

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144477.5 - phase: 1 /pseudo
(206 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
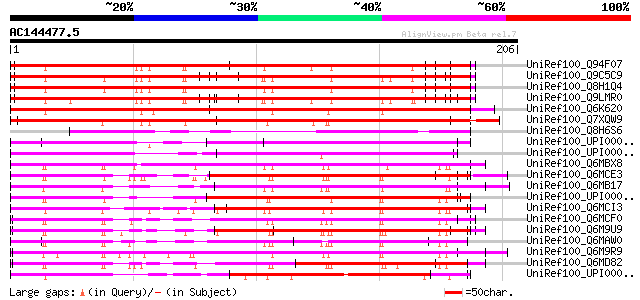
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q94F07 Hypothetical protein F7H2.8 [Arabidopsis thaliana] 191 8e-48
UniRef100_Q9C5C9 Hypothetical protein At1g15740 [Arabidopsis tha... 191 8e-48
UniRef100_Q8H1Q4 Hypothetical protein At1g15740 [Arabidopsis tha... 191 8e-48
UniRef100_Q9LMR0 F7H2.8 protein [Arabidopsis thaliana] 184 1e-45
UniRef100_Q6K620 Leucine-rich repeat-like protein [Oryza sativa] 184 1e-45
UniRef100_Q7XQW9 OSJNBa0014K14.6 protein [Oryza sativa] 182 5e-45
UniRef100_Q8H6S6 CTV.1 [Poncirus trifoliata] 96 6e-19
UniRef100_UPI00002BE69C UPI00002BE69C UniRef100 entry 91 2e-17
UniRef100_UPI00003282C4 UPI00003282C4 UniRef100 entry 87 3e-16
UniRef100_Q6MBX8 Hypothetical protein [Parachlamydia sp.] 83 4e-15
UniRef100_Q6MCE3 Hypothetical protein [Parachlamydia sp.] 79 7e-14
UniRef100_Q6MB17 Hypothetical protein [Parachlamydia sp.] 79 7e-14
UniRef100_UPI000033B186 UPI000033B186 UniRef100 entry 78 2e-13
UniRef100_Q6MCI3 Hypothetical protein [Parachlamydia sp.] 77 2e-13
UniRef100_Q6MCF0 Hypothetical protein [Parachlamydia sp.] 77 4e-13
UniRef100_Q6M9U9 Hypothetical protein [Parachlamydia sp.] 76 5e-13
UniRef100_Q6MAW0 Hypothetical protein [Parachlamydia sp.] 76 6e-13
UniRef100_Q6M9R9 Hypothetical protein [Parachlamydia sp.] 75 8e-13
UniRef100_Q6MD82 Hypothetical protein [Parachlamydia sp.] 75 1e-12
UniRef100_UPI00003634F6 UPI00003634F6 UniRef100 entry 75 1e-12
>UniRef100_Q94F07 Hypothetical protein F7H2.8 [Arabidopsis thaliana]
Length = 332
Score = 191 bits (486), Expect = 8e-48
Identities = 109/192 (56%), Positives = 144/192 (74%), Gaps = 5/192 (2%)
Query: 1 LQKLSTLNVEGCS-ITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPF-LQV 58
L KL+ LN+EGC +TAAC + ++ALA L LNLNRC SD G EKFS L ++ L +
Sbjct: 31 LNKLNLLNLEGCRHVTAACLDTLTALAGLMYLNLNRCNFSDSGCEKFSDLINLKILNLGM 90
Query: 59 LQA*RG*VLHL---TRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYIS 115
++HL T+L++ +IGDEGLV+L+G+ LKSL LSDTEVG++G+R++S
Sbjct: 91 NNITNSCLVHLKGLTKLESLNLDSCRIGDEGLVHLSGMLELKSLELSDTEVGSNGLRHLS 150
Query: 116 GLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFG 175
GL+ LE +NLSFT VTD+GL++L GLT+L++LNLDAR +TDAGL+ LTSL+GL LDLFG
Sbjct: 151 GLSNLESINLSFTVVTDSGLRKLSGLTSLRTLNLDARHVTDAGLSALTSLTGLTHLDLFG 210
Query: 176 ARITDSGTTYLR 187
ARITDSGT +LR
Sbjct: 211 ARITDSGTNHLR 222
Score = 70.9 bits (172), Expect = 2e-11
Identities = 53/177 (29%), Positives = 86/177 (47%), Gaps = 4/177 (2%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPF-LQVL 59
L L LN+ +IT +C ++ L L LNL+ C + D+G S + + L
Sbjct: 80 LINLKILNLGMNNITNSCLVHLKGLTKLESLNLDSCRIGDEGLVHLSGMLELKSLELSDT 139
Query: 60 QA*RG*VLHLT---RLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISG 116
+ + HL+ L++ + D GL L+GLT L++L L V ++G+ ++
Sbjct: 140 EVGSNGLRHLSGLSNLESINLSFTVVTDSGLRKLSGLTSLRTLNLDARHVTDAGLSALTS 199
Query: 117 LNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDL 173
L L L+L +TD+G L L L+SL + +TD G+ N+ LS L L+L
Sbjct: 200 LTGLTHLDLFGARITDSGTNHLRNLKKLQSLEICGGGLTDTGVKNIKDLSSLTLLNL 256
Score = 60.5 bits (145), Expect = 3e-08
Identities = 52/182 (28%), Positives = 89/182 (48%), Gaps = 5/182 (2%)
Query: 3 KLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHM----FPFLQV 58
+L +L + + + ++S L+ L +NL+ ++D G K S L + V
Sbjct: 130 ELKSLELSDTEVGSNGLRHLSGLSNLESINLSFTVVTDSGLRKLSGLTSLRTLNLDARHV 189
Query: 59 LQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLN 118
A + LT L F +I D G +L L L+SL + + ++G++ I L+
Sbjct: 190 TDAGLSALTSLTGLTHLDLFGARITDSGTNHLRNLKKLQSLEICGGGLTDTGVKNIKDLS 249
Query: 119 KLEDLNLSFTS-VTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGAR 177
L LNLS S +TD L+ + GLT L SLN+ +++ +GL +L L L +L L +
Sbjct: 250 SLTLLNLSQNSNLTDKTLELISGLTGLVSLNVSNSRVSSSGLRHLKPLKNLRSLTLESCK 309
Query: 178 IT 179
++
Sbjct: 310 LS 311
Score = 53.9 bits (128), Expect = 3e-06
Identities = 40/125 (32%), Positives = 61/125 (48%), Gaps = 25/125 (20%)
Query: 90 LTGLTLLKSLVLSDTEVGNSGIRYISGLNKLE-------------------------DLN 124
L+ LT L+SL + +++ + GI Y+ GLNKL LN
Sbjct: 4 LSVLTNLRSLQICCSKITDIGISYLKGLNKLNLLNLEGCRHVTAACLDTLTALAGLMYLN 63
Query: 125 LSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDSGTT 184
L+ + +D+G ++ L NLK LNL IT++ L +L L+ L +L+L RI D G
Sbjct: 64 LNRCNFSDSGCEKFSDLINLKILNLGMNNITNSCLVHLKGLTKLESLNLDSCRIGDEGLV 123
Query: 185 YLRCM 189
+L M
Sbjct: 124 HLSGM 128
Score = 51.2 bits (121), Expect = 2e-05
Identities = 48/176 (27%), Positives = 89/176 (50%), Gaps = 7/176 (3%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQ---HMFPF-L 56
L L ++N+ +T + +S L +L LNL+ ++D G + L H+ F
Sbjct: 152 LSNLESINLSFTVVTDSGLRKLSGLTSLRTLNLDARHVTDAGLSALTSLTGLTHLDLFGA 211
Query: 57 QVLQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLS-DTEVGNSGIRYIS 115
++ + + +L +LQ+ + D G+ N+ L+ L L LS ++ + + + IS
Sbjct: 212 RITDSGTNHLRNLKKLQSLEICGGGLTDTGVKNIKDLSSLTLLNLSQNSNLTDKTLELIS 271
Query: 116 GLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANL--TSLSGLI 169
GL L LN+S + V+ +GL+ L L NL+SL L++ +++ + L T L L+
Sbjct: 272 GLTGLVSLNVSNSRVSSSGLRHLKPLKNLRSLTLESCKLSANDIRKLQATDLPNLV 327
>UniRef100_Q9C5C9 Hypothetical protein At1g15740 [Arabidopsis thaliana]
Length = 585
Score = 191 bits (486), Expect = 8e-48
Identities = 109/192 (56%), Positives = 144/192 (74%), Gaps = 5/192 (2%)
Query: 1 LQKLSTLNVEGCS-ITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPF-LQV 58
L KL+ LN+EGC +TAAC + ++ALA L LNLNRC SD G EKFS L ++ L +
Sbjct: 284 LNKLNLLNLEGCRHVTAACLDTLTALAGLMYLNLNRCNFSDSGCEKFSDLINLKILNLGM 343
Query: 59 LQA*RG*VLHL---TRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYIS 115
++HL T+L++ +IGDEGLV+L+G+ LKSL LSDTEVG++G+R++S
Sbjct: 344 NNITNSCLVHLKGLTKLESLNLDSCRIGDEGLVHLSGMLELKSLELSDTEVGSNGLRHLS 403
Query: 116 GLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFG 175
GL+ LE +NLSFT VTD+GL++L GLT+L++LNLDAR +TDAGL+ LTSL+GL LDLFG
Sbjct: 404 GLSNLESINLSFTVVTDSGLRKLSGLTSLRTLNLDARHVTDAGLSALTSLTGLTHLDLFG 463
Query: 176 ARITDSGTTYLR 187
ARITDSGT +LR
Sbjct: 464 ARITDSGTNHLR 475
Score = 70.9 bits (172), Expect = 2e-11
Identities = 53/177 (29%), Positives = 86/177 (47%), Gaps = 4/177 (2%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPF-LQVL 59
L L LN+ +IT +C ++ L L LNL+ C + D+G S + + L
Sbjct: 333 LINLKILNLGMNNITNSCLVHLKGLTKLESLNLDSCRIGDEGLVHLSGMLELKSLELSDT 392
Query: 60 QA*RG*VLHLT---RLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISG 116
+ + HL+ L++ + D GL L+GLT L++L L V ++G+ ++
Sbjct: 393 EVGSNGLRHLSGLSNLESINLSFTVVTDSGLRKLSGLTSLRTLNLDARHVTDAGLSALTS 452
Query: 117 LNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDL 173
L L L+L +TD+G L L L+SL + +TD G+ N+ LS L L+L
Sbjct: 453 LTGLTHLDLFGARITDSGTNHLRNLKKLQSLEICGGGLTDTGVKNIKDLSSLTLLNL 509
Score = 60.5 bits (145), Expect = 3e-08
Identities = 52/182 (28%), Positives = 89/182 (48%), Gaps = 5/182 (2%)
Query: 3 KLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHM----FPFLQV 58
+L +L + + + ++S L+ L +NL+ ++D G K S L + V
Sbjct: 383 ELKSLELSDTEVGSNGLRHLSGLSNLESINLSFTVVTDSGLRKLSGLTSLRTLNLDARHV 442
Query: 59 LQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLN 118
A + LT L F +I D G +L L L+SL + + ++G++ I L+
Sbjct: 443 TDAGLSALTSLTGLTHLDLFGARITDSGTNHLRNLKKLQSLEICGGGLTDTGVKNIKDLS 502
Query: 119 KLEDLNLSFTS-VTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGAR 177
L LNLS S +TD L+ + GLT L SLN+ +++ +GL +L L L +L L +
Sbjct: 503 SLTLLNLSQNSNLTDKTLELISGLTGLVSLNVSNSRVSSSGLRHLKPLKNLRSLTLESCK 562
Query: 178 IT 179
++
Sbjct: 563 LS 564
Score = 59.3 bits (142), Expect = 6e-08
Identities = 42/112 (37%), Positives = 63/112 (55%), Gaps = 2/112 (1%)
Query: 78 FI*KIGDEGLVNLTGLTLLKSLVL-SDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLK 136
F +I + GLV+L+GL+ L SL + + G+R +S L L+ L+L D GL
Sbjct: 171 FCDQISNRGLVHLSGLSNLTSLSFRRNAAITAQGMRALSNLVNLKKLDLEKCPGIDGGLV 230
Query: 137 RLLGLTNLKSLNLD-ARQITDAGLANLTSLSGLITLDLFGARITDSGTTYLR 187
L LT L+SLN+ ITDA + L+ L+ L L + ++ITD G +YL+
Sbjct: 231 HLRALTKLESLNIKWCNCITDADMEPLSVLTNLRRLQICCSKITDIGISYLK 282
Score = 57.4 bits (137), Expect = 2e-07
Identities = 59/191 (30%), Positives = 85/191 (43%), Gaps = 22/191 (11%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCG-LSDDGFEKFSVLQHMFPFLQVL 59
L L L++E C ++ AL L LN+ C ++D E SVL +
Sbjct: 211 LVNLKKLDLEKCPGIDGGLVHLRALTKLESLNIKWCNCITDADMEPLSVLTN-------- 262
Query: 60 QA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTE-VGNSGIRYISGLN 118
L RLQ KI D G+ L GL L L L V + + ++ L
Sbjct: 263 ---------LRRLQICCS---KITDIGISYLKGLNKLNLLNLEGCRHVTAACLDTLTALA 310
Query: 119 KLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARI 178
L LNL+ + +D+G ++ L NLK LNL IT++ L +L L+ L +L+L RI
Sbjct: 311 GLMYLNLNRCNFSDSGCEKFSDLINLKILNLGMNNITNSCLVHLKGLTKLESLNLDSCRI 370
Query: 179 TDSGTTYLRCM 189
D G +L M
Sbjct: 371 GDEGLVHLSGM 381
Score = 52.8 bits (125), Expect = 6e-06
Identities = 34/94 (36%), Positives = 56/94 (59%), Gaps = 2/94 (2%)
Query: 82 IGDEGLVNLTGLTLLKSLVLSDTE-VGNSGIRYISGLNKLEDLNLSFTS-VTDNGLKRLL 139
I D GLV+L G T L+SL + + + N G+ ++SGL+ L L+ + +T G++ L
Sbjct: 150 ITDSGLVSLKGCTNLESLNFNFCDQISNRGLVHLSGLSNLTSLSFRRNAAITAQGMRALS 209
Query: 140 GLTNLKSLNLDARQITDAGLANLTSLSGLITLDL 173
L NLK L+L+ D GL +L +L+ L +L++
Sbjct: 210 NLVNLKKLDLEKCPGIDGGLVHLRALTKLESLNI 243
Score = 51.2 bits (121), Expect = 2e-05
Identities = 33/94 (35%), Positives = 53/94 (56%), Gaps = 1/94 (1%)
Query: 85 EGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTS-VTDNGLKRLLGLTN 143
+G+ L+ L LK L L + G+ ++ L KLE LN+ + + +TD ++ L LTN
Sbjct: 203 QGMRALSNLVNLKKLDLEKCPGIDGGLVHLRALTKLESLNIKWCNCITDADMEPLSVLTN 262
Query: 144 LKSLNLDARQITDAGLANLTSLSGLITLDLFGAR 177
L+ L + +ITD G++ L L+ L L+L G R
Sbjct: 263 LRRLQICCSKITDIGISYLKGLNKLNLLNLEGCR 296
Score = 51.2 bits (121), Expect = 2e-05
Identities = 48/176 (27%), Positives = 89/176 (50%), Gaps = 7/176 (3%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQ---HMFPF-L 56
L L ++N+ +T + +S L +L LNL+ ++D G + L H+ F
Sbjct: 405 LSNLESINLSFTVVTDSGLRKLSGLTSLRTLNLDARHVTDAGLSALTSLTGLTHLDLFGA 464
Query: 57 QVLQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLS-DTEVGNSGIRYIS 115
++ + + +L +LQ+ + D G+ N+ L+ L L LS ++ + + + IS
Sbjct: 465 RITDSGTNHLRNLKKLQSLEICGGGLTDTGVKNIKDLSSLTLLNLSQNSNLTDKTLELIS 524
Query: 116 GLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANL--TSLSGLI 169
GL L LN+S + V+ +GL+ L L NL+SL L++ +++ + L T L L+
Sbjct: 525 GLTGLVSLNVSNSRVSSSGLRHLKPLKNLRSLTLESCKLSANDIRKLQATDLPNLV 580
Score = 48.9 bits (115), Expect = 8e-05
Identities = 34/98 (34%), Positives = 52/98 (52%), Gaps = 2/98 (2%)
Query: 94 TLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTS-VTDNGLKRLLGLTNLKSLNLDAR 152
T L S+ S +++ +SG+ + G LE LN +F +++ GL L GL+NL SL+
Sbjct: 138 TSLLSVDFSGSDITDSGLVSLKGCTNLESLNFNFCDQISNRGLVHLSGLSNLTSLSFRRN 197
Query: 153 Q-ITDAGLANLTSLSGLITLDLFGARITDSGTTYLRCM 189
IT G+ L++L L LDL D G +LR +
Sbjct: 198 AAITAQGMRALSNLVNLKKLDLEKCPGIDGGLVHLRAL 235
>UniRef100_Q8H1Q4 Hypothetical protein At1g15740 [Arabidopsis thaliana]
Length = 585
Score = 191 bits (486), Expect = 8e-48
Identities = 109/192 (56%), Positives = 144/192 (74%), Gaps = 5/192 (2%)
Query: 1 LQKLSTLNVEGCS-ITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPF-LQV 58
L KL+ LN+EGC +TAAC + ++ALA L LNLNRC SD G EKFS L ++ L +
Sbjct: 284 LNKLNLLNLEGCRHVTAACLDTLTALAGLMYLNLNRCNFSDSGCEKFSDLINLKILNLGM 343
Query: 59 LQA*RG*VLHL---TRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYIS 115
++HL T+L++ +IGDEGLV+L+G+ LKSL LSDTEVG++G+R++S
Sbjct: 344 NNITNSCLVHLKGLTKLESLNLDSCRIGDEGLVHLSGMLELKSLELSDTEVGSNGLRHLS 403
Query: 116 GLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFG 175
GL+ LE +NLSFT VTD+GL++L GLT+L++LNLDAR +TDAGL+ LTSL+GL LDLFG
Sbjct: 404 GLSNLESINLSFTVVTDSGLRKLSGLTSLRTLNLDARHVTDAGLSALTSLTGLTHLDLFG 463
Query: 176 ARITDSGTTYLR 187
ARITDSGT +LR
Sbjct: 464 ARITDSGTNHLR 475
Score = 70.9 bits (172), Expect = 2e-11
Identities = 53/177 (29%), Positives = 86/177 (47%), Gaps = 4/177 (2%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPF-LQVL 59
L L LN+ +IT +C ++ L L LNL+ C + D+G S + + L
Sbjct: 333 LINLKILNLGMNNITNSCLVHLKGLTKLESLNLDSCRIGDEGLVHLSGMLELKSLELSDT 392
Query: 60 QA*RG*VLHLT---RLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISG 116
+ + HL+ L++ + D GL L+GLT L++L L V ++G+ ++
Sbjct: 393 EVGSNGLRHLSGLSNLESINLSFTVVTDSGLRKLSGLTSLRTLNLDARHVTDAGLSALTS 452
Query: 117 LNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDL 173
L L L+L +TD+G L L L+SL + +TD G+ N+ LS L L+L
Sbjct: 453 LTGLTHLDLFGARITDSGTNHLRNLKKLQSLEICGGGLTDTGVKNIKDLSSLTLLNL 509
Score = 60.5 bits (145), Expect = 3e-08
Identities = 52/182 (28%), Positives = 89/182 (48%), Gaps = 5/182 (2%)
Query: 3 KLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHM----FPFLQV 58
+L +L + + + ++S L+ L +NL+ ++D G K S L + V
Sbjct: 383 ELKSLELSDTEVGSNGLRHLSGLSNLESINLSFTVVTDSGLRKLSGLTSLRTLNLDARHV 442
Query: 59 LQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLN 118
A + LT L F +I D G +L L L+SL + + ++G++ I L+
Sbjct: 443 TDAGLSALTSLTGLTHLDLFGARITDSGTNHLRNLKKLQSLEICGGGLTDTGVKNIKDLS 502
Query: 119 KLEDLNLSFTS-VTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGAR 177
L LNLS S +TD L+ + GLT L SLN+ +++ +GL +L L L +L L +
Sbjct: 503 SLTLLNLSQNSNLTDKTLELISGLTGLVSLNVSNSRVSSSGLRHLKPLKNLRSLTLESCK 562
Query: 178 IT 179
++
Sbjct: 563 LS 564
Score = 56.6 bits (135), Expect = 4e-07
Identities = 57/191 (29%), Positives = 86/191 (44%), Gaps = 22/191 (11%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCG-LSDDGFEKFSVLQHMFPFLQVL 59
L L L++E C ++ AL L LN+ C ++D E SVL ++ LQ+
Sbjct: 211 LVNLKKLDLEKCPGIDGGLVHLRALTKLESLNIKWCNCITDADMEPLSVLTNLRS-LQIC 269
Query: 60 QA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTE-VGNSGIRYISGLN 118
+ KI D G+ L GL L L L V + + ++ L
Sbjct: 270 CS-------------------KITDIGISYLKGLNKLNLLNLEGCRHVTAACLDTLTALA 310
Query: 119 KLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARI 178
L LNL+ + +D+G ++ L NLK LNL IT++ L +L L+ L +L+L RI
Sbjct: 311 GLMYLNLNRCNFSDSGCEKFSDLINLKILNLGMNNITNSCLVHLKGLTKLESLNLDSCRI 370
Query: 179 TDSGTTYLRCM 189
D G +L M
Sbjct: 371 GDEGLVHLSGM 381
Score = 51.2 bits (121), Expect = 2e-05
Identities = 48/176 (27%), Positives = 89/176 (50%), Gaps = 7/176 (3%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQ---HMFPF-L 56
L L ++N+ +T + +S L +L LNL+ ++D G + L H+ F
Sbjct: 405 LSNLESINLSFTVVTDSGLRKLSGLTSLRTLNLDARHVTDAGLSALTSLTGLTHLDLFGA 464
Query: 57 QVLQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLS-DTEVGNSGIRYIS 115
++ + + +L +LQ+ + D G+ N+ L+ L L LS ++ + + + IS
Sbjct: 465 RITDSGTNHLRNLKKLQSLEICGGGLTDTGVKNIKDLSSLTLLNLSQNSNLTDKTLELIS 524
Query: 116 GLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANL--TSLSGLI 169
GL L LN+S + V+ +GL+ L L NL+SL L++ +++ + L T L L+
Sbjct: 525 GLTGLVSLNVSNSRVSSSGLRHLKPLKNLRSLTLESCKLSANDIRKLQATDLPNLV 580
Score = 40.8 bits (94), Expect = 0.022
Identities = 27/65 (41%), Positives = 39/65 (59%), Gaps = 2/65 (3%)
Query: 120 LEDLNLSFTSVTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANLTSLSGLITLDL-FGAR 177
L ++ S + +TD+GL L G TNL+SLN + QI++ GL +L+ LS L +L A
Sbjct: 140 LLSVDFSGSDITDSGLVSLKGCTNLESLNFNFCDQISNRGLVHLSGLSNLTSLSFRRNAA 199
Query: 178 ITDSG 182
IT G
Sbjct: 200 ITAQG 204
>UniRef100_Q9LMR0 F7H2.8 protein [Arabidopsis thaliana]
Length = 568
Score = 184 bits (468), Expect = 1e-45
Identities = 109/199 (54%), Positives = 144/199 (71%), Gaps = 12/199 (6%)
Query: 1 LQKLSTLNVEGCS-ITAACFEYIS-------ALAALACLNLNRCGLSDDGFEKFSVLQHM 52
L KL+ LN+EGC +TAAC + ++ ALA L LNLNRC SD G EKFS L ++
Sbjct: 260 LNKLNLLNLEGCRHVTAACLDTLTGLYRHPHALAGLMYLNLNRCNFSDSGCEKFSDLINL 319
Query: 53 FPF-LQVLQA*RG*VLHL---TRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGN 108
L + ++HL T+L++ +IGDEGLV+L+G+ LKSL LSDTEVG+
Sbjct: 320 KILNLGMNNITNSCLVHLKGLTKLESLNLDSCRIGDEGLVHLSGMLELKSLELSDTEVGS 379
Query: 109 SGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGL 168
+G+R++SGL+ LE +NLSFT VTD+GL++L GLT+L++LNLDAR +TDAGL+ LTSL+GL
Sbjct: 380 NGLRHLSGLSNLESINLSFTVVTDSGLRKLSGLTSLRTLNLDARHVTDAGLSALTSLTGL 439
Query: 169 ITLDLFGARITDSGTTYLR 187
LDLFGARITDSGT +LR
Sbjct: 440 THLDLFGARITDSGTNHLR 458
Score = 70.9 bits (172), Expect = 2e-11
Identities = 53/177 (29%), Positives = 86/177 (47%), Gaps = 4/177 (2%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPF-LQVL 59
L L LN+ +IT +C ++ L L LNL+ C + D+G S + + L
Sbjct: 316 LINLKILNLGMNNITNSCLVHLKGLTKLESLNLDSCRIGDEGLVHLSGMLELKSLELSDT 375
Query: 60 QA*RG*VLHLT---RLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISG 116
+ + HL+ L++ + D GL L+GLT L++L L V ++G+ ++
Sbjct: 376 EVGSNGLRHLSGLSNLESINLSFTVVTDSGLRKLSGLTSLRTLNLDARHVTDAGLSALTS 435
Query: 117 LNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDL 173
L L L+L +TD+G L L L+SL + +TD G+ N+ LS L L+L
Sbjct: 436 LTGLTHLDLFGARITDSGTNHLRNLKKLQSLEICGGGLTDTGVKNIKDLSSLTLLNL 492
Score = 60.5 bits (145), Expect = 3e-08
Identities = 52/182 (28%), Positives = 89/182 (48%), Gaps = 5/182 (2%)
Query: 3 KLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHM----FPFLQV 58
+L +L + + + ++S L+ L +NL+ ++D G K S L + V
Sbjct: 366 ELKSLELSDTEVGSNGLRHLSGLSNLESINLSFTVVTDSGLRKLSGLTSLRTLNLDARHV 425
Query: 59 LQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLN 118
A + LT L F +I D G +L L L+SL + + ++G++ I L+
Sbjct: 426 TDAGLSALTSLTGLTHLDLFGARITDSGTNHLRNLKKLQSLEICGGGLTDTGVKNIKDLS 485
Query: 119 KLEDLNLSFTS-VTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGAR 177
L LNLS S +TD L+ + GLT L SLN+ +++ +GL +L L L +L L +
Sbjct: 486 SLTLLNLSQNSNLTDKTLELISGLTGLVSLNVSNSRVSSSGLRHLKPLKNLRSLTLESCK 545
Query: 178 IT 179
++
Sbjct: 546 LS 547
Score = 52.8 bits (125), Expect = 6e-06
Identities = 34/94 (36%), Positives = 56/94 (59%), Gaps = 2/94 (2%)
Query: 82 IGDEGLVNLTGLTLLKSLVLSDTE-VGNSGIRYISGLNKLEDLNLSFTS-VTDNGLKRLL 139
I D GLV+L G T L+SL + + + N G+ ++SGL+ L L+ + +T G++ L
Sbjct: 150 ITDSGLVSLKGCTNLESLNFNFCDQISNRGLVHLSGLSNLTSLSFRRNAAITAQGMRALS 209
Query: 140 GLTNLKSLNLDARQITDAGLANLTSLSGLITLDL 173
L NLK L+L+ D GL +L +L+ L +L++
Sbjct: 210 NLVNLKKLDLEKCPGIDGGLVHLRALTKLESLNI 243
Score = 52.0 bits (123), Expect = 1e-05
Identities = 39/102 (38%), Positives = 56/102 (54%), Gaps = 2/102 (1%)
Query: 78 FI*KIGDEGLVNLTGLTLLKSLVL-SDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLK 136
F +I + GLV+L+GL+ L SL + + G+R +S L L+ L+L D GL
Sbjct: 171 FCDQISNRGLVHLSGLSNLTSLSFRRNAAITAQGMRALSNLVNLKKLDLEKCPGIDGGLV 230
Query: 137 RLLGLTNLKSLNLD-ARQITDAGLANLTSLSGLITLDLFGAR 177
L LT L+SLN+ ITDA + L+ L+ L L+L G R
Sbjct: 231 HLRALTKLESLNIKWCNCITDADMEPLSGLNKLNLLNLEGCR 272
Score = 51.2 bits (121), Expect = 2e-05
Identities = 48/176 (27%), Positives = 89/176 (50%), Gaps = 7/176 (3%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQ---HMFPF-L 56
L L ++N+ +T + +S L +L LNL+ ++D G + L H+ F
Sbjct: 388 LSNLESINLSFTVVTDSGLRKLSGLTSLRTLNLDARHVTDAGLSALTSLTGLTHLDLFGA 447
Query: 57 QVLQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLS-DTEVGNSGIRYIS 115
++ + + +L +LQ+ + D G+ N+ L+ L L LS ++ + + + IS
Sbjct: 448 RITDSGTNHLRNLKKLQSLEICGGGLTDTGVKNIKDLSSLTLLNLSQNSNLTDKTLELIS 507
Query: 116 GLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANL--TSLSGLI 169
GL L LN+S + V+ +GL+ L L NL+SL L++ +++ + L T L L+
Sbjct: 508 GLTGLVSLNVSNSRVSSSGLRHLKPLKNLRSLTLESCKLSANDIRKLQATDLPNLV 563
Score = 48.9 bits (115), Expect = 8e-05
Identities = 34/98 (34%), Positives = 52/98 (52%), Gaps = 2/98 (2%)
Query: 94 TLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTS-VTDNGLKRLLGLTNLKSLNLDAR 152
T L S+ S +++ +SG+ + G LE LN +F +++ GL L GL+NL SL+
Sbjct: 138 TSLLSVDFSGSDITDSGLVSLKGCTNLESLNFNFCDQISNRGLVHLSGLSNLTSLSFRRN 197
Query: 153 Q-ITDAGLANLTSLSGLITLDLFGARITDSGTTYLRCM 189
IT G+ L++L L LDL D G +LR +
Sbjct: 198 AAITAQGMRALSNLVNLKKLDLEKCPGIDGGLVHLRAL 235
Score = 48.5 bits (114), Expect = 1e-04
Identities = 37/107 (34%), Positives = 57/107 (52%), Gaps = 9/107 (8%)
Query: 85 EGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTS-VTDNGLKRLLGLTN 143
+G+ L+ L LK L L + G+ ++ L KLE LN+ + + +TD ++ L GL
Sbjct: 203 QGMRALSNLVNLKKLDLEKCPGIDGGLVHLRALTKLESLNIKWCNCITDADMEPLSGLNK 262
Query: 144 LKSLNLD-ARQITDAGLANLT-------SLSGLITLDLFGARITDSG 182
L LNL+ R +T A L LT +L+GL+ L+L +DSG
Sbjct: 263 LNLLNLEGCRHVTAACLDTLTGLYRHPHALAGLMYLNLNRCNFSDSG 309
Score = 48.1 bits (113), Expect = 1e-04
Identities = 42/139 (30%), Positives = 63/139 (45%), Gaps = 33/139 (23%)
Query: 84 DEGLVNLTGLTLLKSLVLSDTE-VGNSGIRYISGLNKLE--------------------- 121
D GLV+L LT L+SL + + ++ + +SGLNKL
Sbjct: 226 DGGLVHLRALTKLESLNIKWCNCITDADMEPLSGLNKLNLLNLEGCRHVTAACLDTLTGL 285
Query: 122 -----------DLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLIT 170
LNL+ + +D+G ++ L NLK LNL IT++ L +L L+ L +
Sbjct: 286 YRHPHALAGLMYLNLNRCNFSDSGCEKFSDLINLKILNLGMNNITNSCLVHLKGLTKLES 345
Query: 171 LDLFGARITDSGTTYLRCM 189
L+L RI D G +L M
Sbjct: 346 LNLDSCRIGDEGLVHLSGM 364
>UniRef100_Q6K620 Leucine-rich repeat-like protein [Oryza sativa]
Length = 582
Score = 184 bits (468), Expect = 1e-45
Identities = 103/191 (53%), Positives = 132/191 (68%), Gaps = 4/191 (2%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHM----FPFL 56
L KL+ LN+EGC +TAAC E IS LA+L LNL+RCG+ +G E F L+ + F
Sbjct: 282 LSKLTQLNLEGCPVTAACLEAISGLASLVVLNLSRCGIYGEGCENFQGLKKLKVLNLGFN 341
Query: 57 QVLQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISG 116
+ + L L++ K+GDEGL++L GL LLKSL LSDTEVG+SG++++SG
Sbjct: 342 NITDDCLAHLKELINLESLNLDSCKVGDEGLLHLRGLMLLKSLELSDTEVGSSGLQHLSG 401
Query: 117 LNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGA 176
L LE +NLSFT VTD G+K++ L +LKS+NLD RQITD GLA LTSL+GL LDLFGA
Sbjct: 402 LRNLESINLSFTLVTDTGMKKISALNSLKSVNLDNRQITDVGLAALTSLTGLTHLDLFGA 461
Query: 177 RITDSGTTYLR 187
RITD GT+ R
Sbjct: 462 RITDYGTSCFR 472
Score = 63.2 bits (152), Expect = 4e-09
Identities = 55/202 (27%), Positives = 96/202 (47%), Gaps = 5/202 (2%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHM----FPFL 56
L L +L + + ++ +++S L L +NL+ ++D G +K S L +
Sbjct: 378 LMLLKSLELSDTEVGSSGLQHLSGLRNLESINLSFTLVTDTGMKKISALNSLKSVNLDNR 437
Query: 57 QVLQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISG 116
Q+ + LT L F +I D G L+SL + + ++G++ I
Sbjct: 438 QITDVGLAALTSLTGLTHLDLFGARITDYGTSCFRFFKNLESLEVCGGLITDAGVKNIKD 497
Query: 117 LNKLEDLNLSFT-SVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFG 175
L L+ LNLS ++TD L+ + GLT L SLN+ ++++AGL +L L L +L L
Sbjct: 498 LKALKQLNLSQNVNLTDKTLELISGLTALVSLNVSNTRVSNAGLRHLKDLQNLRSLSLDS 557
Query: 176 ARITDSGTTYLRCMYIYQLQAI 197
R+T S L+ + L ++
Sbjct: 558 CRVTTSEVKKLQATVLPNLISV 579
Score = 60.1 bits (144), Expect = 4e-08
Identities = 59/193 (30%), Positives = 85/193 (43%), Gaps = 6/193 (3%)
Query: 1 LQKLSTLNVEGCS-ITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPFLQ-- 57
L L++L+ + ITA E + L L L+L RC G L+++
Sbjct: 184 LSNLTSLSFKSSDGITAEAMEAFANLVNLVNLDLERCLKIHGGLVHLKGLRNLESLNMRY 243
Query: 58 ---VLQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYI 114
+ + + LT L+ +I D G+ L GL+ L L L V + + I
Sbjct: 244 CNNIADSDIKYLSDLTNLKELQLACCRITDLGVSYLRGLSKLTQLNLEGCPVTAACLEAI 303
Query: 115 SGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLF 174
SGL L LNLS + G + GL LK LNL ITD LA+L L L +L+L
Sbjct: 304 SGLASLVVLNLSRCGIYGEGCENFQGLKKLKVLNLGFNNITDDCLAHLKELINLESLNLD 363
Query: 175 GARITDSGTTYLR 187
++ D G +LR
Sbjct: 364 SCKVGDEGLLHLR 376
Score = 50.8 bits (120), Expect = 2e-05
Identities = 38/108 (35%), Positives = 55/108 (50%), Gaps = 2/108 (1%)
Query: 82 IGDEGLVNLTGLTLLKSLVLSDTE-VGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLG 140
I + GL L+GL+ L SL ++ + + + L L +L+L GL L G
Sbjct: 173 ISEHGLGILSGLSNLTSLSFKSSDGITAEAMEAFANLVNLVNLDLERCLKIHGGLVHLKG 232
Query: 141 LTNLKSLNLD-ARQITDAGLANLTSLSGLITLDLFGARITDSGTTYLR 187
L NL+SLN+ I D+ + L+ L+ L L L RITD G +YLR
Sbjct: 233 LRNLESLNMRYCNNIADSDIKYLSDLTNLKELQLACCRITDLGVSYLR 280
>UniRef100_Q7XQW9 OSJNBa0014K14.6 protein [Oryza sativa]
Length = 581
Score = 182 bits (462), Expect = 5e-45
Identities = 107/203 (52%), Positives = 137/203 (66%), Gaps = 6/203 (2%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHM----FPFL 56
L KL+ LN+EGC++TAAC E IS LA+L LNL+RCG+ D+G E L + F
Sbjct: 281 LSKLAHLNLEGCAVTAACLEVISGLASLVLLNLSRCGVYDEGCEHLEGLVKLKVLNLGFN 340
Query: 57 QVLQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISG 116
+ A + L L+ KIGDEGL +L GL L+SL LSDTEVG++G+R++SG
Sbjct: 341 YITDACLVHLKELINLECLNLDSCKIGDEGLAHLKGLLKLRSLELSDTEVGSNGLRHLSG 400
Query: 117 LNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGA 176
L L+ +NLSFT VTD GLK++ GL +L+SLNLD RQITD GLA LT L+GL LDLFGA
Sbjct: 401 LRNLQSINLSFTLVTDIGLKKISGLNSLRSLNLDNRQITDNGLAALTCLTGLTHLDLFGA 460
Query: 177 RITDSGTTYLRCMYIYQLQAIVV 199
RITD+GT L+ Y LQ++ V
Sbjct: 461 RITDAGTNCLK--YFKNLQSLEV 481
Score = 68.2 bits (165), Expect = 1e-10
Identities = 61/185 (32%), Positives = 80/185 (42%), Gaps = 21/185 (11%)
Query: 4 LSTLNVEGCSITAACFEYISALAALACLNLNRC-GLSDDGFEKFSVLQHMFPFLQVLQA* 62
L +L++E C ++ L L LNL C G++D + S
Sbjct: 211 LGSLDLERCPKIHGGLVHLKGLRKLEKLNLRYCNGITDSDMKHLS--------------- 255
Query: 63 RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLED 122
LT L+ KI D G+ L GL+ L L L V + + ISGL L
Sbjct: 256 -----DLTNLRELQLSCCKISDLGVSYLRGLSKLAHLNLEGCAVTAACLEVISGLASLVL 310
Query: 123 LNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDSG 182
LNLS V D G + L GL LK LNL ITDA L +L L L L+L +I D G
Sbjct: 311 LNLSRCGVYDEGCEHLEGLVKLKVLNLGFNYITDACLVHLKELINLECLNLDSCKIGDEG 370
Query: 183 TTYLR 187
+L+
Sbjct: 371 LAHLK 375
Score = 57.4 bits (137), Expect = 2e-07
Identities = 36/96 (37%), Positives = 52/96 (53%), Gaps = 1/96 (1%)
Query: 85 EGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSF-TSVTDNGLKRLLGLTN 143
EG + L SL L + G+ ++ GL KLE LNL + +TD+ +K L LTN
Sbjct: 200 EGAKAFANMVNLGSLDLERCPKIHGGLVHLKGLRKLEKLNLRYCNGITDSDMKHLSDLTN 259
Query: 144 LKSLNLDARQITDAGLANLTSLSGLITLDLFGARIT 179
L+ L L +I+D G++ L LS L L+L G +T
Sbjct: 260 LRELQLSCCKISDLGVSYLRGLSKLAHLNLEGCAVT 295
Score = 53.5 bits (127), Expect = 3e-06
Identities = 60/232 (25%), Positives = 94/232 (39%), Gaps = 53/232 (22%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPF-LQVL 59
L KL LN+ IT AC ++ L L CLNL+ C + D+G L + L
Sbjct: 329 LVKLKVLNLGFNYITDACLVHLKELINLECLNLDSCKIGDEGLAHLKGLLKLRSLELSDT 388
Query: 60 QA*RG*VLHLT---RLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSD------------- 103
+ + HL+ LQ+ + D GL ++GL L+SL L +
Sbjct: 389 EVGSNGLRHLSGLRNLQSINLSFTLVTDIGLKKISGLNSLRSLNLDNRQITDNGLAALTC 448
Query: 104 --------------TEVGNSGIRYISGLNKLED---------------------LNLSFT 128
T+ G + ++Y L LE LNLS
Sbjct: 449 LTGLTHLDLFGARITDAGTNCLKYFKNLQSLEVCGGLITDAGVKNIKDLKALTLLNLSQN 508
Query: 129 -SVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARIT 179
++TD L+ + LT L SLN+ +++++GL +L L L +L L ++T
Sbjct: 509 GNLTDKSLELISRLTALVSLNVSNSRVSNSGLHHLKPLQNLRSLSLESCKVT 560
Score = 39.7 bits (91), Expect = 0.050
Identities = 46/182 (25%), Positives = 76/182 (41%), Gaps = 39/182 (21%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHM-------- 52
L+ L ++N+ +T + IS L +L LNL+ ++D+G + L +
Sbjct: 401 LRNLQSINLSFTLVTDIGLKKISGLNSLRSLNLDNRQITDNGLAALTCLTGLTHLDLFGA 460
Query: 53 ------------FPFLQVLQA*RG*V-------------LHLTRLQTHA*FI*KIGDEGL 87
F LQ L+ G + L L L + + D+ L
Sbjct: 461 RITDAGTNCLKYFKNLQSLEVCGGLITDAGVKNIKDLKALTLLNLSQNG----NLTDKSL 516
Query: 88 VNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKR--LLGLTNLK 145
++ LT L SL +S++ V NSG+ ++ L L L+L VT +K+ L L NL
Sbjct: 517 ELISRLTALVSLNVSNSRVSNSGLHHLKPLQNLRSLSLESCKVTAIEIKKLQLAALPNLV 576
Query: 146 SL 147
S+
Sbjct: 577 SV 578
Score = 33.9 bits (76), Expect = 2.7
Identities = 24/94 (25%), Positives = 48/94 (50%), Gaps = 2/94 (2%)
Query: 96 LKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTS-VTDNGLKRLLGLTNLKSLNL-DARQ 153
L S+ +S ++V + G+ + L+ L+ ++ ++++GLK L GL+N+ SL+
Sbjct: 137 LLSVDISCSDVTDGGLNQLKDCINLQSLSCNYCDQISEHGLKTLSGLSNVTSLSFKKCSA 196
Query: 154 ITDAGLANLTSLSGLITLDLFGARITDSGTTYLR 187
+T G ++ L +LDL G +L+
Sbjct: 197 VTAEGAKAFANMVNLGSLDLERCPKIHGGLVHLK 230
Score = 33.9 bits (76), Expect = 2.7
Identities = 20/53 (37%), Positives = 33/53 (61%), Gaps = 1/53 (1%)
Query: 120 LEDLNLSFTSVTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANLTSLSGLITL 171
L +++S + VTD GL +L NL+SL+ + QI++ GL L+ LS + +L
Sbjct: 137 LLSVDISCSDVTDGGLNQLKDCINLQSLSCNYCDQISEHGLKTLSGLSNVTSL 189
>UniRef100_Q8H6S6 CTV.1 [Poncirus trifoliata]
Length = 205
Score = 95.9 bits (237), Expect = 6e-19
Identities = 73/163 (44%), Positives = 81/163 (48%), Gaps = 52/163 (31%)
Query: 25 LAALACLNLNRCGLSDDGFEKFSVLQHMFPFLQVLQA*RG*VLHLTRLQTHA*FI*KIGD 84
L +L LNLNRC LSDDG EKFS L + L V HL IGD
Sbjct: 1 LGSLFYLNLNRCQLSDDGCEKFSNLGFNEITDECL------VYHLDSCG--------IGD 46
Query: 85 EGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNL 144
EGLVN LSFT ++D L++L GL++L
Sbjct: 47 EGLVNY----------------------------------LSFTGISDGSLRKLAGLSSL 72
Query: 145 KSLNLDARQITDAGLANLTSLSGLITLDLFGARITDSGTTYLR 187
KSLNLDARQITD GLA LTS LDLFGARITDSG YLR
Sbjct: 73 KSLNLDARQITDTGLAALTSAH----LDLFGARITDSGAAYLR 111
>UniRef100_UPI00002BE69C UPI00002BE69C UniRef100 entry
Length = 281
Score = 90.5 bits (223), Expect = 2e-17
Identities = 64/187 (34%), Positives = 96/187 (51%), Gaps = 20/187 (10%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPFLQVLQ 60
L++LS L++ IT + ++ L L L+L ++D+G ++ + LQ +
Sbjct: 88 LKELSYLSLGETKITDEGLKEVAKLQKLDGLSLGETKITDEGLKEVAKLQKL-------- 139
Query: 61 A*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKL 120
G L+ T +I DEGL + L LK L L T+V ++G++ ++ L KL
Sbjct: 140 --DGLYLNFT----------EITDEGLKEVAKLQKLKRLTLDATKVTDAGLKELAKLQKL 187
Query: 121 EDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARITD 180
E L L T VTD GLK L L L+ L LDA ++TDAGL + L LI L L G ++T
Sbjct: 188 ERLTLDATKVTDAGLKELAKLQKLERLTLDATKVTDAGLKEVAKLQNLIFLSLNGTKVTK 247
Query: 181 SGTTYLR 187
G L+
Sbjct: 248 EGAAELK 254
Score = 75.5 bits (184), Expect = 8e-13
Identities = 54/173 (31%), Positives = 87/173 (50%), Gaps = 4/173 (2%)
Query: 14 ITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPF----LQVLQA*RG*VLHL 69
IT E ++ L L LNL+ ++D+G ++ + LQ ++ ++ V L
Sbjct: 5 ITFEGLEEVAKLTNLRLLNLHFTEITDEGLKEVAKLQQLYWLSLWDTKITDEGLKDVAKL 64
Query: 70 TRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTS 129
L + KI DEGL + L L L L +T++ + G++ ++ L KL+ L+L T
Sbjct: 65 KELSYLSLGETKITDEGLKEVAKLKELSYLSLGETKITDEGLKEVAKLQKLDGLSLGETK 124
Query: 130 VTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDSG 182
+TD GLK + L L L L+ +ITD GL + L L L L ++TD+G
Sbjct: 125 ITDEGLKEVAKLQKLDGLYLNFTEITDEGLKEVAKLQKLKRLTLDATKVTDAG 177
Score = 62.0 bits (149), Expect = 9e-09
Identities = 40/102 (39%), Positives = 56/102 (54%)
Query: 81 KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLG 140
KI EGL + LT L+ L L TE+ + G++ ++ L +L L+L T +TD GLK +
Sbjct: 4 KITFEGLEEVAKLTNLRLLNLHFTEITDEGLKEVAKLQQLYWLSLWDTKITDEGLKDVAK 63
Query: 141 LTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDSG 182
L L L+L +ITD GL + L L L L +ITD G
Sbjct: 64 LKELSYLSLGETKITDEGLKEVAKLKELSYLSLGETKITDEG 105
Score = 52.8 bits (125), Expect = 6e-06
Identities = 31/79 (39%), Positives = 43/79 (54%)
Query: 104 TEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLT 163
T++ G+ ++ L L LNL FT +TD GLK + L L L+L +ITD GL ++
Sbjct: 3 TKITFEGLEEVAKLTNLRLLNLHFTEITDEGLKEVAKLQQLYWLSLWDTKITDEGLKDVA 62
Query: 164 SLSGLITLDLFGARITDSG 182
L L L L +ITD G
Sbjct: 63 KLKELSYLSLGETKITDEG 81
>UniRef100_UPI00003282C4 UPI00003282C4 UniRef100 entry
Length = 246
Score = 87.0 bits (214), Expect = 3e-16
Identities = 62/180 (34%), Positives = 90/180 (49%), Gaps = 19/180 (10%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPFLQVLQ 60
LQKL L++ G IT + ++ L L L LNR ++D+G ++ S LQ+
Sbjct: 85 LQKLKMLSLSGTQITDEGLKEVAKLQKLKTLILNRTKITDEGLKEVSKLQN--------- 135
Query: 61 A*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKL 120
L RL H +I DEGL L L LK L L +T++ + G++ ++ L +L
Sbjct: 136 --------LERLDFHD--CQQITDEGLKELAKLQKLKRLYLRNTQITDKGLKEMANLQQL 185
Query: 121 EDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARITD 180
L+L T +TD GLK L L L+ L L ITD L + L L TL+L +IT+
Sbjct: 186 THLDLGITQITDEGLKELAKLQKLEELTLYRTHITDESLKEVAKLKDLKTLNLPNTQITN 245
Score = 76.3 bits (186), Expect = 5e-13
Identities = 60/183 (32%), Positives = 90/183 (48%), Gaps = 21/183 (11%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPFLQVLQ 60
L K+ +L + G A + ++ L L L L R ++D+G ++ + LQ +
Sbjct: 37 LAKIDSLYLYGTQAIDAGLKEVAMLLNLEYLFLGRTKITDEGLKEVAKLQKLK------- 89
Query: 61 A*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKL 120
+L L+ Q I DEGL + L LK+L+L+ T++ + G++ +S L L
Sbjct: 90 -----MLSLSGTQ--------ITDEGLKEVAKLQKLKTLILNRTKITDEGLKEVSKLQNL 136
Query: 121 EDLNL-SFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARIT 179
E L+ +TD GLK L L LK L L QITD GL + +L L LDL +IT
Sbjct: 137 ERLDFHDCQQITDEGLKELAKLQKLKRLYLRNTQITDKGLKEMANLQQLTHLDLGITQIT 196
Query: 180 DSG 182
D G
Sbjct: 197 DEG 199
Score = 62.4 bits (150), Expect = 7e-09
Identities = 39/98 (39%), Positives = 53/98 (53%)
Query: 85 EGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNL 144
EG + L + SL L T+ ++G++ ++ L LE L L T +TD GLK + L L
Sbjct: 29 EGELTEADLAKIDSLYLYGTQAIDAGLKEVAMLLNLEYLFLGRTKITDEGLKEVAKLQKL 88
Query: 145 KSLNLDARQITDAGLANLTSLSGLITLDLFGARITDSG 182
K L+L QITD GL + L L TL L +ITD G
Sbjct: 89 KMLSLSGTQITDEGLKEVAKLQKLKTLILNRTKITDEG 126
>UniRef100_Q6MBX8 Hypothetical protein [Parachlamydia sp.]
Length = 666
Score = 83.2 bits (204), Expect = 4e-15
Identities = 74/205 (36%), Positives = 111/205 (54%), Gaps = 13/205 (6%)
Query: 1 LQKLSTLNVEGCS-ITAACFEYISALAALACLNLNRC-GLSDDGFEKFSVL---QHMFPF 55
L+ L L++ CS +T +++ L AL L LN C L+D G ++L QH+
Sbjct: 258 LKALQHLDLSQCSKLTDDGLAHLTPLTALQHLGLNYCENLTDAGLAHLTLLTGLQHL-DL 316
Query: 56 LQVLQA*RG*VLHLTRLQT----HA*FI*KIGDEGLVNLTGLTLLKSLVLSDTE-VGNSG 110
+ HLT L + K+ D GL +LT LT L+ L LS+ + + ++G
Sbjct: 317 SNCKNLTDAGLAHLTSLMALQHLDLSWCLKLTDAGLAHLTSLTGLQHLDLSNCKNLTDAG 376
Query: 111 IRYISGLNKLEDLNLSFT-SVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLI 169
+ +++ L L+ LNLS+ +TD GL L LT L+ LNL +T AGLA+LTSL+GL
Sbjct: 377 LAHLTSLMALQHLNLSWCLKLTDAGLAHLTPLTALQHLNLSRYNLTYAGLAHLTSLTGLQ 436
Query: 170 TLDLFGAR-ITDSGTTYLRCMYIYQ 193
LDL G+R + D+G +LR + Q
Sbjct: 437 HLDLSGSRKLIDAGLAHLRPLVALQ 461
Score = 70.1 bits (170), Expect = 3e-11
Identities = 64/192 (33%), Positives = 99/192 (51%), Gaps = 24/192 (12%)
Query: 1 LQKLSTLNVEGC-SITAACFEYISALAALACLNLNRCG-LSDDGFEKFSVLQHMFPFLQV 58
L+ L TL + C ++T A ++ L AL L+L+ C L+D G L H+ P + +
Sbjct: 482 LKALQTLGLSWCQNLTGAGLAHLKPLVALQYLDLSNCNNLTDAG------LAHLRPLVAL 535
Query: 59 LQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLS-DTEVGNSGIRYISGL 117
L+LT K+ D GL +LT L L+ L LS ++ ++G+ ++ L
Sbjct: 536 QH------LNLTGCW-------KLTDAGLAHLTSLMALQHLNLSWCLKLTDAGLAHLKPL 582
Query: 118 NKLEDLNLS-FTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGA 176
L+ L+LS ++TD GL L L L+ LNL +TD GLA+LT L+ L LDL
Sbjct: 583 VALQHLDLSNCNNLTDEGLTHLRPLVALQHLNLSRYNLTDDGLAHLTPLTTLQYLDLSSC 642
Query: 177 -RITDSGTTYLR 187
+TD+G + +
Sbjct: 643 YNLTDAGLAHFK 654
Score = 62.4 bits (150), Expect = 7e-09
Identities = 65/205 (31%), Positives = 105/205 (50%), Gaps = 13/205 (6%)
Query: 1 LQKLSTLNVEGC-SITAACFEYISALAALACLNLNRC-GLSDDGFEKFSVL---QHMFPF 55
L L L++ C ++T A ++++L AL L+L+ C L+D G + L QH+
Sbjct: 308 LTGLQHLDLSNCKNLTDAGLAHLTSLMALQHLDLSWCLKLTDAGLAHLTSLTGLQHL-DL 366
Query: 56 LQVLQA*RG*VLHLTRLQT----HA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGI 111
+ HLT L + + K+ D GL +LT LT L+ L LS + +G+
Sbjct: 367 SNCKNLTDAGLAHLTSLMALQHLNLSWCLKLTDAGLAHLTPLTALQHLNLSRYNLTYAGL 426
Query: 112 RYISGLNKLEDLNLSFT-SVTDNGLKRLLGLTNLKSLNLDAR-QITDAGLANLTSLSGLI 169
+++ L L+ L+LS + + D GL L L L+ LNL ++TDAGLA+L+ L L
Sbjct: 427 AHLTSLTGLQHLDLSGSRKLIDAGLAHLRPLVALQHLNLTGCWKLTDAGLAHLSPLKALQ 486
Query: 170 TLDL-FGARITDSGTTYLRCMYIYQ 193
TL L + +T +G +L+ + Q
Sbjct: 487 TLGLSWCQNLTGAGLAHLKPLVALQ 511
Score = 45.1 bits (105), Expect = 0.001
Identities = 29/59 (49%), Positives = 38/59 (64%), Gaps = 2/59 (3%)
Query: 130 VTDNGLKRLLGLTNLKSLNLDARQ-ITDAGLANLTSLSGLITLDLFG-ARITDSGTTYL 186
+TD GL L LT+L+ LNL ITDAGLA+LT+L L LDL +++TD G +L
Sbjct: 222 ITDAGLAHLTPLTSLQRLNLSKLWCITDAGLAHLTTLKALQHLDLSQCSKLTDDGLAHL 280
Score = 36.6 bits (83), Expect = 0.42
Identities = 28/66 (42%), Positives = 37/66 (55%), Gaps = 2/66 (3%)
Query: 130 VTDNGLKRLLGLTNLKSLNL-DARQITDAGLANLTSLSGLITLDLFGA-RITDSGTTYLR 187
+TD L L NLK L+ + R ITDAGLA+LT L+ L L+L ITD+G +L
Sbjct: 197 LTDAHLLALKNCKNLKILHFKNCRVITDAGLAHLTPLTSLQRLNLSKLWCITDAGLAHLT 256
Query: 188 CMYIYQ 193
+ Q
Sbjct: 257 TLKALQ 262
>UniRef100_Q6MCE3 Hypothetical protein [Parachlamydia sp.]
Length = 734
Score = 79.0 bits (193), Expect = 7e-14
Identities = 66/192 (34%), Positives = 102/192 (52%), Gaps = 11/192 (5%)
Query: 1 LQKLSTLNVEGCS-ITAACFEYISALAALACLNLNRCGLSDDGFEKFSVL---QHMFPFL 56
L L L + C +T A +++ L AL L+L+ C ++D G + L QH+ +
Sbjct: 523 LTGLQRLVLSWCDKLTDAGLAHLTPLTALQYLDLSCCEITDAGLAHLTPLTGLQHLV-LV 581
Query: 57 QVLQA*RG*VLHLTRLQT----HA*FI*KIGDEGLVNLTGLTLLKSLVLSDT-EVGNSGI 111
Q + HLT L T + ++ D GL +L LT L+ L L+D ++ ++G+
Sbjct: 582 YCWQLTDAGLAHLTPLTTLQYLYLGSCNRLTDAGLAHLAPLTALQHLALNDCRKLTDTGL 641
Query: 112 RYISGLNKLEDLNLS-FTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLIT 170
+++ L L+ L L+ +TD+GL L L L+ L+L +ITDAGLA+LT L L
Sbjct: 642 AHLTPLTALQHLTLNRCEKLTDDGLAHLKPLAALQYLDLSYCEITDAGLAHLTHLMALQR 701
Query: 171 LDLFGARITDSG 182
LDL+G ITD G
Sbjct: 702 LDLYGREITDDG 713
Score = 77.0 bits (188), Expect = 3e-13
Identities = 71/192 (36%), Positives = 106/192 (54%), Gaps = 25/192 (13%)
Query: 1 LQKLSTLNVEGCS-ITAACFEYISALAALACLNLNRCG-LSDDGFEKFSVLQHMFPFLQV 58
L L L++ CS +T A +++ L AL LNLNRC L D G L H+ P +
Sbjct: 298 LTGLQHLDLSWCSSLTDAGLAHLTPLTALQHLNLNRCEYLKDAG------LAHLTPLTGL 351
Query: 59 LQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDT-EVGNSGIRYISGL 117
L+L R + + D GL +L LT L+ L LS+ ++ ++G+ +++ L
Sbjct: 352 QH------LNLNRCK-------DLTDAGLSHLKPLTALQHLNLSECWKLTDAGLAHLTPL 398
Query: 118 NKLEDLNLS-FTSVTDNGLKRLLGLTNLKSLNL-DARQITDAGLANLTSLSGLITLDLFG 175
L+ L+LS S+TD GL L LT L+ L+L D + TDAGLA+LTSL+GL L+L
Sbjct: 399 TALQHLDLSRCNSLTDAGLAHLTPLTALQHLDLSDCQNFTDAGLAHLTSLTGLQYLNLSE 458
Query: 176 AR-ITDSGTTYL 186
+ +TD+G +L
Sbjct: 459 YKNLTDAGLAHL 470
Score = 72.0 bits (175), Expect = 9e-12
Identities = 73/213 (34%), Positives = 111/213 (51%), Gaps = 13/213 (6%)
Query: 1 LQKLSTLNVEGC-SITAACFEYISALAALACLNLNRCG-LSDDGFEKFS---VLQHM--F 53
L L LN+ C +T A +++ L AL L+L+RC L+D G + LQH+
Sbjct: 373 LTALQHLNLSECWKLTDAGLAHLTPLTALQHLDLSRCNSLTDAGLAHLTPLTALQHLDLS 432
Query: 54 PFLQVLQA*RG*VLHLTRLQ-THA*FI*KIGDEGLVNLTGLTLLKSLVLSDT-EVGNSGI 111
A + LT LQ + + D GL +LT LT L+ L L + + ++G+
Sbjct: 433 DCQNFTDAGLAHLTSLTGLQYLNLSEYKNLTDAGLAHLTPLTALQHLNLCNCRKFTDNGL 492
Query: 112 RYISGLNKLEDLNLSFT-SVTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANLTSLSGLI 169
+++ L L+ L+LS ++TD+GL L LT L+ L L ++TDAGLA+LT L+ L
Sbjct: 493 AHLTPLTALQHLDLSHCKNLTDDGLAHLAPLTGLQRLVLSWCDKLTDAGLAHLTPLTALQ 552
Query: 170 TLDLFGARITDSGTTYLRCMYIYQLQAIVVFLC 202
LDL ITD+G +L + LQ +V+ C
Sbjct: 553 YLDLSCCEITDAGLAHL--TPLTGLQHLVLVYC 583
Score = 68.2 bits (165), Expect = 1e-10
Identities = 47/110 (42%), Positives = 73/110 (65%), Gaps = 4/110 (3%)
Query: 82 IGDEGLVNLTGLTLLKSLVLSDTE-VGNSGIRYISGLNKLEDLNLSF-TSVTDNGLKRLL 139
+ D GL +LT LT L+ L LSD E + + G+ +++ L L+ L+LS+ +S+TD GL L
Sbjct: 262 VTDAGLAHLTPLTTLQYLDLSDCEKLTDDGLAHLTPLTGLQHLDLSWCSSLTDAGLAHLT 321
Query: 140 GLTNLKSLNLD-ARQITDAGLANLTSLSGLITLDLFGAR-ITDSGTTYLR 187
LT L+ LNL+ + DAGLA+LT L+GL L+L + +TD+G ++L+
Sbjct: 322 PLTALQHLNLNRCEYLKDAGLAHLTPLTGLQHLNLNRCKDLTDAGLSHLK 371
Score = 61.6 bits (148), Expect = 1e-08
Identities = 58/181 (32%), Positives = 91/181 (50%), Gaps = 8/181 (4%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCG-LSDDGFEKFS---VLQHMF--P 54
L L L++ C IT A +++ L L L L C L+D G + LQ+++
Sbjct: 548 LTALQYLDLSCCEITDAGLAHLTPLTGLQHLVLVYCWQLTDAGLAHLTPLTTLQYLYLGS 607
Query: 55 FLQVLQA*RG*VLHLTRLQTHA*F-I*KIGDEGLVNLTGLTLLKSLVLSDTE-VGNSGIR 112
++ A + LT LQ A K+ D GL +LT LT L+ L L+ E + + G+
Sbjct: 608 CNRLTDAGLAHLAPLTALQHLALNDCRKLTDTGLAHLTPLTALQHLTLNRCEKLTDDGLA 667
Query: 113 YISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLD 172
++ L L+ L+LS+ +TD GL L L L+ L+L R+ITD GL +L+ L+
Sbjct: 668 HLKPLAALQYLDLSYCEITDAGLAHLTHLMALQRLDLYGREITDDGLERFETLAASFNLE 727
Query: 173 L 173
+
Sbjct: 728 I 728
>UniRef100_Q6MB17 Hypothetical protein [Parachlamydia sp.]
Length = 657
Score = 79.0 bits (193), Expect = 7e-14
Identities = 75/209 (35%), Positives = 110/209 (51%), Gaps = 27/209 (12%)
Query: 1 LQKLSTLNVEGC-SITAACFEYISALAALACLNLNRC-GLSDDGFEKFSVLQHMFPFLQV 58
L L LN+ GC ++T ++S+L AL L LN C L+D G L H+ P +
Sbjct: 408 LAALQHLNLFGCENLTGDGLTHLSSLVALQHLGLNFCRNLTDAG------LAHLAPLVT- 460
Query: 59 LQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLS-DTEVGNSGIRYISGL 117
L L + F + D GL +LT L L+ L L + ++G+ ++S L
Sbjct: 461 ----------LQHLDLN--FCDNLTDTGLAHLTSLVTLQHLNLGWCRNLTDAGLVHLSPL 508
Query: 118 NKLEDLNLSFT-SVTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANLTSLSGLITLDLFG 175
L+ L+L+ ++TD GL L L L+ LNL R++TDAGLA+LT L L LDLFG
Sbjct: 509 ENLQHLDLNDCYNLTDAGLAHLTPLVALQHLNLRRCRKLTDAGLAHLTPLVALQYLDLFG 568
Query: 176 AR-ITDSGTTYLRCMYIYQLQAIVVFLCS 203
R +TD+G T+L + LQ + + LC+
Sbjct: 569 CRNLTDAGLTHL--TPLIALQHLYLGLCN 595
Score = 55.8 bits (133), Expect = 7e-07
Identities = 61/188 (32%), Positives = 94/188 (49%), Gaps = 25/188 (13%)
Query: 1 LQKLSTLNVEGC-SITAACFEYISALAALACLNLNRC-GLSDDGFEKFSVLQHMFPFLQV 58
L L LN+ C ++T A ++S L L L+LN C L+D G L H+ P + +
Sbjct: 483 LVTLQHLNLGWCRNLTDAGLVHLSPLENLQHLDLNDCYNLTDAG------LAHLTPLVAL 536
Query: 59 LQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTE-VGNSGIRYISGL 117
L+L R + K+ D GL +LT L L+ L L + ++G+ +++ L
Sbjct: 537 QH------LNLRRCR-------KLTDAGLAHLTPLVALQYLDLFGCRNLTDAGLTHLTPL 583
Query: 118 NKLEDLNLSF-TSVTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANLTSLSGLITLDLFG 175
L+ L L ++TD GL L L L+ L+L +T+AGL +L+ L L LDL G
Sbjct: 584 IALQHLYLGLCNNLTDRGLAHLTPLAVLQRLDLSFCSNLTNAGLRHLSPLVALKYLDLSG 643
Query: 176 A-RITDSG 182
+TD+G
Sbjct: 644 CENLTDAG 651
Score = 51.6 bits (122), Expect = 1e-05
Identities = 41/114 (35%), Positives = 65/114 (56%), Gaps = 4/114 (3%)
Query: 84 DEGLVNLTGLTLLKSLVLSDTE-VGNSGIRYISGLNKLEDLNL-SFTSVTDNGLKRLLGL 141
D L+ L LK+L L + + ++G+ ++S L L+ L+L ++TD GL L L
Sbjct: 324 DAHLLVLKNCKNLKALYLEGCKNLTDTGLAHLSPLVALQHLSLFDCENLTDAGLAYLSPL 383
Query: 142 TNLKSLNLD-ARQITDAGLANLTSLSGLITLDLFGA-RITDSGTTYLRCMYIYQ 193
NL+ LNL ++ T+AGLA+L+ L+ L L+LFG +T G T+L + Q
Sbjct: 384 ENLQHLNLSHSKHFTNAGLAHLSPLAALQHLNLFGCENLTGDGLTHLSSLVALQ 437
>UniRef100_UPI000033B186 UPI000033B186 UniRef100 entry
Length = 310
Score = 77.8 bits (190), Expect = 2e-13
Identities = 69/188 (36%), Positives = 99/188 (51%), Gaps = 27/188 (14%)
Query: 1 LQKLSTLNVEGC-SITAACFEYISALAALACLNLNRCG-LSDDGFEKFSVLQHMFPFLQV 58
L L TLN+ C IT A ++ L L LNL+ C ++D+G L+H+
Sbjct: 68 LNSLQTLNLNWCRKITDAGLYHLQELNTLQTLNLSTCDKITDEG------LKHL------ 115
Query: 59 LQA*RG*VLHLTRLQT-HA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGL 117
LT LQT + +I ++GL +L LT L++L+ ++ + G+ ++ L
Sbjct: 116 --------KELTNLQTLNLRACKRITNQGLKHLRKLTSLQTLICGG-DITDEGLEHLKEL 166
Query: 118 NKLEDLNLSFTS-VTDNGLKRLLGLTNLKSLNL-DARQITDAGLANLTSLSGLITLDLFG 175
L LNL + S +TD GLK L LTNL+SLNL QITD GL +L LS L +L+L
Sbjct: 167 TNLRSLNLMYCSQITDEGLKHLKELTNLRSLNLMYCSQITDEGLKHLKELSNLRSLNLTH 226
Query: 176 A-RITDSG 182
+ITD G
Sbjct: 227 CNKITDVG 234
Score = 65.9 bits (159), Expect = 7e-10
Identities = 47/111 (42%), Positives = 69/111 (61%), Gaps = 4/111 (3%)
Query: 81 KIGDEGLVNLTGLTLLKSLVLSDT-EVGNSGIRYISGLNKLEDLNLSFTS-VTDNGLKRL 138
KI DEGL +L L L++L LS ++ ++G+ ++ L L L L S +TD GLK L
Sbjct: 6 KITDEGLKHLKKLNSLQTLNLSTCFKITDAGLEHLKVLTSLRSLKLKRCSQITDEGLKHL 65
Query: 139 LGLTNLKSLNLD-ARQITDAGLANLTSLSGLITLDLFGA-RITDSGTTYLR 187
L +L++LNL+ R+ITDAGL +L L+ L TL+L +ITD G +L+
Sbjct: 66 KKLNSLQTLNLNWCRKITDAGLYHLQELNTLQTLNLSTCDKITDEGLKHLK 116
Score = 56.6 bits (135), Expect = 4e-07
Identities = 62/191 (32%), Positives = 99/191 (51%), Gaps = 28/191 (14%)
Query: 1 LQKLSTLNVEGCS--ITAACFEYISALAALACLNLNRCG-LSDDGFEKFSVLQHMFPFLQ 57
L+KL++L C IT E++ L L LNL C ++D+G L+H+
Sbjct: 140 LRKLTSLQTLICGGDITDEGLEHLKELTNLRSLNLMYCSQITDEG------LKHL----- 188
Query: 58 VLQA*RG*VLHLTRLQT-HA*FI*KIGDEGLVNLTGLTLLKSLVLSD-TEVGNSGIRYIS 115
LT L++ + + +I DEGL +L L+ L+SL L+ ++ + G+ +
Sbjct: 189 ---------KELTNLRSLNLMYCSQITDEGLKHLKELSNLRSLNLTHCNKITDVGLYDLK 239
Query: 116 GLNKLEDLNLSFTS-VTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANLTSLSGLITLDL 173
L L+ LNL +TD GL L L NL++L+L ++ITD GL L+ L+ L +L+L
Sbjct: 240 ELKNLQTLNLCCCGKITDAGLYHLKELKNLQTLDLQHCKEITDEGLKWLSELACLRSLNL 299
Query: 174 -FGARITDSGT 183
+ +ITD T
Sbjct: 300 TYCDKITDGRT 310
>UniRef100_Q6MCI3 Hypothetical protein [Parachlamydia sp.]
Length = 583
Score = 77.4 bits (189), Expect = 2e-13
Identities = 49/106 (46%), Positives = 67/106 (62%), Gaps = 2/106 (1%)
Query: 84 DEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTS-VTDNGLKRLLGLT 142
D GL +LT LT L+ L L ++G+ +++ L L+ LNLSF S TD GL L LT
Sbjct: 291 DVGLAHLTPLTALQHLDLRGCYFTDAGLAHLTPLTALQHLNLSFCSNATDAGLAHLTPLT 350
Query: 143 NLKSLNLDARQITDAGLANLTSLSGLITLDLFGAR-ITDSGTTYLR 187
L+ L+L +TDAGLA+LT L+GL LDL G + +TD+G +LR
Sbjct: 351 ALQHLDLRGCYLTDAGLAHLTPLTGLQHLDLIGCKDLTDAGLAHLR 396
Score = 76.3 bits (186), Expect = 5e-13
Identities = 72/200 (36%), Positives = 106/200 (53%), Gaps = 27/200 (13%)
Query: 1 LQKLSTLNVEGCS-ITAACFEYISALAALACLNLNRC-GLSDDGFEKFSVLQHMFPFLQV 58
L L L++ GC +T A ++ L AL LNLN C L+D G L H+ P
Sbjct: 373 LTGLQHLDLIGCKDLTDAGLAHLRPLTALQHLNLNWCRNLTDAG------LAHLTP---- 422
Query: 59 LQA*RG*VLHLTRLQ-THA*FI*KIGDEGLVNLTGLTLLKSLVLSDT-EVGNSGIRYISG 116
LT LQ F I D+GL +LT LT L+ L LS ++ ++G+ +++
Sbjct: 423 ----------LTALQHLDLSFCSNITDDGLAHLTLLTTLQHLNLSGCYKLTDAGLAHLTL 472
Query: 117 LNKLEDLNLS-FTSVTDNGLKRLLGLTNLKSLNL-DARQITDAGLANLTSLSGLITLDLF 174
L L+ LNL+ + ++TD GL L L L+ L L D + +TDAGLA+LT L+ L L+L
Sbjct: 473 LTGLQHLNLNWYKNLTDAGLAHLTPLAGLQYLALTDCKNLTDAGLAHLTPLTALQHLNLS 532
Query: 175 GA-RITDSGTTYLRCMYIYQ 193
G ++TD+G +L + Q
Sbjct: 533 GCYKLTDAGLAHLTSLTALQ 552
Score = 76.3 bits (186), Expect = 5e-13
Identities = 76/203 (37%), Positives = 110/203 (53%), Gaps = 24/203 (11%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPF--LQV 58
L L L++ GC T A +++ L AL LNL+ C + D + L H+ P LQ
Sbjct: 300 LTALQHLDLRGCYFTDAGLAHLTPLTALQHLNLSFCSNATD-----AGLAHLTPLTALQH 354
Query: 59 LQA*RG*VL------HLTRLQTHA*FI*KIG-----DEGLVNLTGLTLLKSLVLS-DTEV 106
L RG L HLT L T + IG D GL +L LT L+ L L+ +
Sbjct: 355 LDL-RGCYLTDAGLAHLTPL-TGLQHLDLIGCKDLTDAGLAHLRPLTALQHLNLNWCRNL 412
Query: 107 GNSGIRYISGLNKLEDLNLSFTS-VTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANLTS 164
++G+ +++ L L+ L+LSF S +TD+GL L LT L+ LNL ++TDAGLA+LT
Sbjct: 413 TDAGLAHLTPLTALQHLDLSFCSNITDDGLAHLTLLTTLQHLNLSGCYKLTDAGLAHLTL 472
Query: 165 LSGLITLDL-FGARITDSGTTYL 186
L+GL L+L + +TD+G +L
Sbjct: 473 LTGLQHLNLNWYKNLTDAGLAHL 495
Score = 49.3 bits (116), Expect = 6e-05
Identities = 39/100 (39%), Positives = 57/100 (57%), Gaps = 7/100 (7%)
Query: 89 NLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLS-FTSVTDNGLKRLLGLTNLKSL 147
NL L L L ++D G+ +++ L L+ L+LS ++TD GL L LT L+ L
Sbjct: 252 NLKVLHLEACLAITD-----DGLAHLAPLVALQHLDLSDCENLTDVGLAHLTPLTALQHL 306
Query: 148 NLDARQITDAGLANLTSLSGLITLDL-FGARITDSGTTYL 186
+L TDAGLA+LT L+ L L+L F + TD+G +L
Sbjct: 307 DLRGCYFTDAGLAHLTPLTALQHLNLSFCSNATDAGLAHL 346
>UniRef100_Q6MCF0 Hypothetical protein [Parachlamydia sp.]
Length = 695
Score = 76.6 bits (187), Expect = 4e-13
Identities = 69/195 (35%), Positives = 102/195 (51%), Gaps = 30/195 (15%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRC-GLSDDGFEKFSV---LQHMFPFL 56
L L LN+ C T A ++S L L LNL C L+D G + LQH
Sbjct: 374 LTALQRLNLSNCWHTGAGLSHLSPLTGLQHLNLYECINLTDAGLVHLKLLTGLQH----- 428
Query: 57 QVLQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSD-TEVGNSGIRYIS 115
L+L+ + ++ D GLV+L LT L+ L LS+ + ++G+ ++
Sbjct: 429 ----------LNLS-------YCDELTDAGLVHLKLLTGLQHLNLSNCNNLTDAGLVHLK 471
Query: 116 GLNKLEDLNLSF-TSVTDNGLKRLLGLTNLKSLNL-DARQITDAGLANLTSLSGLITLDL 173
L L+ LNLS+ +TD GL L LT L+ LNL + +TDAGLA+LT L+GL LDL
Sbjct: 472 FLTGLQHLNLSYCDELTDAGLVHLKLLTGLQHLNLSNCNNLTDAGLAHLTPLTGLQHLDL 531
Query: 174 -FGARITDSGTTYLR 187
+ +++TD G +L+
Sbjct: 532 SYCSKLTDDGLAHLK 546
Score = 70.9 bits (172), Expect = 2e-11
Identities = 70/188 (37%), Positives = 101/188 (53%), Gaps = 25/188 (13%)
Query: 1 LQKLSTLNVEGCS-ITAACFEYISALAALACLNLNRCG-LSDDGFEKFSVLQHMFPFLQV 58
L L LN+ C+ +T A +++ L L L+L+ C L+DDG L H+ P L
Sbjct: 498 LTGLQHLNLSNCNNLTDAGLAHLTPLTGLQHLDLSYCSKLTDDG------LAHLKP-LTA 550
Query: 59 LQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTE-VGNSGIRYISGL 117
LQ L+L+ + + D GLV+L LT L+ L LSD + + + G+ ++ L
Sbjct: 551 LQC-----LNLSNCRN-------LTDAGLVHLKLLTGLQHLNLSDYKNLTDDGLIHLMPL 598
Query: 118 NKLEDLNL-SFTSVTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANLTSLSGLITLDLFG 175
L L L ++TD GL L LT L+ LNL +TDAGLA+LTSL+GL L+L G
Sbjct: 599 MALRHLELLGCENLTDAGLVHLTPLTALQHLNLSHCDDLTDAGLAHLTSLTGLQHLELLG 658
Query: 176 A-RITDSG 182
+TD+G
Sbjct: 659 CENLTDAG 666
Score = 63.5 bits (153), Expect = 3e-09
Identities = 64/193 (33%), Positives = 102/193 (52%), Gaps = 24/193 (12%)
Query: 2 QKLSTLNVEGC-SITAACFEYISALAALACLNLNRC-GLSDDGFEKFSVLQHMFPFLQVL 59
+ L L++E C ++T +++ L AL LNL+ L+D G L H+ P L L
Sbjct: 250 KNLKVLHLEKCRALTDDGLAHLTPLTALQYLNLSASYNLTDAG------LVHLAP-LTAL 302
Query: 60 QA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTE-VGNSGIRYISGLN 118
Q L+L R ++ D GL +L LT L+ L LS E + + G+ ++ L
Sbjct: 303 QK-----LNLGRYN-------QLTDAGLAHLKPLTALQRLDLSFCEDLTDDGLAHLRPLT 350
Query: 119 KLEDLNLSFTS-VTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGA- 176
L+ L+L + +TD+GL L LT L+ LNL T AGL++L+ L+GL L+L+
Sbjct: 351 ALQRLDLRYCEKLTDDGLVHLRPLTALQRLNLSNCWHTGAGLSHLSPLTGLQHLNLYECI 410
Query: 177 RITDSGTTYLRCM 189
+TD+G +L+ +
Sbjct: 411 NLTDAGLVHLKLL 423
>UniRef100_Q6M9U9 Hypothetical protein [Parachlamydia sp.]
Length = 761
Score = 76.3 bits (186), Expect = 5e-13
Identities = 71/192 (36%), Positives = 104/192 (53%), Gaps = 25/192 (13%)
Query: 1 LQKLSTLNVEGCS-ITAACFEYISALAALACLNLNRCG-LSDDGFEKFSVLQHMFPFLQV 58
L L LN+ C+ +T A E+++ L +L LNL+ C L+D G L H+ P L
Sbjct: 343 LTALQYLNLSRCNKLTDAGLEHLALLTSLQHLNLSSCKKLTDAG------LAHLTP-LMA 395
Query: 59 LQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTE-VGNSGIRYISGL 117
LQ HL + K+ D GL +L LT L+ L LS + + N+G+ ++ L
Sbjct: 396 LQ-------HLDLSICN-----KLTDRGLTHLNPLTALQYLNLSQCDNITNAGLEHLIPL 443
Query: 118 NKLEDLNLS-FTSVTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANLTSLSGLITLDLFG 175
L+ LNLS +TD GL+ L LT L+ L+L ++TDAG A+LT L+GL LDL
Sbjct: 444 TALQYLNLSQCEKLTDAGLEHLTPLTALQQLDLSWCYKLTDAGFAHLTPLTGLQYLDLSH 503
Query: 176 A-RITDSGTTYL 186
++TD+G +L
Sbjct: 504 CNKLTDAGLAHL 515
Score = 75.5 bits (184), Expect = 8e-13
Identities = 71/199 (35%), Positives = 105/199 (52%), Gaps = 16/199 (8%)
Query: 2 QKLSTLNVEGC-SITAACFEYISALAALACLNLNRC-GLSDDGFE--------KFSVLQH 51
+ L L++E C ++T E+++ L AL LNL+RC L+D G ++ L H
Sbjct: 219 KNLKALHLEACQALTDDGLEHLTLLTALQHLNLSRCKNLTDAGLAHLTPLTGLQYLDLSH 278
Query: 52 MFPFLQVLQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDT-EVGNSG 110
F A + L L KI D GL +LT L L+ L LS + ++G
Sbjct: 279 CNKFTDAGLAYLEILTALQHLDLRG--CDKITDAGLSHLTPLVALQYLSLSQCWNLTDAG 336
Query: 111 IRYISGLNKLEDLNLS-FTSVTDNGLKRLLGLTNLKSLNLDA-RQITDAGLANLTSLSGL 168
+ ++ L L+ LNLS +TD GL+ L LT+L+ LNL + +++TDAGLA+LT L L
Sbjct: 337 LIHLKPLTALQYLNLSRCNKLTDAGLEHLALLTSLQHLNLSSCKKLTDAGLAHLTPLMAL 396
Query: 169 ITLDL-FGARITDSGTTYL 186
LDL ++TD G T+L
Sbjct: 397 QHLDLSICNKLTDRGLTHL 415
Score = 72.4 bits (176), Expect = 7e-12
Identities = 68/195 (34%), Positives = 108/195 (54%), Gaps = 25/195 (12%)
Query: 1 LQKLSTLNVEGCS-ITAACFEYISALAALACLNLNRC-GLSDDGFEKFSVLQHMFPFLQV 58
L L L++ C+ +T A +++ L AL L+L+ C L+DDG L H+ P L
Sbjct: 493 LTGLQYLDLSHCNKLTDAGLAHLTPLTALQYLDLSNCIKLTDDG------LAHLTP-LMA 545
Query: 59 LQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTE-VGNSGIRYISGL 117
LQ HL + K+ D G +L+ LT L+ L LS + + ++ + +++ L
Sbjct: 546 LQ-------HLNLSSCY-----KLTDAGFAHLSPLTALQRLDLSYCQNLTDAELAHLTPL 593
Query: 118 NKLEDLNLSFT-SVTDNGLKRLLGLTNLKSLNL-DARQITDAGLANLTSLSGLITLDLFG 175
L+ L+L + ++TD GL L LT+L+ LNL +TDAGLA+LT+LSGL LDL
Sbjct: 594 TALQRLDLRYCENLTDAGLVHLKLLTDLQYLNLRGCGYLTDAGLAHLTTLSGLQHLDLSS 653
Query: 176 A-RITDSGTTYLRCM 189
++TD+G +L+ +
Sbjct: 654 CEKLTDAGLVHLKLL 668
Score = 66.6 bits (161), Expect = 4e-10
Identities = 71/210 (33%), Positives = 106/210 (49%), Gaps = 22/210 (10%)
Query: 1 LQKLSTLNVEGC-SITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPFLQVL 59
L L LN+ C +T A F ++S L AL L+L+ C D + L H+ P +
Sbjct: 543 LMALQHLNLSSCYKLTDAGFAHLSPLTALQRLDLSYCQNLTD-----AELAHLTPLTALQ 597
Query: 60 QA*RG*VLHLT-------RLQTHA*FI*KIG-----DEGLVNLTGLTLLKSLVLSDTE-V 106
+ +LT +L T ++ G D GL +LT L+ L+ L LS E +
Sbjct: 598 RLDLRYCENLTDAGLVHLKLLTDLQYLNLRGCGYLTDAGLAHLTTLSGLQHLDLSSCEKL 657
Query: 107 GNSGIRYISGLNKLEDLNLS-FTSVTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANLTS 164
++G+ ++ L L+ LNLS ++TD GL L LT L+ L L +TDAGLA+LT
Sbjct: 658 TDAGLVHLKLLTDLQYLNLSRCENLTDEGLALLTPLTALQHLKLRYCINLTDAGLAHLTP 717
Query: 165 LSGLITLDLFGA-RITDSGTTYLRCMYIYQ 193
L+GL LDL +TD+G +L+ + Q
Sbjct: 718 LTGLQRLDLSQCWNLTDAGLIHLKLLTALQ 747
Score = 53.1 bits (126), Expect = 4e-06
Identities = 58/184 (31%), Positives = 86/184 (46%), Gaps = 24/184 (13%)
Query: 1 LQKLSTLNVEGC-SITAACFEYISALAALACLNLNRCG-LSDDGFEKFSVLQHMFPFLQV 58
L L L++ C ++T A ++ L L LNL CG L+D G + L +
Sbjct: 593 LTALQRLDLRYCENLTDAGLVHLKLLTDLQYLNLRGCGYLTDAGLAHLTTLSGL------ 646
Query: 59 LQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTE-VGNSGIRYISGL 117
HL K+ D GLV+L LT L+ L LS E + + G+ ++ L
Sbjct: 647 --------QHLDLSSCE-----KLTDAGLVHLKLLTDLQYLNLSRCENLTDEGLALLTPL 693
Query: 118 NKLEDLNLSFT-SVTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANLTSLSGLITLDLFG 175
L+ L L + ++TD GL L LT L+ L+L +TDAGL +L L+ L L+L
Sbjct: 694 TALQHLKLRYCINLTDAGLAHLTPLTGLQRLDLSQCWNLTDAGLIHLKLLTALQHLNLSD 753
Query: 176 ARIT 179
I+
Sbjct: 754 TNIS 757
Score = 51.2 bits (121), Expect = 2e-05
Identities = 43/108 (39%), Positives = 66/108 (60%), Gaps = 4/108 (3%)
Query: 84 DEGLVNLTGLTLLKSLVLSDTE-VGNSGIRYISGLNKLEDLNLSFTS-VTDNGLKRLLGL 141
D+GL +LT LT L+ L LS + + ++G+ +++ L L+ L+LS + TD GL L L
Sbjct: 234 DDGLEHLTLLTALQHLNLSRCKNLTDAGLAHLTPLTGLQYLDLSHCNKFTDAGLAYLEIL 293
Query: 142 TNLKSLNL-DARQITDAGLANLTSLSGLITLDLFGA-RITDSGTTYLR 187
T L+ L+L +ITDAGL++LT L L L L +TD+G +L+
Sbjct: 294 TALQHLDLRGCDKITDAGLSHLTPLVALQYLSLSQCWNLTDAGLIHLK 341
Score = 50.8 bits (120), Expect = 2e-05
Identities = 35/89 (39%), Positives = 53/89 (59%), Gaps = 3/89 (3%)
Query: 108 NSGIRYISGLNKLEDLNL-SFTSVTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANLTSL 165
N+ + + L+ L+L + ++TD+GL+ L LT L+ LNL + +TDAGLA+LT L
Sbjct: 209 NAHLLALKDCKNLKALHLEACQALTDDGLEHLTLLTALQHLNLSRCKNLTDAGLAHLTPL 268
Query: 166 SGLITLDLFGA-RITDSGTTYLRCMYIYQ 193
+GL LDL + TD+G YL + Q
Sbjct: 269 TGLQYLDLSHCNKFTDAGLAYLEILTALQ 297
>UniRef100_Q6MAW0 Hypothetical protein [Parachlamydia sp.]
Length = 1143
Score = 75.9 bits (185), Expect = 6e-13
Identities = 73/192 (38%), Positives = 99/192 (51%), Gaps = 25/192 (13%)
Query: 1 LQKLSTLNVEGC-SITAACFEYISALAALACLNLNRC-GLSDDGFEKFSVLQHMFPFLQV 58
L L L++ GC IT + Y+S L AL LNLNRC L+DDG + L H+ LQ
Sbjct: 828 LVALQHLDLGGCYEITDSGLTYLSRLVALQHLNLNRCVCLTDDG---LAYLSHLVA-LQY 883
Query: 59 LQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLS-DTEVGNSGIRYISGL 117
L R KI D GL +L+ L L+ L L + +SG+ ++S L
Sbjct: 884 LDLDR---------------CWKITDRGLAHLSSLLALQHLNLGCCNNLTDSGLAHLSHL 928
Query: 118 NKLEDLNL-SFTSVTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANLTSLSGLITLDLFG 175
L+ L+L +TD+GL L L NL+ LNL+ +TD GLA+L+ L L LDL
Sbjct: 929 TSLKHLDLRDCAKLTDSGLAHLSLLVNLQYLNLNRCNNLTDRGLAHLSHLVALQHLDLGE 988
Query: 176 A-RITDSGTTYL 186
+ITDSG +L
Sbjct: 989 CYKITDSGLAHL 1000
Score = 68.6 bits (166), Expect = 1e-10
Identities = 70/192 (36%), Positives = 99/192 (51%), Gaps = 25/192 (13%)
Query: 1 LQKLSTLNVEGC-SITAACFEYISALAALACLNLNRC-GLSDDGFEKFSVLQHMFPFLQV 58
L L L++ GC IT + Y+S L AL LNLN C L+DDG + L H L
Sbjct: 238 LVALQHLDLGGCYKITDSGLTYLSRLVALQHLNLNCCVCLTDDG---LAYLSH----LVA 290
Query: 59 LQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLS-DTEVGNSGIRYISGL 117
LQ HL + + KI D GL +L+ L L+ L L + +SG+ ++S L
Sbjct: 291 LQ-------HLDLGECY-----KITDSGLAHLSSLLALQHLNLGCCNNLTDSGLAHLSHL 338
Query: 118 NKLEDLNL-SFTSVTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANLTSLSGLITLDL-F 174
L+ L+L +TD+GL L L NL+ LNL+ +TD GL++L+ L L LDL
Sbjct: 339 TSLKHLDLRDCAKLTDSGLAHLSLLVNLQYLNLNRCYNLTDRGLSHLSHLVALQYLDLGL 398
Query: 175 GARITDSGTTYL 186
++T SG +L
Sbjct: 399 CKKLTSSGLAHL 410
Score = 60.5 bits (145), Expect = 3e-08
Identities = 65/192 (33%), Positives = 97/192 (49%), Gaps = 25/192 (13%)
Query: 1 LQKLSTLNVEGCS-ITAACFEYISALAALACLNLNRCG-LSDDGFEKFSVLQHMFPFLQV 58
L L L++ C+ +T + ++S L L LNLNRC L+D G S L
Sbjct: 928 LTSLKHLDLRDCAKLTDSGLAHLSLLVNLQYLNLNRCNNLTDRGLAHLS-------HLVA 980
Query: 59 LQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTE-VGNSGIRYISGL 117
LQ HL + + KI D GL +L+ L L+ L L+ + + + G+ ++S L
Sbjct: 981 LQ-------HLDLGECY-----KITDSGLAHLSLLVNLQYLNLNRCDNLTDRGLAHLSRL 1028
Query: 118 NKLEDLNLSF-TSVTDNGLKRLLGLTNLKSLNL-DARQITDAGLANLTSLSGLITLDL-F 174
L+ LNL+ +TD+GL L L L+ LNL +T AGLA+LT L L L+L +
Sbjct: 1029 VTLQHLNLNCCVCLTDDGLAYLSPLVALRHLNLRSCDNLTSAGLAHLTPLIALQYLNLSY 1088
Query: 175 GARITDSGTTYL 186
+ D+G T+L
Sbjct: 1089 CDSLNDNGLTHL 1100
Score = 60.5 bits (145), Expect = 3e-08
Identities = 66/192 (34%), Positives = 98/192 (50%), Gaps = 25/192 (13%)
Query: 1 LQKLSTLNVEGCS-ITAACFEYISALAALACLNLNRC-GLSDDGFEKFSVLQHMFPFLQV 58
L L L++ C+ +T + ++S L L LNLNRC L+D G S L H+ LQ
Sbjct: 338 LTSLKHLDLRDCAKLTDSGLAHLSLLVNLQYLNLNRCYNLTDRGL---SHLSHLVA-LQY 393
Query: 59 LQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDT-EVGNSGIRYISGL 117
L L L + K+ GL +L+ L L+ L L E+ + G+ ++S L
Sbjct: 394 LD------LGLCK---------KLTSSGLAHLSPLVALQYLDLDRCGEITDRGLAHLSRL 438
Query: 118 NKLEDLNLSFTS-VTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANLTSLSGLITLDL-F 174
L+ LNL+ + +TD+GL L L L+ LNL +T AGLA+LT L L L+L +
Sbjct: 439 VALQHLNLNCCACLTDDGLAYLSPLVALRHLNLRCCGNLTSAGLAHLTPLIALQYLNLSY 498
Query: 175 GARITDSGTTYL 186
+ D+G T+L
Sbjct: 499 CDSLNDNGLTHL 510
Score = 54.7 bits (130), Expect = 1e-06
Identities = 52/164 (31%), Positives = 84/164 (50%), Gaps = 29/164 (17%)
Query: 14 ITAACFEYISALAALACLNLNRCG-LSDDGFEKFS---VLQHMFPFLQVLQA*RG*VLHL 69
+T++ ++S L AL L+L+RCG ++D G S LQH+ L+
Sbjct: 402 LTSSGLAHLSPLVALQYLDLDRCGEITDRGLAHLSRLVALQHLN-------------LNC 448
Query: 70 TRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDT-EVGNSGIRYISGLNKLEDLNLSFT 128
T D+GL L+ L L+ L L + ++G+ +++ L L+ LNLS+
Sbjct: 449 CACLT---------DDGLAYLSPLVALRHLNLRCCGNLTSAGLAHLTPLIALQYLNLSYC 499
Query: 129 -SVTDNGLKRLLGLTNLKSLNL-DARQITDAGLANLTSLSGLIT 170
S+ DNGL L L +LK L+L + TD+GLA+ T+L+ +T
Sbjct: 500 DSLNDNGLTHLTRLASLKHLDLSECPYFTDSGLAHFTALATSLT 543
Score = 46.6 bits (109), Expect = 4e-04
Identities = 34/73 (46%), Positives = 43/73 (58%), Gaps = 4/73 (5%)
Query: 116 GLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNL-DARQITDAGLANLTSLSGLITLDLF 174
G+ L+ N ++ +TD L L NLK L L + R TDAGLA+L+ L L LDL
Sbjct: 190 GIESLDFSNNAY--LTDAHLLALKDCKNLKVLRLHECRNFTDAGLAHLSRLVALQHLDLG 247
Query: 175 GA-RITDSGTTYL 186
G +ITDSG TYL
Sbjct: 248 GCYKITDSGLTYL 260
Score = 43.1 bits (100), Expect = 0.005
Identities = 30/58 (51%), Positives = 36/58 (61%), Gaps = 2/58 (3%)
Query: 131 TDNGLKRLLGLTNLKSLNLDA-RQITDAGLANLTSLSGLITLDLFGA-RITDSGTTYL 186
TD GL L L L+ L+L +ITD+GLA+L+ L L LDL G ITDSG TYL
Sbjct: 793 TDAGLAHLSPLVALQHLDLGGCYKITDSGLAHLSRLVALQHLDLGGCYEITDSGLTYL 850
Score = 42.7 bits (99), Expect = 0.006
Identities = 32/73 (43%), Positives = 42/73 (56%), Gaps = 4/73 (5%)
Query: 116 GLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNL-DARQITDAGLANLTSLSGLITLDLF 174
G+ L+ N ++ +TD L L NLK L L + R TDAGLA+L+ L L LDL
Sbjct: 755 GIESLDFSNNAY--LTDAHLLALKDCKNLKVLRLHECRNFTDAGLAHLSPLVALQHLDLG 812
Query: 175 GA-RITDSGTTYL 186
G +ITDSG +L
Sbjct: 813 GCYKITDSGLAHL 825
Score = 37.7 bits (86), Expect = 0.19
Identities = 30/82 (36%), Positives = 46/82 (55%), Gaps = 3/82 (3%)
Query: 108 NSGIRYISGLNKLEDLNLSFT-SVTDNGLKRLLGLTNLKSLNL-DARQITDAGLANLTSL 165
++G+ ++S L L+ L+L +TD+GL L L L+ L+L +ITD+GL L+ L
Sbjct: 794 DAGLAHLSPLVALQHLDLGGCYKITDSGLAHLSRLVALQHLDLGGCYEITDSGLTYLSRL 853
Query: 166 SGLITLDLFG-ARITDSGTTYL 186
L L+L +TD G YL
Sbjct: 854 VALQHLNLNRCVCLTDDGLAYL 875
Score = 32.7 bits (73), Expect = 6.1
Identities = 37/120 (30%), Positives = 55/120 (45%), Gaps = 22/120 (18%)
Query: 1 LQKLSTLNVEGCS-ITAACFEYISALAALACLNLNRCG-LSDDGFEKFSVLQHMFPFLQV 58
L L LN+ C+ +T Y+S L AL LNL CG L+ G L H+ P + +
Sbjct: 438 LVALQHLNLNCCACLTDDGLAYLSPLVALRHLNLRCCGNLTSAG------LAHLTPLIAL 491
Query: 59 LQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTE-VGNSGIRYISGL 117
L+L+ + + D GL +LT L LK L LS+ +SG+ + + L
Sbjct: 492 Q------YLNLS-------YCDSLNDNGLTHLTRLASLKHLDLSECPYFTDSGLAHFTAL 538
>UniRef100_Q6M9R9 Hypothetical protein [Parachlamydia sp.]
Length = 659
Score = 75.5 bits (184), Expect = 8e-13
Identities = 73/204 (35%), Positives = 112/204 (54%), Gaps = 12/204 (5%)
Query: 2 QKLSTLNVEGC-SITAACFEYISALAALACLNLNRCG-LSDDGF---EKFSVLQHMF--P 54
+ L L+++ C ++T A +++ L AL LNLN C L++ G + LQH+
Sbjct: 250 KNLKELHLQECRNLTDAGLVHLAPLVALKHLNLNFCDKLTNTGLAHLRPLTALQHLNLGN 309
Query: 55 FLQVLQA*RG*VLHLTRLQ-THA*FI*KIGDEGLVNLTGLTLLKSLVLSDTE-VGNSGIR 112
+ A + LT LQ + F K+ D GLV L+ LT L+ L LSD E + ++G+
Sbjct: 310 CRNLTDAGLAHLTPLTALQHLNLNFCDKLTDTGLVRLSPLTALQHLDLSDCENLTDAGLV 369
Query: 113 YISGLNKLEDLNLSFT-SVTDNGLKRLLGLTNLKSLNL-DARQITDAGLANLTSLSGLIT 170
++ L L+ LNLS ++TD GL L L L+ L+L D +TDAGLA+LT L+ L
Sbjct: 370 HLKPLVALQHLNLSCCENLTDAGLVHLKLLVALQHLDLSDCNNLTDAGLAHLTPLTALQY 429
Query: 171 LDL-FGARITDSGTTYLRCMYIYQ 193
LDL + +TD+G +L+ + Q
Sbjct: 430 LDLSYCNNLTDAGLVHLKFLTALQ 453
Score = 63.2 bits (152), Expect = 4e-09
Identities = 70/211 (33%), Positives = 104/211 (49%), Gaps = 33/211 (15%)
Query: 1 LQKLSTLNVEGC-SITAACFEYISALAALACLNLNRCG-LSDDGFEKFS---VLQHMFPF 55
L L LN+ C ++T A +++ L AL LNLN C L+D G + S LQH
Sbjct: 299 LTALQHLNLGNCRNLTDAGLAHLTPLTALQHLNLNFCDKLTDTGLVRLSPLTALQH---- 354
Query: 56 LQVLQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTE-VGNSGIRYI 114
L L+ + + D GLV+L L L+ L LS E + ++G+ ++
Sbjct: 355 -----------LDLSDCEN-------LTDAGLVHLKPLVALQHLNLSCCENLTDAGLVHL 396
Query: 115 SGLNKLEDLNLS-FTSVTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANLTSLSGLITLD 172
L L+ L+LS ++TD GL L LT L+ L+L +TDAGL +L L+ L LD
Sbjct: 397 KLLVALQHLDLSDCNNLTDAGLAHLTPLTALQYLDLSYCNNLTDAGLVHLKFLTALQHLD 456
Query: 173 LFGA-RITDSGTTYLRCMYIYQLQAIVVFLC 202
L G ++ D G +L + LQA+ + C
Sbjct: 457 LRGCDKVADDGLAHL--TPLTALQALSLSQC 485
Score = 57.8 bits (138), Expect = 2e-07
Identities = 63/195 (32%), Positives = 98/195 (49%), Gaps = 14/195 (7%)
Query: 1 LQKLSTLNVEGCSITAAC-FEYISALAALACLNLNRC-GLSDDGFEKFSVLQHMFPFLQV 58
L L L++ GC A +++ L AL L+L++C L+D G +L + +L++
Sbjct: 449 LTALQHLDLRGCDKVADDGLAHLTPLTALQALSLSQCRNLTDAGLGHLKLLTAL-QYLRL 507
Query: 59 LQA*R---G*VLHLTRL----QTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTE-VGNSG 110
Q ++HL L + + D GLV+LT L L+ L L+ E + G
Sbjct: 508 SQCWNLTDAGLIHLRPLVALQHLDLSYCGNLTDVGLVHLTPLMALQHLDLNYCENLTGDG 567
Query: 111 IRYISGLNKLEDLNLSFT-SVTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANLTSLSGL 168
+ ++ L L+ L+L+ ++TD GL L LT L+ L+L TD GL +LTSL L
Sbjct: 568 LAHLRSLTTLQHLSLNQCWNLTDAGLVHLEPLTALQHLDLSYCGNFTDVGLVHLTSLMAL 627
Query: 169 ITLDLFGA-RITDSG 182
L+L G R+TD G
Sbjct: 628 QHLNLRGCDRVTDVG 642
>UniRef100_Q6MD82 Hypothetical protein [Parachlamydia sp.]
Length = 765
Score = 75.1 bits (183), Expect = 1e-12
Identities = 69/206 (33%), Positives = 101/206 (48%), Gaps = 15/206 (7%)
Query: 1 LQKLSTLNVEGC-SITAACFEYISALAALACLNLNRCG-LSDDGFEKFSVL--------Q 50
L L L++ C + T A ++ L AL LNL+ CG L+D G +L
Sbjct: 331 LAALQHLDLSHCRNFTDAGLAHLKLLVALQHLNLSHCGKLTDAGLAHLKLLVALQHLDLS 390
Query: 51 HMFPFLQVLQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTE-VGNS 109
H F A ++ L L + + D GL +LT L L+ L L+ + ++
Sbjct: 391 HCRNFTDAGLAHLKLLVALQHLNLS--YCGNLTDAGLAHLTPLMALQHLDLNGCHNLTDA 448
Query: 110 GIRYISGLNKLEDLNLSFT-SVTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANLTSLSG 167
G+ +++ L L+ LNLS+ + TD GL L L L+ LNL TDAGLA+LTSL+
Sbjct: 449 GLTHLTSLVVLQYLNLSWNYNFTDAGLAHLTPLMALQHLNLSYCGNFTDAGLAHLTSLAA 508
Query: 168 LITLDLFGARITDSGTTYLRCMYIYQ 193
L LDL G +TD G +L+ + Q
Sbjct: 509 LKHLDLIGCELTDDGLAHLKLLVALQ 534
Score = 74.7 bits (182), Expect = 1e-12
Identities = 70/197 (35%), Positives = 104/197 (52%), Gaps = 24/197 (12%)
Query: 2 QKLSTLNVEGC-SITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPFLQVLQ 60
+ L LN++ C ++T A +++ LAAL L+L+ C L+DDG L H+ P L LQ
Sbjct: 283 ENLKVLNLQACHNLTDAGLAHLTPLAALKHLDLSGCELTDDG------LVHLTP-LAALQ 335
Query: 61 A*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDT-EVGNSGIRYISGLNK 119
L L+ + D GL +L L L+ L LS ++ ++G+ ++ L
Sbjct: 336 H-----LDLSHCRNFT-------DAGLAHLKLLVALQHLNLSHCGKLTDAGLAHLKLLVA 383
Query: 120 LEDLNLSF-TSVTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANLTSLSGLITLDLFGA- 176
L+ L+LS + TD GL L L L+ LNL +TDAGLA+LT L L LDL G
Sbjct: 384 LQHLDLSHCRNFTDAGLAHLKLLVALQHLNLSYCGNLTDAGLAHLTPLMALQHLDLNGCH 443
Query: 177 RITDSGTTYLRCMYIYQ 193
+TD+G T+L + + Q
Sbjct: 444 NLTDAGLTHLTSLVVLQ 460
Score = 72.4 bits (176), Expect = 7e-12
Identities = 73/216 (33%), Positives = 102/216 (46%), Gaps = 35/216 (16%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCG-LSDDGFEKFSVL---QHM---- 52
L L L++ GC +T ++ L AL LNL+ CG L+DDG +L QH+
Sbjct: 506 LAALKHLDLIGCELTDDGLAHLKLLVALQHLNLSYCGKLTDDGLAHLKLLVALQHLDLSG 565
Query: 53 -----------FPFLQVLQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVL 101
FL LQ L+L+ K+ D+GLVNLT L L+ L L
Sbjct: 566 CDKLTGAGLAHLKFLVALQH-----LNLSHCG-------KLTDDGLVNLTPLAALRHLDL 613
Query: 102 SDT-EVGNSGIRYISGLNKLEDLNLSFT-SVTDNGLKRLLGLTNLKSLNLD-ARQITDAG 158
S ++ +G+ ++ L L+ LNLS +TD GL L L L+ L+L +TDAG
Sbjct: 614 SHCGKLTGAGLAHLKFLVALQHLNLSHCGKLTDAGLVNLSPLMALQHLDLSHCGNLTDAG 673
Query: 159 LANLTSLSGLITLDL-FGARITDSGTTYLRCMYIYQ 193
L NL+ L L LDL +TD G L+ + Q
Sbjct: 674 LVNLSPLMALQHLDLSHCGNLTDDGLVNLKFLVALQ 709
Score = 60.8 bits (146), Expect = 2e-08
Identities = 66/198 (33%), Positives = 98/198 (49%), Gaps = 13/198 (6%)
Query: 1 LQKLSTLNVEGC-SITAACFEYISALAALACLNLNRC-GLSDDGFEKFS---VLQHMFPF 55
L L LN+ C ++T A +++ L AL L+LN C L+D G + VLQ++
Sbjct: 406 LVALQHLNLSYCGNLTDAGLAHLTPLMALQHLDLNGCHNLTDAGLTHLTSLVVLQYLNLS 465
Query: 56 LQVLQA*RG*VLHLTRLQT----HA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGI 111
G + HLT L + + D GL +LT L LK L L E+ + G+
Sbjct: 466 WNYNFTDAG-LAHLTPLMALQHLNLSYCGNFTDAGLAHLTSLAALKHLDLIGCELTDDGL 524
Query: 112 RYISGLNKLEDLNLSFT-SVTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANLTSLSGLI 169
++ L L+ LNLS+ +TD+GL L L L+ L+L ++T AGLA+L L L
Sbjct: 525 AHLKLLVALQHLNLSYCGKLTDDGLAHLKLLVALQHLDLSGCDKLTGAGLAHLKFLVALQ 584
Query: 170 TLDL-FGARITDSGTTYL 186
L+L ++TD G L
Sbjct: 585 HLNLSHCGKLTDDGLVNL 602
Score = 56.2 bits (134), Expect = 5e-07
Identities = 35/72 (48%), Positives = 45/72 (61%), Gaps = 2/72 (2%)
Query: 117 LNKLEDLNLSFTS-VTDNGLKRLLGLTNLKSLNLDA-RQITDAGLANLTSLSGLITLDLF 174
LN++E+LN S + +TD L L NLK LNL A +TDAGLA+LT L+ L LDL
Sbjct: 257 LNEIEELNFSKNAHLTDAHLLALKNCENLKVLNLQACHNLTDAGLAHLTPLAALKHLDLS 316
Query: 175 GARITDSGTTYL 186
G +TD G +L
Sbjct: 317 GCELTDDGLVHL 328
Score = 50.4 bits (119), Expect = 3e-05
Identities = 68/222 (30%), Positives = 98/222 (43%), Gaps = 61/222 (27%)
Query: 1 LQKLSTLNVEGCS-ITAACFEYISALAALACLNLNRCG-LSDDGFEKFS---VLQHM--- 52
L L L++ GC +T A ++ L AL LNL+ CG L+DDG + L+H+
Sbjct: 555 LVALQHLDLSGCDKLTGAGLAHLKFLVALQHLNLSHCGKLTDDGLVNLTPLAALRHLDLS 614
Query: 53 ------------FPFLQVLQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLV 100
FL LQ L+L+ K+ D GLVNL+ L L+ L
Sbjct: 615 HCGKLTGAGLAHLKFLVALQH-----LNLSHCG-------KLTDAGLVNLSPLMALQHLD 662
Query: 101 LSDT-EVGNSGIRYISGLNKLEDLNLSFT-SVTDNGLKRLLGLTNLKSLNLDA------- 151
LS + ++G+ +S L L+ L+LS ++TD+GL L L L+ L+L
Sbjct: 663 LSHCGNLTDAGLVNLSPLMALQHLDLSHCGNLTDDGLVNLKFLVALQHLDLSHCGNLTDD 722
Query: 152 -------------------RQITD-AGLANLTSLSGLITLDL 173
+TD +GLA+LTSL L LDL
Sbjct: 723 GLAHLSPLIALQHLDRSKYNNLTDGSGLAHLTSLVDLQHLDL 764
Score = 45.4 bits (106), Expect = 0.001
Identities = 31/82 (37%), Positives = 44/82 (52%), Gaps = 2/82 (2%)
Query: 114 ISGLNKLEDLNLSFT-SVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLD 172
+ L+ LNL ++TD GL L L LK L+L ++TD GL +LT L+ L LD
Sbjct: 279 LKNCENLKVLNLQACHNLTDAGLAHLTPLAALKHLDLSGCELTDDGLVHLTPLAALQHLD 338
Query: 173 LFGAR-ITDSGTTYLRCMYIYQ 193
L R TD+G +L+ + Q
Sbjct: 339 LSHCRNFTDAGLAHLKLLVALQ 360
>UniRef100_UPI00003634F6 UPI00003634F6 UniRef100 entry
Length = 857
Score = 74.7 bits (182), Expect = 1e-12
Identities = 62/188 (32%), Positives = 97/188 (50%), Gaps = 21/188 (11%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPFLQVLQ 60
L L TLN++G ++ + E++++ L+ L+L ++D Q LQ
Sbjct: 610 LSGLQTLNLDGTDVSESGLEHLASHPLLSSLSLAGISVADGN--------------QALQ 655
Query: 61 A*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSD-TEVGNSGIRYISGLNK 119
G L+LT+L + D GL L+ LTLL L L+D T+V + G+ ++ + +
Sbjct: 656 IISG--LNLTQLTLPGRR--SVTDGGLSFLSRLTLLTELDLTDYTQVTDQGVAQLASMRR 711
Query: 120 LEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARIT 179
L+ L+LS T VTD GL L GL L+ L LD +T G+A+L ++GL L + G T
Sbjct: 712 LKKLSLSNTQVTDAGLPPLRGLQELQELCLDRTAVTSRGVADL--ITGLPHLQVLGLACT 769
Query: 180 DSGTTYLR 187
G T +R
Sbjct: 770 QVGDTVVR 777
Score = 49.3 bits (116), Expect = 6e-05
Identities = 56/191 (29%), Positives = 93/191 (48%), Gaps = 26/191 (13%)
Query: 1 LQKLSTLNVEGCS-ITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPFLQVL 59
L +L LN+ CS +T +C ++I+ L +L L+L++ ++D G + H P
Sbjct: 512 LVRLQYLNLASCSKLTDSCLQHITGLKSLCFLSLDQTKVTDAGMVLYL---HSAPSC--- 565
Query: 60 QA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLT-LLKSLVLSDTEVGNSGIRYISGLN 118
+ L+ QT + + LV L L+ L + T+V + + ++GL+
Sbjct: 566 ------LSQLSLNQT------AVTEATLVVLPSCAPQLRLLSIKQTKVRD--VAALAGLS 611
Query: 119 KLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSG--LITLDLFGA 176
L+ LNL T V+++GL+ L L SL+L + D A L +SG L L L G
Sbjct: 612 GLQTLNLDGTDVSESGLEHLASHPLLSSLSLAGISVADGNQA-LQIISGLNLTQLTLPGR 670
Query: 177 R-ITDSGTTYL 186
R +TD G ++L
Sbjct: 671 RSVTDGGLSFL 681
Score = 48.9 bits (115), Expect = 8e-05
Identities = 32/83 (38%), Positives = 54/83 (64%), Gaps = 2/83 (2%)
Query: 90 LTGLTLLKSLVLSDTEVGNSGIRY-ISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLN 148
+TGL L+ L L+ T+VG++ +R + ++L LNLS T +TDNGLK L +L +N
Sbjct: 755 ITGLPHLQVLGLACTQVGDTVVRKGLLRCSQLVKLNLSRTRITDNGLK-FLKRMHLAHVN 813
Query: 149 LDARQITDAGLANLTSLSGLITL 171
LD ++ AG+A+L +L+ + ++
Sbjct: 814 LDGTGVSLAGVADLLTLTNISSI 836
Score = 43.5 bits (101), Expect = 0.003
Identities = 38/125 (30%), Positives = 61/125 (48%), Gaps = 25/125 (20%)
Query: 81 KIGDEGLVNLTGLTLLKSLVLSDTEVGNSG-IRYI-SGLNKLEDLNLSFTSVTDNGL--- 135
K+ D L ++TGL L L L T+V ++G + Y+ S + L L+L+ T+VT+ L
Sbjct: 525 KLTDSCLQHITGLKSLCFLSLDQTKVTDAGMVLYLHSAPSCLSQLSLNQTAVTEATLVVL 584
Query: 136 --------------------KRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFG 175
L GL+ L++LNLD ++++GL +L S L +L L G
Sbjct: 585 PSCAPQLRLLSIKQTKVRDVAALAGLSGLQTLNLDGTDVSESGLEHLASHPLLSSLSLAG 644
Query: 176 ARITD 180
+ D
Sbjct: 645 ISVAD 649
Score = 42.0 bits (97), Expect = 0.010
Identities = 40/129 (31%), Positives = 63/129 (48%), Gaps = 26/129 (20%)
Query: 84 DEGLVNLTGLTLLKSLVLSD-TEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLL--G 140
D GL L+ L L+ L L+ +++ +S +++I+GL L L+L T VTD G+ L
Sbjct: 503 DSGLSVLSSLVRLQYLNLASCSKLTDSCLQHITGLKSLCFLSLDQTKVTDAGMVLYLHSA 562
Query: 141 LTNLKSLNLDARQITDAGL-----------------------ANLTSLSGLITLDLFGAR 177
+ L L+L+ +T+A L A L LSGL TL+L G
Sbjct: 563 PSCLSQLSLNQTAVTEATLVVLPSCAPQLRLLSIKQTKVRDVAALAGLSGLQTLNLDGTD 622
Query: 178 ITDSGTTYL 186
+++SG +L
Sbjct: 623 VSESGLEHL 631
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.335 0.148 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 280,052,951
Number of Sequences: 2790947
Number of extensions: 9787232
Number of successful extensions: 47208
Number of sequences better than 10.0: 874
Number of HSP's better than 10.0 without gapping: 334
Number of HSP's successfully gapped in prelim test: 555
Number of HSP's that attempted gapping in prelim test: 43381
Number of HSP's gapped (non-prelim): 2913
length of query: 206
length of database: 848,049,833
effective HSP length: 121
effective length of query: 85
effective length of database: 510,345,246
effective search space: 43379345910
effective search space used: 43379345910
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 72 (32.3 bits)
Medicago: description of AC144477.5


