

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144484.11 - phase: 0
(796 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
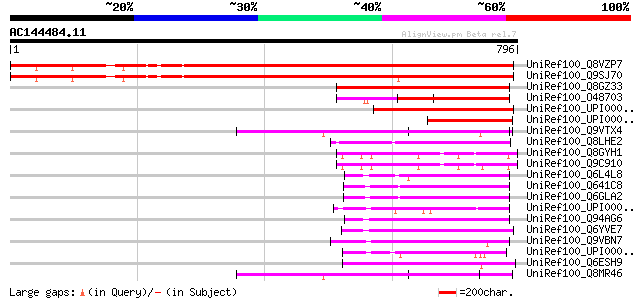
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8VZP7 Putative mitochondrial carrier protein [Arabido... 994 0.0
UniRef100_Q9SJ70 Probable calcium-binding mitochondrial carrier ... 983 0.0
UniRef100_Q8GZ33 Putative mitochondrial carrier protein [Arabido... 379 e-103
UniRef100_O48703 Putative mitochondrial carrier protein [Arabido... 256 2e-66
UniRef100_UPI0000291028 UPI0000291028 UniRef100 entry 204 6e-51
UniRef100_UPI000028DB4C UPI000028DB4C UniRef100 entry 135 4e-30
UniRef100_Q9VTX4 CG32103-PA, isoform A [Drosophila melanogaster] 134 1e-29
UniRef100_Q8LHE2 P0458E05.7 protein [Oryza sativa] 134 1e-29
UniRef100_Q8GYH1 Putative mitochondrial carrier protein [Arabido... 131 7e-29
UniRef100_Q9C910 Putative mitochondrial carrier protein; 35518-3... 131 9e-29
UniRef100_Q6L4L8 Putative mitochondrial carrier protein [Oryza s... 130 1e-28
UniRef100_Q641C8 MGC82075 protein [Xenopus laevis] 128 6e-28
UniRef100_Q6GLA2 MGC69323 protein [Xenopus tropicalis] 128 8e-28
UniRef100_UPI000023D6B1 UPI000023D6B1 UniRef100 entry 124 8e-27
UniRef100_Q94AG6 Putative mitochondrial carrier protein [Arabido... 124 8e-27
UniRef100_Q6YVE7 Mitochondrial aspartate-glutamate carrier prote... 123 2e-26
UniRef100_Q9VBN7 CG4743-PA [Drosophila melanogaster] 122 4e-26
UniRef100_UPI000042F384 UPI000042F384 UniRef100 entry 120 2e-25
UniRef100_Q6ESH9 Mitochondrial substrate carrier protein-like [O... 119 4e-25
UniRef100_Q8MR46 GH25190p [Drosophila melanogaster] 118 6e-25
>UniRef100_Q8VZP7 Putative mitochondrial carrier protein [Arabidopsis thaliana]
Length = 823
Score = 994 bits (2571), Expect = 0.0
Identities = 538/833 (64%), Positives = 639/833 (76%), Gaps = 65/833 (7%)
Query: 1 MVLDNDPVESFFNSIQVMKES-LSPLEVGFRKAAKDFEHCF--------------AKNKT 45
MV ND +E+ FNSIQ++K+ L P+E+G +KAA+D E+C+ +N+
Sbjct: 1 MVSKNDHIETLFNSIQLVKDVVLLPIELGVKKAARDIENCWISKERDLGLVFRSSGRNRK 60
Query: 46 QGVCLIAQVKDGGDFQI-CDV---KKKKGLSMK-VPLKAFLGKFSQN--SEKLNKTQVV- 97
+ + + D + C V ++KKGLS+K +P+K+ G FS N SEKL++ V
Sbjct: 61 KRIVATPEFDDNATNNVQCVVVTDERKKGLSIKKIPVKSLFGMFSPNLVSEKLSRGNDVV 120
Query: 98 ---------KENESSCSNCLKFSVTWSLLVSGFIQSLPIPFKSVKKRGQKVCDEDSHKEK 148
K+++ SC++C KF++TWSLLVSGF+ + PIPFK KKR K+ D+++ K
Sbjct: 121 VAKKDKSLEKKDDDSCTDCFKFAMTWSLLVSGFVHAFPIPFKIGKKRIHKMGDDENSLRK 180
Query: 149 CSCMKPSLSPCEMKHNESKGRTIKEKVVK-------RKDGKEHVSLECVIGFIFDQLSHT 201
+SK + K V+ K+G S+EC +GF+ + L+
Sbjct: 181 HGL-------------KSKAAFVSRKEVRCQSVESVEKEGNPF-SIECAVGFVVEMLAQN 226
Query: 202 LQSLDQGINGLQEKNDELECGK-ASLDSAPFGHVNAFTSFLEGHKVDVNGFLGNLNFAKV 260
LQ LDQ I E +E C K AS + +P + E K+DVNGFLGNL FA+V
Sbjct: 227 LQKLDQFIQDSSE--NESCCSKEASSNDSPL-----IFNIWEARKLDVNGFLGNLMFARV 279
Query: 261 GGVPSSVAGEEIASQNEMGDSANDET--KEESVGISAQKVASNIFSIPLTNVERLKTTLS 318
G V S + G + +E GD +N T KEES S Q +A+ + SIPL+NVERLK+TLS
Sbjct: 280 GDVASGIGGLT-SHISEDGDESNVSTAGKEESAVDSPQNLATGLLSIPLSNVERLKSTLS 338
Query: 319 TVSLTELIEMLPQLGKTTKDHPDKKKLFSVQDFFRYTESEGRRFFEELDRDGDGQVTLED 378
T+SLTELIE+LPQLG+ ++DHPDKKKL SVQDFFRYTESEGRRFFEELDRDGDG+VTLED
Sbjct: 339 TISLTELIELLPQLGRPSRDHPDKKKLISVQDFFRYTESEGRRFFEELDRDGDGKVTLED 398
Query: 379 LEIAMRRRKLPRRYAKEFMSRTRSHLFSRSFGWKQFLSFMEQKEPTILRAYTSLCLTKSG 438
LEIAMRRRKLPRRYAKEFM R RSHLFS+SFGWKQFLS MEQKEPTILRAYTSLCLTKSG
Sbjct: 399 LEIAMRRRKLPRRYAKEFMRRARSHLFSKSFGWKQFLSLMEQKEPTILRAYTSLCLTKSG 458
Query: 439 TLKKSEILESLKNSGLPANEDNAAAMMRFLNADTEESISYGHFRNFMLLLPSDRLQEDPR 498
TLKKSEIL SL N+GLPANE+NA AMMRFL ADTEESISYGHFRNFM+LLP +RLQ+DPR
Sbjct: 459 TLKKSEILASLNNAGLPANEENAIAMMRFLKADTEESISYGHFRNFMVLLPYERLQDDPR 518
Query: 499 SIWFEAATVVAVPPSVEIPAGSVLRSALAGGLSCALSCALLHPVDSIKTRVQASSMSFPE 558
+IWFEAATVVAV P V +PAG VL+SALAGGL+ ALS +L+HP+D+IKTRVQAS++SFPE
Sbjct: 519 NIWFEAATVVAVAPPVALPAGDVLKSALAGGLASALSTSLMHPIDTIKTRVQASTLSFPE 578
Query: 559 IIAKLPEIGTRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASF 618
+IAKLPEIG RG+YRGSIPAILGQFSSHGLRTGIFEASKLVL+N APNLPE+QVQSIASF
Sbjct: 579 VIAKLPEIGVRGVYRGSIPAILGQFSSHGLRTGIFEASKLVLINFAPNLPEIQVQSIASF 638
Query: 619 CSTFLGTAVRIPCEVLKQRLQAGLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYV 678
CST LGTAVRIPCEVLKQRLQAG+FNNVGEA+VGTW+QDG GFFRGTGATLCREVP YV
Sbjct: 639 CSTLLGTAVRIPCEVLKQRLQAGMFNNVGEAIVGTWKQDGPSGFFRGTGATLCREVPLYV 698
Query: 679 AGMGLYAESKKGVQKLLGRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTA-QGRSV 737
GMGLYAESKK V + LGRELEAWETIAVGA+SGG+AAVVTTPFDVMKTRMMTA GR +
Sbjct: 699 VGMGLYAESKKMVAQALGRELEAWETIAVGAVSGGIAAVVTTPFDVMKTRMMTATPGRPI 758
Query: 738 SMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAMNKNDEA 790
SMS+V SILR+EGPLGLFKGAVPRFFW+APLGAMNFAGYELA+KAM KN++A
Sbjct: 759 SMSMVVVSILRNEGPLGLFKGAVPRFFWVAPLGAMNFAGYELAKKAMQKNEDA 811
>UniRef100_Q9SJ70 Probable calcium-binding mitochondrial carrier At2g35800
[Arabidopsis thaliana]
Length = 844
Score = 983 bits (2541), Expect = 0.0
Identities = 539/854 (63%), Positives = 639/854 (74%), Gaps = 86/854 (10%)
Query: 1 MVLDNDPVESFFNSIQVMKES-LSPLEVGFRKAAKDFEHCF--------------AKNKT 45
MV ND +E+ FNSIQ++K+ L P+E+G +KAA+D E+C+ +N+
Sbjct: 1 MVSKNDHIETLFNSIQLVKDVVLLPIELGVKKAARDIENCWISKERDLGLVFRSSGRNRK 60
Query: 46 QGVCLIAQVKDGGDFQI-CDV---KKKKGLSMK-VPLKAFLGKFSQN--SEKLNKTQVV- 97
+ + + D + C V ++KKGLS+K +P+K+ G FS N SEKL++ V
Sbjct: 61 KRIVATPEFDDNATNNVQCVVVTDERKKGLSIKKIPVKSLFGMFSPNLVSEKLSRGNDVV 120
Query: 98 ---------KENESSCSNCLKFSVTWSLLVSGFIQSLPIPFKSVKKRGQKVCDEDSHKEK 148
K+++ SC++C KF++TWSLLVSGF+ + PIPFK KKR K+ D+++ K
Sbjct: 121 VAKKDKSLEKKDDDSCTDCFKFAMTWSLLVSGFVHAFPIPFKIGKKRIHKMGDDENSLRK 180
Query: 149 CSCMKPSLSPCEMKHNESKGRTIKEKVVK-------RKDGKEHVSLECVIGFIFDQLSHT 201
+SK + K V+ K+G S+EC +GF+ + L+
Sbjct: 181 HGL-------------KSKAAFVSRKEVRCQSVESVEKEGNPF-SIECAVGFVVEMLAQN 226
Query: 202 LQSLDQGINGLQEKNDELECGK-ASLDSAPFGHVNAFTSFLEGHKVDVNGFLGNLNFAKV 260
LQ LDQ I E +E C K AS + +P + E K+DVNGFLGNL FA+V
Sbjct: 227 LQKLDQFIQDSSE--NESCCSKEASSNDSPL-----IFNIWEARKLDVNGFLGNLMFARV 279
Query: 261 GGVPSSVAGEEIASQNEMGDSANDET--KEESVGISAQKVASNIFSIPLTNVERLKTTLS 318
G V S + G + +E GD +N T KEES S Q +A+ + SIPL+NVERLK+TLS
Sbjct: 280 GDVASGIGGLT-SHISEDGDESNVSTAGKEESAVDSPQNLATGLLSIPLSNVERLKSTLS 338
Query: 319 TVSLTELIEMLPQLGKTTKDHPDKKKLFSVQDFFRYTESEGRRFFEELDRDGDGQVTLED 378
T+SLTELIE+LPQLG+ ++DHPDKKKL SVQDFFRYTESEGRRFFEELDRDGDG+VTLED
Sbjct: 339 TISLTELIELLPQLGRPSRDHPDKKKLISVQDFFRYTESEGRRFFEELDRDGDGKVTLED 398
Query: 379 LEIAMRRRKLPRRYAKEFMSRTRSHLFSRSFGWKQFLSFMEQKEPTILRAYTSLCLTKSG 438
LEIAMRRRKLPRRYAKEFM R RSHLFS+SFGWKQFLS MEQKEPTILRAYTSLCLTKSG
Sbjct: 399 LEIAMRRRKLPRRYAKEFMRRARSHLFSKSFGWKQFLSLMEQKEPTILRAYTSLCLTKSG 458
Query: 439 TLKKSEILESLKNSGLPANEDNAAAMMRFLNADTEESISYGHFRNFMLLLPSDRLQEDPR 498
TLKKSEIL SL N+GLPANE+NA AMMRFL ADTEESISYGHFRNFM+LLP +RLQ+DPR
Sbjct: 459 TLKKSEILASLNNAGLPANEENAIAMMRFLKADTEESISYGHFRNFMVLLPYERLQDDPR 518
Query: 499 SIWFEAATVVAVPPSVEIPAGSVLRSALAGGLSCALSCALLHPVDSIKTRVQASSMSFPE 558
+IWFEAATVVAV P V +PAG VL+SALAGGL+ ALS +L+HP+D+IKTRVQAS++SFPE
Sbjct: 519 NIWFEAATVVAVAPPVALPAGDVLKSALAGGLASALSTSLMHPIDTIKTRVQASTLSFPE 578
Query: 559 IIAKLPEIGTRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPE--------- 609
+IAKLPEIG RG+YRGSIPAILGQFSSHGLRTGIFEASKLVL+N APNLPE
Sbjct: 579 VIAKLPEIGVRGVYRGSIPAILGQFSSHGLRTGIFEASKLVLINFAPNLPEIQVIITLYS 638
Query: 610 ------------LQVQSIASFCSTFLGTAVRIPCEVLKQRLQAGLFNNVGEALVGTWQQD 657
LQVQSIASFCST LGTAVRIPCEVLKQRLQAG+FNNVGEA+VGTW+QD
Sbjct: 639 LFGWFRQDSNFVLQVQSIASFCSTLLGTAVRIPCEVLKQRLQAGMFNNVGEAIVGTWKQD 698
Query: 658 GLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEAWETIAVGALSGGLAAV 717
G GFFRGTGATLCREVP YV GMGLYAESKK V + LGRELEAWETIAVGA+SGG+AAV
Sbjct: 699 GPSGFFRGTGATLCREVPLYVVGMGLYAESKKMVAQALGRELEAWETIAVGAVSGGIAAV 758
Query: 718 VTTPFDVMKTRMMTA-QGRSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAG 776
VTTPFDVMKTRMMTA GR +SMS+V SILR+EGPLGLFKGAVPRFFW+APLGAMNFAG
Sbjct: 759 VTTPFDVMKTRMMTATPGRPISMSMVVVSILRNEGPLGLFKGAVPRFFWVAPLGAMNFAG 818
Query: 777 YELARKAMNKNDEA 790
YELA+KAM KN++A
Sbjct: 819 YELAKKAMQKNEDA 832
>UniRef100_Q8GZ33 Putative mitochondrial carrier protein [Arabidopsis thaliana]
Length = 369
Score = 379 bits (973), Expect = e-103
Identities = 191/273 (69%), Positives = 226/273 (81%), Gaps = 2/273 (0%)
Query: 514 VEIPAGSVLRSALAGGLSCALSCALLHPVDSIKTRVQASS-MSFPEIIAKLPEIGTRGLY 572
V + G +L+SALAGG+SCA S L+HPVD++KT+VQAS+ +SF EI++K+PEIG RGLY
Sbjct: 86 VGLDVGHLLKSALAGGISCAFSAFLMHPVDTVKTQVQASTTLSFLEILSKIPEIGARGLY 145
Query: 573 RGSIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCE 632
+GSIPA++GQF+SHGLRT I+EASKL L VAP L ++QVQSIASF T LGT +RIPCE
Sbjct: 146 KGSIPAVVGQFASHGLRTSIYEASKLALPLVAPTLLDIQVQSIASFIGTVLGTTLRIPCE 205
Query: 633 VLKQRLQAGLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQ 692
VLKQRLQA F+N+ EA V TW Q+GLKG FRGTG TL REVPFYVAGMGLY +SKK V+
Sbjct: 206 VLKQRLQANQFDNIVEATVSTWHQEGLKGLFRGTGVTLLREVPFYVAGMGLYNQSKKVVE 265
Query: 693 KLLGRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTA-QGRSVSMSIVAFSILRHEG 751
+ LGRELE WE IAVGALSGG AV+TTPFDV+KTRMMTA QG +SM + A+SIL HEG
Sbjct: 266 RQLGRELEPWEAIAVGALSGGFTAVLTTPFDVIKTRMMTAPQGVELSMLMAAYSILTHEG 325
Query: 752 PLGLFKGAVPRFFWIAPLGAMNFAGYELARKAM 784
PL +KGAVPRFFW APLGA+N AGYEL +KAM
Sbjct: 326 PLAFYKGAVPRFFWTAPLGALNLAGYELLQKAM 358
>UniRef100_O48703 Putative mitochondrial carrier protein [Arabidopsis thaliana]
Length = 303
Score = 256 bits (653), Expect = 2e-66
Identities = 128/176 (72%), Positives = 145/176 (81%), Gaps = 1/176 (0%)
Query: 610 LQVQSIASFCSTFLGTAVRIPCEVLKQRLQAGLFNNVGEALVGTWQQDGLKGFFRGTGAT 669
++VQSIASF T LGT +RIPCEVLKQRLQA F+N+ EA V TW Q+GLKG FRGTG T
Sbjct: 117 VKVQSIASFIGTVLGTTLRIPCEVLKQRLQANQFDNIVEATVSTWHQEGLKGLFRGTGVT 176
Query: 670 LCREVPFYVAGMGLYAESKKGVQKLLGRELEAWETIAVGALSGGLAAVVTTPFDVMKTRM 729
L REVPFYVAGMGLY +SKK V++ LGRELE WE IAVGALSGG AV+TTPFDV+KTRM
Sbjct: 177 LLREVPFYVAGMGLYNQSKKVVERQLGRELEPWEAIAVGALSGGFTAVLTTPFDVIKTRM 236
Query: 730 MTA-QGRSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAM 784
MTA QG +SM + A+SIL HEGPL +KGAVPRFFW APLGA+N AGYEL +KAM
Sbjct: 237 MTAPQGVELSMLMAAYSILTHEGPLAFYKGAVPRFFWTAPLGALNLAGYELLQKAM 292
Score = 48.5 bits (114), Expect = 8e-04
Identities = 44/182 (24%), Positives = 80/182 (43%), Gaps = 31/182 (17%)
Query: 514 VEIPAGSVLRSALAGGLSCALSCALLHPVDSIKTRVQASSMS------------------ 555
V + G +L+SALAGG+SCA S L+HPVD++K + AS +
Sbjct: 86 VGLDVGHLLKSALAGGISCAFSAFLMHPVDTVKVQSIASFIGTVLGTTLRIPCEVLKQRL 145
Query: 556 ----FPEII----AKLPEIGTRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLV-NVAPN 606
F I+ + + G +GL+RG+ +L + + G++ SK V+ +
Sbjct: 146 QANQFDNIVEATVSTWHQEGLKGLFRGTGVTLLREVPFYVAGMGLYNQSKKVVERQLGRE 205
Query: 607 LPELQVQSIASFCSTFLGTAVRIPCEVLKQRLQA---GLFNNVGEALVGTWQQDGLKGFF 663
L + ++ + F + P +V+K R+ G+ ++ A +G F+
Sbjct: 206 LEPWEAIAVGALSGGFT-AVLTTPFDVIKTRMMTAPQGVELSMLMAAYSILTHEGPLAFY 264
Query: 664 RG 665
+G
Sbjct: 265 KG 266
>UniRef100_UPI0000291028 UPI0000291028 UniRef100 entry
Length = 252
Score = 204 bits (520), Expect = 6e-51
Identities = 115/225 (51%), Positives = 143/225 (63%), Gaps = 5/225 (2%)
Query: 571 LYRGSIPAILGQFSSHGLRTGIFEASKLVLVNVA--PNLPELQVQSIASFCSTFLGTAVR 628
+Y+G IP++ G F+ HGLRT +E + L P + E +Q + S T LGT VR
Sbjct: 1 MYQGIIPSVTGNFAGHGLRTATYEVVCIALAPAMALPMVTETTIQGLGSGLGTLLGTCVR 60
Query: 629 IPCEVLKQRLQAGLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESK 688
IPCEVLKQRLQ + NV AL G + + + + GT ATL RE+PFYV G+ +Y K
Sbjct: 61 IPCEVLKQRLQTDQYPNVPAALKGIVKTNP-RALYAGTAATLTREIPFYVTGLMIYENLK 119
Query: 689 KGVQKLLG-RELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFS-I 746
KG L G RELE ++ I VGA +G L +V+ PFDVMKTR MT VA + I
Sbjct: 120 KGAVALKGGRELENYQVIMVGACAGALGSVMVNPFDVMKTRTMTGSVPLGQPLWVAMAHI 179
Query: 747 LRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAMNKNDEAK 791
++ EGPL L KGA+PR WIAPLGAMNFAGYELA+KAMN DEAK
Sbjct: 180 VKTEGPLALMKGAIPRMAWIAPLGAMNFAGYELAKKAMNTADEAK 224
>UniRef100_UPI000028DB4C UPI000028DB4C UniRef100 entry
Length = 257
Score = 135 bits (341), Expect = 4e-30
Identities = 70/135 (51%), Positives = 88/135 (64%), Gaps = 1/135 (0%)
Query: 656 QDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEAWETIAVGALSGGLA 715
++G +G F GT ATL REVPFYV G+ Y + KK + ++L AWETI +G LSG +A
Sbjct: 6 KNGPRGLFAGTAATLSREVPFYVIGLVAYEKLKKAATAMKRKDLTAWETIGLGGLSGAIA 65
Query: 716 AVVTTPFDVMKTRMMT-AQGRSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNF 774
A TTP DV+KTR MT A ++ + +I+ EG L KG PR WIAPLGAMNF
Sbjct: 66 AAATTPADVLKTRAMTGASPAGEAIWVTVKNIIAKEGGGALMKGCFPRMLWIAPLGAMNF 125
Query: 775 AGYELARKAMNKNDE 789
AGYELA++AM DE
Sbjct: 126 AGYELAKRAMEAADE 140
>UniRef100_Q9VTX4 CG32103-PA, isoform A [Drosophila melanogaster]
Length = 583
Score = 134 bits (337), Expect = 1e-29
Identities = 111/466 (23%), Positives = 215/466 (45%), Gaps = 38/466 (8%)
Query: 356 ESEGRRFFEELDRDGDGQVTLEDLEIAMRRRKLPRRYAKEFMSRTRSHLFSRSFGWKQFL 415
E R F +LDRDGDG++ + DL A+ L YA++F+ ++ S + G+ +FL
Sbjct: 114 EERLERIFNKLDRDGDGRIDIHDLSAALHEFGLSSVYAEKFLQQSDKDQ-SGNVGFAEFL 172
Query: 416 SFMEQKEPTILRAYTSLCLTKSGTLKKSEILESLKNSGLPANEDNAAAMMRFLNADTEES 475
++ + E ++ ++ L + G + E++ + K+ GL + D A ++ ++ D +
Sbjct: 173 HYVREHEKNLVLQFSHLDKNRDGKVDLEELISAFKDLGLDIDMDEARNLLTRMDKDGSLN 232
Query: 476 ISYGHFRNFMLLLPS----DRLQEDPRSIWFEAATVVAVPPSV---EIPAGSVLRSALAG 528
IS+ +R+FMLL PS D ++ S + + + VP E+ G R +AG
Sbjct: 233 ISFNEWRDFMLLAPSTDIHDLIKFWRHSTYLDIGEDMNVPDDFTQKEMQTGLWWRHLVAG 292
Query: 529 GLSCALSCALLHPVDSIKT--RVQASSMSFPEII-AKLPEIGTRGLYRGSIPAILGQFSS 585
G++ A+S P+D IK +VQ M E + L E G+R ++RG+ +L
Sbjct: 293 GIAGAVSRTCTAPLDRIKVYLQVQTQRMGISECMHIMLNEGGSRSMWRGNGINVLKIAPE 352
Query: 586 HGLRTGIFEASK-LVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLKQRL---QAG 641
+ +E K L+ + + + A + + + P EVLK RL + G
Sbjct: 353 TAFKFAAYEQMKRLIRGDDGSRQMSIVERFYAGAAAGGISQTIIYPMEVLKTRLALRRTG 412
Query: 642 LFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKG-VQKLLGRELE 700
+ + +A V ++Q+G++ F+RG + +P+ + +Y K+ + E
Sbjct: 413 QYAGIADAAVKIYKQEGVRSFYRGYVPNILGILPYAGIDLAVYETLKRRYIANHDNNEQP 472
Query: 701 AW-ETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSV---------------------S 738
++ +A G+ S L + + P +++TR+ ++ +
Sbjct: 473 SFLVLLACGSTSSTLGQLCSYPLALVRTRLQAQAAETIANQKRKTQIPLKSSDAHSGEET 532
Query: 739 MSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAM 784
M+ + I+R EG GL++G P F + P ++++ YE +A+
Sbjct: 533 MTGLFRKIVRQEGLTGLYRGITPNFLKVLPAVSISYVVYEYTSRAL 578
Score = 56.2 bits (134), Expect = 4e-06
Identities = 37/165 (22%), Positives = 74/165 (44%), Gaps = 8/165 (4%)
Query: 627 VRIPCEVLKQRLQAGLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAE 686
+++ +V QR+ + E + + G + +RG G + + P Y +
Sbjct: 309 IKVYLQVQTQRM------GISECMHIMLNEGGSRSMWRGNGINVLKIAPETAFKFAAYEQ 362
Query: 687 SKKGVQKLLG-RELEAWETIAVGALSGGLAAVVTTPFDVMKTRM-MTAQGRSVSMSIVAF 744
K+ ++ G R++ E GA +GG++ + P +V+KTR+ + G+ ++ A
Sbjct: 363 MKRLIRGDDGSRQMSIVERFYAGAAAGGISQTIIYPMEVLKTRLALRRTGQYAGIADAAV 422
Query: 745 SILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAMNKNDE 789
I + EG ++G VP I P ++ A YE ++ N +
Sbjct: 423 KIYKQEGVRSFYRGYVPNILGILPYAGIDLAVYETLKRRYIANHD 467
Score = 37.0 bits (84), Expect = 2.3
Identities = 21/90 (23%), Positives = 41/90 (45%), Gaps = 1/90 (1%)
Query: 702 WETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFSILRHEGPLGLFKGAVP 761
W + G ++G ++ T P D +K + Q + + +S +L G +++G
Sbjct: 286 WRHLVAGGIAGAVSRTCTAPLDRIKVYLQV-QTQRMGISECMHIMLNEGGSRSMWRGNGI 344
Query: 762 RFFWIAPLGAMNFAGYELARKAMNKNDEAK 791
IAP A FA YE ++ + +D ++
Sbjct: 345 NVLKIAPETAFKFAAYEQMKRLIRGDDGSR 374
>UniRef100_Q8LHE2 P0458E05.7 protein [Oryza sativa]
Length = 360
Score = 134 bits (336), Expect = 1e-29
Identities = 97/290 (33%), Positives = 142/290 (48%), Gaps = 13/290 (4%)
Query: 504 AATVVAVPPSVEIPAGSVLRSALAGGLSCALSCALLHPVDSIKTRVQASSM---SFPEII 560
AA PP+ P A AG + A + A L P+D++KTR+QA + S+ +
Sbjct: 56 AAVAATAPPT---PLARAAIGACAGAAAGAFTYAALLPIDAVKTRIQAGAAAGGSWQVFL 112
Query: 561 AKLPEIGTRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCS 620
L G GLYRG ILG SS + G E +K +L P+LP V +A
Sbjct: 113 DILRTDGPLGLYRGLSAVILGSASSSAVYFGTCELAKSLL---RPHLPPFLVPPLAGASG 169
Query: 621 TFLGTAVRIPCEVLKQRLQAGLFNNVG-EALVGTWQQDGLKGFFRGTGATLCREVPFYVA 679
+A+ +P E++ QRLQ+G + L+ Q DG G + G ATL R +P V
Sbjct: 170 NVSSSAIMVPKELITQRLQSGAAKGRSWQVLLQILQTDGFFGLYAGYAATLLRNLPAGVL 229
Query: 680 GMGLYAESKKGVQKLLGRE-LEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVS 738
+ K K +E L E++ GAL+G ++A +TTP DV+KTR+MT G S
Sbjct: 230 SYSSFEYLKAFTLKQRNKESLTPGESVLCGALAGAISAALTTPLDVVKTRLMTRVGTEGS 289
Query: 739 MSIVAF--SILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAMNK 786
++V ++ EG +GL +G PR A A+ + +E AR A+ K
Sbjct: 290 RTVVGTMREVVAEEGLMGLSRGIGPRVLHSACFAALGYCAFETARLAILK 339
>UniRef100_Q8GYH1 Putative mitochondrial carrier protein [Arabidopsis thaliana]
Length = 364
Score = 131 bits (330), Expect = 7e-29
Identities = 100/339 (29%), Positives = 155/339 (45%), Gaps = 60/339 (17%)
Query: 513 SVEIPAGS----VLRSALAGGLSCALSCALLHPVDSIKTRVQA-----SSMSFPEIIAKL 563
SVEI A V R L GG++ A ++HPVD++KTR+Q+ ++ I+ L
Sbjct: 20 SVEIKATHDQFFVWREFLWGGIAGAFGEGMMHPVDTLKTRLQSQIIMNATQRQKSILQML 79
Query: 564 PEI----GTRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFC 619
+ G +G YRG P + G ++ G E++K + P+L IA
Sbjct: 80 RTVWVGDGLKGFYRGIAPGVTGSLATGATYFGFIESTKKWIEESHPSLAGHWAHFIAGAV 139
Query: 620 STFLGTAVRIPCEVLKQRLQA-------------------------GLFNNVGEALVGTW 654
LG+ + +PCEV+KQR+Q G + + +A W
Sbjct: 140 GDTLGSFIYVPCEVIKQRMQIQGTSSSWSSYISRNSVPVQPRGDMYGYYTGMFQAGCSIW 199
Query: 655 QQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEAW---------ETI 705
++ G KG + G +TL R+VPF GL +G++ L + + + E +
Sbjct: 200 KEQGPKGLYAGYWSTLARDVPF----AGLMVVFYEGLKDLTDQGKKKFPQYGVNSSIEGL 255
Query: 706 AVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMS---IVAFSILRHEGPLGLFKGAVPR 762
+G L+GGL+A +TTP DV+KTR+ QG ++ I R EGP G F+G+VPR
Sbjct: 256 VLGGLAGGLSAYLTTPLDVVKTRLQ-VQGSTIKYKGWLDAVGQIWRKEGPQGFFRGSVPR 314
Query: 763 FFWIAPLGAMNFAGYELAR-----KAMNKNDEAKTGNLE 796
W P A+ F E R K+ N N+ ++E
Sbjct: 315 VMWYLPASALTFMAVEFLRDNFREKSNNNNNVVSNLSIE 353
>UniRef100_Q9C910 Putative mitochondrial carrier protein; 35518-32968 [Arabidopsis
thaliana]
Length = 367
Score = 131 bits (329), Expect = 9e-29
Identities = 100/342 (29%), Positives = 156/342 (45%), Gaps = 63/342 (18%)
Query: 513 SVEIPAGS----VLRSALAGGLSCALSCALLHPVDSIKTRVQA-----SSMSFPEIIAKL 563
SVEI A V R L GG++ A ++HPVD++KTR+Q+ ++ I+ L
Sbjct: 20 SVEIKATHDQFFVWREFLWGGIAGAFGEGMMHPVDTLKTRLQSQIIMNATQRQKSILQML 79
Query: 564 PEI----GTRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFC 619
+ G +G YRG P + G ++ G E++K + P+L IA
Sbjct: 80 RTVWVGDGLKGFYRGIAPGVTGSLATGATYFGFIESTKKWIEESHPSLAGHWAHFIAGAV 139
Query: 620 STFLGTAVRIPCEVLKQRLQA-------------------------GLFNNVGEALVGTW 654
LG+ + +PCEV+KQR+Q G + + +A W
Sbjct: 140 GDTLGSFIYVPCEVIKQRMQIQGTSSSWSSYISRNSVPVQPRGDMYGYYTGMFQAGCSIW 199
Query: 655 QQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEAW---------ETI 705
++ G KG + G +TL R+VPF GL +G++ L + + + E +
Sbjct: 200 KEQGPKGLYAGYWSTLARDVPF----AGLMVVFYEGLKDLTDQGKKKFPQYGVNSSIEGL 255
Query: 706 AVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSI------VAFSILRHEGPLGLFKGA 759
+G L+GGL+A +TTP DV+KTR+ QG ++ + I R EGP G F+G+
Sbjct: 256 VLGGLAGGLSAYLTTPLDVVKTRLQ-VQGSTIKYASYKGWLDAVGQIWRKEGPQGFFRGS 314
Query: 760 VPRFFWIAPLGAMNFAGYELAR-----KAMNKNDEAKTGNLE 796
VPR W P A+ F E R K+ N N+ ++E
Sbjct: 315 VPRVMWYLPASALTFMAVEFLRDNFREKSNNNNNVVSNLSIE 356
>UniRef100_Q6L4L8 Putative mitochondrial carrier protein [Oryza sativa]
Length = 288
Score = 130 bits (328), Expect = 1e-28
Identities = 84/265 (31%), Positives = 135/265 (50%), Gaps = 18/265 (6%)
Query: 526 LAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAKLPEIGTRGLYRGSIPAILGQFSS 585
+AGG + + L+P+D+IKTR+QA+ +I +GLY G I G +
Sbjct: 22 IAGGTAGVVVETALYPIDTIKTRLQAARGG--------SQIQWKGLYSGLAGNIAGVLPA 73
Query: 586 HGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLG----TAVRIPCEVLKQRLQAG 641
+ GI+E +K L+ P + ++A F + +G + +R+P EV+KQR+Q G
Sbjct: 74 SAVFVGIYEPTKRKLLETFPE----NLSAVAHFTAGAIGGIAASLIRVPTEVVKQRMQTG 129
Query: 642 LFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEA 701
F + +A+ ++G +G + G G+ L R++PF +Y + + G + + REL
Sbjct: 130 QFRSAPDAVRLIVGKEGFRGLYAGYGSFLLRDLPFDAIQFCIYEQLRIGYKVVAKRELND 189
Query: 702 WETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIV--AFSILRHEGPLGLFKGA 759
E +GA +G + +TTP DVMKTR+M + IV A +ILR EGP KG
Sbjct: 190 PENALIGAFAGAITGAITTPLDVMKTRLMVQGSANQYSGIVSCAQTILREEGPGAFLKGI 249
Query: 760 VPRFFWIAPLGAMNFAGYELARKAM 784
PR WI G++ F E + +
Sbjct: 250 EPRVLWIGIGGSIFFGVLEKTKSML 274
>UniRef100_Q641C8 MGC82075 protein [Xenopus laevis]
Length = 266
Score = 128 bits (322), Expect = 6e-28
Identities = 87/265 (32%), Positives = 138/265 (51%), Gaps = 12/265 (4%)
Query: 524 SALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAKLPEIGTRGLYRGSIPAILGQF 583
S LAGG + +L P+D+IKTR+Q S + F + G RG+Y G +G F
Sbjct: 9 SLLAGGAAGMSVDLILFPLDTIKTRLQ-SPLGFSK------SGGFRGIYAGVPSTAVGSF 61
Query: 584 SSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLKQRLQAGLF 643
+ +E++K L + + L + + A+F + +R+P EV+KQR Q
Sbjct: 62 PNAAAFFVTYESAKRFLGSDSSYLSPI-IHMAAAFLGELVACLIRVPSEVIKQRAQVSPS 120
Query: 644 NNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEAWE 703
+ + L T +++G+KG +RG +T+ RE+PF + L+ K GR ++ W+
Sbjct: 121 STTYQMLSVTLREEGIKGLYRGYKSTVLREIPFSLVQFPLWEFLKNLWSWKQGRAVDCWQ 180
Query: 704 TIAVGALSGGLAAVVTTPFDVMKTRMMTAQ-GRSVSMSIVAFS---ILRHEGPLGLFKGA 759
+ GA +GG AA VTTP DV KTR+M A+ G V+ V F+ I R +G +GLF G
Sbjct: 181 SAVCGAFAGGFAAAVTTPLDVAKTRIMLAKAGSGVANGNVLFALHEIWRTQGIMGLFAGV 240
Query: 760 VPRFFWIAPLGAMNFAGYELARKAM 784
+PR I+ G + Y+ R ++
Sbjct: 241 IPRMTMISLGGFIFLGAYDKVRSSL 265
>UniRef100_Q6GLA2 MGC69323 protein [Xenopus tropicalis]
Length = 269
Score = 128 bits (321), Expect = 8e-28
Identities = 87/265 (32%), Positives = 138/265 (51%), Gaps = 12/265 (4%)
Query: 524 SALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAKLPEIGTRGLYRGSIPAILGQF 583
S LAGG + +L P+D+IKTR+Q S + F + G RG+Y G +G F
Sbjct: 9 SLLAGGTAGMCVDLILFPLDTIKTRLQ-SPLGFSK------SGGFRGIYAGVPSTAVGSF 61
Query: 584 SSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLKQRLQAGLF 643
+ +E++K +L + + L + + A+ + +R+P EV+KQR Q
Sbjct: 62 PNAAAFFVTYESAKQLLRSDSSYLSPI-IHMAAASLGEVVACLIRVPSEVIKQRAQVSPS 120
Query: 644 NNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEAWE 703
+ + L T +Q+G+KG +RG +T+ RE+PF + L+ K GR +++W+
Sbjct: 121 STTYQMLSATLRQEGIKGLYRGYKSTVLREIPFSLVQFPLWESLKDLWSWKQGRAVDSWQ 180
Query: 704 TIAVGALSGGLAAVVTTPFDVMKTRMMTAQ-GRSVSMSIVAFS---ILRHEGPLGLFKGA 759
+ GA +GG AA +TTP DV KTR+M A+ G V+ V F+ I R +G +GLF G
Sbjct: 181 SAVCGAFAGGFAAALTTPLDVAKTRIMLAKAGSGVASGNVLFALHEIWRTQGIMGLFAGV 240
Query: 760 VPRFFWIAPLGAMNFAGYELARKAM 784
+PR I+ G + Y+ R M
Sbjct: 241 IPRMTAISLGGFIFLGAYDKVRTLM 265
>UniRef100_UPI000023D6B1 UPI000023D6B1 UniRef100 entry
Length = 280
Score = 124 bits (312), Expect = 8e-27
Identities = 96/290 (33%), Positives = 141/290 (48%), Gaps = 28/290 (9%)
Query: 509 AVPPSVEIPAGSVLRSALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAKLPEIGT 568
A PPS + + LAG L+ L P+D++KTR+Q+S+ FP G
Sbjct: 3 ADPPSFQ-------SALLAGALAGTTVDLSLFPLDTLKTRLQSSAGFFPSG-------GF 48
Query: 569 RGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNV----APNLPELQVQSIASFCSTFLG 624
G+YRG A++G +E+ K +L + AP A+
Sbjct: 49 SGIYRGIGSALVGSAPGAAFFFCTYESVKGLLADKDNTSAPGWKAPVTHMAAASAGEIAA 108
Query: 625 TAVRIPCEVLKQRLQAGLFNNVG----EALVGTWQQDGL----KGFFRGTGATLCREVPF 676
AVR+P EV+KQR QAG A++ + G + +RG G T+ REVPF
Sbjct: 109 CAVRVPTEVVKQRAQAGHHGGSSAAALRAILSKYSSHGFVPMWRELYRGWGITVFREVPF 168
Query: 677 YVAGMGLYAESKK-GVQKLLGRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGR 735
V L+ K G ++ GRE+ A E+ G+++GG +A +TTP DV+KTR+M ++
Sbjct: 169 TVIQFPLWEAMKSWGRRRRDGREVTAAESALYGSMAGGFSAALTTPLDVLKTRVMLSK-E 227
Query: 736 SVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAMN 785
SVS+S + I+R EG F G PR WI+ GA+ Y+ A MN
Sbjct: 228 SVSVSRIFSQIMREEGSKAFFAGLAPRVTWISIGGAIFLGSYQWAINTMN 277
>UniRef100_Q94AG6 Putative mitochondrial carrier protein [Arabidopsis thaliana]
Length = 325
Score = 124 bits (312), Expect = 8e-27
Identities = 81/261 (31%), Positives = 125/261 (47%), Gaps = 10/261 (3%)
Query: 526 LAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAKLPEIGTRGLYRGSIPAILGQFSS 585
+AGG + + L+P+D+IKTR+QA+ +I +GLY G I G +
Sbjct: 59 IAGGTAGVVVETALYPIDTIKTRLQAARGG--------GKIVLKGLYSGLAGNIAGVLPA 110
Query: 586 HGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLKQRLQAGLFNN 645
L G++E +K L+ P+ A + +R+P EV+KQR+Q G F +
Sbjct: 111 SALFVGVYEPTKQKLLKTFPDHLSAVAHLTAGAIGGLAASLIRVPTEVVKQRMQTGQFTS 170
Query: 646 VGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEAWETI 705
A+ ++G +G + G + L R++PF +Y + G +K REL E
Sbjct: 171 APSAVRMIASKEGFRGLYAGYRSFLLRDLPFDAIQFCIYEQLCLGYKKAARRELSDPENA 230
Query: 706 AVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIV--AFSILRHEGPLGLFKGAVPRF 763
+GA +G L VTTP DV+KTR+M IV +I+R EG L KG PR
Sbjct: 231 LIGAFAGALTGAVTTPLDVIKTRLMVQGSAKQYQGIVDCVQTIVREEGAPALLKGIGPRV 290
Query: 764 FWIAPLGAMNFAGYELARKAM 784
WI G++ F E ++ +
Sbjct: 291 LWIGIGGSIFFGVLESTKRTL 311
Score = 35.4 bits (80), Expect = 6.7
Identities = 28/89 (31%), Positives = 43/89 (47%), Gaps = 6/89 (6%)
Query: 524 SALAGGLSCALSCALLHPVDSIKTR--VQASSMSFPEII----AKLPEIGTRGLYRGSIP 577
+AL G + AL+ A+ P+D IKTR VQ S+ + I+ + E G L +G P
Sbjct: 229 NALIGAFAGALTGAVTTPLDVIKTRLMVQGSAKQYQGIVDCVQTIVREEGAPALLKGIGP 288
Query: 578 AILGQFSSHGLRTGIFEASKLVLVNVAPN 606
+L + G+ E++K L PN
Sbjct: 289 RVLWIGIGGSIFFGVLESTKRTLAQRRPN 317
>UniRef100_Q6YVE7 Mitochondrial aspartate-glutamate carrier protein-like [Oryza
sativa]
Length = 284
Score = 123 bits (308), Expect = 2e-26
Identities = 78/273 (28%), Positives = 131/273 (47%), Gaps = 10/273 (3%)
Query: 521 VLRSALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAKLPEIGTRGLYRGSIPAIL 580
+ +AGG + + L+P+D+IKTR+QA+ +I +GLY G I
Sbjct: 16 LFEGVIAGGAAGVVVETALYPIDTIKTRLQAAKGG--------SKIQWKGLYAGLGGNIA 67
Query: 581 GQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLKQRLQA 640
G + + G++E +K L+ + P A + +R+P EV+KQR+Q
Sbjct: 68 GVLPASAIFIGVYEPTKRKLLEMFPENLSAVAHLTAGAIGGAASSLIRVPTEVVKQRMQM 127
Query: 641 GLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELE 700
F +A+ +++G KG + G G+ L R++PF +Y + + G + R+L+
Sbjct: 128 SQFKTAPDAVRLIIRKEGFKGLYAGYGSFLLRDLPFDAIQFCIYEQLRIGYKLAAKRDLK 187
Query: 701 AWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIV--AFSILRHEGPLGLFKG 758
E +GA +G + +TTP DV+KTR+M + I+ A +ILR EG KG
Sbjct: 188 DGENALIGAFAGAITGAITTPLDVLKTRLMVQGQANQYRGIISCAQTILREEGAGAFLKG 247
Query: 759 AVPRFFWIAPLGAMNFAGYELARKAMNKNDEAK 791
PR WI G++ F E + + + + K
Sbjct: 248 IEPRVLWIGIGGSIFFGVLEKTKSILAERNSRK 280
>UniRef100_Q9VBN7 CG4743-PA [Drosophila melanogaster]
Length = 297
Score = 122 bits (306), Expect = 4e-26
Identities = 91/288 (31%), Positives = 139/288 (47%), Gaps = 14/288 (4%)
Query: 504 AATVVAVPPSVEIPAGSVLRSALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAKL 563
AA VA+ + + +AGG++ + L P+D++KTR+Q S + F
Sbjct: 10 AAGSVAIKMQEPVNKLKFFHALVAGGVAGMVVDIALFPIDTVKTRLQ-SELGFWRAG--- 65
Query: 564 PEIGTRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFL 623
G RG+Y+G PA G + L +E K L +V V A+ + L
Sbjct: 66 ---GFRGIYKGLAPAAAGSAPTAALFFCTYECGKQFLSSVTQTKDSPYVHMAAASAAEVL 122
Query: 624 GTAVRIPCEVLKQRLQAGLFNNVG--EALVGTWQQDGLK-GFFRGTGATLCREVPFYVAG 680
+R+P E+ KQR Q N + L+ ++ +GLK G +RG G+T+ RE+PF +
Sbjct: 123 ACLIRVPVEIAKQRSQTLQGNKQSGLQILLRAYRTEGLKRGLYRGFGSTIMREIPFSLIQ 182
Query: 681 MGLYAESKKGVQKLLGRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMS 740
L+ K L G + + GA++GG++A +TTP DV+KTR+M A+ S++
Sbjct: 183 FPLWEYFKLQWTPLTGFDSTPFSVALCGAVAGGISAGLTTPLDVVKTRIMLAERESLNRR 242
Query: 741 IVAFSILR----HEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAM 784
A IL G GLF G VPR WI GA F Y+L + +
Sbjct: 243 RSARRILHGIYLERGFSGLFAGFVPRVLWITLGGAFFFGFYDLTTRIL 290
Score = 36.6 bits (83), Expect = 3.0
Identities = 22/94 (23%), Positives = 41/94 (43%), Gaps = 11/94 (11%)
Query: 698 ELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFSILRHEGPLGLFK 757
+L+ + + G ++G + + P D +KTR+ + G R G G++K
Sbjct: 24 KLKFFHALVAGGVAGMVVDIALFPIDTVKTRLQSELG-----------FWRAGGFRGIYK 72
Query: 758 GAVPRFFWIAPLGAMNFAGYELARKAMNKNDEAK 791
G P AP A+ F YE ++ ++ + K
Sbjct: 73 GLAPAAAGSAPTAALFFCTYECGKQFLSSVTQTK 106
>UniRef100_UPI000042F384 UPI000042F384 UniRef100 entry
Length = 307
Score = 120 bits (301), Expect = 2e-25
Identities = 94/293 (32%), Positives = 147/293 (50%), Gaps = 49/293 (16%)
Query: 523 RSALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAKLPEIGTRGLYRGSIPAILGQ 582
R+ ++G +S + P+D++KTR+Q+S+ + G +G+YRG LG
Sbjct: 15 RALISGAISGLSVDFMFFPLDTVKTRIQSSAGFWSSG-------GFKGVYRGVGSVGLGS 67
Query: 583 FSSHGLRTGIFEASKLVLVNVAPNLPELQV--------QSIASFCSTFLGTAVRIPCEVL 634
+EA K LP+ QV +A+ + ++ +R+P EV+
Sbjct: 68 APGASAFFVTYEALK-------KRLPKYQVFANNSSLTHMVAASGAEYVSCLIRVPTEVV 120
Query: 635 KQRLQAGLFNNVGEAL---VGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGV 691
K R QAG + +L + T + +G++GF+RG G TL RE+PF LY K +
Sbjct: 121 KSRTQAGAYGQGKSSLHSAISTMKYEGIRGFYRGFGITLTREIPFTSIQFPLYEFFKSFL 180
Query: 692 QK--LLGRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMM-------TAQGRSV----- 737
+ L G+ ++E G+L+GG+AA TTP DV+KTR+M +A G +V
Sbjct: 181 SQHYLGGKRPTSYEAALCGSLAGGIAAACTTPLDVVKTRVMLEARVSASASGANVVNDVL 240
Query: 738 -----SMSIVAF-----SILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELA 780
S S+++F +ILR EGP LF+G VPR F I+ GA+ Y+LA
Sbjct: 241 PPKQPSPSVLSFPPRLLNILRTEGPTALFRGWVPRTFAISMGGAVFLGIYDLA 293
>UniRef100_Q6ESH9 Mitochondrial substrate carrier protein-like [Oryza sativa]
Length = 618
Score = 119 bits (298), Expect = 4e-25
Identities = 87/283 (30%), Positives = 131/283 (45%), Gaps = 11/283 (3%)
Query: 523 RSALAGGLSCALSCALLHPVDSIKTRVQASSM---SFPEIIAK-LPEIGTRGLYRGSIPA 578
R A+AG L+ + LHP+D++KT +Q +S SF + + L E G GLY G
Sbjct: 335 RHAVAGALAGTVVSVSLHPIDTVKTIIQVNSSRRSSFYHTLRRALVERGVLGLYGGLASK 394
Query: 579 ILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLKQRL 638
I + T +E K L+ + P A CS+ + V P E +KQ++
Sbjct: 395 IACSAPISAIYTLTYEIVKGSLLPILPKEYHSIAHCTAGGCSSIATSFVFTPSECIKQQM 454
Query: 639 QAGL-FNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLL-- 695
Q G + N +AL+G ++ G+ + G GA LCR +P V Y K+ + K
Sbjct: 455 QVGSQYQNCWDALLGCLRKGGITSLYAGWGAVLCRNIPHSVIKFYTYESLKQFMLKSAPA 514
Query: 696 GRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVS----MSIVAFSILRHEG 751
L++ +T+ G +G AA+ TTPFDV+KTR+ +S + I +HEG
Sbjct: 515 NANLDSGQTLFCGGFAGSTAALCTTPFDVVKTRVQLQALSPISKYDGVLHALKEIFQHEG 574
Query: 752 PLGLFKGAVPRFFWIAPLGAMNFAGYELARKAMNKNDEAKTGN 794
GL++G PR GA+ F YE + M E N
Sbjct: 575 LQGLYRGLAPRLAMYISQGAIFFTSYEFLKTIMFSEQELHARN 617
>UniRef100_Q8MR46 GH25190p [Drosophila melanogaster]
Length = 520
Score = 118 bits (296), Expect = 6e-25
Identities = 97/398 (24%), Positives = 188/398 (46%), Gaps = 17/398 (4%)
Query: 356 ESEGRRFFEELDRDGDGQVTLEDLEIAMRRRKLPRRYAKEFMSRTRSHLFSRSFGWKQFL 415
E R F +LDRDGDG++ + DL A+ L YA++F+ ++ S + G+ +FL
Sbjct: 114 EERLERIFNKLDRDGDGRIDIHDLSAALHEFGLSSVYAEKFLQQSDKDQ-SGNVGFAEFL 172
Query: 416 SFMEQKEPTILRAYTSLCLTKSGTLKKSEILESLKNSGLPANEDNAAAMMRFLNADTEES 475
++ + E ++ ++ L + G + E++ + K+ GL + D A ++ ++ D +
Sbjct: 173 HYVREHEKNLVLQFSHLDKNRDGKVDLEELISAFKDLGLDIDMDEARNLLTRMDKDGSLN 232
Query: 476 ISYGHFRNFMLLLPS----DRLQEDPRSIWFEAATVVAVPPSV---EIPAGSVLRSALAG 528
IS+ +R+FMLL PS D ++ S + + + VP E+ G R +AG
Sbjct: 233 ISFNEWRDFMLLAPSTDIHDLIKFWRHSTYLDIGEDMNVPDDFTQKEMQTGLWWRHLVAG 292
Query: 529 GLSCALSCALLHPVDSIKT--RVQASSMSFPEII-AKLPEIGTRGLYRGSIPAILGQFSS 585
G++ A+S P+D IK +VQ M E + L E G+R ++RG+ +L
Sbjct: 293 GIAGAVSRTCTAPLDRIKVYLQVQTQRMGISECMHIMLNEGGSRSMWRGNGINVLKIAPE 352
Query: 586 HGLRTGIFEASK-LVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLKQRL---QAG 641
+ +E K L+ + + + A + + + P EVLK RL + G
Sbjct: 353 TAFKFAAYEQMKRLIRGDDGSRQMSIVERFYAGAAAGGISQTIIYPMEVLKTRLALRRTG 412
Query: 642 LFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKG-VQKLLGRELE 700
+ + +A V ++Q+G++ F+RG + +P+ + +Y K+ + E
Sbjct: 413 QYAGIADAAVKIYKQEGVRSFYRGYVPNILGILPYAGIDLAVYETLKRRYIANHDNNEQP 472
Query: 701 AW-ETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSV 737
++ +A G+ S L + + P +++TR+ + +
Sbjct: 473 SFLVLLACGSTSSTLGQLCSYPLALVRTRLQAQAAKQL 510
Score = 56.2 bits (134), Expect = 4e-06
Identities = 37/165 (22%), Positives = 74/165 (44%), Gaps = 8/165 (4%)
Query: 627 VRIPCEVLKQRLQAGLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAE 686
+++ +V QR+ + E + + G + +RG G + + P Y +
Sbjct: 309 IKVYLQVQTQRM------GISECMHIMLNEGGSRSMWRGNGINVLKIAPETAFKFAAYEQ 362
Query: 687 SKKGVQKLLG-RELEAWETIAVGALSGGLAAVVTTPFDVMKTRM-MTAQGRSVSMSIVAF 744
K+ ++ G R++ E GA +GG++ + P +V+KTR+ + G+ ++ A
Sbjct: 363 MKRLIRGDDGSRQMSIVERFYAGAAAGGISQTIIYPMEVLKTRLALRRTGQYAGIADAAV 422
Query: 745 SILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAMNKNDE 789
I + EG ++G VP I P ++ A YE ++ N +
Sbjct: 423 KIYKQEGVRSFYRGYVPNILGILPYAGIDLAVYETLKRRYIANHD 467
Score = 37.0 bits (84), Expect = 2.3
Identities = 21/90 (23%), Positives = 41/90 (45%), Gaps = 1/90 (1%)
Query: 702 WETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFSILRHEGPLGLFKGAVP 761
W + G ++G ++ T P D +K + Q + + +S +L G +++G
Sbjct: 286 WRHLVAGGIAGAVSRTCTAPLDRIKVYLQV-QTQRMGISECMHIMLNEGGSRSMWRGNGI 344
Query: 762 RFFWIAPLGAMNFAGYELARKAMNKNDEAK 791
IAP A FA YE ++ + +D ++
Sbjct: 345 NVLKIAPETAFKFAAYEQMKRLIRGDDGSR 374
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.318 0.134 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,274,742,021
Number of Sequences: 2790947
Number of extensions: 53640618
Number of successful extensions: 149657
Number of sequences better than 10.0: 1926
Number of HSP's better than 10.0 without gapping: 496
Number of HSP's successfully gapped in prelim test: 1433
Number of HSP's that attempted gapping in prelim test: 139398
Number of HSP's gapped (non-prelim): 5459
length of query: 796
length of database: 848,049,833
effective HSP length: 136
effective length of query: 660
effective length of database: 468,481,041
effective search space: 309197487060
effective search space used: 309197487060
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 79 (35.0 bits)
Medicago: description of AC144484.11


