

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC143339.7 - phase: 0
(358 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
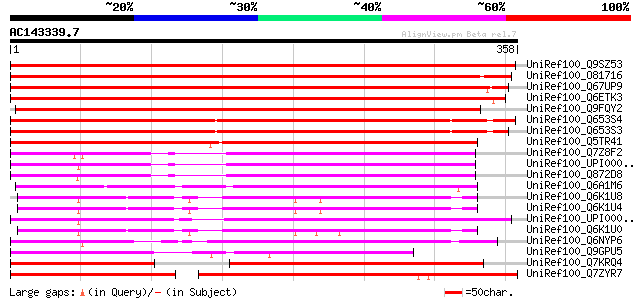
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SZ53 Protein phosphatase 2C-like protein [Arabidopsi... 536 e-151
UniRef100_O81716 Hypothetical protein At2g25070 [Arabidopsis tha... 524 e-147
UniRef100_Q67UP9 Hypothetical protein P0453H04.39-1 [Oryza sativa] 523 e-147
UniRef100_Q6ETK3 Hypothetical protein P0544B02.31 [Oryza sativa] 516 e-145
UniRef100_Q9FQY2 Protein phosphatase type-2C [Zea mays] 508 e-142
UniRef100_Q653S4 Hypothetical protein OJ1065_E04.2-2 [Oryza sativa] 470 e-131
UniRef100_Q653S3 Hypothetical protein OJ1065_E04.2-1 [Oryza sativa] 470 e-131
UniRef100_Q5TR41 ENSANGP00000028924 [Anopheles gambiae str. PEST] 275 1e-72
UniRef100_Q7Z8F2 Putative serine/threonine phosphatase 2C ptc2 [... 272 9e-72
UniRef100_UPI000023D430 UPI000023D430 UniRef100 entry 271 3e-71
UniRef100_Q872D8 Probable protein phosphatase 2C [Neurospora cra... 251 2e-65
UniRef100_Q6A1M6 Protein phosphatase 2C [Euplotes vannus] 231 2e-59
UniRef100_Q6K1U8 Hypothetical protein OSJNBa0038P01.3 [Oryza sat... 217 3e-55
UniRef100_Q6K1U4 Hypothetical protein OSJNBa0038P01.17 [Oryza sa... 217 3e-55
UniRef100_UPI0000498F52 UPI0000498F52 UniRef100 entry 211 3e-53
UniRef100_Q6K1U0 Hypothetical protein OSJNBa0038P01.32 [Oryza sa... 209 9e-53
UniRef100_Q6NYP6 Protein phosphatase type 2C alpha 2 [Brachydani... 207 5e-52
UniRef100_Q9GPU5 Phosphatase 2C [Oxytricha trifallax] 206 1e-51
UniRef100_Q7KRQ4 CG10417-PA, isoform A [Drosophila melanogaster] 191 3e-47
UniRef100_Q7ZYR7 Ppm1g-prov protein [Xenopus laevis] 189 1e-46
>UniRef100_Q9SZ53 Protein phosphatase 2C-like protein [Arabidopsis thaliana]
Length = 357
Score = 536 bits (1380), Expect = e-151
Identities = 254/357 (71%), Positives = 302/357 (84%)
Query: 1 MGTTLSIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVDSSTSFFGVYDGHG 60
MG LS PKT+KFSEDGEN LRYGLSSMQGWRA+MEDAHAA LD+D +TSF GVYDGHG
Sbjct: 1 MGIYLSTPKTDKFSEDGENHKLRYGLSSMQGWRASMEDAHAAILDLDDNTSFLGVYDGHG 60
Query: 61 GKAVAKFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKG 120
GK V+KFCAK+LHQQVL E Y AGDVGTSL KAF RMDEMM+GQRGWRELAVLGDK+
Sbjct: 61 GKVVSKFCAKYLHQQVLSDEAYAAGDVGTSLQKAFFRMDEMMQGQRGWRELAVLGDKINK 120
Query: 121 FNGIIEGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGD 180
F+G+IEGLI SPRS D+ ++ D WAFE+GPHSDF GPNSGSTACVA++R+ LFVANAGD
Sbjct: 121 FSGMIEGLIWSPRSGDSANKPDAWAFEEGPHSDFAGPNSGSTACVAVVRDKQLFVANAGD 180
Query: 181 SRCVISRNGQAYNLSRDHKPELVIEKERIYKAGGFIHAGRINGSLNLARAIGDVDFKNNR 240
SRCVISR QAYNLSRDHKP+L EKERI KAGGFIHAGR+NGSLNL+RAIGD++FK N+
Sbjct: 181 SRCVISRKNQAYNLSRDHKPDLEAEKERILKAGGFIHAGRVNGSLNLSRAIGDMEFKQNK 240
Query: 241 FLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEV 300
FL +EKQ+VTA+PD+N V+L D+D+F+V+ACDGIWDC++SQQLVDF+ ++L ETKLS V
Sbjct: 241 FLPSEKQIVTASPDVNTVELCDDDDFLVLACDGIWDCMTSQQLVDFIHEQLNSETKLSVV 300
Query: 301 CERVLDRCLAPSLAVGDGCDNMTMILVQFKKPLQTSAPAQEQSSSNEQDASEPTLEN 357
CE+VLDRCLAP+ + G+GCDNMTMILV+FK P + + ++S E + EP+ N
Sbjct: 301 CEKVLDRCLAPNTSGGEGCDNMTMILVRFKNPTPSETELKPEASQAEGNHDEPSSSN 357
>UniRef100_O81716 Hypothetical protein At2g25070 [Arabidopsis thaliana]
Length = 355
Score = 524 bits (1349), Expect = e-147
Identities = 250/354 (70%), Positives = 292/354 (81%), Gaps = 2/354 (0%)
Query: 1 MGTTLSIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVDSSTSFFGVYDGHG 60
MGT LS PKTEK SEDGEND LR+GLSSMQGWRATMEDAHAA LD+D TSFFGVYDGHG
Sbjct: 1 MGTYLSSPKTEKLSEDGENDKLRFGLSSMQGWRATMEDAHAAILDLDDKTSFFGVYDGHG 60
Query: 61 GKAVAKFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKG 120
GK VAKFCAK+LHQQV+ +E Y GDV TSL +AF RMD+MM+GQRGWRELAVLGDK+
Sbjct: 61 GKVVAKFCAKYLHQQVISNEAYKTGDVETSLRRAFFRMDDMMQGQRGWRELAVLGDKMNK 120
Query: 121 FNGIIEGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGD 180
F+G+IEG I SPRS D +Q D W E GPHSDF GP SG TACVA+I++ LFVANAGD
Sbjct: 121 FSGMIEGFIWSPRSGDTNNQPDSWPLEDGPHSDFTGPTSGCTACVALIKDKKLFVANAGD 180
Query: 181 SRCVISRNGQAYNLSRDHKPELVIEKERIYKAGGFIHAGRINGSLNLARAIGDVDFKNNR 240
SRCVISR QAYNLS+DHKP+L +EKERI KAGGFIHAGRINGSLNL RAIGD++FK N+
Sbjct: 181 SRCVISRKSQAYNLSKDHKPDLEVEKERILKAGGFIHAGRINGSLNLTRAIGDMEFKQNK 240
Query: 241 FLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEV 300
FL +EKQ+VTA+PDIN +DL D+D+F+V+ACDGIWDC+SSQ+LVDF+ ++L ETKLS V
Sbjct: 241 FLPSEKQMVTADPDINTIDLCDDDDFLVVACDGIWDCMSSQELVDFIHEQLKSETKLSTV 300
Query: 301 CERVLDRCLAPSLAVGDGCDNMTMILVQFKKPLQTSAPAQEQSSSNEQDASEPT 354
CE+V+DRCLAP A G+GCDNMT+ILVQFKKP + + + S E EP+
Sbjct: 301 CEKVVDRCLAPDTATGEGCDNMTIILVQFKKP--NPSETEPEDSKPEPSEDEPS 352
>UniRef100_Q67UP9 Hypothetical protein P0453H04.39-1 [Oryza sativa]
Length = 368
Score = 523 bits (1347), Expect = e-147
Identities = 248/355 (69%), Positives = 300/355 (83%), Gaps = 4/355 (1%)
Query: 1 MGTTLSIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVDSSTSFFGVYDGHG 60
MG LS PKTEK SEDGEND L++GLSSMQGWRATMEDAH+A LD+D+ TSFFGV+DGHG
Sbjct: 1 MGVYLSTPKTEKLSEDGENDKLKFGLSSMQGWRATMEDAHSALLDIDNDTSFFGVFDGHG 60
Query: 61 GKAVAKFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKG 120
G+ VAKFCAK+LH++VL+SE Y AGD+G + KAF RMDEMMRGQRGWREL LGDK+
Sbjct: 61 GRVVAKFCAKYLHREVLRSEAYSAGDLGNAAHKAFFRMDEMMRGQRGWRELQALGDKINQ 120
Query: 121 FNGIIEGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGD 180
+G+IEGLI SPR +D+ DQ DDWAFE+GPHSDF GP GSTACVAI+RN+ L VANAGD
Sbjct: 121 ISGMIEGLIWSPRGSDSNDQHDDWAFEEGPHSDFAGPTCGSTACVAIVRNSQLVVANAGD 180
Query: 181 SRCVISRNGQAYNLSRDHKPELVIEKERIYKAGGFIHAGRINGSLNLARAIGDVDFKNNR 240
SRCVISRNGQAYNLSRDHKPEL E+ERI KAGG+I GR+NG++NL+RAIGD++FK N+
Sbjct: 181 SRCVISRNGQAYNLSRDHKPELEAERERILKAGGYIQMGRVNGTINLSRAIGDIEFKQNK 240
Query: 241 FLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEV 300
FLS +KQ++TANPDIN V+L D+D+F+V+ACDGIWDC+SSQQLVDF+ + + E+ LS V
Sbjct: 241 FLSPDKQMLTANPDINTVELCDDDDFLVLACDGIWDCMSSQQLVDFIHEHINTESSLSAV 300
Query: 301 CERVLDRCLAPSLAVGDGCDNMTMILVQFKKPLQTS---APAQEQSSSNEQDASE 352
CERVLDRCLAPS G+GCDNMTMILVQFKKP+ + +PA EQS++++Q +
Sbjct: 301 CERVLDRCLAPSTLGGEGCDNMTMILVQFKKPISQNKNVSPA-EQSAADKQPTGD 354
>UniRef100_Q6ETK3 Hypothetical protein P0544B02.31 [Oryza sativa]
Length = 362
Score = 516 bits (1330), Expect = e-145
Identities = 246/352 (69%), Positives = 292/352 (82%), Gaps = 2/352 (0%)
Query: 1 MGTTLSIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVDSSTSFFGVYDGHG 60
MG LS PKT+KFSEDGEND L+ GLSSMQGWRA MEDAH+A L++D+ TSFFGV+DGHG
Sbjct: 1 MGIYLSTPKTDKFSEDGENDKLKLGLSSMQGWRANMEDAHSALLNLDNETSFFGVFDGHG 60
Query: 61 GKAVAKFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKG 120
G+ VAKFCAK+LH QVL+SE Y AGD+GT++ +AF RMDEMMRGQRGWREL+ LGDK+
Sbjct: 61 GRVVAKFCAKYLHSQVLRSEAYSAGDLGTAVHRAFFRMDEMMRGQRGWRELSALGDKINK 120
Query: 121 FNGIIEGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGD 180
G+IEGLI SPR +D+ + DDW+FE+GPHSDF GP G TACVA+IRNN L VANAGD
Sbjct: 121 IGGMIEGLIWSPRGSDSNNGQDDWSFEEGPHSDFAGPTCGCTACVALIRNNQLVVANAGD 180
Query: 181 SRCVISRNGQAYNLSRDHKPELVIEKERIYKAGGFIHAGRINGSLNLARAIGDVDFKNNR 240
SRCVISR GQAYNLSRDHKPEL E++RI KAGGFIH GRINGSLNL RAIGD++FK N+
Sbjct: 181 SRCVISRAGQAYNLSRDHKPELEAERDRIVKAGGFIHMGRINGSLNLTRAIGDMEFKQNK 240
Query: 241 FLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEV 300
FL EKQ+VTANPDIN+V+L D+D+F+V+ACDGIWDC+SSQQLVDF+ + + E+ LS V
Sbjct: 241 FLPPEKQIVTANPDINVVELCDDDDFLVLACDGIWDCMSSQQLVDFIHEHIQKESSLSAV 300
Query: 301 CERVLDRCLAPSLAVGDGCDNMTMILVQFKKPLQTSAPAQ--EQSSSNEQDA 350
CERVLDRCLAPS G+GCDNMTM+LVQFKKP+ + A EQS ++A
Sbjct: 301 CERVLDRCLAPSTIGGEGCDNMTMVLVQFKKPITQNKKADVGEQSVKGVEEA 352
>UniRef100_Q9FQY2 Protein phosphatase type-2C [Zea mays]
Length = 366
Score = 508 bits (1307), Expect = e-142
Identities = 238/328 (72%), Positives = 282/328 (85%)
Query: 5 LSIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVDSSTSFFGVYDGHGGKAV 64
LS PKT+K SEDGEND L++GLSSMQGWRATMEDAH+A LD+D+ T+ FGV+DGHGGK V
Sbjct: 5 LSTPKTDKVSEDGENDKLKFGLSSMQGWRATMEDAHSALLDLDNDTASFGVFDGHGGKVV 64
Query: 65 AKFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFNGI 124
AKFCAK+LH++VL +E Y AGD+G ++ +A+LRMDEMMRGQRGWREL LGDK+ F GI
Sbjct: 65 AKFCAKYLHREVLHTEAYAAGDLGAAVHRAYLRMDEMMRGQRGWRELQALGDKINQFTGI 124
Query: 125 IEGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGDSRCV 184
IEGLI SP+++D+ D+ DDWAFE+GPHSDF GPN GSTACVA++RN L VANAGDSRCV
Sbjct: 125 IEGLIWSPKASDSNDRHDDWAFEEGPHSDFTGPNCGSTACVALVRNRQLVVANAGDSRCV 184
Query: 185 ISRNGQAYNLSRDHKPELVIEKERIYKAGGFIHAGRINGSLNLARAIGDVDFKNNRFLSA 244
ISRNGQAYNLSRDHKPEL E+ERI AGG+I G +NGSLNL+RAIGD++ K N+FLS
Sbjct: 185 ISRNGQAYNLSRDHKPELEAERERIQSAGGYIKMGHVNGSLNLSRAIGDMELKQNKFLSP 244
Query: 245 EKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEVCERV 304
+KQ++TANPDINIV+L D+DEFIV+ACDGIWDC+SSQQLVDF+R+ + E LS VCE V
Sbjct: 245 DKQILTANPDINIVELCDDDEFIVLACDGIWDCMSSQQLVDFIREHINTEESLSAVCEGV 304
Query: 305 LDRCLAPSLAVGDGCDNMTMILVQFKKP 332
LDRCLAPS G+GCDNMTMILVQFKKP
Sbjct: 305 LDRCLAPSTMGGEGCDNMTMILVQFKKP 332
>UniRef100_Q653S4 Hypothetical protein OJ1065_E04.2-2 [Oryza sativa]
Length = 362
Score = 470 bits (1210), Expect = e-131
Identities = 231/357 (64%), Positives = 284/357 (78%), Gaps = 6/357 (1%)
Query: 1 MGTTLSIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVDSSTSFFGVYDGHG 60
MG LS PKTEK+S +G ND LRYGL+SMQGWR TMEDAH A +D TSFFGVYDGHG
Sbjct: 1 MGVYLSTPKTEKYSGEGGNDRLRYGLASMQGWRTTMEDAHTALPRLDECTSFFGVYDGHG 60
Query: 61 GKAVAKFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKG 120
GKAV+KFCAKHLH QVLK+E Y +GD+ TS+ K+F RMDEMM+GQRGWRELA LGDK +
Sbjct: 61 GKAVSKFCAKHLHLQVLKNEAYSSGDLATSVLKSFFRMDEMMKGQRGWRELAELGDKGQK 120
Query: 121 FNGIIEGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGD 180
F G++EG+I SP+ ++ D W E+GPHS F GP SGSTACVAIIRN+ L VANAGD
Sbjct: 121 FTGMLEGIIWSPKPGESDKPEDTWT-EEGPHSHFPGPTSGSTACVAIIRNDELIVANAGD 179
Query: 181 SRCVISRNGQAYNLSRDHKPELVIEKERIYKAGGFIHAGRINGSLNLARAIGDVDFKNNR 240
SRCV+SR G+AY+LS+DHKP+L EKERI AGGFI AGR+NGSLNLARAIGD++ K N
Sbjct: 180 SRCVLSRKGRAYDLSKDHKPDLDAEKERILNAGGFIVAGRVNGSLNLARAIGDMELKQNE 239
Query: 241 FLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEV 300
FL AE+Q+VTA P++N V L ++DEFIV+ACDGIWDC+SSQ++VDFV +E+ E LS V
Sbjct: 240 FLPAERQIVTAEPELNTVKLSEDDEFIVLACDGIWDCMSSQEVVDFVHKEMNTEDSLSAV 299
Query: 301 CERVLDRCLAPSLAVGDGCDNMTMILVQFKKPLQTSAPAQEQSSSNEQDASEPTLEN 357
CE++LD CLAP ++ GDGCDNMT+I+V+FKKP +++A SS+N+ +SE N
Sbjct: 300 CEKLLDHCLAP-VSGGDGCDNMTVIIVKFKKPSKSAA----TSSTNQSVSSEEMRPN 351
>UniRef100_Q653S3 Hypothetical protein OJ1065_E04.2-1 [Oryza sativa]
Length = 352
Score = 470 bits (1209), Expect = e-131
Identities = 230/352 (65%), Positives = 283/352 (80%), Gaps = 6/352 (1%)
Query: 1 MGTTLSIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVDSSTSFFGVYDGHG 60
MG LS PKTEK+S +G ND LRYGL+SMQGWR TMEDAH A +D TSFFGVYDGHG
Sbjct: 1 MGVYLSTPKTEKYSGEGGNDRLRYGLASMQGWRTTMEDAHTALPRLDECTSFFGVYDGHG 60
Query: 61 GKAVAKFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKG 120
GKAV+KFCAKHLH QVLK+E Y +GD+ TS+ K+F RMDEMM+GQRGWRELA LGDK +
Sbjct: 61 GKAVSKFCAKHLHLQVLKNEAYSSGDLATSVLKSFFRMDEMMKGQRGWRELAELGDKGQK 120
Query: 121 FNGIIEGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGD 180
F G++EG+I SP+ ++ D W E+GPHS F GP SGSTACVAIIRN+ L VANAGD
Sbjct: 121 FTGMLEGIIWSPKPGESDKPEDTWT-EEGPHSHFPGPTSGSTACVAIIRNDELIVANAGD 179
Query: 181 SRCVISRNGQAYNLSRDHKPELVIEKERIYKAGGFIHAGRINGSLNLARAIGDVDFKNNR 240
SRCV+SR G+AY+LS+DHKP+L EKERI AGGFI AGR+NGSLNLARAIGD++ K N
Sbjct: 180 SRCVLSRKGRAYDLSKDHKPDLDAEKERILNAGGFIVAGRVNGSLNLARAIGDMELKQNE 239
Query: 241 FLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEV 300
FL AE+Q+VTA P++N V L ++DEFIV+ACDGIWDC+SSQ++VDFV +E+ E LS V
Sbjct: 240 FLPAERQIVTAEPELNTVKLSEDDEFIVLACDGIWDCMSSQEVVDFVHKEMNTEDSLSAV 299
Query: 301 CERVLDRCLAPSLAVGDGCDNMTMILVQFKKPLQTSAPAQEQSSSNEQDASE 352
CE++LD CLAP ++ GDGCDNMT+I+V+FKKP +++A SS+N+ +SE
Sbjct: 300 CEKLLDHCLAP-VSGGDGCDNMTVIIVKFKKPSKSAA----TSSTNQSVSSE 346
>UniRef100_Q5TR41 ENSANGP00000028924 [Anopheles gambiae str. PEST]
Length = 338
Score = 275 bits (703), Expect = 1e-72
Identities = 157/336 (46%), Positives = 209/336 (61%), Gaps = 7/336 (2%)
Query: 1 MGTTLSIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVDSSTSFFGVYDGHG 60
MG LS P T K S D N+ L G SSMQGWR + EDAH L+ D SFF VYDGHG
Sbjct: 1 MGAYLSEPLTTKDSSDESNEFLASGSSSMQGWRISQEDAHNCILNFDDQCSFFAVYDGHG 60
Query: 61 GKAVAKFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKG 120
G VA++C+ HL + E Y + +L +AFL D + ++ EL VL DK G
Sbjct: 61 GAEVAQYCSLHLPTFLKTVEAYGRKEFEKALKEAFLGFDATLLQEKVIEELKVLSDKTNG 120
Query: 121 FNGIIEGLIRSPRSNDNKDQ---SDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVAN 177
G E P + + +D AF K +D G +SG TA VA++ L+VAN
Sbjct: 121 EEGSAEEEEEEPDEYADMGEYINEEDAAFMK-TITDEPGKDSGCTAVVALLHGKDLYVAN 179
Query: 178 AGDSRCVISRNGQAYNLSRDHKPELVIEKERIYKAGGFIHA-GRINGSLNLARAIGDVDF 236
AGDSRCV+ RNG+A +S DHKPE +E +RI KAGG + GR+NG LNL+RAIGD +
Sbjct: 180 AGDSRCVVCRNGKALEMSFDHKPEDTVEYQRIEKAGGRVTLDGRVNGGLNLSRAIGDHGY 239
Query: 237 KNNRFLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLE-T 295
K N+ L AE+Q+++A PDI + + EDEF+V+ACDGIW+ ++S+Q+V FV++ +
Sbjct: 240 KMNKSLPAEEQMISALPDIEKITVGPEDEFMVLACDGIWNFMTSEQVVQFVQERINKPGM 299
Query: 296 KLSEVCERVLDRCLAP-SLAVGDGCDNMTMILVQFK 330
KLS++CE + D CLAP + G GCDNMT I+VQFK
Sbjct: 300 KLSKICEELFDHCLAPHTRGDGTGCDNMTAIIVQFK 335
>UniRef100_Q7Z8F2 Putative serine/threonine phosphatase 2C ptc2 [Trichoderma reesei]
Length = 438
Score = 272 bits (696), Expect = 9e-72
Identities = 153/340 (45%), Positives = 197/340 (57%), Gaps = 57/340 (16%)
Query: 1 MGTTLSIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHL-----DVDSST----- 50
MG TLS P EK SE GE+D L YG+S+MQGWR +MEDAH A L D D+ T
Sbjct: 1 MGQTLSEPVVEKTSEKGEDDRLIYGVSAMQGWRISMEDAHTAELNLPPPDNDTKTHPDRL 60
Query: 51 SFFGVYDGHGGKAVAKFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRE 110
SFFGV+DGHGG VA F +++H V K E + +GD L FL D
Sbjct: 61 SFFGVFDGHGGDKVALFAGENIHNIVFKQESFKSGDYAQGLKDGFLATDR---------- 110
Query: 111 LAVLGDKVKGFNGIIEGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRN 170
A+L D ++ SG TACV +I
Sbjct: 111 -AILNDP-----------------------------------KYEEEVSGCTACVTLIAG 134
Query: 171 NLLFVANAGDSRCVISRNGQAYNLSRDHKPELVIEKERIYKAGGFIHAGRINGSLNLARA 230
N L+VANAGDSR V+ G+A LS DHKP+L EK RI AGGF+ GR+NG+L L+RA
Sbjct: 135 NKLYVANAGDSRSVLGIKGRAKPLSNDHKPQLETEKNRITAAGGFVDFGRVNGNLALSRA 194
Query: 231 IGDVDFKNNRFLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQE 290
IGD +FK + LS E Q+VTA PD+ + +L +EDEF+VIACDGIWDC SSQ +V+FVR+
Sbjct: 195 IGDFEFKKSAELSPENQIVTAFPDVEVHELTEEDEFLVIACDGIWDCQSSQAVVEFVRRG 254
Query: 291 LLLETKLSEVCERVLDRCLAPSLAVGD-GCDNMTMILVQF 329
+ + L ++CE ++D CLA + G GCDNMTM+++ F
Sbjct: 255 IAAKQDLDKICENMMDNCLASNSETGGVGCDNMTMVIIGF 294
>UniRef100_UPI000023D430 UPI000023D430 UniRef100 entry
Length = 430
Score = 271 bits (692), Expect = 3e-71
Identities = 150/336 (44%), Positives = 197/336 (57%), Gaps = 53/336 (15%)
Query: 1 MGTTLSIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVD------SSTSFFG 54
MG TLS P EK SE GE++ L YG+S+MQGWR +MEDAH LD+D S SFFG
Sbjct: 1 MGQTLSEPVVEKTSEKGEDERLIYGVSAMQGWRISMEDAHTTVLDLDTAKTHDSKLSFFG 60
Query: 55 VYDGHGGKAVAKFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVL 114
V+DGHGG VA F +++H + K + + +GD L FL D A+L
Sbjct: 61 VFDGHGGDKVALFTGQNIHNIIFKQDTFKSGDYAQGLKDGFLATDR-----------AIL 109
Query: 115 GDKVKGFNGIIEGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLF 174
D ++ SG TACV++I N L+
Sbjct: 110 NDP-----------------------------------KYEEEVSGCTACVSLIAGNKLY 134
Query: 175 VANAGDSRCVISRNGQAYNLSRDHKPELVIEKERIYKAGGFIHAGRINGSLNLARAIGDV 234
VANAGDSR V+ G+A LS+DHKP+L EK RI AGGF+ GR+NG+L L+RAIGD
Sbjct: 135 VANAGDSRGVLGIKGRAKPLSQDHKPQLENEKNRITAAGGFVDFGRVNGNLALSRAIGDF 194
Query: 235 DFKNNRFLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLE 294
+FK + LS E Q+VTA PD+ DL DEDEF+V+ACDGIWDC SSQ +V+FVR+ + +
Sbjct: 195 EFKKSAELSPENQIVTAYPDVEQHDLTDEDEFLVLACDGIWDCQSSQAVVEFVRRGIAAK 254
Query: 295 TKLSEVCERVLDRCLAPSLAVGD-GCDNMTMILVQF 329
+L ++CE ++D CLA + G GCDNMTM ++ F
Sbjct: 255 QELEKICENMMDNCLASNSETGGVGCDNMTMCIIGF 290
>UniRef100_Q872D8 Probable protein phosphatase 2C [Neurospora crassa]
Length = 439
Score = 251 bits (642), Expect = 2e-65
Identities = 143/341 (41%), Positives = 191/341 (55%), Gaps = 58/341 (17%)
Query: 1 MGTTLSIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDV-----------DSS 49
MG TLS P EK S G ++ L YG+S+MQGWR +MEDAH LD+
Sbjct: 1 MGQTLSEPVVEKASATGGDERLIYGVSAMQGWRISMEDAHTTVLDLLANNPKEAKDHSQK 60
Query: 50 TSFFGVYDGHGGKAVAKFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWR 109
SFFGV+DGHGG VA F ++H + K + + G+ +L FL D
Sbjct: 61 LSFFGVFDGHGGDKVALFAGANIHDIIAKQDTFKTGNYEQALKDGFLATDR--------- 111
Query: 110 ELAVLGDKVKGFNGIIEGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIR 169
A+L D ++ SG TACV +I
Sbjct: 112 --AILNDP-----------------------------------KYEEEVSGCTACVGLIT 134
Query: 170 NNLLFVANAGDSRCVISRNGQAYNLSRDHKPELVIEKERIYKAGGFIHAGRINGSLNLAR 229
+ +FVANAGDSR V+ G+A LS DHKP+ EK RI AGGF+ GR+NG+L L+R
Sbjct: 135 DEKIFVANAGDSRSVLGVKGRAKPLSFDHKPQNEGEKARITAAGGFVDFGRVNGNLALSR 194
Query: 230 AIGDVDFKNNRFLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQ 289
AIGD +FK + L+ E+Q+VTA PD+ + DL D+DEF+V+ACDGIWDC SSQ +V+FVR+
Sbjct: 195 AIGDFEFKKSAELAPEQQIVTAYPDVMVHDLADDDEFLVLACDGIWDCQSSQAVVEFVRR 254
Query: 290 ELLLETKLSEVCERVLDRCLAPSLAVGD-GCDNMTMILVQF 329
+ + L ++CE ++D CLA + G GCDNMTMI+V F
Sbjct: 255 GIAAKQDLDKICENMMDNCLASNSETGGVGCDNMTMIIVGF 295
>UniRef100_Q6A1M6 Protein phosphatase 2C [Euplotes vannus]
Length = 327
Score = 231 bits (590), Expect = 2e-59
Identities = 130/329 (39%), Positives = 192/329 (57%), Gaps = 12/329 (3%)
Query: 5 LSIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVDSSTSFFGVYDGHGGKAV 64
L P EK + G+ND L YG MQGWR MEDA A+LD D TS FGV+DGHGGK
Sbjct: 8 LESPNKEKHVQHGQNDKLSYGACEMQGWRLGMEDAVIANLDFDEDTSLFGVFDGHGGKEA 67
Query: 65 AKFCAKHLHQQVLKSEE-YIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFNG 123
++ K ++++LK + Y GD L +F D + + G +E+ + D N
Sbjct: 68 SQV-VKDNYERILKGDSSYKDGDCQKGLYDSFKGTDVFLGSKTGKQEMKAVADS----NP 122
Query: 124 IIEGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGDSRC 183
++ + + KD+ E+ D + G TA V +I+ + ++ ANAGDSRC
Sbjct: 123 EVKNPLLKILGEEAKDKPAGERDEESYMLD----SKGCTANVVLIKGSAIYCANAGDSRC 178
Query: 184 VISRNGQAYNLSRDHKPELVIEKERIYKAGGFIHAGRINGSLNLARAIGDVDFKNNRFLS 243
V+SR G+A NLS DHKPE IE+ERI KAG I GR++G+LNL+R++GD+ K L
Sbjct: 179 VLSREGKAVNLSGDHKPENEIERERIRKAGSEIVDGRVDGNLNLSRSLGDLKHKQKPGLK 238
Query: 244 AEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETKLSEVCER 303
E+Q +T PDI + + D+FIV+ACDGIW+ SSQ +VDF+ + L + KL+++
Sbjct: 239 PEEQPITCVPDITVDKIKPGDDFIVMACDGIWEVKSSQDVVDFISERLKKDMKLTDIIGE 298
Query: 304 VLDRCLAPSLAV--GDGCDNMTMILVQFK 330
+ + ++P G GCDNM+ I+++ +
Sbjct: 299 LFEDIISPDYTATQGLGCDNMSCIIIKLR 327
>UniRef100_Q6K1U8 Hypothetical protein OSJNBa0038P01.3 [Oryza sativa]
Length = 521
Score = 217 bits (553), Expect = 3e-55
Identities = 139/354 (39%), Positives = 191/354 (53%), Gaps = 58/354 (16%)
Query: 6 SIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVD--SSTSFFGVYDGHGGKA 63
S+P KF+ + END ++Y +SSMQGW MEDAHAA L++D +STSFFGVYDGHGG
Sbjct: 4 SLPVESKFTFEEENDRIKYVVSSMQGWGEKMEDAHAAILNLDDATSTSFFGVYDGHGGAE 63
Query: 64 VAKFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFNG 123
VA +CAK H ++ E+Y D+ +L FL MDE ++ WREL + D NG
Sbjct: 64 VALYCAKQFHIELCNHEDY-HNDLINALDNVFLSMDENLQQSDAWRELVIPHD-----NG 117
Query: 124 II----EGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPN-SGSTACVAIIRNNLLFVANA 178
+ G+ P P + + GP GSTACV +IR N + V +
Sbjct: 118 CMYFLKAGVCAKPF----------------PQATYTGPAYEGSTACVVVIRGNQMIVGHV 161
Query: 179 GDSRCVISRNGQ-AYNLSRDHKP--ELVIEKERIYKAGGFIHA----------------- 218
GDSRCV+SR G A +LS DHKP E+ER+ AGG
Sbjct: 162 GDSRCVLSRQGGLAIDLSFDHKPCTRTESERERVQNAGGRSLGLRCEQVMGNYVVKEQWV 221
Query: 219 -GRINGSLNLARAIGDVDFKNNRFLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDC 277
G G + ++R+IGD FK N+ L EKQ++ +PDI D+ D+ EF+VIA G+W C
Sbjct: 222 LGDFGGGVTISRSIGDFAFKKNKDLDREKQMLVCDPDILADDITDDMEFLVIASQGLWSC 281
Query: 278 LSSQQLVDFVRQELLLE-TKLSEVCERVLDRCLAPSLAVGDGCDNMTMILVQFK 330
+ S +V ++ L +E +L +CE V++ LA +N T+ILVQFK
Sbjct: 282 VDSADVVSYIHDRLSVEGAELRVICEEVVEFGLASG-------ENTTVILVQFK 328
>UniRef100_Q6K1U4 Hypothetical protein OSJNBa0038P01.17 [Oryza sativa]
Length = 521
Score = 217 bits (553), Expect = 3e-55
Identities = 139/354 (39%), Positives = 191/354 (53%), Gaps = 58/354 (16%)
Query: 6 SIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVD--SSTSFFGVYDGHGGKA 63
S+P KF+ + END ++Y +SSMQGW MEDAHAA L++D +STSFFGVYDGHGG
Sbjct: 4 SLPVESKFTFEEENDRIKYVVSSMQGWGEKMEDAHAAILNLDDATSTSFFGVYDGHGGAE 63
Query: 64 VAKFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFNG 123
VA +CAK H ++ E+Y D+ +L FL MDE ++ WREL + D NG
Sbjct: 64 VALYCAKQFHIELCNHEDY-HNDLINALDNVFLSMDENLQQSDAWRELVIPHD-----NG 117
Query: 124 II----EGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPN-SGSTACVAIIRNNLLFVANA 178
+ G+ P P + + GP GSTACV +IR N + V +
Sbjct: 118 CMYFLKAGVCAKPF----------------PQATYTGPAYEGSTACVVVIRGNQMIVGHV 161
Query: 179 GDSRCVISRNGQ-AYNLSRDHKP--ELVIEKERIYKAGGFIHA----------------- 218
GDSRCV+SR G A +LS DHKP E+ER+ AGG
Sbjct: 162 GDSRCVLSRQGGLAIDLSFDHKPCTRTESERERVQNAGGRSLGLRCEQVMGNYVVKEQWV 221
Query: 219 -GRINGSLNLARAIGDVDFKNNRFLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDC 277
G G + ++R+IGD FK N+ L EKQ++ +PDI D+ D+ EF+VIA G+W C
Sbjct: 222 LGDFGGGVTISRSIGDFAFKKNKDLDREKQMLVCDPDILADDITDDMEFLVIASQGLWSC 281
Query: 278 LSSQQLVDFVRQELLLE-TKLSEVCERVLDRCLAPSLAVGDGCDNMTMILVQFK 330
+ S +V ++ L +E +L +CE V++ LA +N T+ILVQFK
Sbjct: 282 VDSADVVSYIHDRLSVEGAELRVICEEVVEFGLASG-------ENTTVILVQFK 328
>UniRef100_UPI0000498F52 UPI0000498F52 UniRef100 entry
Length = 334
Score = 211 bits (536), Expect = 3e-53
Identities = 126/358 (35%), Positives = 188/358 (52%), Gaps = 29/358 (8%)
Query: 1 MGTTLSIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVD----SSTSFFGVY 56
MG+ LS+P TE+ S + + + L G +SMQGWR TMEDAH ++ SFFGV+
Sbjct: 1 MGSLLSVPVTEQQSGETKGEFLDCGYTSMQGWRRTMEDAHIVDIEFTCENGKKASFFGVF 60
Query: 57 DGHGGKAVAKFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGD 116
DGHGG VA++C+K + +L ++ + AG+ +L +++DE MR L LG
Sbjct: 61 DGHGGDQVAEYCSKIYVETLLNTDAFKAGNYQQALIDTNIKIDEQMRTPAVNDLLKTLGS 120
Query: 117 KVKGFNGIIEGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVA 176
G + I EG+ D G T+ V +I +N ++
Sbjct: 121 ---GSSNIYEGMF----------------------GDLVADGMGCTSVVILIIDNTIYCG 155
Query: 177 NAGDSRCVISRNGQAYNLSRDHKPELVIEKERIYKAGGFIHAGRINGSLNLARAIGDVDF 236
NAGDSR V+ + LS DHKP L E +RI AGG I GR+NG+LNL R IGD+ +
Sbjct: 156 NAGDSRSVMLKGDSVIPLSIDHKPSLQSEIDRITLAGGTIDNGRVNGNLNLTRTIGDLMY 215
Query: 237 KNNRFLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFVRQELLLETK 296
K L + Q+++ PD+ + ++ I++ACDGIWD L+++Q V+ V + L
Sbjct: 216 KRQSELPPQSQIISCYPDVKQEAIDGTEKLIILACDGIWDVLTNEQCVEKVVEYLKQGNS 275
Query: 297 LSEVCERVLDRCLAPSLAVGDGCDNMTMILVQFKKPLQTSAPAQEQSSSNEQDASEPT 354
L E CE++ + CL+ G DNMT+++V+FK +SS E+D T
Sbjct: 276 LKETCEKIANDCLSKEPYSKPGWDNMTLLVVKFKSFNPQPDITTVVTSSQEKDEGSVT 333
>UniRef100_Q6K1U0 Hypothetical protein OSJNBa0038P01.32 [Oryza sativa]
Length = 735
Score = 209 bits (532), Expect = 9e-53
Identities = 136/346 (39%), Positives = 189/346 (54%), Gaps = 50/346 (14%)
Query: 6 SIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVD--SSTSFFGVYDGHGGKA 63
S+P KF+ + END ++Y +SSMQGW MEDAHAA L++D +STSFFGVYDGHGG
Sbjct: 28 SLPVESKFTFEEENDRIKYVVSSMQGWGEKMEDAHAAILNLDDTTSTSFFGVYDGHGGAE 87
Query: 64 VAKFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKGFNG 123
VA +CAK H ++ E+Y D+ +L FL MDE ++ WREL + D NG
Sbjct: 88 VALYCAKQFHIELCNHEDY-HNDLINALDNVFLSMDENLQQSDAWRELVIPHD-----NG 141
Query: 124 II----EGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPN-SGSTACVAIIRNNLLFVANA 178
+ G+ P P + + GP GSTACV +IR N + V +
Sbjct: 142 CMYFLKAGVCAKPF----------------PQATYTGPAYEGSTACVVVIRGNQMIVGHV 185
Query: 179 GDSRCVISRNGQ-AYNLSRDHKP--ELVIEKERIYKAGGF---IHAGRINGSLNLARAI- 231
GDSRCV+SR G A +LS DHKP E+ER+ AGG + ++ G+ +
Sbjct: 186 GDSRCVLSRQGGLAIDLSFDHKPCTRTESERERVQNAGGRSLGLRCEQVMGNYVVKEQWV 245
Query: 232 ------GDVDFKNNRFLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVD 285
GD FK N+ L EKQ++ +PDI D+ D+ EF+VIA G+W C+ S +V
Sbjct: 246 LGDFGGGDFAFKKNKDLDREKQMLVCDPDILADDITDDMEFLVIASQGLWSCVDSADVVS 305
Query: 286 FVRQELLLE-TKLSEVCERVLDRCLAPSLAVGDGCDNMTMILVQFK 330
++ L +E +L +CE V++ LA +N T+ILVQFK
Sbjct: 306 YIHDRLSVEGAELRVICEEVVEFGLASG-------ENTTVILVQFK 344
>UniRef100_Q6NYP6 Protein phosphatase type 2C alpha 2 [Brachydanio rerio]
Length = 384
Score = 207 bits (526), Expect = 5e-52
Identities = 126/348 (36%), Positives = 184/348 (52%), Gaps = 39/348 (11%)
Query: 1 MGTTLSIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVDSST---SFFGVYD 57
MG L PK EK + G+ ++LRYGLSSMQGWR MEDAH A + + +S SFF VYD
Sbjct: 1 MGAFLDKPKMEKHNAHGDGNSLRYGLSSMQGWRVEMEDAHTAVIGLPNSLDLWSFFAVYD 60
Query: 58 GHGGKAVAKFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDK 117
GH G VA++C +HL + + + ++ G G G + D
Sbjct: 61 GHAGSQVARYCCEHLLEHITSNPDFQGGGGG------------------GGPAVEPSVDS 102
Query: 118 VKGFNGIIEGLIRSPRSNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVAN 177
VK +GI G + Q DD + SGSTA +I ++ N
Sbjct: 103 VK--SGIRTGFL----------QIDDHMRQISEKKHGGADRSGSTAVGVMISPRHIYFIN 150
Query: 178 AGDSRCVISRNGQAYNLSRDHKPELVIEKERIYKAGGFIHAGRINGSLNLARAIGDVDFK 237
GDSR ++SR G + ++DHKP +EKERI AGG + R+NGSL ++RA+GD D+K
Sbjct: 151 CGDSRGLLSRGGAVHFFTQDHKPSNPLEKERIQNAGGSVMIQRVNGSLAVSRALGDFDYK 210
Query: 238 NNRFLSAEKQVVTANPDINIVDLHD-EDEFIVIACDGIWDCLSSQQLVDFVRQELLLETK 296
+Q+V+ P++ ++ + EDEFIV+ACDGIWD +++++L DFVR L +
Sbjct: 211 CVHGKGPTEQLVSPEPEVCAIERSEAEDEFIVLACDGIWDVMANEELCDFVRSRLEVTDD 270
Query: 297 LSEVCERVLDRCLAPSLAVGDGCDNMTMILVQFKKPLQTSAPAQEQSS 344
L VC ++D CL DNM+++LV F + S A ++ +
Sbjct: 271 LERVCNEIVDTCLYKG-----SRDNMSVVLVCFVSAPKVSPEAVKREA 313
>UniRef100_Q9GPU5 Phosphatase 2C [Oxytricha trifallax]
Length = 280
Score = 206 bits (523), Expect = 1e-51
Identities = 115/290 (39%), Positives = 162/290 (55%), Gaps = 36/290 (12%)
Query: 1 MGTTLSIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVDSSTSFFGVYDGHG 60
MG LS+P K SE+G++ + +G ++MQGWR T EDAH A LD+ S F V+DGHG
Sbjct: 1 MGDYLSVPDKNKHSEEGKDHRIAFGATTMQGWRKTQEDAHIARLDIGDGNSLFAVFDGHG 60
Query: 61 GKAVAKFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDKVKG 120
G VAK+ K + Q++LK + Y D SL + FL++DE+M
Sbjct: 61 GDQVAKYAEKTMVQELLKLQSYKDKDYKKSLEEVFLKIDELM------------------ 102
Query: 121 FNGIIEGLIRSPRSNDNKDQS--DDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANA 178
++ R N + +S DD++ + S+ SG T+ V +I + ++ ANA
Sbjct: 103 --------LQHIRQNGSSGRSRFDDYSADPNDLSE-----SGCTSNVILITKDKIYCANA 149
Query: 179 GDSR---CVISRNGQAYNLSRDHKPELVIEKERIYKAGGFIHAGRINGSLNLARAIGDVD 235
GDSR CV + LS DHKP+ EK+RI A GF+ GR NG ++L+RA+GD D
Sbjct: 150 GDSRAVMCVFGSGPETVELSHDHKPDNETEKQRIVNADGFVQMGRTNGVISLSRALGDFD 209
Query: 236 FKNNRFLSAEKQVVTANPDINIVDLHDEDEFIVIACDGIWDCLSSQQLVD 285
+K EKQ TA PD++ DL + +FIV ACDGIWDCL+S + VD
Sbjct: 210 YKKKADFPPEKQATTAFPDVSEHDLTENCQFIVQACDGIWDCLTSPEAVD 259
>UniRef100_Q7KRQ4 CG10417-PA, isoform A [Drosophila melanogaster]
Length = 662
Score = 191 bits (485), Expect = 3e-47
Identities = 99/181 (54%), Positives = 131/181 (71%), Gaps = 2/181 (1%)
Query: 156 GPNSGSTACVAIIRNNLLFVANAGDSRCVISRNGQAYNLSRDHKPELVIEKERIYKAGGF 215
G +SG TA V +++ L+VANAGDSRCVISR+GQA +S DHKPE E RI KAGG
Sbjct: 389 GKDSGCTAVVCLLQGRDLYVANAGDSRCVISRSGQAIEMSIDHKPEDDEEASRIIKAGGR 448
Query: 216 IHA-GRINGSLNLARAIGDVDFKNNRFLSAEKQVVTANPDINIVDLHDEDEFIVIACDGI 274
+ GR+NG LNL+RA+GD +K N L AE+Q+++A PDI + + EDEF+V+ACDGI
Sbjct: 449 VTLDGRVNGGLNLSRALGDHAYKTNVTLPAEEQMISALPDIKKLIITPEDEFMVLACDGI 508
Query: 275 WDCLSSQQLVDFVRQELLLETKLSEVCERVLDRCLAP-SLAVGDGCDNMTMILVQFKKPL 333
W+ +SS+++V+FVR L KLS +CE + D CLAP ++ G GCDNMT ++VQFKK L
Sbjct: 509 WNYMSSEEVVEFVRCRLKDNKKLSTICEELFDNCLAPNTMGDGTGCDNMTAVIVQFKKKL 568
Query: 334 Q 334
Q
Sbjct: 569 Q 569
Score = 95.9 bits (237), Expect = 2e-18
Identities = 50/102 (49%), Positives = 64/102 (62%)
Query: 1 MGTTLSIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVDSSTSFFGVYDGHG 60
MG LS PKT+K S D N+ L G SSMQGWR + EDAH + L+ D++TSFF VYDGHG
Sbjct: 1 MGAYLSHPKTDKTSTDQFNELLAVGASSMQGWRNSQEDAHNSILNFDNNTSFFAVYDGHG 60
Query: 61 GKAVAKFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMM 102
G VA++CA L + E Y G +L +AFL D+ +
Sbjct: 61 GAEVAQYCADKLPHFLKNLETYKNGQFEVALKEAFLGFDKTL 102
>UniRef100_Q7ZYR7 Ppm1g-prov protein [Xenopus laevis]
Length = 544
Score = 189 bits (479), Expect = 1e-46
Identities = 107/233 (45%), Positives = 146/233 (61%), Gaps = 8/233 (3%)
Query: 134 SNDNKDQSDDWAFEKGPHSDFDGPNSGSTACVAIIRNNLLFVANAGDSRCVISRNGQAYN 193
S D +++ +D + G +SG+TA VA+IR L VANAGDSRCV+S G+A +
Sbjct: 305 SEDAEEEDEDMMLPGMEGKEEPGSDSGTTAVVALIRGQQLIVANAGDSRCVVSEGGKAVD 364
Query: 194 LSRDHKPELVIEKERIYKAGGFIHA-GRINGSLNLARAIGDVDFKNNRFLSAEKQVVTAN 252
+S DHKPE +E RI AGG + GR+NG LNL+RAIGD +K N+ L E+Q+++A
Sbjct: 365 MSYDHKPEDELELSRIKNAGGKVTMDGRVNGGLNLSRAIGDHFYKRNKNLPPEEQMISAL 424
Query: 253 PDINIVDLHDEDEFIVIACDGIWDCLSSQQLVDFV--RQELLLE----TKLSEVCERVLD 306
PDI ++ L +E EF+VIACDGIW+ +SSQ++VDFV R+E L+ LS + E +LD
Sbjct: 425 PDIKVLTLSEEHEFMVIACDGIWNVMSSQEVVDFVHERRESQLQKGDTLSLSSIVEELLD 484
Query: 307 RCLAPSLA-VGDGCDNMTMILVQFKKPLQTSAPAQEQSSSNEQDASEPTLENS 358
+CLAP + G GCDNMT I+V F+ Q P + D S T NS
Sbjct: 485 QCLAPDTSGDGTGCDNMTCIIVGFQPYSQCGGPEVGKRKVENTDESAETCSNS 537
Score = 103 bits (258), Expect = 6e-21
Identities = 51/117 (43%), Positives = 73/117 (61%)
Query: 1 MGTTLSIPKTEKFSEDGENDNLRYGLSSMQGWRATMEDAHAAHLDVDSSTSFFGVYDGHG 60
MG LS P T+K S +G + L YG S+MQGWR +MEDAH ++DS T+ F VYDGHG
Sbjct: 1 MGAYLSQPNTDKSSGEGGSQRLTYGYSAMQGWRVSMEDAHNCIPELDSQTAMFSVYDGHG 60
Query: 61 GKAVAKFCAKHLHQQVLKSEEYIAGDVGTSLTKAFLRMDEMMRGQRGWRELAVLGDK 117
G+ VA +CAK+L + + EY G + +L AFL +D+ + + +ELA + +
Sbjct: 61 GEEVALYCAKYLPEVIKSQREYKDGKLQKALEDAFLAIDQKLTREEVIKELAQMAGR 117
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.134 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 609,606,931
Number of Sequences: 2790947
Number of extensions: 25792160
Number of successful extensions: 59815
Number of sequences better than 10.0: 847
Number of HSP's better than 10.0 without gapping: 658
Number of HSP's successfully gapped in prelim test: 189
Number of HSP's that attempted gapping in prelim test: 57326
Number of HSP's gapped (non-prelim): 1535
length of query: 358
length of database: 848,049,833
effective HSP length: 128
effective length of query: 230
effective length of database: 490,808,617
effective search space: 112885981910
effective search space used: 112885981910
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 75 (33.5 bits)
Medicago: description of AC143339.7


