678
L + + + L +N ++D + KM+ETQ++Q+A V +
Sbjct: 216 DALSKKLAANDKILENINVQLDGFTSSFQNQLIFNKMIETQLAQLASLVPANEIGRIPRR 275
Query: 679 GQTETNPKAHVNAISLGGNKLEETITKAKSVKGEIVKLLGEKNAIKTPLDKNKTLNPLRL 738
G T + + N G N + + S E+ EK + D P R
Sbjct: 276 GGKSTRDRPYPNPA--GTNGVAREAPSSDSANKEVQP---EKAVPQEYYDTRLLSFPQRS 330
Query: 739 TKLNLEAQFAKFLNILKRICIKIPFAEALSRMPLYAKFLREIFSKKKAIDHKETIALTRE 798
K +++ QFA+F+ ++++I I + +A+ ++P YA++L++I + K+ + E I LT +
Sbjct: 331 RKPSVDEQFARFVEVIQKIHINVSLFDAM-QVPTYARYLKDILNNKRPLPTTEVIELTEQ 389
Query: 799 -SSAIVKKQPQKLRDPGSFAIPCVIGKETVDKALCHLGASVSLLPLSLFKRMGIGELKPT 857
S+ I+ K +K + PG I C IG + ++AL LG SVS++P+ +F ++ L PT
Sbjct: 390 CSNVILHKLSEKRKHPGCPTITCSIGAQQFNQALRDLGVSVSVIPMDVFDKLNFTVLAPT 449
Query: 858 EMILKLADRS 867
M L+LAD S
Sbjct: 450 PMRLQLADSS 459
>UniRef100_Q7XLB4 OSJNBa0011K22.15 protein [Oryza sativa]
Length = 406
Score = 181 bits (460), Expect = 7e-44
Identities = 126/359 (35%), Positives = 177/359 (49%), Gaps = 32/359 (8%)
Query: 322 LARPFSGSPTEDPNLHISSFLRLSGTIK---ENQEAVRLHLFPFSLRDRASAWFHSLEVG 378
+A PF G P ED N H+ FL + T + +AVRL LF FSL RA WF++
Sbjct: 1 MASPFCGKPNEDANAHLQQFLEICSTYTIKGVSPDAVRLRLFLFSLLGRAKQWFYANRA- 59
Query: 379 SITSWDDMRRAFLARFSPPSKTAKLRDQITRFNQKDGESLYEA*ERFKEMLRLCPHHGLE 438
+I +WD AFL++F P KT LR +I+ F Q ES+ EA ER +E + CP HG++
Sbjct: 60 AINTWDKCSTAFLSKFFPMGKTNALRGRISSFQQTRDESIPEAWERLQEYVAACPLHGMD 119
Query: 439 KWLIVHTFYNGLSYTTKMSVDAAAGGALMNKNYTEAYALIEDMAQNHYQWTNERAVTTPT 498
WLI+ FYNGL+ ++ +D AAGGA +K A LIE M N W+ ER T
Sbjct: 120 DWLILLNFYNGLTLMSRDHLDTAAGGAFFSKTVQGAVDLIEKMVSN-MGWSEERLQT--- 175
Query: 499 PSKKEAGIYEVSEYNHLAAKVEALTQKI---EKLNVNAAQPSPASPTCEVCGITGHTGVD 555
+ G++ V E LAAK++ L +++ EK + TCEVC T H+G D
Sbjct: 176 ---HQQGMHIVKETELLAAKLDLLIKRLDDHEKQPQGTVKALDYHITCEVCSNTSHSGND 232
Query: 556 CQLGSAANIEQLNYAQYNQGMRPNQNFYKNPQGSYGQTAPPGYTNNQRVAQKSSLEILLE 615
C E + N G RP N Y Q G NN + +SSL+ L
Sbjct: 233 C---PETREEAMYMGNNNNGYRPQGGQRWNQPRPYYQ----GGNNNGNFSNQSSLKDL-- 283
Query: 616 NCMMNQNKQLQELKNQTGSLNDSLSKLNTKVDSIA-------THTKMLETQISQVAQQV 667
+ Q K L + + ++ L +N K+D A + KM+ET ++Q+A V
Sbjct: 284 --VFAQAKTTDALSKKLAANDNILENINVKLDGFASTFQNQLSFNKMIETHLAQLASLV 340
>UniRef100_Q6UUB7 Hypothetical protein [Oryza sativa]
Length = 450
Score = 173 bits (438), Expect = 2e-41
Identities = 140/429 (32%), Positives = 219/429 (50%), Gaps = 62/429 (14%)
Query: 446 FYNGLSYTTKMSVDAAAGGALMNKNYTEAYALIEDMAQNHYQWTNERAVTTPTPSKKEAG 505
FYNGL+ ++ +DAAAGGA +K A L+E M N W+ E+ T + G
Sbjct: 74 FYNGLTPMSRDHLDAAAGGAFFSKMVQGAVDLVEKMVSN-MGWSEEQLQTC------QRG 126
Query: 506 IYEVSEYNHLAAKVEALTQKI---EKLNVNAAQPSPASPTCEVCGITGHTGVDCQLGSAA 562
++ V E LAAK++ L +++ EK + + TCEVC TGH+G DC
Sbjct: 127 MHTVKETELLAAKLDLLMKRLDNHEKRPQGTVKALDSHVTCEVCSSTGHSGNDCP----- 181
Query: 563 NIEQLNYAQYNQGMRPNQNFYKNPQGSYG--QTAP--PGYTNNQRVAQKSSLEILLENCM 618
E A Y N N+Y+ PQG G Q P G NN + + SL+ L+
Sbjct: 182 --ETREEAMY----MGNNNWYR-PQGGQGWSQLRPYYQGGNNNGNFSNQPSLKDLV---- 230
Query: 619 MNQNKQLQELKNQTGSLNDSLSKLNTKVDSIATHTKMLETQ-ISQVAQQVAISSQTPGVF 677
+Q K D+LSK +AT+ K+LE ++Q+A V ++ G
Sbjct: 231 FSQAKT-----------TDALSK------KLATNDKILENNNLAQLASLVPVNET--GRI 271
Query: 678 PGQTETNPKAHVNAISLGGNKLEETITKAKSVKGEIVKLLGEKNAIKTPLDKNKTLNPLR 737
PGQ +++ + +V AI++ G ++ K EK + D L P R
Sbjct: 272 PGQPDSSIE-NVKAITMRGASSSDSADKEDQP---------EKTVPQEYCDTWLLLFPQR 321
Query: 738 LTKLNLEAQFAKFLNILKRICIKIPFAEALSRMPLYAKFLREIFSKKKAIDHKETIALTR 797
+ K ++ QFA+F+ ++++I I +P +A+ +MP YA++L+ I + K+ + E + LT
Sbjct: 322 MRKPLVDEQFARFVEVIQKIHINVPLLDAM-QMPTYARYLKGILNNKRPLPTTEVVKLTE 380
Query: 798 E-SSAIVKKQPQKLRDPGSFAIPCVIGKETVDKALCHLGASVSLLPLSLFKRMGIGELKP 856
+ S+ I+ K P+K +DPG I C IG + D+ALC LGASVS++P ++F ++ L P
Sbjct: 381 QWSNVILHKLPEKKKDPGCPTITCSIGAQQFDQALCDLGASVSVMPKNVFDKLNFTMLAP 440
Query: 857 TEMILKLAD 865
T M L+LAD
Sbjct: 441 TPMHLQLAD 449
>UniRef100_Q8S618 Hypothetical protein OSJNBb0048O22.17 [Oryza sativa]
Length = 412
Score = 163 bits (412), Expect = 3e-38
Identities = 107/286 (37%), Positives = 150/286 (52%), Gaps = 26/286 (9%)
Query: 309 DSETKIRLPKVRSLARPFSGSPTEDPNLHISSFLRLSGTIK---ENQEAVRLHLFPFSLR 365
D + K L K+ A PF P ED N + FL + T + +A++L LFPFSL
Sbjct: 29 DFDLKSSLIKMAQ-ASPFCDKPNEDANAPLQQFLEICNTYAIKGVSPDAIKLRLFPFSLI 87
Query: 366 DRASAWFHSLEVGSITSWDDMRRAFLARFSPPSKTAKLRDQITRFNQKDGESLYEA*ERF 425
+A WF++ I +WD AF ++F P KT LR +I+ F Q ES+ EA ER
Sbjct: 88 GKAKRWFYANRA-MINTWDKCSTAFPSKFFPMGKTNALRGRISSFQQTRDESIPEAWERL 146
Query: 426 KEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSVDAAAGGALMNKNYTEAYALIEDMAQNH 485
+E + +CPHHG++ WLI+ FYNGL+ ++ +DAAAGGA +K A LIE M N
Sbjct: 147 QEYVAVCPHHGMDDWLILQNFYNGLTLMSRDHLDAAAGGAFFSKTVRGAIDLIEKMVSN- 205
Query: 486 YQWTNERAVTTPTPSKKEAGIYEVSEYNHLAAKVEALTQKI---EKLNVNAAQPSPASPT 542
W+ ER T + G++ V E LAAK++ L +++ +K + + T
Sbjct: 206 MGWSEERLQT------HQQGMHTVKETEMLAAKLDLLMKRLDDHDKRPQGTVKALDSHVT 259
Query: 543 CEVCGITGHTGVDCQLGSAANIEQLNYAQYNQGMRPNQNFYKNPQG 588
CEVCG TGH+G D +E A Y M N N Y+ PQG
Sbjct: 260 CEVCGSTGHSGNDF-------LETREEAMY---MGSNNNGYR-PQG 294
>UniRef100_Q9LH13 Similarity to Athila retroelement ORF1 protein [Arabidopsis
thaliana]
Length = 583
Score = 162 bits (411), Expect = 3e-38
Identities = 127/501 (25%), Positives = 242/501 (47%), Gaps = 37/501 (7%)
Query: 456 MSVDAAAGGALMNKNYTEAYALIEDMAQNHYQWTNERAVTTPTPSKKEAGIYEVSEYNHL 515
M++D A+ G M K TEA LIE++A ++ + V ++ G E ++ L
Sbjct: 1 MALDTASNGEFMTKTETEATKLIENLAGSN----SNHNVDYDRSNRGGGG--ESKQFAEL 54
Query: 516 AAKVEALTQKIEKLNVNAAQPSPASPTCEVCGITGHTGVDCQLG---SAANIEQLNYAQY 572
+AKVE L ++ +K +VN + S + G + ++ + N+E Y
Sbjct: 55 SAKVEQLMRRDQK-SVNFCEDSSKGMVHQEFSGDGSEDLQAEINFVNGSTNVENPQDQVY 113
Query: 573 -----NQGMRPNQNFYKNPQGSYGQTAPPGYTNNQRVAQKSSLEILLENCMMNQNKQLQE 627
+QG + ++N GQ PP T + + ++++++ + +Q K +
Sbjct: 114 PTQAGSQGQKFQPYGFQNKGNYQGQFQPPAGTGHASSFGDNEMKLMMQQVLEDQKKNAAD 173
Query: 628 LKNQTGSLNDSLSKLNTKVDSIATHTKMLETQISQVAQQVAISSQTPGVFPGQTETNPKA 687
+ + S+ + L N K ++++H K LE Q+SQ+ V+ S + G G+ + K
Sbjct: 174 INVKVDSMYNDL---NGKFATLSSHVKTLENQVSQI---VSASMRPDGTHSGKVKPKGKE 227
Query: 688 HVNAISLGGNKLEETITKAKSVKGEIVKLLGEKN--------AIKTPLDKNKTLNPLRLT 739
AI + E + K +V+ L E +++ P K P R
Sbjct: 228 QCYAIMIQEELREIVVAKQVETNVVVVETLVEDKIVEDDEPLSVEPPPYVPKLPFPGRER 287
Query: 740 KLNLEAQFAKFLNILKRICIKIPFAEALSRMPLYAKFLREIFSKKKAIDHK-ETIALTRE 798
++ + ++A+F I+K++ +++PF + + +P Y +L+ I S K++I+ + I+ E
Sbjct: 288 QIQRQKEYARFDEIMKQLYVRLPFLQLVLHVPSYRSYLKYILSNKRSIEEGVKLISKGEE 347
Query: 799 SSAIVKKQPQKLRDPGSFAIPCVIGKETVDKALCHLGASVSLLPLSLFKRMGIGELKPTE 858
+ +V+ Q Q+ + + + GKE V+ ++ P ++ K++GI KP+
Sbjct: 348 HAQLVESQRQQKEAQLTNVVEMLAGKEMVNTCS-------AIPPATIPKKLGITNFKPSR 400
Query: 859 MILKLADRSTIPLAGYIEDIPVKIEGIYIPTDFVVVDIEEDHDVPIILGRPFLATAGAII 918
+ L LADRS G E++ ++ YIPT+FVV++++++ P+ LGRPFL T AII
Sbjct: 401 ISLILADRSVQFPMGLAENVHARVGNFYIPTNFVVLELDKEPHDPLNLGRPFLNTVEAII 460
Query: 919 DVQSGRIVFQASDAMIGFELE 939
DV+ I Q D + F+++
Sbjct: 461 DVRRSTINLQIGDYALEFDIK 481
>UniRef100_Q8S6J2 Hypothetical protein OSJNBa0019N10.34 [Oryza sativa]
Length = 237
Score = 155 bits (392), Expect = 5e-36
Identities = 85/195 (43%), Positives = 126/195 (64%), Gaps = 2/195 (1%)
Query: 735 PLRLTKLNLEAQFAKFLNILKRICIKIPFAEALSRMPLYAKFLREIFSKKKAIDHKETIA 794
P R K +++ QFA F+ ++++I I +P +A+ ++P YA +L++I + K+ + E +
Sbjct: 36 PQRSRKPSVDEQFAHFVEVIQKIHIDVPLLDAM-QVPTYACYLKDILNNKRPLPTTEVVK 94
Query: 795 LTRE-SSAIVKKQPQKLRDPGSFAIPCVIGKETVDKALCHLGASVSLLPLSLFKRMGIGE 853
LT + S+ I+ K P+K +DPG I C IG + D+ALC LGASVS++P +F ++
Sbjct: 95 LTLQCSNVILHKLPEKKKDPGCPTITCSIGAQQFDQALCDLGASVSVMPKYVFDKLNFTV 154
Query: 854 LKPTEMILKLADRSTIPLAGYIEDIPVKIEGIYIPTDFVVVDIEEDHDVPIILGRPFLAT 913
L PT M L+LAD S AG ED+PVKI +I DFVV+D++ ++ IILGRPFL T
Sbjct: 155 LAPTPMRLQLADSSVRYPAGIAEDVPVKIRDFFILVDFVVLDMDTGKEMSIILGRPFLGT 214
Query: 914 AGAIIDVQSGRIVFQ 928
AGA I V +G I FQ
Sbjct: 215 AGANIHVGTGSIRFQ 229
>UniRef100_Q96467 Hypothetical protein [Hordeum vulgare]
Length = 337
Score = 155 bits (391), Expect = 7e-36
Identities = 83/219 (37%), Positives = 131/219 (58%), Gaps = 16/219 (7%)
Query: 326 FSGSPTEDPNLHISSFLRLSGTIKE---NQEAVRLHLFPFSLRDRASAWFHSLEVGSITS 382
FSG P+ED H+++F+ L K+ + + ++L LFPFSLRDRA WF SL SI S
Sbjct: 47 FSGLPSEDVASHLNTFIELCDMQKKKDVDNDVIKLKLFPFSLRDRAKTWFSSLPKSSIDS 106
Query: 383 WDDMRRAFLARFSPPSKTAKLRDQITRFNQKDGESLYEA*ERFKEMLRLCPHHGLEKWLI 442
WD + A+++++ PP+K LR+ I F Q D E + +A ER K M+R CP +GL W+I
Sbjct: 107 WDKCKDAYISKYFPPAKIISLRNDIMNFKQLDHEHVAQAWERMKLMIRNCPANGLSLWMI 166
Query: 443 VHTFYNGLSYTTKMSVDAAAGGALMNKNYTEAYALIEDMAQNHYQWTNERAVTTPTPSKK 502
+ FY GL++ ++ +D+A GG M EA L++++ N+ QW ER+ T SKK
Sbjct: 167 IQIFYAGLNFASRNILDSATGGTFMEITLGEATKLLDNIMTNYSQWHTERSPT----SKK 222
Query: 503 EAGIYEVSEYNHLAAKVEALTQKIEKLNVNAAQPSPASP 541
++ + E N L++K++ E +N+ A++ +P P
Sbjct: 223 ---VHVIEEINSLSSKMD------ELMNLVASRSAPLDP 252
>UniRef100_O22104 Vicia faba mRNA expressed from retrotransposon-like gene [Vicia
faba]
Length = 238
Score = 150 bits (380), Expect = 1e-34
Identities = 80/201 (39%), Positives = 121/201 (59%), Gaps = 3/201 (1%)
Query: 744 EAQFAKFLNILKRICIKIPFAEALSRMPLYAKFLREIFSKKKAIDHKETIALTRESSAIV 803
E F FL + K++ + I EAL +MP YAKF+++I SKK+ ID + I T S+I+
Sbjct: 11 EKNFEIFLEMFKKLELNILVLEALEKMPTYAKFMKDIISKKRTID-TDPIIHTETCSSIL 69
Query: 804 K--KQPQKLRDPGSFAIPCVIGKETVDKALCHLGASVSLLPLSLFKRMGIGELKPTEMIL 861
+ K P K +D G PC IG + K +LGASVSLLP ++K++GI ++ T M L
Sbjct: 70 QGMKIPVKKKDRGFVTTPCTIGDRSFKKYFINLGASVSLLPFYIYKKLGISNVQNTRMTL 129
Query: 862 KLADRSTIPLAGYIEDIPVKIEGIYIPTDFVVVDIEEDHDVPIILGRPFLATAGAIIDVQ 921
+ AD S G +ED+ VKI+ P DFVV+++ ED ++ +ILGRPFL T +ID++
Sbjct: 130 RFADHSVKIPYGIVEDVLVKIDKFVFPVDFVVLEMPEDEEITLILGRPFLETRRCLIDIE 189
Query: 922 SGRIVFQASDAMIGFELENVM 942
G + + D + +++N M
Sbjct: 190 EGTVTLKVYDEELKIDVQNTM 210
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.332 0.142 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,402,393,150
Number of Sequences: 2790947
Number of extensions: 55521488
Number of successful extensions: 204007
Number of sequences better than 10.0: 667
Number of HSP's better than 10.0 without gapping: 321
Number of HSP's successfully gapped in prelim test: 349
Number of HSP's that attempted gapping in prelim test: 202853
Number of HSP's gapped (non-prelim): 1193
length of query: 942
length of database: 848,049,833
effective HSP length: 137
effective length of query: 805
effective length of database: 465,690,094
effective search space: 374880525670
effective search space used: 374880525670
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 80 (35.4 bits)
Medicago: description of AC144538.1


