

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144345.5 - phase: 0
(131 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
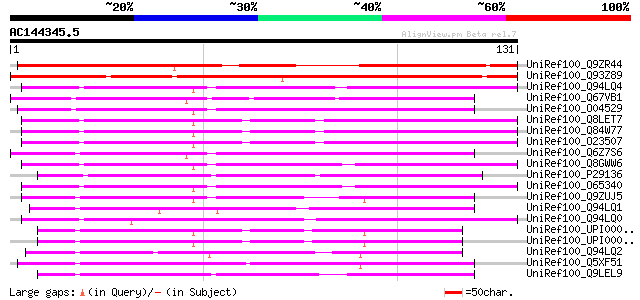
Sequences producing significant alignments: (bits) Value
UniRef100_Q9ZR44 MtN9 protein [Medicago truncatula] 136 9e-32
UniRef100_Q93Z89 Matrix metalloproteinase MMP2 [Glycine max] 127 4e-29
UniRef100_Q94LQ4 Putative metalloproteinase [Oryza sativa] 82 3e-15
UniRef100_Q67VB1 Putative metalloproteinase [Oryza sativa] 76 2e-13
UniRef100_O04529 F20P5.11 protein [Arabidopsis thaliana] 75 3e-13
UniRef100_Q8LET7 Proteinase like protein [Arabidopsis thaliana] 73 1e-12
UniRef100_Q84W77 Hypothetical protein At4g16640 [Arabidopsis tha... 73 1e-12
UniRef100_O23507 Proteinase like protein [Arabidopsis thaliana] 73 1e-12
UniRef100_Q6Z7S6 Putative zinc metalloproteinase [Oryza sativa] 73 2e-12
UniRef100_Q8GWW6 Putative metalloproteinase [Arabidopsis thaliana] 71 5e-12
UniRef100_P29136 Metalloendoproteinase 1 precursor [Glycine max] 71 5e-12
UniRef100_O65340 Metalloproteinase [Arabidopsis thaliana] 70 8e-12
UniRef100_Q9ZUJ5 T2K10.2 protein [Arabidopsis thaliana] 66 2e-10
UniRef100_Q94LQ1 Putative metalloproteinase [Oryza sativa] 63 1e-09
UniRef100_Q94LQ0 Putative metalloproteinase [Oryza sativa] 62 3e-09
UniRef100_UPI0000362B3E UPI0000362B3E UniRef100 entry 62 4e-09
UniRef100_UPI0000318A69 UPI0000318A69 UniRef100 entry 60 1e-08
UniRef100_Q94LQ2 Putative metalloproteinase [Oryza sativa] 60 1e-08
UniRef100_Q5XF51 At1g24140 [Arabidopsis thaliana] 59 2e-08
UniRef100_Q9LEL9 Matrix metalloproteinase [Cucumis sativus] 57 9e-08
>UniRef100_Q9ZR44 MtN9 protein [Medicago truncatula]
Length = 191
Score = 136 bits (343), Expect = 9e-32
Identities = 81/130 (62%), Positives = 88/130 (67%), Gaps = 22/130 (16%)
Query: 3 NVFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINY-IDGVDDVVVGDTVIKLGSNV 61
NVFRNAFTRWSQTTRVL FSE TSYDDA+IKIGFYNI+Y V DVVV D I N+
Sbjct: 26 NVFRNAFTRWSQTTRVLKFSEATSYDDADIKIGFYNISYNSKEVIDVVVSDFFI----NL 81
Query: 62 NSGFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPAIL 121
S IRL ASK W DLET AMHQIGHLLGLDHS D E +MYP I+
Sbjct: 82 RSFTIRLEASKVW----------------DLETVAMHQIGHLLGLDHSSDVESIMYPTIV 125
Query: 122 PLQQQRKLQI 131
PL Q+K+QI
Sbjct: 126 PL-HQKKVQI 134
>UniRef100_Q93Z89 Matrix metalloproteinase MMP2 [Glycine max]
Length = 357
Score = 127 bits (320), Expect = 4e-29
Identities = 67/133 (50%), Positives = 93/133 (69%), Gaps = 5/133 (3%)
Query: 1 MTNVFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVDDVVVGDTVIKLGSN 60
MT VFR++F RW+Q + VLN +ETT YD+A+I++GFYN Y+ G+D V G ++I L +
Sbjct: 183 MTKVFRDSFARWAQASGVLNLTETT-YDNADIQVGFYNFTYL-GIDIEVYGGSLIFLQPD 240
Query: 61 VNSGFIRLI--ASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYP 118
+ L+ +K W LP++N SW+ G DLE+AAMH+IGHLLGLDHS ++ VMYP
Sbjct: 241 STKKGVILLDGTNKLWALPSENGRLSWEEGVLDLESAAMHEIGHLLGLDHSNKEDSVMYP 300
Query: 119 AILPLQQQRKLQI 131
ILP QRK+Q+
Sbjct: 301 CILP-SHQRKVQL 312
>UniRef100_Q94LQ4 Putative metalloproteinase [Oryza sativa]
Length = 355
Score = 81.6 bits (200), Expect = 3e-15
Identities = 50/129 (38%), Positives = 76/129 (58%), Gaps = 7/129 (5%)
Query: 4 VFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVD-DVVVGDTVIKLGSNVN 62
VF+ AF RW++T V F ET Y+ A+IK+GFY N+ DGV D +G ++ +
Sbjct: 180 VFQRAFARWARTIPV-GFVETDDYEAADIKVGFYAGNHGDGVPFDGPLG--ILGHAFSPK 236
Query: 63 SGFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPAILP 122
+G + L AS++W + D + DLE+ A H+IGH+LGL HS VMYP+I P
Sbjct: 237 NGRLHLDASEHWAVDFDVDATA---SAIDLESVATHEIGHVLGLGHSASPRAVMYPSIKP 293
Query: 123 LQQQRKLQI 131
+++ +L +
Sbjct: 294 REKKVRLTV 302
>UniRef100_Q67VB1 Putative metalloproteinase [Oryza sativa]
Length = 371
Score = 75.9 bits (185), Expect = 2e-13
Identities = 51/121 (42%), Positives = 70/121 (57%), Gaps = 5/121 (4%)
Query: 1 MTNVFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDG-VDDVVVGDTVIKLGS 59
++ VF +AF RWS T LNF+E S DA+I IGFY ++ DG D +G T+ S
Sbjct: 184 LSKVFASAFARWSAAT-TLNFTEAASAADADITIGFYGGDHGDGEAFDGPLG-TLAHAFS 241
Query: 60 NVNSGFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPA 119
N G + L AS+ WV D S N DLE+ A+H+IGH+LGL HS + +M+P
Sbjct: 242 PTN-GRLHLDASEAWVAGGDVTRAS-SNAAVDLESVAVHEIGHILGLGHSSAADSIMFPT 299
Query: 120 I 120
+
Sbjct: 300 L 300
>UniRef100_O04529 F20P5.11 protein [Arabidopsis thaliana]
Length = 378
Score = 75.1 bits (183), Expect = 3e-13
Identities = 48/119 (40%), Positives = 67/119 (55%), Gaps = 4/119 (3%)
Query: 3 NVFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVD-DVVVGDTVIKLGSNV 61
+VF AF RWS T LNF+ + S+ ++I IGFY ++ DG D V+G + +
Sbjct: 187 SVFSRAFGRWSDVT-ALNFTLSESFSTSDITIGFYTGDHGDGEPFDGVLG--TLAHAFSP 243
Query: 62 NSGFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPAI 120
SG L A + WV+ D + DLE+ A+H+IGHLLGL HS +E +MYP I
Sbjct: 244 PSGKFHLDADENWVVSGDLDSFLSVTAAVDLESVAVHEIGHLLGLGHSSVEESIMYPTI 302
>UniRef100_Q8LET7 Proteinase like protein [Arabidopsis thaliana]
Length = 364
Score = 73.2 bits (178), Expect = 1e-12
Identities = 46/129 (35%), Positives = 71/129 (54%), Gaps = 6/129 (4%)
Query: 4 VFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVD-DVVVGDTVIKLGSNVN 62
VFR AF++WS V +F E + A++KIGFY ++ DG+ D V+G
Sbjct: 185 VFRRAFSQWSSVIPV-SFEEVDDFTTADLKIGFYAGDHGDGLPFDGVLGTLAHAFAPE-- 241
Query: 63 SGFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPAILP 122
+G + L A++ W++ D + DLE+ A H+IGHLLGL HS + VMYP++ P
Sbjct: 242 NGRLHLDAAETWIVDDD--LKGSSEVAVDLESVATHEIGHLLGLGHSSQESAVMYPSLRP 299
Query: 123 LQQQRKLQI 131
++ L +
Sbjct: 300 RTKKVDLTV 308
>UniRef100_Q84W77 Hypothetical protein At4g16640 [Arabidopsis thaliana]
Length = 364
Score = 73.2 bits (178), Expect = 1e-12
Identities = 46/129 (35%), Positives = 71/129 (54%), Gaps = 6/129 (4%)
Query: 4 VFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVD-DVVVGDTVIKLGSNVN 62
VFR AF++WS V +F E + A++KIGFY ++ DG+ D V+G
Sbjct: 185 VFRRAFSQWSSVIPV-SFEEVDDFTTADLKIGFYAGDHGDGLPFDGVLGTLAHAFAPE-- 241
Query: 63 SGFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPAILP 122
+G + L A++ W++ D + DLE+ A H+IGHLLGL HS + VMYP++ P
Sbjct: 242 NGRLHLDAAETWIVDDD--LKGSSEVAVDLESVATHEIGHLLGLGHSSQESAVMYPSLRP 299
Query: 123 LQQQRKLQI 131
++ L +
Sbjct: 300 RTKKVDLTV 308
>UniRef100_O23507 Proteinase like protein [Arabidopsis thaliana]
Length = 364
Score = 73.2 bits (178), Expect = 1e-12
Identities = 46/129 (35%), Positives = 71/129 (54%), Gaps = 6/129 (4%)
Query: 4 VFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVD-DVVVGDTVIKLGSNVN 62
VFR AF++WS V +F E + A++KIGFY ++ DG+ D V+G
Sbjct: 185 VFRRAFSQWSSVIPV-SFEEVDDFTTADLKIGFYAGDHGDGLPFDGVLGTLAHAFAPE-- 241
Query: 63 SGFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPAILP 122
+G + L A++ W++ D + DLE+ A H+IGHLLGL HS + VMYP++ P
Sbjct: 242 NGRLHLDAAETWIVDDD--LKGSSEVAVDLESVATHEIGHLLGLGHSSQESAVMYPSLRP 299
Query: 123 LQQQRKLQI 131
++ L +
Sbjct: 300 RTKKVDLTV 308
>UniRef100_Q6Z7S6 Putative zinc metalloproteinase [Oryza sativa]
Length = 372
Score = 72.8 bits (177), Expect = 2e-12
Identities = 47/121 (38%), Positives = 68/121 (55%), Gaps = 4/121 (3%)
Query: 1 MTNVFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDG-VDDVVVGDTVIKLGS 59
++ VF AF+RW+ TR L F+E +S +A+I IGFY+ ++ DG D +G +
Sbjct: 184 LSAVFARAFSRWAAATR-LQFTEVSSASNADITIGFYSGDHGDGEAFDGPLG--TLAHAF 240
Query: 60 NVNSGFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPA 119
+ G L A++ WV D S DLE+ A+H+IGHLLGL HS + +MYP
Sbjct: 241 SPTDGRFHLDAAEAWVASGDVSTSSSFGTAVDLESVAVHEIGHLLGLGHSSVPDSIMYPT 300
Query: 120 I 120
I
Sbjct: 301 I 301
>UniRef100_Q8GWW6 Putative metalloproteinase [Arabidopsis thaliana]
Length = 342
Score = 71.2 bits (173), Expect = 5e-12
Identities = 48/129 (37%), Positives = 72/129 (55%), Gaps = 7/129 (5%)
Query: 4 VFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVD-DVVVGDTVIKLGSNVN 62
VFR AF +W+ V +F ET Y A+IKIGF+N ++ DG D V+G V+ +
Sbjct: 163 VFRRAFGKWASVIPV-SFIETEDYVIADIKIGFFNGDHGDGEPFDGVLG--VLAHTFSPE 219
Query: 63 SGFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPAILP 122
+G + L ++ W + D S DLE+ A+H+IGH+LGL HS K+ MYP + P
Sbjct: 220 NGRLHLDKAETWAVDFDEEKSSVA---VDLESVAVHEIGHVLGLGHSSVKDAAMYPTLKP 276
Query: 123 LQQQRKLQI 131
++ L +
Sbjct: 277 RSKKVNLNM 285
>UniRef100_P29136 Metalloendoproteinase 1 precursor [Glycine max]
Length = 305
Score = 71.2 bits (173), Expect = 5e-12
Identities = 46/115 (40%), Positives = 61/115 (53%), Gaps = 3/115 (2%)
Query: 8 AFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVDDVVVGDTVIKLGSNVNSGFIR 67
AF++W+ + F ETTSY+ ANIKI F + N+ D G ++ G
Sbjct: 171 AFSKWTPVVNIA-FQETTSYETANIKILFASKNHGDPYPFDGPGG-ILGHAFAPTDGRCH 228
Query: 68 LIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPAILP 122
A +YWV D S FDLE+ A+H+IGHLLGL HS D +MYP+I P
Sbjct: 229 FDADEYWVASGD-VTKSPVTSAFDLESVAVHEIGHLLGLGHSSDLRAIMYPSIPP 282
>UniRef100_O65340 Metalloproteinase [Arabidopsis thaliana]
Length = 341
Score = 70.5 bits (171), Expect = 8e-12
Identities = 48/129 (37%), Positives = 71/129 (54%), Gaps = 7/129 (5%)
Query: 4 VFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVD-DVVVGDTVIKLGSNVN 62
VFR AF W+ V +F ET Y A+IKIGF+N ++ DG D V+G V+ +
Sbjct: 162 VFRRAFGEWASVIPV-SFIETEDYVIADIKIGFFNGDHGDGEPFDGVLG--VLAHTFSPE 218
Query: 63 SGFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPAILP 122
+G + L ++ W + D S DLE+ A+H+IGH+LGL HS K+ MYP + P
Sbjct: 219 NGRLHLDKAETWAVDFDEEKSSVA---VDLESVAVHEIGHVLGLGHSSVKDAAMYPTLKP 275
Query: 123 LQQQRKLQI 131
++ L +
Sbjct: 276 RSKKVNLNM 284
>UniRef100_Q9ZUJ5 T2K10.2 protein [Arabidopsis thaliana]
Length = 360
Score = 65.9 bits (159), Expect = 2e-10
Identities = 46/129 (35%), Positives = 69/129 (52%), Gaps = 24/129 (18%)
Query: 4 VFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVD-DVVVGDTVIKLGSNVN 62
VF AFTRW++ T LNF+ + S A+I IGF++ + DG D +G + S+
Sbjct: 176 VFSRAFTRWAEVTP-LNFTRSESILRADIVIGFFSGEHGDGEPFDGAMG--TLAHASSPP 232
Query: 63 SGFIRLIASKYWVLPTDNYMWSWQNGEF-----------DLETAAMHQIGHLLGLDHSFD 111
+G + L + W++ NGE DLE+ A+H+IGHLLGL HS
Sbjct: 233 TGMLHLDGDEDWLI---------SNGEISRRILPVTTVVDLESVAVHEIGHLLGLGHSSV 283
Query: 112 KEYVMYPAI 120
++ +M+PAI
Sbjct: 284 EDAIMFPAI 292
>UniRef100_Q94LQ1 Putative metalloproteinase [Oryza sativa]
Length = 289
Score = 63.2 bits (152), Expect = 1e-09
Identities = 47/127 (37%), Positives = 59/127 (46%), Gaps = 16/127 (12%)
Query: 6 RNAFTRWSQTTRVLNFSETTSYDDANIKIGFY--------NINYIDGVDDVVVGD----T 53
R+AF RW+ + F E YD A+IK+GFY ID DD GD
Sbjct: 84 RSAFARWADVIP-MRFLEAERYDAADIKVGFYLYTDGRCDGCACIDSDDDDDDGDDCEGV 142
Query: 54 VIKLGSNVNSGFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKE 113
+ SG I L A+ W T N DLE+ A H+IGH+LGLDHS +
Sbjct: 143 LAHSSMPEKSGQIHLHAAHRW---TVNLAADTAPLAVDLESVAAHEIGHVLGLDHSSSRS 199
Query: 114 YVMYPAI 120
+MYP I
Sbjct: 200 SMMYPFI 206
>UniRef100_Q94LQ0 Putative metalloproteinase [Oryza sativa]
Length = 261
Score = 62.0 bits (149), Expect = 3e-09
Identities = 43/133 (32%), Positives = 65/133 (48%), Gaps = 9/133 (6%)
Query: 4 VFRNAFTRWSQTTRVLNFSETTSYDDA-----NIKIGFYNINYIDGVDDVVVGDTVIKLG 58
VFR+AF RW++ V +F+E T+ DDA +I++GFY V
Sbjct: 87 VFRSAFARWAEVIPV-SFAEITTEDDAAAAEADIRVGFYGAGEHGDGHPFDGPLNVYAHA 145
Query: 59 SNVNSGFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYP 118
+ G I A++ W + + DLET A H+IGH LGLDHS + VMYP
Sbjct: 146 TGPEDGRIDFDAAERWAV---DLAADASPAAVDLETVATHEIGHALGLDHSTSESSVMYP 202
Query: 119 AILPLQQQRKLQI 131
+ +++ +L +
Sbjct: 203 YVGTRERKVRLTV 215
>UniRef100_UPI0000362B3E UPI0000362B3E UniRef100 entry
Length = 295
Score = 61.6 bits (148), Expect = 4e-09
Identities = 42/120 (35%), Positives = 61/120 (50%), Gaps = 15/120 (12%)
Query: 8 AFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVD-------DVVVGDTVIKLGSN 60
AF WS + +F E + ++A+IKIGFY IN+ D + D + G+
Sbjct: 22 AFGLWSDVSP-FSFREVPADEEADIKIGFYPINHTDCLQSYLHHCFDGITGELAHAFFPQ 80
Query: 61 VNSGFIRLIASKYWVLPTDNYMWSWQNGEF---DLETAAMHQIGHLLGLDHSFDKEYVMY 117
+G I S+YW+L N +SW+ G DL A H+IGH LGL HS D + +M+
Sbjct: 81 --TGEIHFDDSEYWIL--GNMRFSWKKGSVWLTDLVHVATHEIGHALGLMHSMDPKAIMH 136
>UniRef100_UPI0000318A69 UPI0000318A69 UniRef100 entry
Length = 316
Score = 60.1 bits (144), Expect = 1e-08
Identities = 39/120 (32%), Positives = 61/120 (50%), Gaps = 15/120 (12%)
Query: 8 AFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVD-------DVVVGDTVIKLGSN 60
AF WS + +F E + ++A+IKIGFY +N+ D + D + G+
Sbjct: 43 AFALWSDVSP-FSFREVPADEEADIKIGFYPVNHTDCLQSYLHHCFDGITGELAHAFFPQ 101
Query: 61 VNSGFIRLIASKYWVLPTDNYMWSWQNGEF---DLETAAMHQIGHLLGLDHSFDKEYVMY 117
+G I +YW+L N +SW+ G DL A H+IGH+LG+ HS D + +M+
Sbjct: 102 --TGEIHFDDHEYWIL--GNMRFSWKKGRVWLTDLVHVATHEIGHVLGIMHSMDPKAIMH 157
>UniRef100_Q94LQ2 Putative metalloproteinase [Oryza sativa]
Length = 266
Score = 59.7 bits (143), Expect = 1e-08
Identities = 40/124 (32%), Positives = 63/124 (50%), Gaps = 17/124 (13%)
Query: 5 FRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVDDVVV------GDTVIKLG 58
FR+A RW++ T L F+E Y++A+I++GFY ++ DG D G+ +
Sbjct: 68 FRSALARWAEVTP-LRFAEAARYEEADIRVGFY-LHTADGKCDACGCVCKGGGEEALAHA 125
Query: 59 SNVNSGFIRLIASKYWVLPTDNYMWSWQNGE-----FDLETAAMHQIGHLLGLDHSFDKE 113
G I L A++ W + + G+ DLE+ A+H+IGH LGL HS +
Sbjct: 126 HPPQDGRIHLHAARKWAVTNV----AGAGGDAPPLAVDLESVAVHEIGHALGLGHSSSES 181
Query: 114 YVMY 117
+MY
Sbjct: 182 SMMY 185
>UniRef100_Q5XF51 At1g24140 [Arabidopsis thaliana]
Length = 384
Score = 59.3 bits (142), Expect = 2e-08
Identities = 41/119 (34%), Positives = 60/119 (49%), Gaps = 3/119 (2%)
Query: 3 NVFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVDDVVVGDTVIKLGSNVN 62
+VF AFTRW + T L F+ + ++I IGFY+ + DG T+ S
Sbjct: 191 SVFSRAFTRWEEVTP-LTFTRVERFSTSDISIGFYSGEHGDGEPFDGPMRTLAHAFSPP- 248
Query: 63 SGFIRLIASKYWVLPTDNYMWSWQNGE-FDLETAAMHQIGHLLGLDHSFDKEYVMYPAI 120
+G L + W++ + E DLE+ A+H+IGHLLGL HS + +MYP I
Sbjct: 249 TGHFHLDGEENWIVSGEGGDGFISVSEAVDLESVAVHEIGHLLGLGHSSVEGSIMYPTI 307
>UniRef100_Q9LEL9 Matrix metalloproteinase [Cucumis sativus]
Length = 320
Score = 57.0 bits (136), Expect = 9e-08
Identities = 38/113 (33%), Positives = 54/113 (47%), Gaps = 9/113 (7%)
Query: 8 AFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVDDVVVGDTVIKLGSNVNSGFIR 67
AF++WS T FS Y A+IKI F + D VG V+ G +
Sbjct: 195 AFSKWSLNTH-FKFSHVADYRKADIKISFERGEHGDNAPFDGVGG-VLAHAYAPTDGRLH 252
Query: 68 LIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPAI 120
W + + G FD+ET A+H+IGH+LGL HS +E +M+P+I
Sbjct: 253 FDGDDAWSVGAIS-------GYFDVETVALHEIGHILGLQHSTIEEAIMFPSI 298
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.322 0.138 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 232,850,779
Number of Sequences: 2790947
Number of extensions: 9462917
Number of successful extensions: 24640
Number of sequences better than 10.0: 511
Number of HSP's better than 10.0 without gapping: 370
Number of HSP's successfully gapped in prelim test: 141
Number of HSP's that attempted gapping in prelim test: 24211
Number of HSP's gapped (non-prelim): 522
length of query: 131
length of database: 848,049,833
effective HSP length: 107
effective length of query: 24
effective length of database: 549,418,504
effective search space: 13186044096
effective search space used: 13186044096
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC144345.5


