

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144345.11 - phase: 0
(339 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
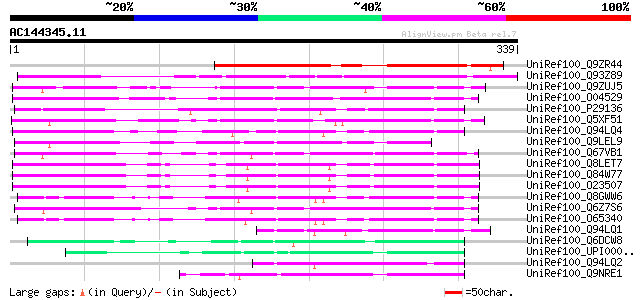
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9ZR44 MtN9 protein [Medicago truncatula] 216 9e-55
UniRef100_Q93Z89 Matrix metalloproteinase MMP2 [Glycine max] 187 5e-46
UniRef100_Q9ZUJ5 T2K10.2 protein [Arabidopsis thaliana] 134 5e-30
UniRef100_O04529 F20P5.11 protein [Arabidopsis thaliana] 128 2e-28
UniRef100_P29136 Metalloendoproteinase 1 precursor [Glycine max] 122 2e-26
UniRef100_Q5XF51 At1g24140 [Arabidopsis thaliana] 119 9e-26
UniRef100_Q94LQ4 Putative metalloproteinase [Oryza sativa] 117 4e-25
UniRef100_Q9LEL9 Matrix metalloproteinase [Cucumis sativus] 116 1e-24
UniRef100_Q67VB1 Putative metalloproteinase [Oryza sativa] 113 6e-24
UniRef100_Q8LET7 Proteinase like protein [Arabidopsis thaliana] 113 8e-24
UniRef100_Q84W77 Hypothetical protein At4g16640 [Arabidopsis tha... 112 1e-23
UniRef100_O23507 Proteinase like protein [Arabidopsis thaliana] 112 1e-23
UniRef100_Q8GWW6 Putative metalloproteinase [Arabidopsis thaliana] 111 3e-23
UniRef100_Q6Z7S6 Putative zinc metalloproteinase [Oryza sativa] 110 5e-23
UniRef100_O65340 Metalloproteinase [Arabidopsis thaliana] 108 2e-22
UniRef100_Q94LQ1 Putative metalloproteinase [Oryza sativa] 88 4e-16
UniRef100_Q6DCW8 Mmp24-prov protein [Xenopus laevis] 74 6e-12
UniRef100_UPI00003AA134 UPI00003AA134 UniRef100 entry 72 2e-11
UniRef100_Q94LQ2 Putative metalloproteinase [Oryza sativa] 72 2e-11
UniRef100_Q9NRE1 Matrix metalloproteinase-26 precursor [Homo sap... 67 7e-10
>UniRef100_Q9ZR44 MtN9 protein [Medicago truncatula]
Length = 191
Score = 216 bits (549), Expect = 9e-55
Identities = 125/198 (63%), Positives = 135/198 (68%), Gaps = 29/198 (14%)
Query: 138 KWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRVLNFSETTSYDDADIKIGFY 197
KWFPKGTK+LTYGFLP K S+D + FRNAFTRWSQTTRVL FSE TSYDDADIKIGFY
Sbjct: 1 KWFPKGTKELTYGFLPESKISIDKVNVFRNAFTRWSQTTRVLKFSEATSYDDADIKIGFY 60
Query: 198 HI-YNNDIVDDVVVGDSFISRNLDSNVKSGMIRLERSKFWVSPTTTYFKKWELQEFDLET 256
+I YN+ V DVVV D FI N++S IRLE SK W DLET
Sbjct: 61 NISYNSKEVIDVVVSDFFI------NLRSFTIRLEASKVW----------------DLET 98
Query: 257 VVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDSDNQAIQQLYTNTAKANPNSDH 316
V MHQIGHLLGLDHSSD ESIMYP I PL +KKVQIT SDNQAIQQLYT + N + D
Sbjct: 99 VAMHQIGHLLGLDHSSDVESIMYPTIVPLHQKKVQITVSDNQAIQQLYTK--QTNQDRDE 156
Query: 317 SGCF----KLFESSMFLL 330
G F FESS LL
Sbjct: 157 LGFFDYSGDFFESSSGLL 174
>UniRef100_Q93Z89 Matrix metalloproteinase MMP2 [Glycine max]
Length = 357
Score = 187 bits (474), Expect = 5e-46
Identities = 124/336 (36%), Positives = 174/336 (50%), Gaps = 59/336 (17%)
Query: 6 LSQIKKHLSTFGYFRQFLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKISLPR 65
+S IK +LS +GY + + E ISAIKTYQQ++ LQ TG LN ETLQ++
Sbjct: 78 VSLIKDYLSNYGYIESSGPLSNSMDQETIISAIKTYQQYYCLQPTGKLNNETLQQM---- 133
Query: 66 CGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRYEYGF 125
S LRCG+PD+ +Y F
Sbjct: 134 --------------------------------------------SFLRCGVPDINIDYNF 149
Query: 126 DVGSDVSFPK-GNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRVLNFSET 184
++S+PK G++WFP LTYGFLP + +M + FR++F RW+Q + VLN +ET
Sbjct: 150 -TDDNMSYPKAGHRWFPH--TNLTYGFLPENQIPANMTKVFRDSFARWAQASGVLNLTET 206
Query: 185 TSYDDADIKIGFYHIYNNDIVDDVVVGDSFISRNLDSNVKSGMIRLE-RSKFWVSPTTTY 243
T YD+ADI++GFY+ I D V G S I DS K G+I L+ +K W P+
Sbjct: 207 T-YDNADIQVGFYNFTYLGI-DIEVYGGSLIFLQPDST-KKGVILLDGTNKLWALPSENG 263
Query: 244 FKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDSDNQAIQQL 303
WE DLE+ MH+IGHLLGLDHS+ ++S+MYP I P ++KVQ++ SD +Q
Sbjct: 264 RLSWEEGVLDLESAAMHEIGHLLGLDHSNKEDSVMYPCILPSHQRKVQLSKSDKTNVQHQ 323
Query: 304 YTNTAKANPNSDHSGCFKLFESSMFLLAFCPLVLLL 339
+ N ++ H G + + L F L+LLL
Sbjct: 324 FAN---VEDSAGHVGRLGVSLITTLSLVFAYLLLLL 356
>UniRef100_Q9ZUJ5 T2K10.2 protein [Arabidopsis thaliana]
Length = 360
Score = 134 bits (336), Expect = 5e-30
Identities = 111/329 (33%), Positives = 156/329 (46%), Gaps = 75/329 (22%)
Query: 3 IQGLSQIKKHLSTFGYFR---QFLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQ 59
I GLS++K++ FGY FDDVL SAI TYQ+ FNL+VTG L++ TL+
Sbjct: 57 INGLSKLKQYFRRFGYITTTGNCTDDFDDVLQ----SAINTYQKNFNLKVTGKLDSSTLR 112
Query: 60 KISLPRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDM 119
+I PRCG PD+ VS G K + T+K Y F PGK
Sbjct: 113 QIVKPRCGNPDLI--------DGVSEMNGGKIL-RTTEK--YSFFPGKP----------- 150
Query: 120 RYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRVL 179
+W PK + LTY F P + ++ F AFTRW++ T L
Sbjct: 151 ------------------RW-PKRKRDLTYAFAPQNNLTDEVKRVFSRAFTRWAEVTP-L 190
Query: 180 NFSETTSYDDADIKIGFYHIYNNDIVDDVVVGDSFISRNLDSNVKSGMIRLERSKFW--- 236
NF+ + S ADI IGF+ + + D + + S+ +GM+ L+ + W
Sbjct: 191 NFTRSESILRADIVIGFF---SGEHGDGEPFDGAMGTLAHASSPPTGMLHLDGDEDWLIS 247
Query: 237 -------VSPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKK 289
+ P TT DLE+V +H+IGHLLGL HSS +++IM+P I ++K
Sbjct: 248 NGEISRRILPVTTVV--------DLESVAVHEIGHLLGLGHSSVEDAIMFPAISG-GDRK 298
Query: 290 VQITDSDNQAIQQLYTNTAKANPNSDHSG 318
V++ D + IQ LY NPN D G
Sbjct: 299 VELAKDDIEGIQHLY----GGNPNGDGGG 323
>UniRef100_O04529 F20P5.11 protein [Arabidopsis thaliana]
Length = 378
Score = 128 bits (322), Expect = 2e-28
Identities = 105/313 (33%), Positives = 149/313 (47%), Gaps = 42/313 (13%)
Query: 3 IQGLSQIKKHLSTFGYFRQ-FLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKI 61
+ GL +IKK+ FGY + F F D D+ +A++ YQ FNL VTG L+ T+Q I
Sbjct: 56 VDGLYRIKKYFQRFGYIPETFSGNFTDDFDDILKAAVELYQTNFNLNVTGELDALTIQHI 115
Query: 62 SLPRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRY 121
+PRCG PD+ G+ + K F + + K+++L
Sbjct: 116 VIPRCGNPDVVN------GTSLMHGGRRKTFEVNFSRTHLHAV--KRYTL---------- 157
Query: 122 EYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRVLNF 181
FP +W P+ + LTY F P + ++ F AF RWS T LNF
Sbjct: 158 -----------FPGEPRW-PRNRRDLTYAFDPKNPLTEEVKSVFSRAFGRWSDVT-ALNF 204
Query: 182 SETTSYDDADIKIGFYHIYNNDIVD-DVVVGDSFISRNLDSNVKSGMIRLERSKFWVSPT 240
+ + S+ +DI IGFY + D D V+G + + SG L+ + WV
Sbjct: 205 TLSESFSTSDITIGFYTGDHGDGEPFDGVLGTLAHA----FSPPSGKFHLDADENWVVSG 260
Query: 241 TTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDSDNQAI 300
DLE+V +H+IGHLLGL HSS +ESIMYP I ++KV +T+ D + I
Sbjct: 261 DLDSFLSVTAAVDLESVAVHEIGHLLGLGHSSVEESIMYPTI-TTGKRKVDLTNDDVEGI 319
Query: 301 QQLYTNTAKANPN 313
Q LY ANPN
Sbjct: 320 QYLY----GANPN 328
>UniRef100_P29136 Metalloendoproteinase 1 precursor [Glycine max]
Length = 305
Score = 122 bits (305), Expect = 2e-26
Identities = 93/309 (30%), Positives = 135/309 (43%), Gaps = 67/309 (21%)
Query: 4 QGLSQIKKHLSTFGYFRQFLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKISL 63
+GLS +K + GY FDD D+ +SAIKTYQ+ +NL VTG + TL++I
Sbjct: 52 KGLSNVKNYFHHLGYIPN-APHFDDNFDDTLVSAIKTYQKNYNLNVTGKFDINTLKQIMT 110
Query: 64 PRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDM---- 119
P RCG+PD+
Sbjct: 111 P------------------------------------------------RCGVPDIIINT 122
Query: 120 RYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRVL 179
F + SD +F K + GT +LTY F P + AF++W+ +
Sbjct: 123 NKTTSFGMISDYTFFKDMPRWQAGTTQLTYAFSPEPRLDDTFKSAIARAFSKWTPVVNIA 182
Query: 180 NFSETTSYDDADIKIGFYHIYNNDIVD----DVVVGDSFISRNLDSNVKSGMIRLERSKF 235
F ETTSY+ A+IKI F + D ++G +F + G + ++
Sbjct: 183 -FQETTSYETANIKILFASKNHGDPYPFDGPGGILGHAFAPTD-------GRCHFDADEY 234
Query: 236 WVSPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDS 295
WV+ + K FDLE+V +H+IGHLLGL HSSD +IMYP I P + +KV +
Sbjct: 235 WVA-SGDVTKSPVTSAFDLESVAVHEIGHLLGLGHSSDLRAIMYPSIPP-RTRKVNLAQD 292
Query: 296 DNQAIQQLY 304
D I++LY
Sbjct: 293 DIDGIRKLY 301
>UniRef100_Q5XF51 At1g24140 [Arabidopsis thaliana]
Length = 384
Score = 119 bits (299), Expect = 9e-26
Identities = 102/324 (31%), Positives = 147/324 (44%), Gaps = 51/324 (15%)
Query: 2 KIQGLSQIKKHLSTFGYFRQFLLK--FDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQ 59
K GL +K++ FGY + L F D D+ +A++ YQ+ F L VTG L+ TL+
Sbjct: 57 KYDGLYMLKQYFQHFGYITETNLSGNFTDDFDDILKNAVEMYQRNFQLNVTGVLDELTLK 116
Query: 60 KISLPRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLP-GKKFSLLRCGIPD 118
+ +PRCG PD+ G G K F G++F ++
Sbjct: 117 HVVIPRCGNPDV--------------VNGTSTMHSGRKTFEVSFAGRGQRFHAVK----- 157
Query: 119 MRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRV 178
Y F FP +W P+ + LTY F P + ++ F AFTRW + T
Sbjct: 158 ---HYSF-------FPGEPRW-PRNRRDLTYAFDPRNALTEEVKSVFSRAFTRWEEVTP- 205
Query: 179 LNFSETTSYDDADIKIGFYHIYNNDIVDDVVVGDSFIS--RNLDS--NVKSGMIRLERSK 234
L F+ + +DI IGFY + D G+ F R L + +G L+ +
Sbjct: 206 LTFTRVERFSTSDISIGFYSGEHGD-------GEPFDGPMRTLAHAFSPPTGHFHLDGEE 258
Query: 235 FWVSPTTTYFKKWELQE-FDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQIT 293
W+ + E DLE+V +H+IGHLLGL HSS + SIMYP I +KV +T
Sbjct: 259 NWIVSGEGGDGFISVSEAVDLESVAVHEIGHLLGLGHSSVEGSIMYPTI-RTGRRKVDLT 317
Query: 294 DSDNQAIQQLYTNTAKANPNSDHS 317
D + +Q LY ANPN + S
Sbjct: 318 TDDVEGVQYLY----GANPNFNGS 337
>UniRef100_Q94LQ4 Putative metalloproteinase [Oryza sativa]
Length = 355
Score = 117 bits (293), Expect = 4e-25
Identities = 94/316 (29%), Positives = 143/316 (44%), Gaps = 68/316 (21%)
Query: 3 IQGLSQIKKHLSTFGYFRQFLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKIS 62
+ GL+++K++L+ FGY + D DE A++ YQ F+L VTG L+ TL +I
Sbjct: 51 VTGLAELKRYLARFGYMAKPGRDTTDAFDEHLEVAVRRYQTRFSLPVTGRLDNATLDQIM 110
Query: 63 LPRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRYE 122
PRCG+ D DV P ++ PG + ++
Sbjct: 111 SPRCGVGD----------DDVERP------------VSVALSPGAQGGVV---------- 138
Query: 123 YGFDVGSDVSFPKGNKWFPKGTKKL----------TYGFLPGKKFSLDMIEGFRNAFTRW 172
S +F KG + + + T G+LP F+ AF RW
Sbjct: 139 ------SRFTFFKGEPRWTRSDPPIVLSYAVSPTATVGYLPPAAVRAV----FQRAFARW 188
Query: 173 SQTTRVLNFSETTSYDDADIKIGFYHIYNNDIVDDV----VVGDSFISRNLDSNVKSGMI 228
++T V F ET Y+ ADIK+GFY + D V ++G +F +N G +
Sbjct: 189 ARTIPV-GFVETDDYEAADIKVGFYAGNHGDGVPFDGPLGILGHAFSPKN-------GRL 240
Query: 229 RLERSKFWVSPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEK 288
L+ S+ W + DLE+V H+IGH+LGL HS+ ++MYP I P +EK
Sbjct: 241 HLDASEHWA---VDFDVDATASAIDLESVATHEIGHVLGLGHSASPRAVMYPSIKP-REK 296
Query: 289 KVQITDSDNQAIQQLY 304
KV++T D + +Q LY
Sbjct: 297 KVRLTVDDVEGVQALY 312
>UniRef100_Q9LEL9 Matrix metalloproteinase [Cucumis sativus]
Length = 320
Score = 116 bits (290), Expect = 1e-24
Identities = 91/285 (31%), Positives = 127/285 (43%), Gaps = 53/285 (18%)
Query: 4 QGLSQIKKHLSTFGYFRQFLLK-----FDDVLDEETISAIKTYQQFFNLQVTGNLNTETL 58
QG+ QIKK+L FGY + K FDD D SA+KTYQ NL +G L++ T+
Sbjct: 61 QGIHQIKKYLQRFGYITTNIQKHSNPIFDDTFDHILESALKTYQTNHNLAPSGILDSNTI 120
Query: 59 QKISLPRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPD 118
+I++PRCG+ D+ K TKK F
Sbjct: 121 AQIAMPRCGVQDVIKN-------------------KKTKKRNQNFTNNGHTH-------- 153
Query: 119 MRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRV 178
F S +F +GN +P L+YGFLP + +D I+ AF++WS T
Sbjct: 154 ------FHKVSHFTFFEGNLKWPSSKLHLSYGFLPN--YPIDAIKPVSRAFSKWSLNTH- 204
Query: 179 LNFSETTSYDDADIKIGFYHIYNNDIVDDVVVGDSFISRNLDSNVKSGMIRLERSKFW-V 237
FS Y ADIKI F + D VG ++ G + + W V
Sbjct: 205 FKFSHVADYRKADIKISFERGEHGDNAPFDGVGGVLAHAYAPTD---GRLHFDGDDAWSV 261
Query: 238 SPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLI 282
+ YF D+ETV +H+IGH+LGL HS+ +E+IM+P I
Sbjct: 262 GAISGYF--------DVETVALHEIGHILGLQHSTIEEAIMFPSI 298
>UniRef100_Q67VB1 Putative metalloproteinase [Oryza sativa]
Length = 371
Score = 113 bits (283), Expect = 6e-24
Identities = 99/318 (31%), Positives = 139/318 (43%), Gaps = 59/318 (18%)
Query: 4 QGLSQIKKHLSTFGYFR-----QFLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETL 58
QGL ++K +L FGY F+D+ D + AIK YQ F L VTG+L+ T+
Sbjct: 60 QGLGKLKDYLWHFGYLSYPSSSSLSPSFNDLFDADMELAIKMYQGNFGLDVTGDLDAATV 119
Query: 59 QKISLPRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPD 118
++ PRCG+ D+ GT + G G
Sbjct: 120 SQMMAPRCGVADV---------------------VNGTSTMGGG------------GGVR 146
Query: 119 MRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLD---MIEGFRNAFTRWSQT 175
R Y + FP +W P+ L Y + S+D + + F +AF RWS
Sbjct: 147 GRGLYSY-------FPGSPRW-PRSRTTLRYAITATSQTSIDRATLSKVFASAFARWSAA 198
Query: 176 TRVLNFSETTSYDDADIKIGFYHIYNNDIVDDVVVGDSFISRNLDSNVKSGMIRLERSKF 235
T LNF+E S DADI IGFY D D + + +G + L+ S+
Sbjct: 199 T-TLNFTEAASAADADITIGFY---GGDHGDGEAFDGPLGTLAHAFSPTNGRLHLDASEA 254
Query: 236 WVSPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDS 295
WV+ + DLE+V +H+IGH+LGL HSS +SIM+P + + KKV +
Sbjct: 255 WVAGGDVT-RASSNAAVDLESVAVHEIGHILGLGHSSAADSIMFPTLTS-RTKKVNLATD 312
Query: 296 DNQAIQQLYTNTAKANPN 313
D IQ LY N NPN
Sbjct: 313 DVAGIQGLYGN----NPN 326
>UniRef100_Q8LET7 Proteinase like protein [Arabidopsis thaliana]
Length = 364
Score = 113 bits (282), Expect = 8e-24
Identities = 99/320 (30%), Positives = 139/320 (42%), Gaps = 66/320 (20%)
Query: 3 IQGLSQIKKHLSTFGYFRQFLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKIS 62
+ G+S++K++L FGY + F DV D SAI YQ+ L +TG L+T T+ +S
Sbjct: 67 VSGVSELKRYLHRFGYVKDGSEIFSDVFDGPLESAISLYQENLGLPITGRLDTSTVTLMS 126
Query: 63 LPRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRYE 122
LPRCG+ D D F T TY GK
Sbjct: 127 LPRCGVXDTHMTINND-------------FLHTTAHYTY--FNGK--------------- 156
Query: 123 YGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKF----SLDMIEGFRNAFTRWSQTTRV 178
PK N+ LTY K S D+ FR AF++WS V
Sbjct: 157 -----------PKWNR------DTLTYAISKTHKLDYLTSEDVKTVFRRAFSQWSSVIPV 199
Query: 179 LNFSETTSYDDADIKIGFYHIYNND-IVDDVVVGD---SFISRNLDSNVKSGMIRLERSK 234
+F E + AD+KIGFY + D + D V+G +F N G + L+ ++
Sbjct: 200 -SFEEVDDFTTADLKIGFYAGDHGDGLPFDGVLGTLAHAFAPEN-------GRLHLDAAE 251
Query: 235 FWVSPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITD 294
W+ K DLE+V H+IGHLLGL HSS + ++MYP + P + KKV +T
Sbjct: 252 TWIVDDD--LKGSSEVAVDLESVATHEIGHLLGLGHSSQESAVMYPSLRP-RTKKVDLTV 308
Query: 295 SDNQAIQQLYTNTAKANPNS 314
D + +LY K +S
Sbjct: 309 DDVAGVLKLYGPNPKLRLDS 328
>UniRef100_Q84W77 Hypothetical protein At4g16640 [Arabidopsis thaliana]
Length = 364
Score = 112 bits (281), Expect = 1e-23
Identities = 99/320 (30%), Positives = 138/320 (42%), Gaps = 66/320 (20%)
Query: 3 IQGLSQIKKHLSTFGYFRQFLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKIS 62
+ G+S++K++L FGY F DV D SAI YQ+ L +TG L+T T+ +S
Sbjct: 67 VSGVSELKRYLHRFGYVNDGSEIFSDVFDGPLESAISLYQENLGLPITGRLDTSTVTLMS 126
Query: 63 LPRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRYE 122
LPRCG+ D D F T TY GK
Sbjct: 127 LPRCGVSDTHMTINND-------------FLHTTAHYTY--FNGK--------------- 156
Query: 123 YGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKF----SLDMIEGFRNAFTRWSQTTRV 178
PK N+ LTY K S D+ FR AF++WS V
Sbjct: 157 -----------PKWNR------DTLTYAISKTHKLDYLTSEDVKTVFRRAFSQWSSVIPV 199
Query: 179 LNFSETTSYDDADIKIGFYHIYNND-IVDDVVVGD---SFISRNLDSNVKSGMIRLERSK 234
+F E + AD+KIGFY + D + D V+G +F N G + L+ ++
Sbjct: 200 -SFEEVDDFTTADLKIGFYAGDHGDGLPFDGVLGTLAHAFAPEN-------GRLHLDAAE 251
Query: 235 FWVSPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITD 294
W+ K DLE+V H+IGHLLGL HSS + ++MYP + P + KKV +T
Sbjct: 252 TWIVDDD--LKGSSEVAVDLESVATHEIGHLLGLGHSSQESAVMYPSLRP-RTKKVDLTV 308
Query: 295 SDNQAIQQLYTNTAKANPNS 314
D + +LY K +S
Sbjct: 309 DDVAGVLKLYGPNPKLRLDS 328
>UniRef100_O23507 Proteinase like protein [Arabidopsis thaliana]
Length = 364
Score = 112 bits (281), Expect = 1e-23
Identities = 99/320 (30%), Positives = 138/320 (42%), Gaps = 66/320 (20%)
Query: 3 IQGLSQIKKHLSTFGYFRQFLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKIS 62
+ G+S++K++L FGY F DV D SAI YQ+ L +TG L+T T+ +S
Sbjct: 67 VSGVSELKRYLHRFGYVNDGSEIFSDVFDGPLESAISLYQENLGLPITGRLDTSTVTLMS 126
Query: 63 LPRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRYE 122
LPRCG+ D D F T TY GK
Sbjct: 127 LPRCGVSDTHMTINND-------------FLHTTAHYTY--FNGK--------------- 156
Query: 123 YGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKF----SLDMIEGFRNAFTRWSQTTRV 178
PK N+ LTY K S D+ FR AF++WS V
Sbjct: 157 -----------PKWNR------DTLTYAISKTHKLDYLTSEDVKTVFRRAFSQWSSVIPV 199
Query: 179 LNFSETTSYDDADIKIGFYHIYNND-IVDDVVVGD---SFISRNLDSNVKSGMIRLERSK 234
+F E + AD+KIGFY + D + D V+G +F N G + L+ ++
Sbjct: 200 -SFEEVDDFTTADLKIGFYAGDHGDGLPFDGVLGTLAHAFAPEN-------GRLHLDAAE 251
Query: 235 FWVSPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITD 294
W+ K DLE+V H+IGHLLGL HSS + ++MYP + P + KKV +T
Sbjct: 252 TWIVDDD--LKGSSEVAVDLESVATHEIGHLLGLGHSSQESAVMYPSLRP-RTKKVDLTV 308
Query: 295 SDNQAIQQLYTNTAKANPNS 314
D + +LY K +S
Sbjct: 309 DDVAGVLKLYGPNPKLRLDS 328
>UniRef100_Q8GWW6 Putative metalloproteinase [Arabidopsis thaliana]
Length = 342
Score = 111 bits (277), Expect = 3e-23
Identities = 102/316 (32%), Positives = 144/316 (45%), Gaps = 73/316 (23%)
Query: 6 LSQIKKHLSTFGYFRQFLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKISLPR 65
+ +IK+HL +GY Q + DDV E+ A+ YQ+ L +TG +++TL +I LPR
Sbjct: 50 IPEIKRHLQQYGYLPQNK-ESDDVSFEQ---ALVRYQKNLGLPITGKPDSDTLSQILLPR 105
Query: 66 CGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRYEYGF 125
CG PD DV PK T F GKK+
Sbjct: 106 CGFPD-----------DVE--------PK-----TAPFHTGKKY---------------- 125
Query: 126 DVGSDVSFPKGNKWFPKGTKKLTYGF----LPGKKFSLDMIEGFRNAFTRWSQTTRVLNF 181
V FP +W KLTY F L D+ FR AF +W+ V +F
Sbjct: 126 -----VYFPGRPRWTRDVPLKLTYAFSQENLTPYLAPTDIRRVFRRAFGKWASVIPV-SF 179
Query: 182 SETTSYDDADIKIGFYHIYNND--IVDDV--VVGDSFISRNLDSNVKSGMIRLERSKFWV 237
ET Y ADIKIGF++ + D D V V+ +F N G + L++++ W
Sbjct: 180 IETEDYVIADIKIGFFNGDHGDGEPFDGVLGVLAHTFSPEN-------GRLHLDKAETWA 232
Query: 238 SPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDSDN 297
+ ++ DLE+V +H+IGH+LGL HSS K++ MYP + P + KKV + D
Sbjct: 233 ---VDFDEEKSSVAVDLESVAVHEIGHVLGLGHSSVKDAAMYPTLKP-RSKKVNLNMDDV 288
Query: 298 QAIQQLYTNTAKANPN 313
+Q LY NPN
Sbjct: 289 VGVQSLY----GTNPN 300
>UniRef100_Q6Z7S6 Putative zinc metalloproteinase [Oryza sativa]
Length = 372
Score = 110 bits (275), Expect = 5e-23
Identities = 95/316 (30%), Positives = 135/316 (42%), Gaps = 54/316 (17%)
Query: 4 QGLSQIKKHLSTFGYFRQ--FLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKI 61
+GL ++K +LS FGY + D D+ +AI YQ+ F L TG L+T+T+ ++
Sbjct: 60 EGLGRLKDYLSHFGYLPPPPSSSPYSDAFDDSLEAAIAAYQRNFGLNATGELDTDTVDQM 119
Query: 62 SLPRCGIPDM-RYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMR 120
PRCG+ D+ D S + +G R
Sbjct: 120 VAPRCGVADVINGTSTMDRNSSAAALRG-------------------------------R 148
Query: 121 YEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLD---MIEGFRNAFTRWSQTTR 177
+ Y + FP G W P + L Y S+D + F AF+RW+ TR
Sbjct: 149 HLYSY-------FPGGPMW-PPFRRNLRYAITATSATSIDRATLSAVFARAFSRWAAATR 200
Query: 178 VLNFSETTSYDDADIKIGFYHIYNNDIVDDVVVGDSFISRNLDSNVKSGMIRLERSKFWV 237
L F+E +S +ADI IGFY + D D + + G L+ ++ WV
Sbjct: 201 -LQFTEVSSASNADITIGFY---SGDHGDGEAFDGPLGTLAHAFSPTDGRFHLDAAEAWV 256
Query: 238 SPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDSDN 297
+ DLE+V +H+IGHLLGL HSS +SIMYP I +KV + D
Sbjct: 257 ASGDVSTSSSFGTAVDLESVAVHEIGHLLGLGHSSVPDSIMYPTI-RTGTRKVDLESDDV 315
Query: 298 QAIQQLYTNTAKANPN 313
IQ LY NPN
Sbjct: 316 LGIQSLY----GTNPN 327
>UniRef100_O65340 Metalloproteinase [Arabidopsis thaliana]
Length = 341
Score = 108 bits (270), Expect = 2e-22
Identities = 101/315 (32%), Positives = 142/315 (45%), Gaps = 72/315 (22%)
Query: 6 LSQIKKHLSTFGYFRQFLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKISLPR 65
+ +IK+HL +GY Q + DDV E+ A+ YQ+ L +TG +++TL +I LPR
Sbjct: 50 IPEIKRHLQQYGYLPQNK-ESDDVSFEQ---ALVRYQKNLGLPITGKPDSDTLSQILLPR 105
Query: 66 CGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRYEYGF 125
CG PD DV PK T F GKK+
Sbjct: 106 CGFPD-----------DVE--------PK-----TAPFHTGKKY---------------- 125
Query: 126 DVGSDVSFPKGNKWFPKGTKKLTYGFLPGKK---FSLDMIEGFRNAFTRWSQTTRVLNFS 182
V FP +W KLTY F + I FR AF W+ V +F
Sbjct: 126 -----VYFPGRPRWTRDVPLKLTYAFSQENLTPYLAPTDIRVFRRAFGEWASVIPV-SFI 179
Query: 183 ETTSYDDADIKIGFYHIYNND--IVDDV--VVGDSFISRNLDSNVKSGMIRLERSKFWVS 238
ET Y ADIKIGF++ + D D V V+ +F N G + L++++ W
Sbjct: 180 ETEDYVIADIKIGFFNGDHGDGEPFDGVLGVLAHTFSPEN-------GRLHLDKAETWA- 231
Query: 239 PTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDSDNQ 298
+ ++ DLE+V +H+IGH+LGL HSS K++ MYP + P + KKV + D
Sbjct: 232 --VDFDEEKSSVAVDLESVAVHEIGHVLGLGHSSVKDAAMYPTLKP-RSKKVNLNMDDVV 288
Query: 299 AIQQLYTNTAKANPN 313
+Q LY NPN
Sbjct: 289 GVQSLY----GTNPN 299
>UniRef100_Q94LQ1 Putative metalloproteinase [Oryza sativa]
Length = 289
Score = 87.8 bits (216), Expect = 4e-16
Identities = 64/167 (38%), Positives = 84/167 (49%), Gaps = 21/167 (12%)
Query: 166 RNAFTRWSQTTRVLNFSETTSYDDADIKIGFYHIYNN---------DIVDDVVVGDSFIS 216
R+AF RW+ + F E YD ADIK+GFY +Y + D DD GD
Sbjct: 84 RSAFARWADVIP-MRFLEAERYDAADIKVGFY-LYTDGRCDGCACIDSDDDDDDGDDCEG 141
Query: 217 RNLDSNV--KSGMIRLERSKFWVSPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDK 274
S++ KSG I L + W T DLE+V H+IGH+LGLDHSS +
Sbjct: 142 VLAHSSMPEKSGQIHLHAAHRW---TVNLAADTAPLAVDLESVAAHEIGHVLGLDHSSSR 198
Query: 275 ESIMYPLIDPLQEKKVQITDSDNQAIQQLYTNTAKANPNSDHSGCFK 321
S+MYP I +E+KV++T D IQ+LY ANP+ FK
Sbjct: 199 SSMMYPFIS-CRERKVRLTTDDVHGIQELY----GANPHFSFGAYFK 240
>UniRef100_Q6DCW8 Mmp24-prov protein [Xenopus laevis]
Length = 576
Score = 73.9 bits (180), Expect = 6e-12
Identities = 71/300 (23%), Positives = 117/300 (38%), Gaps = 63/300 (21%)
Query: 13 LSTFGYFRQFLLKFDDVLDEETI-SAIKTYQQFFNLQVTGNLNTETLQKISLPRCGIPDM 71
L +GY L+ + +++ SAI Q+F+ L+VTG +++T + + PRCG+PD
Sbjct: 30 LQQYGYLPPGDLRTHTLRSPQSMNSAISAMQKFYGLKVTGTFDSDTEKAMKRPRCGVPD- 88
Query: 72 RYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRYEYGFDVGSDV 131
++G ++ ++V K++++
Sbjct: 89 --KFGAEIKANVR---------------------RKRYAI-------------------- 105
Query: 132 SFPKGNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRVLNFSETTSYD--- 188
+G KW K + P K E R AF W T L F E D
Sbjct: 106 ---QGLKWQHKDITFCIQNYTP-KIGEYSTHEAIRRAFKVWESVT-PLRFQEVRYVDIQD 160
Query: 189 ----DADIKIGFYHIYNNDIVDDVVVGDSFISRNLDSNVKSGMIRLERSKFWVSPTTTYF 244
ADI + F ++ D G G + ++ W +
Sbjct: 161 GYTKHADIMLFFAEGFHGDSTPFDGEGGFLAHAYFPGTGIGGDTHFDSAEPWTA------ 214
Query: 245 KKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDSDNQAIQQLY 304
+ +L+ DL V +H++GH LGL+HS+D +IM P + Q+ D D + IQQLY
Sbjct: 215 RNEDLEGNDLFLVAVHELGHALGLEHSNDPSAIMAPFYQWMDTTNFQLPDDDRRGIQQLY 274
>UniRef100_UPI00003AA134 UPI00003AA134 UniRef100 entry
Length = 467
Score = 72.4 bits (176), Expect = 2e-11
Identities = 69/267 (25%), Positives = 93/267 (33%), Gaps = 68/267 (25%)
Query: 38 IKTYQQFFNLQVTGNLNTETLQKISLPRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTK 97
I+ Q FF L+VTG LN +T+ + PR
Sbjct: 64 IREMQSFFGLEVTGELNRKTMDMMKQPR-------------------------------- 91
Query: 98 KLTYGFLPGKKFSLLRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKF 157
CGIPD+R S +FP+ KW + + P
Sbjct: 92 ----------------CGIPDVR--------SYSTFPRNPKWKKEDVTYRILNYTPDM-L 126
Query: 158 SLDMIEGFRNAFTRWSQTTRVLNFSETTSYDDADIKIGFYHIYNNDIVDDVVVGDSFISR 217
D+ E AF WS T L F S DADI I F +ND + G S
Sbjct: 127 QADVDEAIAKAFQLWSSVTP-LRFIRLYS-GDADIMISFASGCHNDFIPFDGPGGSVAHA 184
Query: 218 NLDSNVKSGMIRLERSKFWVSPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESI 277
G + + W T Y +L VV H++GH LGL HS+D ++
Sbjct: 185 YAPGKDFGGDAHFDEDETWTKSTEGY---------NLFIVVAHELGHSLGLSHSNDPGAL 235
Query: 278 MYPLIDPLQEKKVQITDSDNQAIQQLY 304
MYP K+ + D IQ +Y
Sbjct: 236 MYPNYAYTDPKEFLLPQDDIDGIQAIY 262
>UniRef100_Q94LQ2 Putative metalloproteinase [Oryza sativa]
Length = 266
Score = 72.4 bits (176), Expect = 2e-11
Identities = 51/146 (34%), Positives = 74/146 (49%), Gaps = 9/146 (6%)
Query: 163 EGFRNAFTRWSQTTRVLNFSETTSYDDADIKIGFY-HIYNN--DIVDDVVVGDSFISRNL 219
E FR+A RW++ T L F+E Y++ADI++GFY H + D V G +
Sbjct: 66 EAFRSALARWAEVTP-LRFAEAARYEEADIRVGFYLHTADGKCDACGCVCKGGGEEALAH 124
Query: 220 DSNVKSGMIRLERSKFW-VSPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIM 278
+ G I L ++ W V+ DLE+V +H+IGH LGL HSS + S+M
Sbjct: 125 AHPPQDGRIHLHAARKWAVTNVAGAGGDAPPLAVDLESVAVHEIGHALGLGHSSSESSMM 184
Query: 279 YPLIDPLQEKKVQITDSDNQAIQQLY 304
Y KV +TD D + +Q+LY
Sbjct: 185 Y----RHYRGKVSLTDDDVKGVQELY 206
>UniRef100_Q9NRE1 Matrix metalloproteinase-26 precursor [Homo sapiens]
Length = 261
Score = 67.0 bits (162), Expect = 7e-10
Identities = 56/193 (29%), Positives = 84/193 (43%), Gaps = 24/193 (12%)
Query: 114 CGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFL--PGKKFSLDMIEGFRNAFTR 171
CG+PD GSD S G + K T LTY + P + + NA +
Sbjct: 82 CGVPD---------GSDTSISPGRCKWNKHT--LTYRIINYPHDMKPSAVKDSIYNAVSI 130
Query: 172 WSQTTRVLNFSETTSYDDADIKIGFYHIYNNDIVDDVVVGDSFISRNLDSNVKSGMIRLE 231
WS T ++ + DADIK+ F+ + D G L ++ G++ +
Sbjct: 131 WSNVTPLI--FQQVQNGDADIKVSFWQWAHEDGWPFDGPGGILGHAFLPNSGNPGVVHFD 188
Query: 232 RSKFWVSPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQ 291
+++ W + T Y +L V H+IGH LGL HS ++ SIMYP + Q
Sbjct: 189 KNEHWSASDTGY---------NLFLVATHEIGHSLGLQHSGNQSSIMYPTYWYHDPRTFQ 239
Query: 292 ITDSDNQAIQQLY 304
++ D Q IQ LY
Sbjct: 240 LSADDIQRIQHLY 252
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.140 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 593,330,394
Number of Sequences: 2790947
Number of extensions: 26198705
Number of successful extensions: 57061
Number of sequences better than 10.0: 520
Number of HSP's better than 10.0 without gapping: 460
Number of HSP's successfully gapped in prelim test: 60
Number of HSP's that attempted gapping in prelim test: 56109
Number of HSP's gapped (non-prelim): 918
length of query: 339
length of database: 848,049,833
effective HSP length: 128
effective length of query: 211
effective length of database: 490,808,617
effective search space: 103560618187
effective search space used: 103560618187
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 75 (33.5 bits)
Medicago: description of AC144345.11


