

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144513.2 - phase: 0
(215 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
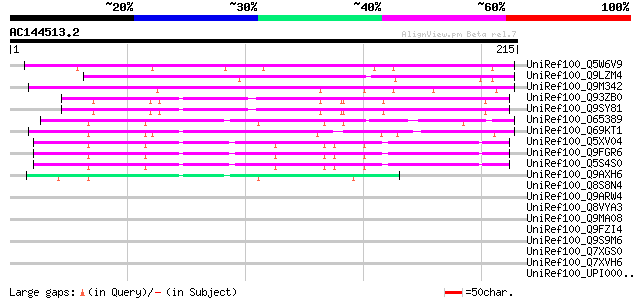
Sequences producing significant alignments: (bits) Value
UniRef100_Q5W6V9 Hypothetical protein OSJNBb0059K16.4 [Oryza sat... 144 2e-33
UniRef100_Q9LZM4 Hypothetical protein T7H20_120 [Arabidopsis tha... 141 1e-32
UniRef100_Q9M342 Protein kinase-like protein [Arabidopsis thaliana] 131 1e-29
UniRef100_Q93ZB0 At1g10380/F14N23_32 [Arabidopsis thaliana] 62 1e-08
UniRef100_Q9SY81 F14N23.27 [Arabidopsis thaliana] 62 1e-08
UniRef100_O65389 F12F1.23 protein [Arabidopsis thaliana] 56 7e-07
UniRef100_Q69KT1 Hypothetical protein OSJNBa0034G14.6 [Oryza sat... 54 2e-06
UniRef100_Q5XV04 Hypothetical protein [Arabidopsis thaliana] 52 1e-05
UniRef100_Q9FGR6 Arabidopsis thaliana genomic DNA, chromosome 5,... 52 1e-05
UniRef100_Q5S4S0 Hypothetical protein [Arabidopsis thaliana] 50 3e-05
UniRef100_Q9AXH6 Wall-associated protein kinase [Oryza sativa] 50 3e-05
UniRef100_Q8S8N4 Putative Ser/Thr protein kinase [Arabidopsis th... 45 0.002
UniRef100_Q9ARW4 Putative wall-associated kinase 4 [Oryza sativa] 44 0.004
UniRef100_Q8VYA3 Wall-associated kinase 2, putative [Arabidopsis... 41 0.019
UniRef100_Q9MA08 F20B17.10 [Arabidopsis thaliana] 41 0.019
UniRef100_Q9FZI4 F1O19.2 protein [Arabidopsis thaliana] 41 0.024
UniRef100_Q9S9M6 T24D18.19 protein [Arabidopsis thaliana] 40 0.042
UniRef100_Q7XGS0 Putative wall-associated protein kinase [Oryza ... 39 0.071
UniRef100_Q7XVH6 OSJNBa0016N04.10 protein [Oryza sativa] 39 0.093
UniRef100_UPI0000498CCD UPI0000498CCD UniRef100 entry 39 0.12
>UniRef100_Q5W6V9 Hypothetical protein OSJNBb0059K16.4 [Oryza sativa]
Length = 640
Score = 144 bits (362), Expect = 2e-33
Identities = 88/226 (38%), Positives = 117/226 (50%), Gaps = 18/226 (7%)
Query: 7 NSLKMVILLLFLLITGLGSVI-----NASPCSNCGDFEVPYPLSTNDDCGDKRYKIYC-- 59
+SL LLLF L + + +A C CG EVPYPLST D CGD YK+ C
Sbjct: 4 SSLSRRALLLFFLEVAVAMALLLPGGHARVCPPCGSTEVPYPLSTADGCGDPEYKVRCAA 63
Query: 60 -----NNDSLEFLSATGTYYKILKIDTSANKLVIKP-PNIFKHTCYSSDLIGG-GLVLDE 112
+L F + GT Y I I ++ +LV+ P P + C S G+ LD
Sbjct: 64 AAAGGTAPTLLFDALNGTSYPITSISPASQRLVVSPAPFVSPGACVSVGAAASRGVQLDP 123
Query: 113 SLPFNISTLNTVMLLNCSDNILQSPLNCSSNSICRQFEEKV-EEGNGCMN-TLCCHYLKD 170
S PFN+S+ NTVMLLNC++ +L+SPLNCSSNS+C + + C LCC ++
Sbjct: 124 SRPFNVSSSNTVMLLNCTELLLRSPLNCSSNSLCHAYAGAAGSTASACAPLPLCCTFVAG 183
Query: 171 SVMNSHKIRLRVGSCTAYTCLVDFKPDEPFKKW--NYGIELQWKPP 214
S++IRL SC+AY V P +P W G+ELQW P
Sbjct: 184 GSSTSYRIRLGPQSCSAYRSFVGLDPSQPPATWGSRLGLELQWATP 229
>UniRef100_Q9LZM4 Hypothetical protein T7H20_120 [Arabidopsis thaliana]
Length = 657
Score = 141 bits (356), Expect = 1e-32
Identities = 74/190 (38%), Positives = 109/190 (56%), Gaps = 9/190 (4%)
Query: 32 CSNCGDFEVPYPLSTNDDCGDKRYKIYCNNDSLEFLSATGTYYKILKIDTSANKLVIKPP 91
C NCG VPYPLST CGD+ Y+I C L F + G+ Y I I++ ++V++PP
Sbjct: 43 CPNCGPMVVPYPLSTGPTCGDQAYRINCVGGKLYFGALHGSSYVITSINSVTQRIVLRPP 102
Query: 92 NIFKH-TCYSSDLIGGGLVLDESLPFNISTLNTVMLLNCSDNILQSPLNCSSNSICRQFE 150
+ +C S+D+ GL LD LPF+I++ NT++LLNCS +LQ+P++CS S+C +
Sbjct: 103 GLASSVSCISADVSKQGLELDPHLPFSITSSNTILLLNCSQAMLQAPIDCSPTSLCYSYI 162
Query: 151 EKVEEGNGCMNT-LCCHYLKDSVMNSHKIRLRVGSCTAYTCLVDFKPDE----PFKKW-N 204
+ + C LCC + D ++ IR+ G C AY V P++ P KKW +
Sbjct: 163 K--NNASPCSKAPLCCTFRTDGSQTAYTIRINGGGCLAYQSFVGLNPNKEVPPPGKKWPD 220
Query: 205 YGIELQWKPP 214
G+ELQW P
Sbjct: 221 TGLELQWALP 230
>UniRef100_Q9M342 Protein kinase-like protein [Arabidopsis thaliana]
Length = 640
Score = 131 bits (330), Expect = 1e-29
Identities = 78/215 (36%), Positives = 111/215 (51%), Gaps = 9/215 (4%)
Query: 9 LKMVILLLFLLITGLGSVINASPCSNCGDFEVPYPLSTNDDCGDKRYKIYCNN-DSLEFL 67
L + L LLI + C NCG VPYPLST DCGD Y+I C+N SL F
Sbjct: 6 LSLTTFTLSLLIYFSSTTQAFKRCPNCGSTRVPYPLSTGLDCGDPGYRIRCDNYGSLWFD 65
Query: 68 SATGTYYKILKIDTSANKLVIKPPNIFKHTCYSSDLIGGGLVLDESLPFNISTLNTVMLL 127
+ G+ I ID S + V++PP ++ C S D+ G+ LD +LPFN+S NTV+++
Sbjct: 66 TLNGSTNPIKTIDPSGQRFVLRPPGFEQNKCVSVDIKYHGIQLDLNLPFNVSCSNTVIIM 125
Query: 128 NCS----DNILQSPLNCSSNSICRQF-EEKVEEGNGCMN-TLCCHYLKDSVMNSHKI-RL 180
NC+ D NCS NS+C +F +E C T CC Y + +N++K+ R
Sbjct: 126 NCTKDGLDAYSSQGFNCSDNSLCHKFLNANLEARGNCRGVTSCCWYKTGASVNTYKVYRA 185
Query: 181 RVGSCTAYTCLVDFKPDEPFKKWNY-GIELQWKPP 214
R C+AY ++ P KW +E+ W+ P
Sbjct: 186 RPDMCSAYQSFMNLDLTIPVSKWGEPAVEILWEAP 220
>UniRef100_Q93ZB0 At1g10380/F14N23_32 [Arabidopsis thaliana]
Length = 305
Score = 62.0 bits (149), Expect = 1e-08
Identities = 55/211 (26%), Positives = 93/211 (44%), Gaps = 25/211 (11%)
Query: 23 LGSVINASPCSN-CGDFEVPYPLSTNDDCGDKRYKIY--CNND--SLEFLSATGTYYKIL 77
L S +++ C CG + YPL T CGD R+ Y C+ D +L + TG+ Y I
Sbjct: 20 LSSHVSSQACQKTCGQIPIKYPLGTGSGCGDPRFTRYITCDPDQQTLTLTTHTGS-YPIT 78
Query: 78 KIDTSANKLVIKPPNIFKHTCYSSDLIGGGLVLDESLPFNISTLNTVMLLNCS--DNILQ 135
+D + ++ + P++ C G LD PF+ LL+CS ++ +
Sbjct: 79 SVDYAKQEIYVTDPSMSTCACTRP---SHGFGLDWDAPFSFHDDTVFTLLDCSVDESPVF 135
Query: 136 SPLN--------C--SSNSICRQFEEKVEEGN--GCMNTLCCHYLKDSVMNSHKIRLRVG 183
+PL+ C S+SIC + + CC Y+ + S ++ L
Sbjct: 136 TPLSNGSGRVSLCDRQSSSICTFLYSNCRAISLINLQVSTCCVYVPLDLGPSFEMDLNKL 195
Query: 184 SCTAYTCLVDFKPDEPF--KKWNYGIELQWK 212
C++Y+ + P + + WNYGI L++K
Sbjct: 196 KCSSYSGFYNLGPGQESHPENWNYGIALKYK 226
>UniRef100_Q9SY81 F14N23.27 [Arabidopsis thaliana]
Length = 333
Score = 62.0 bits (149), Expect = 1e-08
Identities = 55/211 (26%), Positives = 93/211 (44%), Gaps = 25/211 (11%)
Query: 23 LGSVINASPCSN-CGDFEVPYPLSTNDDCGDKRYKIY--CNND--SLEFLSATGTYYKIL 77
L S +++ C CG + YPL T CGD R+ Y C+ D +L + TG+ Y I
Sbjct: 20 LSSHVSSQACQKTCGQIPIKYPLGTGSGCGDPRFTRYITCDPDQQTLTLTTHTGS-YPIT 78
Query: 78 KIDTSANKLVIKPPNIFKHTCYSSDLIGGGLVLDESLPFNISTLNTVMLLNCS--DNILQ 135
+D + ++ + P++ C G LD PF+ LL+CS ++ +
Sbjct: 79 SVDYAKQEIYVTDPSMSTCACTRP---SHGFGLDWDAPFSFHDDTVFTLLDCSVDESPVF 135
Query: 136 SPLN--------C--SSNSICRQFEEKVEEGN--GCMNTLCCHYLKDSVMNSHKIRLRVG 183
+PL+ C S+SIC + + CC Y+ + S ++ L
Sbjct: 136 TPLSNGSGRVSLCDRQSSSICTFLYSNCRAISLINLQVSTCCVYVPLDLGPSFEMDLNKL 195
Query: 184 SCTAYTCLVDFKPDEPF--KKWNYGIELQWK 212
C++Y+ + P + + WNYGI L++K
Sbjct: 196 KCSSYSGFYNLGPGQESHPENWNYGIALKYK 226
>UniRef100_O65389 F12F1.23 protein [Arabidopsis thaliana]
Length = 329
Score = 55.8 bits (133), Expect = 7e-07
Identities = 57/228 (25%), Positives = 93/228 (40%), Gaps = 36/228 (15%)
Query: 14 LLLFLLITGLGSVINASPC-SNCGDFEVPYPLSTNDDCGDKRYK---IYCNNDSLEFLSA 69
LL+ +L+ + ++ C S+CG+ + YP S +D CG Y+ I +ND+ L
Sbjct: 13 LLMTILLQSSTTSSQSNLCRSSCGNIPINYPFSIDDGCGSPYYRHMLICSDNDTKLELRT 72
Query: 70 TGTYYKILKIDTSANKLVIKPPNIFKHTCYSSDLI--GGGLVLDESLPFNISTLNTVMLL 127
Y + I S L++ P F C D +D S F +S N +
Sbjct: 73 PSGKYPVKSISYSDPHLLVSDP--FMWNCQDRDNFRPTRSFSIDSSTHFTVSPQNDYLFF 130
Query: 128 NCSDN---ILQSPLNC------------SSNSICRQFEEKVEEGNGCMNTLCCHYLKDSV 172
NC+ + + PL C SS+ +CR E G + C +Y K
Sbjct: 131 NCNTDKVIVEPKPLFCERFPDRCDSSCDSSSYLCRHLPE-CGSALGSRVSCCSYYPK--- 186
Query: 173 MNSHKIRLRVGSCTAYTCL------VDFKPDEPFKKWNYGIELQWKPP 214
+ +RL + C YT + V+ P + F + YGI + ++ P
Sbjct: 187 -ATQSLRLMLQDCATYTSVYWRSTGVENAPYDQFPE--YGIRVDYEFP 231
>UniRef100_Q69KT1 Hypothetical protein OSJNBa0034G14.6 [Oryza sativa]
Length = 327
Score = 54.3 bits (129), Expect = 2e-06
Identities = 52/232 (22%), Positives = 100/232 (42%), Gaps = 34/232 (14%)
Query: 9 LKMVILLLFLLITGLGSVINASPC-SNCGDFEVPYPLSTNDDCGDKRYK--IYC-NNDSL 64
L + L + +++ LGS C +CG V YPLS +D CG Y+ + C +N +L
Sbjct: 5 LHLHFLAILVVVPMLGSPAAGGLCRDSCGGIPVRYPLSIDDGCGSPYYRNMLTCADNATL 64
Query: 65 EFLSATGTYYKILKIDTSANKLVIKPPNIFKHTCYSSDLIGGGLVLDESLPFNISTLNTV 124
+ +GT Y ++ D + LV+ P+++ + + LD S F++S N
Sbjct: 65 RLRTPSGT-YPVVGADYADPHLVVTDPSMWTCERPFTSVRAAPFSLDTSTRFSLSPRNDY 123
Query: 125 MLLNC-SDNILQSPLNCSSNSICRQFEEKVEEG-----------NGCMNTL------CCH 166
+ +C + ++ P ++C ++ E+ + GC L CC
Sbjct: 124 LFFDCDEERVIVEP----RPAVCDRYPERCDSTCDSAGYLCRNLPGCRGALEENNMSCCA 179
Query: 167 YLKDSVMNSHKIRLRVGSCTAYTCLVDFKPDEPFKKWN----YGIELQWKPP 214
Y + + +RL + C +YT + + F ++ YG+ + ++ P
Sbjct: 180 YRPRA---AESLRLMLRHCESYTSVYWRAVGDKFPPYDQVPAYGVRVDFEIP 228
>UniRef100_Q5XV04 Hypothetical protein [Arabidopsis thaliana]
Length = 303
Score = 52.0 bits (123), Expect = 1e-05
Identities = 57/229 (24%), Positives = 95/229 (40%), Gaps = 33/229 (14%)
Query: 11 MVILLLFLLITGLGSVINASPC-SNCGDFEVPYPLSTNDDCGDKRYK--IYCNNDSLEFL 67
M IL+L L L + C S CG+ V YP + CG Y+ ++C ND L F
Sbjct: 1 MKILILILSFVTLFEICVVDACRSYCGNITVDYPFGIRNGCGHPGYRDLLFCMNDVLMFH 60
Query: 68 SATGTYYKILKIDTSANKLVIKPPNIFKHTCYSSDLIGGGLVLD----ESLPFNISTLNT 123
++G+ Y++L ID + + + P++ C + L G G + + FN ++ N
Sbjct: 61 ISSGS-YRVLDIDYAYQSITLHDPHM--SNCETIVLGGKGNGFEAEDWRTPYFNPTSDNV 117
Query: 124 VMLLNCSDN--ILQS------PLNCSSNSICRQF------------EEKVEEGNGCMNTL 163
ML+ CS I Q P S C ++ + + G+G +
Sbjct: 118 FMLIGCSPKSPIFQGFPEKKVPCRNISGMSCEEYMSCPAWDMVGYRQPGIHSGSG--PPM 175
Query: 164 CCHYLKDSVMNSHKIRLRVGSCTAYTCLVDFKPDEPFKKWNYGIELQWK 212
CC +SV + +L ++ L K P W YGI ++++
Sbjct: 176 CCGVGFESVKAINLSKLECEGYSSAYNLAPLKLRGP-SDWAYGIRVKYE 223
>UniRef100_Q9FGR6 Arabidopsis thaliana genomic DNA, chromosome 5, TAC clone:K6A12
[Arabidopsis thaliana]
Length = 335
Score = 52.0 bits (123), Expect = 1e-05
Identities = 57/229 (24%), Positives = 95/229 (40%), Gaps = 33/229 (14%)
Query: 11 MVILLLFLLITGLGSVINASPC-SNCGDFEVPYPLSTNDDCGDKRYK--IYCNNDSLEFL 67
M IL+L L L + C S CG+ V YP + CG Y+ ++C ND L F
Sbjct: 1 MKILILILSFVTLFEICVVDACRSYCGNITVDYPFGIRNGCGHPGYRDLLFCMNDVLMFH 60
Query: 68 SATGTYYKILKIDTSANKLVIKPPNIFKHTCYSSDLIGGGLVLD----ESLPFNISTLNT 123
++G+ Y++L ID + + + P++ C + L G G + + FN ++ N
Sbjct: 61 ISSGS-YRVLDIDYAYQSITLHDPHM--SNCETIVLGGKGNGFEAEDWRTPYFNPTSDNV 117
Query: 124 VMLLNCSDN--ILQS------PLNCSSNSICRQF------------EEKVEEGNGCMNTL 163
ML+ CS I Q P S C ++ + + G+G +
Sbjct: 118 FMLIGCSPKSPIFQGFPEKKVPCRNISGMSCEEYMSCPAWDMVGYRQPGIHSGSG--PPM 175
Query: 164 CCHYLKDSVMNSHKIRLRVGSCTAYTCLVDFKPDEPFKKWNYGIELQWK 212
CC +SV + +L ++ L K P W YGI ++++
Sbjct: 176 CCGVGFESVKAINLSKLECEGYSSAYNLAPLKLRGP-SDWAYGIRVKYE 223
>UniRef100_Q5S4S0 Hypothetical protein [Arabidopsis thaliana]
Length = 303
Score = 50.4 bits (119), Expect = 3e-05
Identities = 57/229 (24%), Positives = 94/229 (40%), Gaps = 33/229 (14%)
Query: 11 MVILLLFLLITGLGSVINASPC-SNCGDFEVPYPLSTNDDCGDKRYK--IYCNNDSLEFL 67
M IL L L L + C S CG+ V YP + CG Y+ ++C ND L F
Sbjct: 1 MKILSLILSFVTLFEICVVDACRSYCGNITVDYPFGIRNGCGHPGYRDLLFCMNDVLMFH 60
Query: 68 SATGTYYKILKIDTSANKLVIKPPNIFKHTCYSSDLIGGGLVLD----ESLPFNISTLNT 123
++G+ Y++L ID + + + P++ C + L G G + + FN ++ N
Sbjct: 61 ISSGS-YRVLDIDYAYQSITLHDPHM--SNCETIVLGGKGNGFEAEDWRTPYFNPTSDNV 117
Query: 124 VMLLNCSDN--ILQS------PLNCSSNSICRQF------------EEKVEEGNGCMNTL 163
ML+ CS I Q P S C ++ + + G+G +
Sbjct: 118 FMLIGCSPKSPIFQGFPEKKVPCRNISGMSCEEYMSCPAWDMVGYRQPGIHSGSG--PPM 175
Query: 164 CCHYLKDSVMNSHKIRLRVGSCTAYTCLVDFKPDEPFKKWNYGIELQWK 212
CC +SV + +L ++ L K P W YGI ++++
Sbjct: 176 CCGVGFESVKAINLSKLECEGYSSAYNLAPLKLRGP-SDWAYGIRVKYE 223
>UniRef100_Q9AXH6 Wall-associated protein kinase [Oryza sativa]
Length = 722
Score = 50.4 bits (119), Expect = 3e-05
Identities = 43/170 (25%), Positives = 67/170 (39%), Gaps = 15/170 (8%)
Query: 8 SLKMVILLLFLL---ITGLGSVINASPC-SNCGDFEVPYPLSTNDDCGDKRYKIYCNNDS 63
S + V+LLL+LL ++ + C S CGD ++P P D C + + + CN
Sbjct: 12 STQRVVLLLWLLHAPAAADAALTTVAGCPSKCGDVDIPLPFGIGDHCAWESFDVVCNESF 71
Query: 64 LEFLSATGTYYKILKIDTSANKLVIKPPNIFKHTCYSSDLI---GGGLVLDESLPFNIST 120
TG +I +I A ++ + P CY+S G G L+ + PF ++
Sbjct: 72 SPPRPHTGN-IEIKEISVEAGEMRVYTP--VADQCYNSSSTSAPGFGASLELTAPFLLAQ 128
Query: 121 LNTVMLLNCSDNILQSPLNCSSNS-----ICRQFEEKVEEGNGCMNTLCC 165
N + C+ N S S C E + G C CC
Sbjct: 129 SNEFTAIGCNTVAFLDGRNNGSYSTGCITTCGSVEAAAQNGEPCTGLGCC 178
>UniRef100_Q8S8N4 Putative Ser/Thr protein kinase [Arabidopsis thaliana]
Length = 633
Score = 44.7 bits (104), Expect = 0.002
Identities = 53/220 (24%), Positives = 93/220 (42%), Gaps = 26/220 (11%)
Query: 7 NSLKMVILLLFLLITGLGSVINASPCS-----NCGDFEVPYPLSTNDDCGDKRYKIYCNN 61
+S ++LLL L + L S+ + P + CG+F V +P + +++ C N
Sbjct: 10 SSALFLLLLLLLTLQTLTSISLSQPQALRSPEKCGNFSVSFPFQLSSSSSAAAFRLSCEN 69
Query: 62 DSLEFLSATGTYYKILKIDTSANKLVIKPPNIFKHTCYS-SDLIGGGLVLDESLPFNIST 120
S FL Y+I++ T L++ P+ +C +DL + F+IS
Sbjct: 70 SSTLFLHINHQSYRIIEFFTDG--LLVDFPS--SPSCRQFNDL--RSFPFSANQFFSISF 123
Query: 121 LNTVMLLNCSDNILQSPLNCSSNSI--CRQFEEKVEEGNGCMNTLCCHYLKD----SVMN 174
N + L +C D+ L C +N + C EE G + CC+ L D V +
Sbjct: 124 ENVIGLYDCEDSSL-CKFGCETNDLFGCDGREEDETSGG---DIGCCYPLSDHSAWRVGD 179
Query: 175 SHKIRLRVGSCTAYTCLVDFKPDEPFKKWNYGIELQWKPP 214
+ R G C ++ + + K+ G++L+W P
Sbjct: 180 DFSVFSRYG-CRGFSSWLVPRGTNRGKR---GVKLEWAIP 215
>UniRef100_Q9ARW4 Putative wall-associated kinase 4 [Oryza sativa]
Length = 725
Score = 43.5 bits (101), Expect = 0.004
Identities = 43/175 (24%), Positives = 68/175 (38%), Gaps = 14/175 (8%)
Query: 14 LLLFLLITGLGSVINASPC----SNCGDFEVPYPLSTNDDCG-DKRYKIYCNNDSLEFLS 68
+LL + I S I+A P S+CGD E+PYP +C + + IYCN + +
Sbjct: 5 ILLSIAIMAQLSSISAQPAPGCQSHCGDMEIPYPFGIGTECAIEPGFVIYCNKTADGSMK 64
Query: 69 ATGTYYKILKIDTSANKLVIKPPNIFKHTCY---SSDLIGGGLVLDESL-PFNISTL-NT 123
++L I + + N CY + + LD S P+ S L N
Sbjct: 65 PFLINVEVLNISLLHGQ--TRALNALSTYCYNDVTKSMESSRWSLDFSTWPYRFSNLHNK 122
Query: 124 VMLLNCSDNILQSPLNCSSNSICRQFEEKVEEGNGCMNTLCCHYLKDSVMNSHKI 178
+++ C N L N + C K + C CC +NS+ +
Sbjct: 123 FVVIGC--NTLSYIYNGEYTTACASVCAKAPTNDSCDGVGCCQNNIAKGLNSYNV 175
>UniRef100_Q8VYA3 Wall-associated kinase 2, putative [Arabidopsis thaliana]
Length = 769
Score = 41.2 bits (95), Expect = 0.019
Identities = 36/150 (24%), Positives = 60/150 (40%), Gaps = 15/150 (10%)
Query: 13 ILLLFLLITGLGSVINASPCSNCGDFEVPYPLSTNDDCG-DKRYKIYCNNDSLEFLSATG 71
+L LF L+ + + +S CG ++PYP C +K Y+I C N+S+ FLS
Sbjct: 9 LLSLFSLLLIIDLTVASSCPKTCGGIDIPYPFGIGTGCYLEKWYEIICVNNSVPFLSIIN 68
Query: 72 TYYKILK--------IDTSANKLVIKPPNIFKHTCYSSDLIGGGLVLDESLPFNISTLNT 123
+ + + I+ P I C S G L+ PF + N
Sbjct: 69 REVVSISFSDMYRRFFNVGYGSIRIRNP-IASKGCSSGGQEFGSLLNMTGYPFYLGDNNM 127
Query: 124 VMLLNCSD-----NILQSPLNCSSNSICRQ 148
++ + C++ N+ S + C S Q
Sbjct: 128 LIAVGCNNTASLTNVEPSIVGCESTCSTNQ 157
>UniRef100_Q9MA08 F20B17.10 [Arabidopsis thaliana]
Length = 1487
Score = 41.2 bits (95), Expect = 0.019
Identities = 36/150 (24%), Positives = 60/150 (40%), Gaps = 15/150 (10%)
Query: 13 ILLLFLLITGLGSVINASPCSNCGDFEVPYPLSTNDDCG-DKRYKIYCNNDSLEFLSATG 71
+L LF L+ + + +S CG ++PYP C +K Y+I C N+S+ FLS
Sbjct: 9 LLSLFSLLLIIDLTVASSCPKTCGGIDIPYPFGIGTGCYLEKWYEIICVNNSVPFLSIIN 68
Query: 72 TYYKILK--------IDTSANKLVIKPPNIFKHTCYSSDLIGGGLVLDESLPFNISTLNT 123
+ + + I+ P I C S G L+ PF + N
Sbjct: 69 REVVSISFSDMYRRFFNVGYGSIRIRNP-IASKGCSSGGQEFGSLLNMTGYPFYLGDNNM 127
Query: 124 VMLLNCSD-----NILQSPLNCSSNSICRQ 148
++ + C++ N+ S + C S Q
Sbjct: 128 LIAVGCNNTASLTNVEPSIVGCESTCSTNQ 157
>UniRef100_Q9FZI4 F1O19.2 protein [Arabidopsis thaliana]
Length = 332
Score = 40.8 bits (94), Expect = 0.024
Identities = 18/69 (26%), Positives = 35/69 (50%), Gaps = 3/69 (4%)
Query: 35 CGDFEVPYPLSTND---DCGDKRYKIYCNNDSLEFLSATGTYYKILKIDTSANKLVIKPP 91
CG+ +P S + +CG +++CN +++ L + Y +L+ID ++N L +
Sbjct: 39 CGNITAGFPFSGGNRHKECGHPSLELHCNKNNITSLFISNQKYSVLRIDQTSNTLTLAKQ 98
Query: 92 NIFKHTCYS 100
N+ C S
Sbjct: 99 NLLGSFCSS 107
>UniRef100_Q9S9M6 T24D18.19 protein [Arabidopsis thaliana]
Length = 1016
Score = 40.0 bits (92), Expect = 0.042
Identities = 54/209 (25%), Positives = 86/209 (40%), Gaps = 59/209 (28%)
Query: 5 IWNSLKMVILLLFLLITGLGSVINA-SPC-SNCGDFEVPYPLSTNDDCGDKRYKIYCNND 62
I+N L +++LL+ LI + S C S+CG +PYP DC Y NN+
Sbjct: 8 IYNLLSLLVLLIHGLIAASAQDLTVFSSCQSHCGGIAIPYPFGIGKDC-------YLNNN 60
Query: 63 SLEFLSATGTYYKILKIDTSANKL-VIKPPNIFKHTCYSSDLIGGGLVLDESLPFNISTL 121
+Y+++ TS N L V+K N +L+ L D S F ++ +
Sbjct: 61 E---------WYEVICNRTSGNPLPVLKSIN--------RELVNISLPDDSSDVFGLTRI 103
Query: 122 -NTVMLLNCSD--------NILQSP-----------LNCSSNSICRQFEEKVEEGNGCMN 161
N V L CS+ N+ SP + C++N+ ++K++ G GC +
Sbjct: 104 KNPVTSLGCSNMEEISLALNVTGSPFFLTGRNTLVAVGCNNNA--SMTDDKLQIG-GCES 160
Query: 162 TLCCHYLKDSVMNSHKIRLRVGSCTAYTC 190
T + + R R SC Y C
Sbjct: 161 TCDVGFGQ---------RGRNRSCNGYRC 180
>UniRef100_Q7XGS0 Putative wall-associated protein kinase [Oryza sativa]
Length = 693
Score = 39.3 bits (90), Expect = 0.071
Identities = 56/220 (25%), Positives = 86/220 (38%), Gaps = 38/220 (17%)
Query: 15 LLFLLITGLGSVIN---ASPCSNCGDFEVPYPLSTNDDCG-DKRYKIYCN--NDSLEFL- 67
LL LL + SV AS + CGD ++PYP +C K ++I CN NDS E +
Sbjct: 9 LLILLASAAESVAGQPAASCQARCGDIDIPYPFGIGPNCSRGKGFEIACNPRNDSGEMVP 68
Query: 68 ---SATGTYYKILKIDTSANKLVIKPPNIFKHTCYSS-----DLIGGGLVLDESLPFNIS 119
+A GT + + ++ + P ++ CY S D G + L+ + + IS
Sbjct: 69 TLAAANGTIHVQSLLVAPIPEVKVMLPVAYQ--CYYSNNSITDSFYGEVDLNNTGVYRIS 126
Query: 120 -TLNTVMLLNCSDNILQSPLNCSSNSICRQFEEKVEEGNGCMNTLCCHYLKDSVMNSHKI 178
+ N +++ C N L N +S G T C Y DS +
Sbjct: 127 DSRNMFVVIGC--NTLSYTQNGNSGG--------KGPYAGLYYTGCVSYCNDSSSARDSM 176
Query: 179 RLRVGSCTAYTCLVDFKPD-----EPFKKWNYGIELQWKP 213
VG C +D P F W G ++ + P
Sbjct: 177 CAGVGCCH-----IDISPGLSDNVVSFGPWKRGFQVDFSP 211
>UniRef100_Q7XVH6 OSJNBa0016N04.10 protein [Oryza sativa]
Length = 739
Score = 38.9 bits (89), Expect = 0.093
Identities = 41/189 (21%), Positives = 70/189 (36%), Gaps = 33/189 (17%)
Query: 32 CSN--CGDFEVPYPLSTN-DDCGDKRYKIYCNNDSLEFLSATGTYYKILKIDTSANKLVI 88
C N CGD E+PYP ST+ D C ++ CN+ ++L + + +
Sbjct: 31 CQNTKCGDVEIPYPFSTSLDKCAASAFEFDCNDTGNGVYKPFYGNVEVLSVSLQLGQ--V 88
Query: 89 KPPNIFKHTCYS------------SDLIGGGLVLDESLPFNISTLNTVMLLNCSDNILQS 136
+ N +CY+ ++ G +L +S F + T + D + +
Sbjct: 89 RVMNHISSSCYNLSSKEMDSDTWQLNMTGTPFMLSDSNKFTVVGCRTQAYIADQDYVGKY 148
Query: 137 PLNCSSNSICRQFEEKVEEGNGCMNTLCCH------------YLKDSVMNSHKIRLRVGS 184
C S+CR+ + C CC + DS MN+ I R +
Sbjct: 149 MSGCV--SVCRRGDVWKATNGTCSGIGCCQTAIPKGLDYYQAFFDDSSMNTSGIYNR--T 204
Query: 185 CTAYTCLVD 193
+Y L+D
Sbjct: 205 PCSYAVLMD 213
>UniRef100_UPI0000498CCD UPI0000498CCD UniRef100 entry
Length = 819
Score = 38.5 bits (88), Expect = 0.12
Identities = 24/62 (38%), Positives = 36/62 (57%), Gaps = 3/62 (4%)
Query: 82 SANKLVIKPPNIFKHTCYSSDLIGGGLVLDESLPFNISTLNTVMLLNCSDNILQS-PLNC 140
S+NK+ I P ++FK T ++ ++ + LP NISTL+ + LN S N L PL+
Sbjct: 58 SSNKISILPSHLFKITSLKKLILSQNILYE--LPLNISTLSNLTCLNLSQNKLSKIPLSI 115
Query: 141 SS 142
SS
Sbjct: 116 SS 117
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.139 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 391,844,470
Number of Sequences: 2790947
Number of extensions: 16160555
Number of successful extensions: 39020
Number of sequences better than 10.0: 63
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 55
Number of HSP's that attempted gapping in prelim test: 38989
Number of HSP's gapped (non-prelim): 71
length of query: 215
length of database: 848,049,833
effective HSP length: 122
effective length of query: 93
effective length of database: 507,554,299
effective search space: 47202549807
effective search space used: 47202549807
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 72 (32.3 bits)
Medicago: description of AC144513.2


