

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC143339.11 + phase: 0
(678 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
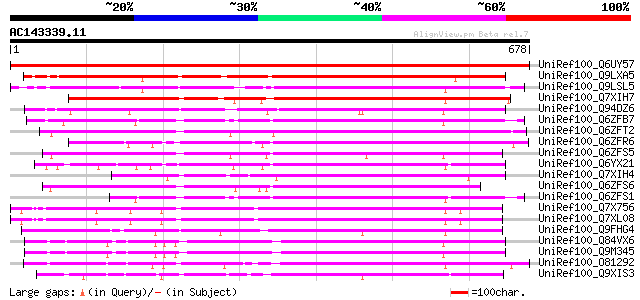
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6UY57 Lectin-like receptor kinase 1;1 [Medicago trunc... 1369 0.0
UniRef100_Q9LXA5 Lectin-like protein kinase-like [Arabidopsis th... 512 e-143
UniRef100_Q9LSL5 Receptor protein kinase-like protein [Arabidops... 459 e-127
UniRef100_Q7XIH7 Putative lectin-like protein kinase [Oryza sativa] 452 e-125
UniRef100_Q94DZ6 Putative lectin-like receptor kinase 1;1 [Oryza... 422 e-116
UniRef100_Q6ZFB7 Putative receptor-type protein kinase LRK1 [Ory... 416 e-114
UniRef100_Q6ZFT2 Putative receptor-type protein kinase LRK1 [Ory... 405 e-111
UniRef100_Q6ZFR6 Putative receptor-type protein kinase LRK1 [Ory... 404 e-111
UniRef100_Q6ZFS5 Putative receptor-type protein kinase LRK1 [Ory... 388 e-106
UniRef100_Q6YX21 Putative receptor kinase Lecrk [Oryza sativa] 383 e-105
UniRef100_Q7XIH4 Putative lectin-like protein kinase [Oryza sativa] 365 3e-99
UniRef100_Q6ZFS6 Putative receptor-type protein kinase LRK1 [Ory... 364 4e-99
UniRef100_Q6ZFS1 Putative receptor-type protein kinase LRK1 [Ory... 362 2e-98
UniRef100_Q7X756 OSJNBb0096E05.1 protein [Oryza sativa] 351 4e-95
UniRef100_Q7XL08 OSJNBa0065J03.1 protein [Oryza sativa] 350 6e-95
UniRef100_Q9FHG4 Serine/threonine-specific kinase like protein [... 346 1e-93
UniRef100_Q84VX6 At3g53810 [Arabidopsis thaliana] 342 2e-92
UniRef100_Q9M345 Serine/threonine-specific kinase like protein [... 342 2e-92
UniRef100_O81292 T14P8.3 protein [Arabidopsis thaliana] 335 3e-90
UniRef100_Q9XIS3 Lectin-like protein kinase [Populus nigra] 333 8e-90
>UniRef100_Q6UY57 Lectin-like receptor kinase 1;1 [Medicago truncatula]
Length = 678
Score = 1369 bits (3544), Expect = 0.0
Identities = 678/678 (100%), Positives = 678/678 (100%)
Query: 1 MLNSPSSCSSNIINQSKNILVLFSLLLVLSYLPKTNSLLFNITNFDDPTVASNMSYQGDG 60
MLNSPSSCSSNIINQSKNILVLFSLLLVLSYLPKTNSLLFNITNFDDPTVASNMSYQGDG
Sbjct: 1 MLNSPSSCSSNIINQSKNILVLFSLLLVLSYLPKTNSLLFNITNFDDPTVASNMSYQGDG 60
Query: 61 KSTNGSIDLNKVSYLFRVGRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGDGF 120
KSTNGSIDLNKVSYLFRVGRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGDGF
Sbjct: 61 KSTNGSIDLNKVSYLFRVGRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGDGF 120
Query: 121 VFYLAPLGYQIPPNSAGGVYGLFNATTNSNFVMNYVVGVEFDTFVGPTDPPMKHVGIDDN 180
VFYLAPLGYQIPPNSAGGVYGLFNATTNSNFVMNYVVGVEFDTFVGPTDPPMKHVGIDDN
Sbjct: 121 VFYLAPLGYQIPPNSAGGVYGLFNATTNSNFVMNYVVGVEFDTFVGPTDPPMKHVGIDDN 180
Query: 181 ALTSVAFGKFDIDKNLGRVCYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSSISYQID 240
ALTSVAFGKFDIDKNLGRVCYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSSISYQID
Sbjct: 181 ALTSVAFGKFDIDKNLGRVCYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSSISYQID 240
Query: 241 LMKKLPEFVNIGFSASTGLSTESNVIHSWEFSSNLEDSNSTTSLVEGNDGKGSLKTVIVV 300
LMKKLPEFVNIGFSASTGLSTESNVIHSWEFSSNLEDSNSTTSLVEGNDGKGSLKTVIVV
Sbjct: 241 LMKKLPEFVNIGFSASTGLSTESNVIHSWEFSSNLEDSNSTTSLVEGNDGKGSLKTVIVV 300
Query: 301 VAVIVPVILVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLDKATIPRRFEYSEL 360
VAVIVPVILVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLDKATIPRRFEYSEL
Sbjct: 301 VAVIVPVILVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLDKATIPRRFEYSEL 360
Query: 361 VAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIISRLI 420
VAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIISRLI
Sbjct: 361 VAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIISRLI 420
Query: 421 HRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKSLAWEVRYKVALGVANALRYLH 480
HRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKSLAWEVRYKVALGVANALRYLH
Sbjct: 421 HRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKSLAWEVRYKVALGVANALRYLH 480
Query: 481 DDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGYLAPEYIN 540
DDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGYLAPEYIN
Sbjct: 481 DDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGYLAPEYIN 540
Query: 541 GGRASKESDMYSFGIVALELATGRRFFQDGDFHVPLMNWVWGLYVEGNLMCAADEKLNME 600
GGRASKESDMYSFGIVALELATGRRFFQDGDFHVPLMNWVWGLYVEGNLMCAADEKLNME
Sbjct: 541 GGRASKESDMYSFGIVALELATGRRFFQDGDFHVPLMNWVWGLYVEGNLMCAADEKLNME 600
Query: 601 FDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPELPLDMHDRAPPIVPFRQS 660
FDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPELPLDMHDRAPPIVPFRQS
Sbjct: 601 FDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPELPLDMHDRAPPIVPFRQS 660
Query: 661 NGPSMAPPMTNSLITSGR 678
NGPSMAPPMTNSLITSGR
Sbjct: 661 NGPSMAPPMTNSLITSGR 678
>UniRef100_Q9LXA5 Lectin-like protein kinase-like [Arabidopsis thaliana]
Length = 651
Score = 512 bits (1318), Expect = e-143
Identities = 289/640 (45%), Positives = 401/640 (62%), Gaps = 32/640 (5%)
Query: 18 NILVLFSLLLVLSYLPKTNSLLFNITNFDDPTVASNMSYQGDGKSTNGSIDLNKVSYLFR 77
N ++LFS +LVL P S+ FNI+ F S ++YQGD ++ NG+++L + Y R
Sbjct: 3 NSILLFSFVLVL---PFVCSVQFNISRFGSDV--SEIAYQGDARA-NGAVELTNIDYTCR 56
Query: 78 VGRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDT--TYGDGFVFYLAPLGYQIPPNS 135
G A Y + + LW+ T+ + F+TRFSF ID N YG GF F+LAP Q+PPNS
Sbjct: 57 AGWATYGKQVPLWNPGTSKPSDFSTRFSFRIDTRNVGYGNYGHGFAFFLAPARIQLPPNS 116
Query: 136 AGGVYGLFNATTNSNFVMNYVVGVEFDTFVGPTDPPM---KHVGIDDNALTSVAFGKFDI 192
AGG GLFN T N + V VEFDTF P P+ HVGI++N+L S + ++
Sbjct: 117 AGGFLGLFNGTNNQSSAFPLVY-VEFDTFTNPEWDPLDVKSHVGINNNSLVSSNYTSWNA 175
Query: 193 DKNLGRVCYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSSISYQIDLMKKLPEFVNIG 252
+ + VLI Y+S + L V W++ NSS+SY IDL K LP V IG
Sbjct: 176 TSHNQDIGRVLIFYDSARRNLSVSWTYD----LTSDPLENSSLSYIIDLSKVLPSEVTIG 231
Query: 253 FSASTGLSTESNVIHSWEFSSNLEDSNSTTSLVEGNDGKGSLKTVIVVVAVIVPVILVFL 312
FSA++G TE N + SWEFSS+LE L++ + K +I+ ++V V+L F
Sbjct: 232 FSATSGGVTEGNRLLSWEFSSSLE-------LIDIKKSQNDKKGMIIGISVSGFVLLTFF 284
Query: 313 IASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLDKATIPRRFEYSELVAATNGFDDDRM 372
I S++ ++ K+++K +E + I DL++ PR+F Y +L +A N F DDR
Sbjct: 285 ITSLIVFLKRKQQKKKAEETENLTSI----NEDLERGAGPRKFTYKDLASAANNFADDRK 340
Query: 373 LGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIISRLIHRNLVQFIGWCH 432
LG GG+G VY+G L+ L +VA+K+ + +R F+ EV+IIS L HRNLVQ IGWCH
Sbjct: 341 LGEGGFGAVYRGYLNSLDMMVAIKKFAGGSKQGKREFVTEVKIISSLRHRNLVQLIGWCH 400
Query: 433 EQDELLLVFEYMPNGSLDTHLFGDKKSLAWEVRYKVALGVANALRYLHDDAEQCVLHRDI 492
E+DE L+++E+MPNGSLD HLFG K LAW VR K+ LG+A+AL YLH++ EQCV+HRDI
Sbjct: 401 EKDEFLMIYEFMPNGSLDAHLFGKKPHLAWHVRCKITLGLASALLYLHEEWEQCVVHRDI 460
Query: 493 KSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGYLAPEYINGGRASKESDMYS 552
K++NV+LD++F+ KLGDFG+A+L+D L Q T + GT+GY+APEYI+ GRASKESD+YS
Sbjct: 461 KASNVMLDSNFNAKLGDFGLARLMDHELGPQTTGLAGTFGYMAPEYISTGRASKESDVYS 520
Query: 553 FGIVALELATGRRFFQDGDFHV-PLMNWV---WGLYVEGNLMCAADEKLNM-EFDVSEMK 607
FG+V LE+ TGR+ V P+ N V W LY +G ++ A DEKL + FD + +
Sbjct: 521 FGVVTLEIVTGRKSVDRRQGRVEPVTNLVEKMWDLYGKGEVITAIDEKLRIGGFDEKQAE 580
Query: 608 SLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPELPLDM 647
L+IVGLWC H + RP + I+VL LE +P LP M
Sbjct: 581 CLMIVGLWCAHPDVNTRPSIKQAIQVLNLEAPVPHLPTKM 620
>UniRef100_Q9LSL5 Receptor protein kinase-like protein [Arabidopsis thaliana]
Length = 675
Score = 459 bits (1180), Expect = e-127
Identities = 276/692 (39%), Positives = 405/692 (57%), Gaps = 61/692 (8%)
Query: 2 LNSPSSCSSNIINQSKNILVLFSLLLVLSYLPKTNSLLFNITNF--DDPTVASNMSYQGD 59
L+S SS S++I+ SL L L ++ +SL FN T+F DP ++ Y GD
Sbjct: 10 LSSSSSMSNSIL--------FLSLFLFLPFV--VDSLYFNFTSFRQGDP---GDIFYHGD 56
Query: 60 GK-STNGSIDLNKVSYLFRVGRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGD 118
+G+++ N +VG YS+ + +W KT + F+T FSF ID N + G
Sbjct: 57 ATPDEDGTVNFNNAEQTSQVGWITYSKKVPIWSHKTGKASDFSTSFSFKIDARNLSADGH 116
Query: 119 GFVFYLAPLGYQIPPNSAGGVYGLFNATTNSNFVMNY-VVGVEFDTFVGPTDPPM---KH 174
G F+LAP+G Q+P S GG LF T +N+ ++ +V VEFDTF P P H
Sbjct: 117 GICFFLAPMGAQLPAYSVGGFLNLF--TRKNNYSSSFPLVHVEFDTFNNPGWDPNDVGSH 174
Query: 175 VGIDDNALTSVAFGKFDIDKNLGRVCYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSS 234
VGI++N+L S + ++ + +C+ I Y+S K L V W+++ +SS
Sbjct: 175 VGINNNSLVSSNYTSWNASSHSQDICHAKISYDSVTKNLSVTWAYE--LTATSDPKESSS 232
Query: 235 ISYQIDLMKKLPEFVNIGFSASTGLSTESNVIHSWEFSSNLED--SNSTTSLVEGNDGKG 292
+SY IDL K LP V GF A+ G +TE + + SWE SS+L+ ++S LV G G
Sbjct: 233 LSYIIDLAKVLPSDVMFGFIAAAGTNTEEHRLLSWELSSSLDSDKADSRIGLVIGISASG 292
Query: 293 SLKTVIVVVAVIVPVILVFL-IASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLDKATI 351
V L F+ I ++V W +RK+K D E I K DL++
Sbjct: 293 F-------------VFLTFMVITTVVVWSRKQRKKKERDI---ENMISINK--DLEREAG 334
Query: 352 PRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFIN 411
PR+F Y +LV+ATN F R LG GG+G VY+G L + +VAVK++ D + F+N
Sbjct: 335 PRKFSYKDLVSATNRFSSHRKLGEGGFGAVYEGNLKEINTMVAVKKLSGDSRQGKNEFLN 394
Query: 412 EVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKSL-AWEVRYKVAL 470
EV+IIS+L HRNLVQ IGWC+E++E LL++E +PNGSL++HLFG + +L +W++RYK+ L
Sbjct: 395 EVKIISKLRHRNLVQLIGWCNEKNEFLLIYELVPNGSLNSHLFGKRPNLLSWDIRYKIGL 454
Query: 471 GVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGT 530
G+A+AL YLH++ +QCVLHRDIK++N++LD++F+ KLGDFG+A+L++ L + T + GT
Sbjct: 455 GLASALLYLHEEWDQCVLHRDIKASNIMLDSEFNVKLGDFGLARLMNHELGSHTTGLAGT 514
Query: 531 YGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQ---------DGDFHVPLMNWVW 581
+GY+APEY+ G ASKESD+YSFGIV LE+ TGR+ + + D L+ VW
Sbjct: 515 FGYMAPEYVMKGSASKESDIYSFGIVLLEIVTGRKSLERTQEDNSDTESDDEKSLVEKVW 574
Query: 582 GLYVEGNLMCA-ADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMAL 640
LY + L+ + D+KL +FD E + LL++GLWC H + RP + I+V+ E L
Sbjct: 575 ELYGKQELITSCVDDKLGEDFDKKEAECLLVLGLWCAHPDKNSRPSIKQGIQVMNFESPL 634
Query: 641 PELPLDMHDRAPPIVPFRQSNGPSMAPPMTNS 672
P+LPL P+ + S S + P NS
Sbjct: 635 PDLPLKR-----PVAMYYISTTTSSSSPSVNS 661
>UniRef100_Q7XIH7 Putative lectin-like protein kinase [Oryza sativa]
Length = 689
Score = 452 bits (1163), Expect = e-125
Identities = 255/592 (43%), Positives = 358/592 (60%), Gaps = 35/592 (5%)
Query: 78 VGRAFYSQPLHLWDKKTNTLTSFTTRFSFTI-DKLNDTTYGDGFVFYLAPLGYQIPPNSA 136
VGRA Y+ P+ LWD T L SFTTRF+F I ND++YG+G F+L+ +P NS
Sbjct: 71 VGRAIYTDPVPLWDSTTGQLASFTTRFTFKIYAPTNDSSYGEGLAFFLSSYPSVVPNNSM 130
Query: 137 GGVYGLFNATTNSNFVMNYVVGVEFDTFVGPTDPPMKHVGIDDNALTSVAFGKFDIDKNL 196
G GLF+ + + + +N +V VEFD+ DP HVGI+ +++ SVA + N
Sbjct: 131 DGYLGLFSNSNDQSDPLNQIVAVEFDSHKNTWDPDGNHVGINIHSIVSVANVTWRSSIND 190
Query: 197 GRVCYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSSISYQIDLMKKLPEFVNIGFSAS 256
GR+ + Y ++ + L VF S++ GNSS+SY +DL K LP+ V+IGFSAS
Sbjct: 191 GRIANAWVTYQANSRNLSVFLSYQDN----PQFSGNSSLSYSVDLSKYLPDKVSIGFSAS 246
Query: 257 TGLSTESNVIHSWEFSSNLEDSNSTTSLVEGNDGKG----SLKTVIVVVAVIVPVILVFL 312
TG E + I WEF S + L++ KG SL T VV + ++ FL
Sbjct: 247 TGKFVELHQILYWEFDS------TDVHLMKTEKTKGILVISLSTSGSVVVCSIGLVCFFL 300
Query: 313 IASIVGWVIVKRKRK----NCDEGLDEYGIPFPKKFDLDKATIPRRFEYSELVAATNGFD 368
+ R+++ +CDE +D + +K PRRF+Y+ELV AT+ F
Sbjct: 301 CFRRIRRTTRSREKEKEKLDCDESIDS---------EFEKGKGPRRFQYNELVVATDNFA 351
Query: 369 DDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIISRLIHRNLVQFI 428
+R LG GG+G VY+G L +A+KR+ + +I+EV+IISRL HRNLVQ +
Sbjct: 352 AERKLGEGGFGAVYQGFLKDQNIEIAIKRVAKGSTQGRKEYISEVKIISRLRHRNLVQLV 411
Query: 429 GWCHEQDELLLVFEYMPNGSLDTHLFGDKKSLAWEVRYKVALGVANALRYLHDDAEQCVL 488
GWCHE E LLV+E+MPN SLD HL+ LAW +R+K+ +GVA+AL YLH++ EQCV+
Sbjct: 412 GWCHEHGEFLLVYEFMPNRSLDKHLYDGGNLLAWPLRFKITIGVASALLYLHEEWEQCVV 471
Query: 489 HRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGYLAPEYINGGRASKES 548
HRD+K +NV+LD+ F+ KLGDFG+A+LVD +Q T + GT GY+APE + G+ASKE+
Sbjct: 472 HRDVKPSNVMLDSGFNAKLGDFGLARLVDHDRGSQTTVIAGTMGYMAPECVTTGKASKET 531
Query: 549 DMYSFGIVALELATGRRFF---QDGDFHVPLMNWVWGLYVEGNLMCAADEKLNMEFDVSE 605
D+YSFGI+ALE+A GRR +D D + L+ WVW LY ++ A D +L+ EF+ E
Sbjct: 532 DVYSFGILALEIACGRRPVVPKEDND-RISLVQWVWDLYGRNEILNAIDGRLDGEFEERE 590
Query: 606 MKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPELPLDMHDR---APPI 654
+ SL++VGLWC H + RP +VI VL+ E LP+LP M APPI
Sbjct: 591 VISLMVVGLWCAHPDYNIRPSIRQVISVLKFEAPLPDLPPKMPVAMYFAPPI 642
>UniRef100_Q94DZ6 Putative lectin-like receptor kinase 1;1 [Oryza sativa]
Length = 696
Score = 422 bits (1085), Expect = e-116
Identities = 257/654 (39%), Positives = 377/654 (57%), Gaps = 47/654 (7%)
Query: 20 LVLFSLLLVLSYLPKTNSLLFNITNFDDPTVASNMSYQGDGKSTNGSIDLNKVSYLFRV- 78
L+L +LLL+L S +N T+ D T S +++QGD N I L + + +
Sbjct: 13 LLLVALLLLLPSPAAAFSFTYNFTSAD--TAPSGIAFQGDA-FFNKFIRLTRDERIGPIT 69
Query: 79 ---GRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTY-GDGFVFYLAPLGYQIPPN 134
GRAF+S+P+ L D + SF+T FSF+I + + GDG F+L+P +P +
Sbjct: 70 SSAGRAFFSRPVPLCDPVSRRRASFSTAFSFSIAAPDPSAASGDGLAFFLSPFPSVLPNS 129
Query: 135 SAGGVYGLFNATTNSNFVMNY---VVGVEFDTFVGPTDPPMKHVGIDDNALTSVAFGKFD 191
SAGG+ GLFN+ + + +V VEFDT+ DP HVG+D + S A +
Sbjct: 130 SAGGLLGLFNSFSRGGAAAAHPRPLVAVEFDTYKNEWDPSDDHVGVDLGGIVSAATVDWP 189
Query: 192 IDKNLGRVCYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSSISYQIDLMKKLPEFVNI 251
GR + + Y+ K L V S+ + + + Y +DLM+ LP+ V +
Sbjct: 190 TSMKDGRRAHARVAYDGQAKNLTVALSYGD--AAAAAALTDPVLWYAVDLMEYLPDAVAV 247
Query: 252 GFSASTGLSTESNVIHSWEFSSNLEDSNSTTSLVEGNDGKGSLKTVIVVVAVIVPVILVF 311
GFSA+TG + E + + WEF+S+++ T ++ VV + ++LV
Sbjct: 248 GFSAATGEAAELHQVLYWEFTSSIDTKEETV--------------ILWVVLGLCGLLLVL 293
Query: 312 LIASIVGWVIVKRKRKNCDEG-LDE---YGIPFPKKFDLDKATIPRRFEYSELVAATNGF 367
+ A ++ +V RK +G +DE Y ++F ++ PRRF YS+L AAT F
Sbjct: 294 VAAGVLWFVSQWRKAGELADGDIDEEMGYDELADEEFFVESG--PRRFRYSDLAAATKNF 351
Query: 368 DDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIISRLIHRNLVQF 427
D+R LG+GG+G VY+G L LG VA+KR+ + + EVRIIS+L HR+LV+
Sbjct: 352 SDERKLGQGGFGAVYRGFLKELGLAVAIKRVSKGSTQGRKEYAAEVRIISQLRHRHLVRL 411
Query: 428 IGWCHE-QDELLLVFEYMPNGSLDTHLFG----DKKS------LAWEVRYKVALGVANAL 476
+GWCHE + + LLV+E MPNGS+D HL+G KK+ L+W RY VALG+A+AL
Sbjct: 412 VGWCHEHRGDFLLVYELMPNGSVDRHLYGGGGGSKKAGGAAPPLSWPTRYNVALGLASAL 471
Query: 477 RYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGYLAP 536
YLH++ QCV+HRDIK +NV+LD FS KLGDFG+AKLV+ + T + GT GYLAP
Sbjct: 472 LYLHEECPQCVVHRDIKPSNVMLDATFSAKLGDFGLAKLVEHGSQPHTTVLAGTLGYLAP 531
Query: 537 EYINGGRASKESDMYSFGIVALELATGRR---FFQDGDFHVPLMNWVWGLYVEGNLMCAA 593
E + GRAS+ESD+YSFG+VALE+A GRR ++ L+ WVW LY + ++ AA
Sbjct: 532 ECVITGRASRESDVYSFGVVALEIACGRRPAELDEEDPSKARLVPWVWELYGKRAILEAA 591
Query: 594 DEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPELPLDM 647
D++LN +FD+ +M+ L++VGLWC H + RP + + VL+ E LP LP M
Sbjct: 592 DQRLNGKFDLEQMERLMVVGLWCAHPDHAHRPSIRQALNVLKFEAPLPSLPPKM 645
>UniRef100_Q6ZFB7 Putative receptor-type protein kinase LRK1 [Oryza sativa]
Length = 676
Score = 416 bits (1068), Expect = e-114
Identities = 263/672 (39%), Positives = 373/672 (55%), Gaps = 58/672 (8%)
Query: 22 LFSLLLVLSYLPKT--NSLLFNITNFDDPTVASNMSYQGDGKSTNGS-IDL-----NKVS 73
L SLL++L LP ++ FN ++F + + N++ QG I+L N +S
Sbjct: 19 LVSLLVMLRCLPSVVATTVSFNYSSFSN--ASKNITLQGSAALAGAEWIELTKGKGNNLS 76
Query: 74 YLFRVGRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGDGFVFYLAPLGYQIPP 133
+GR Y+ P+ LWD T + SFTTRFSF I N + GDG F+L ++P
Sbjct: 77 SGGTMGRMVYTPPVQLWDAATGEVASFTTRFSFNITPKNKSNKGDGMTFFLVSYPSRMPY 136
Query: 134 NSAGGVYGLFNATTNSNFVMNYVVGVEFDT----FVGPTDPPMKHVGIDDNALTSVAFGK 189
GG GL + T ++ + V VEFDT F+ P D H+GID NAL SV
Sbjct: 137 MGYGGALGLTSQTFDNATAGDRFVAVEFDTYNNSFLDP-DATYDHIGIDVNALRSVKTES 195
Query: 190 FDIDKNLGRVCYVLIDYNSDEKMLEV-FWSFKGRFVKGDGSYGNSSISYQIDLMKKLPEF 248
+G + ++DYNS+ ++ V W+ +GS ++S ++DL LPE
Sbjct: 196 LPSYILIGNMT-AIVDYNSNSSIMSVKLWA--------NGSTTPYNLSSKVDLKSALPEK 246
Query: 249 VNIGFSASTGLSTESNVIHSWEFSSNLEDSNSTTSLVEGNDGKGSLKTVIVVVAVIVPVI 308
V +GFSA+TG S E + + SW F+ LE T G +G +V A + ++
Sbjct: 247 VAVGFSAATGSSFEQHQLRSWYFNLTLEQKQPT-----GQHSRGG----VVAGATVGAIL 297
Query: 309 LVFLIASIVGWVIVKRKRKNC--DEGLDEYGIPFPKKFDLDKATIPRRFEYSELVAATNG 366
+ L+ ++V ++ +R+RK +E D G P +++ T PRRF Y LV AT
Sbjct: 298 FIVLLFTMVAILVRRRQRKKMREEEEDDSEGDPI---VEIEMGTGPRRFPYHILVNATKS 354
Query: 367 FDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERV-FINEVRIISRLIHRNLV 425
F + LG+GG+G VY+G L LG VA+KR D R + +E+++ISRL HRNLV
Sbjct: 355 FAAEEKLGQGGFGAVYRGNLRELGLDVAIKRFAKDSSKQGRKEYKSEIKVISRLRHRNLV 414
Query: 426 QFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKSLAWEVRYKVALGVANALRYLHDDAEQ 485
Q IGWCH +DELLLV+E +PN SLD HL G+ L W +R + LG+ NAL YLH++ EQ
Sbjct: 415 QLIGWCHGRDELLLVYELVPNRSLDVHLHGNGTFLTWPMRINIVLGLGNALLYLHEEWEQ 474
Query: 486 CVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQ-RTDVVGTYGYLAPEYINGGRA 544
CV+HRDIK +N++LD F+ KLGDFG+A+L+D + Q T GT GYL PE + G+A
Sbjct: 475 CVVHRDIKPSNIMLDESFNAKLGDFGLARLIDHNVGVQTMTHPSGTPGYLDPECVITGKA 534
Query: 545 SKESDMYSFGIVALELATGRR----FFQDGDFHVPLMNWVWGLYVEGNLMCAADEKLNME 600
S ESD+YSFG+V LE+A GRR + L+ WVW LY +G ++ AADE+LN +
Sbjct: 535 SAESDVYSFGVVLLEVACGRRPMSLLDNQNNSLFRLVEWVWDLYGQGVVLKAADERLNND 594
Query: 601 FDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPELPLDMHDRAPPIVPFRQS 660
+D + M+ ++ VGLWC H + RP + VLQ LP LP M P+
Sbjct: 595 YDATSMECVMAVGLWCAHPDRYARPSIRAAMTVLQSNGPLPVLPSKM-----PV------ 643
Query: 661 NGPSMAPPMTNS 672
P APPM +S
Sbjct: 644 --PIYAPPMASS 653
>UniRef100_Q6ZFT2 Putative receptor-type protein kinase LRK1 [Oryza sativa]
Length = 688
Score = 405 bits (1040), Expect = e-111
Identities = 261/665 (39%), Positives = 365/665 (54%), Gaps = 40/665 (6%)
Query: 40 FNITNFDDPTVASN----MSYQGDGKSTNGSIDLNKVSYLFRVGRAFYSQPLHLWDKKTN 95
F +F+ PT AS+ + +G+ + G ID++ S VGR F+ P+ LWD T
Sbjct: 28 FAALSFNYPTFASSHNQYIEIEGNASVSVGYIDISANSVGNNVGRVFHKPPVQLWDAATG 87
Query: 96 TLTSFTTRFSFTIDKLNDTTYGDGFVFYLAPLGYQIPPNSAGGV-YGLFNATTNSNFVMN 154
+ SFTTRFSF I N + GDG F+L ++P AGG GL N T + N
Sbjct: 88 EVASFTTRFSFNIIPGNRSKKGDGMTFFLTSYPSRLPEGDAGGQNLGLTNQTVGVSTGEN 147
Query: 155 YVVGVEFDTFVGPTDPPMK--HVGIDDNALTSVAFGKFDIDKNLGRVCYVLIDYNSDEKM 212
V VEFDTFV P DP H+GID N++ SV +G + +DYN++ ++
Sbjct: 148 RFVAVEFDTFVNPFDPNATNDHIGIDVNSVVSVTNESLPNFSLIGNMT-ATVDYNNNSRI 206
Query: 213 LEV-FWSFKGRFVKGDGSYGNSSISYQIDLMKKLPEFVNIGFSASTGLSTESNVIHSWEF 271
L V W +GS ++S +DL + LPE V IGFSAS G + E + + SW F
Sbjct: 207 LSVKLWI--------NGSTTPYTLSSMVDLKRALPENVTIGFSASIGSAYEQHQLTSWYF 258
Query: 272 SSNLEDSNSTTSLVEG-------NDGKGSLKTVIVVVAVIVPVILVFLIASIVGWVIVKR 324
S + V + S + +V A + V+ V L+ ++V ++ +R
Sbjct: 259 KSTSSFEQKLAAEVASPPPPSSPSPPPTSRRGGVVAGATVGVVMFVILLFTMVQVLVRRR 318
Query: 325 KRKNCDEGLDE--YGIPFPKK----FDLDKATIPRRFEYSELVAATNGFDDDRMLGRGGY 378
+ K E D +G +++ T PRRF Y +LV AT F + LG+GG+
Sbjct: 319 QSKKRREAQDGSWHGSDDDDDGELIMEIEMGTGPRRFPYHKLVDATKSFAPEEKLGQGGF 378
Query: 379 GQVYKGALSYLGKIVAVKRIFADFENSERV-FINEVRIISRLIHRNLVQFIGWCHEQDEL 437
G VY+G L LG VA+KR + R + +E+++ISRL HRNLVQ IGWCH + EL
Sbjct: 379 GAVYRGYLRELGLAVAIKRFAKNSSKQGRKEYKSEIKVISRLRHRNLVQLIGWCHGRTEL 438
Query: 438 LLVFEYMPNGSLDTHLFGDKKSLAWEVRYKVALGVANALRYLHDDAEQCVLHRDIKSANV 497
LLV+E PN SLD HL G+ L W +R + G+ +AL YLH++ +QCV+HRDIK +NV
Sbjct: 439 LLVYELFPNRSLDVHLHGNGTFLTWPMRINIVHGLGSALLYLHEEWDQCVVHRDIKPSNV 498
Query: 498 LLDTDFSTKLGDFGMAKLVDPMLRTQ-RTDVVGTYGYLAPEYINGGRASKESDMYSFGIV 556
+LD F+ KLGDFG+A+L+D + Q T GT GYL PE + G+AS ESD+YSFGIV
Sbjct: 499 MLDESFNAKLGDFGLARLIDHAVGIQTMTHPSGTPGYLDPECVITGKASAESDVYSFGIV 558
Query: 557 ALELATGRR--FFQD--GDFHVPLMNWVWGLYVEGNLMCAADEKLNMEFDVSEMKSLLIV 612
LE+A GRR QD + L+ WVW LY +G ++ AADE+LN E+D + M+ ++ V
Sbjct: 559 LLEVACGRRPISLQDTQNNCLFRLVEWVWDLYGQGAVLKAADERLNNEYDTTSMECVMAV 618
Query: 613 GLWCTHSNDKERPKAYEVIKVLQLEMALPELPLDM--HDRAPPIVPFRQSNGPSMAPPMT 670
GLWC H + RP V+ VLQ LP LP M APP+ PS + M+
Sbjct: 619 GLWCAHPDRCARPSIRAVMAVLQSNGPLPVLPAKMPVPTYAPPMA--SAEGQPSSSTRMS 676
Query: 671 NSLIT 675
+S +T
Sbjct: 677 SSSLT 681
>UniRef100_Q6ZFR6 Putative receptor-type protein kinase LRK1 [Oryza sativa]
Length = 705
Score = 404 bits (1039), Expect = e-111
Identities = 255/639 (39%), Positives = 356/639 (54%), Gaps = 56/639 (8%)
Query: 78 VGRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGDGFVFYLAPLGYQIPPNSAG 137
+GR YS P+ LW+ T + SFTTRFSF I N GDG F+L ++P + G
Sbjct: 55 MGRVAYSPPVQLWEAATGEVASFTTRFSFNITPTNLDNKGDGMAFFLVGYPSRMPDTADG 114
Query: 138 GVYGLFNATTNSNFVM---NYVVGVEFDTFVGPTDPPMK--HVGIDDNALTSV---AFGK 189
G GL + T ++ VM N V VEFDTF DP H+G+D N++ SV +
Sbjct: 115 GALGLTSRTFDA--VMSGDNRFVAVEFDTFNNSFDPSATYDHIGVDVNSIVSVQTESLPS 172
Query: 190 FDIDKNLGRVCYVLIDYNSDEKMLEVFWSFKGRFVKG--DGSYGNSSISYQIDLMKKLPE 247
F + N+ + +DYNS +L V + VK +GS ++S +DL LPE
Sbjct: 173 FSLTGNMAAI----VDYNSSSSILSV------QLVKTWTNGSTTLYNLSTTVDLKTALPE 222
Query: 248 FVNIGFSASTGLSTESNVIHSWEFSSNLEDSNSTTSLVEGNDGKGSLKTVIVVVAVIVPV 307
V++GFSA+TG S E + +HSW F+S+ + + + L VI +
Sbjct: 223 KVSVGFSAATGSSLELHQLHSWYFNSSFQQNPPPAAQPSPTTSGPGLAGVIAGATAGGAL 282
Query: 308 ILVFLIASIVGWVIVKRKRKNC-----------------DEGLDEYGIPFPKKFDLDKAT 350
+V L A IV V+V+R+R D+ D+ G P +++
Sbjct: 283 FVVLLFAMIV--VLVRRRRSKKRREAEEAEEARHVGLAGDDDDDDDGEPI---VEIEMGM 337
Query: 351 IPRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADF-ENSERVF 409
PR+ Y ELV AT F + LG+GG+G VY+G L G VA+KR D + ++ +
Sbjct: 338 GPRQIPYHELVEATKSFAAEEKLGQGGFGAVYRGYLREQGLAVAIKRFAKDSSKQGKKEY 397
Query: 410 INEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKSLAWEVRYKVA 469
+E+++ISRL HRNLVQ IGWCH +DELLLV+E +PN SLD HL G+ L W +R K+
Sbjct: 398 RSEIKVISRLRHRNLVQLIGWCHGRDELLLVYELVPNRSLDIHLHGNGTFLTWPMRVKIV 457
Query: 470 LGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQ-RTDVV 528
LG+ +AL YLH++ EQCV+HRDIK +NV+LD F+ KLGDFG+A+ +D + Q T V
Sbjct: 458 LGLGSALFYLHEEWEQCVVHRDIKPSNVMLDESFNAKLGDFGLARFIDHAVGMQTMTAVS 517
Query: 529 GTYGYLAPEYINGGRASKESDMYSFGIVALELATGRR--FFQDGDFH--VPLMNWVWGLY 584
GT GY+ PE + GRAS ESD+YSFGIV LE+A GRR QD + L+ WVW L+
Sbjct: 518 GTPGYVDPECVITGRASSESDVYSFGIVLLEVACGRRPMSLQDNQKNGIFRLVEWVWDLH 577
Query: 585 VEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPELP 644
+G+++ AADE+LN ++DVSEM+ ++ VGLWC H + RP + +LQ LP LP
Sbjct: 578 GQGDVISAADERLNGDYDVSEMERVITVGLWCAHPDPSARPSIRAAMAMLQSSGQLPVLP 637
Query: 645 LDM--HDRAPPIVP----FRQSNGPSMAPPMTNSLITSG 677
M APP+ F S G + +S TSG
Sbjct: 638 AKMPVPTYAPPVASVEGLFTSSTGMLSSSATQSSSTTSG 676
>UniRef100_Q6ZFS5 Putative receptor-type protein kinase LRK1 [Oryza sativa]
Length = 683
Score = 388 bits (996), Expect = e-106
Identities = 248/647 (38%), Positives = 351/647 (53%), Gaps = 58/647 (8%)
Query: 44 NFDDPTVASN----MSYQGDGKSTNGSIDLNKVSYLFRVGRAFYSQPLHLWDKKTNTLTS 99
+F+ PT AS+ + +G+ + G ID++ S VGR FY P+ LWD T + S
Sbjct: 32 SFNYPTFASSHNQYIEIEGNASVSVGYIDISANSVGNNVGRVFYKPPVQLWDAATGEVAS 91
Query: 100 FTTRFSFTIDKLNDTTY-GDGFVFYLAPLGYQIPPNSAGGV-YGLFNATT-NSNFVMNYV 156
FTTRFSF I +D + GDG F+L ++P GG GL N T N + N
Sbjct: 92 FTTRFSFNIIAPSDRSKKGDGMAFFLTSYPSRLPVGHEGGENLGLTNQTVGNVSTGQNRF 151
Query: 157 VGVEFDTFVGPTDPPMK--HVGIDDNALTSVAFGKFDIDKNLGRVCYVLIDYNSDEKMLE 214
V VEFDTFV P DP H+GID N++ SV +G + +DYN++ ++L
Sbjct: 152 VAVEFDTFVNPFDPNTTNDHIGIDVNSVVSVTNESLPNFSLIGNMT-ATVDYNNNSRILS 210
Query: 215 V-FWSFKGRFVKGDGSYGNSSISYQIDLMKKLPEFVNIGFSASTGLSTESNVIHSWEFSS 273
V W +GS ++S +DL + LPE + +GFSAS G + E + + SW F S
Sbjct: 211 VKLWI--------NGSTTPYTLSSMVDLKRALPENITVGFSASIGSAYEQHQLTSWYFKS 262
Query: 274 NLEDSNSTTSLVEG-------------------NDGKGSLKTVIVVVAVIVPVILVFLIA 314
+ + V + +G + V AV+ ++L ++A
Sbjct: 263 SSSFEQKLAAKVASPPPPSSPSPPPPSLTPITSHSRRGGVVAGATVGAVMFVILLFAMVA 322
Query: 315 SIVGWVIVKRKRKNCDEGL-----DEYGIPFPKKFDLDKATIPRRFEYSELVAATNGFDD 369
+V K++R+ D G D+ G P +++ PRRF Y ELV AT F
Sbjct: 323 VLVRRRQSKKRREAEDGGWHGSDDDDDGEPI---VEIEMGMGPRRFPYHELVDATKSFAT 379
Query: 370 DRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERV-FINEVRIISRLIHRNLVQFI 428
+ LG+GG+G VY+G L LG VA+KR D R + +E+++ISRL HRNLVQ I
Sbjct: 380 EEKLGQGGFGAVYRGYLRELGLAVAIKRFAKDSSKQGRKEYKSEIKVISRLRHRNLVQLI 439
Query: 429 GWCHEQDELLLVFEYMPNGSLDTHLFGDKKSLAWEV------RYKVALGVANALRYLHDD 482
GWCH + ELLLV+E +PN SLD HL G+ L W + R + G+ +AL YLH++
Sbjct: 440 GWCHGRTELLLVYELVPNRSLDVHLHGNGTFLTWPMRLGCCHRINIVHGLGSALLYLHEE 499
Query: 483 AEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQ-RTDVVGTYGYLAPEYING 541
+QCV+HRDIK +NV+LD F+ KLGDFG+A+L+D + Q T GT GYL PE +
Sbjct: 500 WDQCVVHRDIKPSNVMLDESFNAKLGDFGLARLIDHAVGVQTMTHPSGTPGYLDPECVIT 559
Query: 542 GRASKESDMYSFGIVALELATGRR----FFQDGDFHVPLMNWVWGLYVEGNLMCAADEKL 597
G+AS ESD+YSFG+V LE+A GRR + L+ WVW LY +G ++ AADE+L
Sbjct: 560 GKASAESDVYSFGVVLLEVACGRRPMSLLDNQNNSLFRLVEWVWDLYGQGVVLKAADERL 619
Query: 598 NMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPELP 644
N ++D + M+ ++ VGLWC H + RP + VLQ LP LP
Sbjct: 620 NNDYDATSMECVMAVGLWCVHPDRYARPSIRAAMTVLQSNGPLPVLP 666
>UniRef100_Q6YX21 Putative receptor kinase Lecrk [Oryza sativa]
Length = 750
Score = 383 bits (984), Expect = e-105
Identities = 249/667 (37%), Positives = 362/667 (53%), Gaps = 70/667 (10%)
Query: 33 PKTNSLLFNITNFD-----------DPTVASNMSYQGDG-------KSTNGSIDLNKVSY 74
P +L FN TNF D +++ S+ GDG + +GSI+ ++
Sbjct: 37 PVALALTFNHTNFGPDEQTNIRLEGDAAFSADFSFSGDGGGWVDISANRHGSIEDSR--- 93
Query: 75 LFRVGRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDT----TYGDGFVFYLAPLGYQ 130
GR Y+ P+ LWD T + SFTT FSF I+ G G F+LA +
Sbjct: 94 ----GRVSYALPVPLWDAATGEVASFTTGFSFVINPPKQDGGIDNKGAGMAFFLAGFPSR 149
Query: 131 IPPNSAGGVYGLFNATTNSNFVMN-YVVGVEFDTF----VGPTDPPMKHVGIDDNAL--- 182
+P + + GL N T + + V VEFDTF V D H+G+D N++
Sbjct: 150 LPGSYPYNL-GLTNQTADQVAAGDDRFVAVEFDTFNDTIVHDPDATYDHLGVDVNSVVSK 208
Query: 183 TSVAFGKFDIDKNLGRVCYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSSISYQIDLM 242
T++ F + N+ V ++Y++ +L + G + G + ++SY++DL
Sbjct: 209 TTLTLPSFTLVGNMTAV----VEYDNVSSILAMRLHL-GYGLSGPRHRPDYNLSYKVDLK 263
Query: 243 KKLPEFVNIGFSASTGLSTESNVIHSWEFSSNLEDSNSTTSLVEGNDG----KGSLKTVI 298
LPE V +GFSA+T S E + + SW FSS+LE + + + GS +
Sbjct: 264 SVLPEQVAVGFSAATSTSVELHQLRSWYFSSSLEPKAAPPPVAPPSPSPPPTSGSGSGGV 323
Query: 299 VVVAVIVPVILVFLIASIVGWVIVKRKRKNC----------DEGLDEYGIPFPKKFDLDK 348
V A + + V L+ ++V V++ R+R D+ D G P +++
Sbjct: 324 VAGATVGAALFVVLLLAMVAVVVLVRRRHQRKKMREAEEANDDDDDTEGDPI---MEIEN 380
Query: 349 ATIPRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENS-ER 407
T PRRF Y LV AT F + LG+GG+G VY+G L G VA+KR D N R
Sbjct: 381 GTGPRRFAYHVLVNATKSFAAEEKLGQGGFGAVYRGYLREQGLAVAIKRFIKDSSNQGRR 440
Query: 408 VFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKSLAWEVRYK 467
+ +E+++ISRL HRNLVQ IGWCH DELLLV+E +PN SLD HL G+ L W +R K
Sbjct: 441 EYKSEIKVISRLRHRNLVQLIGWCHGHDELLLVYELVPNRSLDIHLHGNGTFLTWPMRVK 500
Query: 468 VALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQ-RTD 526
+ LG+ +AL YLH++ EQCV+HRDIK +NV+LD F+ KLGDFG+A+ +D ++ Q T
Sbjct: 501 IILGLGSALFYLHEEWEQCVVHRDIKPSNVMLDESFNAKLGDFGLARFIDHIVGMQTMTA 560
Query: 527 VVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFF------QDGDFHVPLMNWV 580
V GT GY+ PE + GRAS ESD+YSFGIV LE+A GRR ++G F L+ W
Sbjct: 561 VSGTPGYVDPECVITGRASAESDVYSFGIVLLEVACGRRPMSLLDSQKNGIFR--LVEWA 618
Query: 581 WGLYVEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMAL 640
W LY +G+++ AADE+LN ++D +EM+ ++++GLWC H + RP + +LQ L
Sbjct: 619 WDLYGKGDILMAADERLNGDYDAAEMERVIVIGLWCAHPDPNARPSIRNAMAMLQSGGQL 678
Query: 641 PELPLDM 647
P LP M
Sbjct: 679 PVLPAKM 685
>UniRef100_Q7XIH4 Putative lectin-like protein kinase [Oryza sativa]
Length = 588
Score = 365 bits (936), Expect = 3e-99
Identities = 217/530 (40%), Positives = 306/530 (56%), Gaps = 27/530 (5%)
Query: 133 PNSAGGVYGLFNATTNSN-FVMNYVVGVEFDTFVGPTDPPMKHVGIDDNALTSVAFGKFD 191
P GG+ G+F +T N +V VEFDTF D H+GID N++ S A K
Sbjct: 3 PLDGGGLLGVFTNSTGMNPSAAAPIVAVEFDTFQNEWDQSSDHIGIDVNSINSTAV-KLL 61
Query: 192 IDKNLGRVCYVLI---DYNSDEKMLEVFWSFKGRFVKGDGSYGNSSISYQIDLMKKLPEF 248
D++L V ++ YN+ +ML V + DG ++ +DL LP
Sbjct: 62 SDRSLSNVTEPMVASVSYNNSTRMLAVMLQMAPQ----DGGK-RYELNSTVDLKSLLPAQ 116
Query: 249 VNIGFSASTGLSTESNVIHSWEFSSNLEDSNSTTSLVEGNDGKGSLKTVIVVVAVIVPVI 308
V IGFSA++G S E + + +W F+S L S N +G ++ V+ V+
Sbjct: 117 VAIGFSAASGWSEERHQVLTWSFNSTLVASEERRE----NATRGRPAAAVLAGVVVASVV 172
Query: 309 LVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDL----DKATIPRRFEYSELVAAT 364
+V ASI +V+++R+R + +EY + FD+ ++ T PRRF YS+L AT
Sbjct: 173 VVG--ASICLFVMIRRRRISRRRTREEYEMGGSDDFDMNDEFEQGTGPRRFLYSQLATAT 230
Query: 365 NGFDDDRMLGRGGYGQVYKGALSY-LGKIVAVKRIFADFENSERVFINEVRIISRLIHRN 423
N F +D LG GG+G VY+G LS G VAVKRI + + + +EV IISRL HRN
Sbjct: 231 NDFSEDGKLGEGGFGSVYRGVLSEPAGVHVAVKRISKTSKQGRKEYASEVSIISRLRHRN 290
Query: 424 LVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKSLAWEVRYKVALGVANALRYLHDDA 483
LVQ +GWCH + + LLV+E +PNGSLD HL+G +L W RY++ALG+ +AL YLH
Sbjct: 291 LVQLVGWCHGRGDFLLVYELVPNGSLDAHLYGGGATLPWPTRYEIALGLGSALLYLHSGY 350
Query: 484 EQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVV-GTYGYLAPEYINGG 542
E+CV+HRDIK +N++LD+ F+ KLGDFG+AKLVD +Q T V+ GT GY+ PEY G
Sbjct: 351 EKCVVHRDIKPSNIMLDSAFAAKLGDFGLAKLVDHGDASQTTAVLAGTMGYMDPEYAASG 410
Query: 543 RASKESDMYSFGIVALELATGRR--FFQDGDFHVPLMNWVWGLYVEGNLMCAADEKL--- 597
+AS SD+YSFGIV LE+ GRR Q+ L+ WVW L+ G ++ AADE+L
Sbjct: 411 KASTASDVYSFGIVLLEMCCGRRPVLLQEQSIRSRLLEWVWDLHGRGAILEAADERLRGG 470
Query: 598 NMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPELPLDM 647
+E D +++ +++VGLWC H + RP + + LQ E LP LP M
Sbjct: 471 ELELDAKQVECVMVVGLWCAHPDRGVRPSIKQALAALQFEAPLPALPPTM 520
>UniRef100_Q6ZFS6 Putative receptor-type protein kinase LRK1 [Oryza sativa]
Length = 681
Score = 364 bits (935), Expect = 4e-99
Identities = 239/614 (38%), Positives = 339/614 (54%), Gaps = 55/614 (8%)
Query: 44 NFDDPTVAS----NMSYQGDGKSTNGSIDL--NKVSYLFR-VGRAFYSQPLHLWDKKTNT 96
+F+ PT AS N+ QG + G +D+ N VS + GR Y+ P+ LWD T
Sbjct: 11 SFNYPTFASSDNQNIDIQGQASVSVGYVDISANSVSGMGNSAGRVVYAPPVQLWDAATGE 70
Query: 97 LTSFTTRFSFTIDKLNDTTY-GDGFVFYLAPLGYQIPPNSAGGV-YGLFNATT-NSNFVM 153
+ SFTTRFSF I +D + GDG F+L ++P GG GL N T N +
Sbjct: 71 VASFTTRFSFNIIAPSDRSKKGDGMAFFLTSYPSRLPVGHEGGENLGLTNQTVGNVSTGQ 130
Query: 154 NYVVGVEFDTFVGPTDPPMK--HVGIDDNALTSVAFGKFDIDKNLGRVCYVLIDYNSDEK 211
N V VEFDTFV P DP H+GID N++ SV +G + +DYN++ +
Sbjct: 131 NRFVAVEFDTFVNPFDPNTTNDHIGIDVNSVVSVTNESLPNFSLIGNMT-ATVDYNNNSR 189
Query: 212 MLEV-FWSFKGRFVKGDGSYGNSSISYQIDLMKKLPEFVNIGFSASTGLSTESNVIHSWE 270
+L + W + + ++S +DL + LPE V +GFSASTG + E + + SW
Sbjct: 190 ILSIKLWI--------NETTTPYTLSSMVDLKRALPENVTVGFSASTGSAFEQHQLTSWY 241
Query: 271 FSSNLEDSNSTTSLVEGNDGKGS---------------LKTVIVVVAVIVPVILVFLIAS 315
F S+ + V S + +V A + V+ V L+ +
Sbjct: 242 FKSSSSFEQKLAAKVASPPPPSSPSPPPPSLTPITSHSRRGGVVAGATLGAVMFVILLFA 301
Query: 316 IVGWVIVKR---KRKNCDEGL------DEYGIPFPKKFDLDKATIPRRFEYSELVAATNG 366
+V ++ +R KR+ ++G D+ G P +++ PRRF Y ELV AT
Sbjct: 302 MVAVLVRRRQSKKRREAEDGGWHGSDDDDDGEPI---VEIEMGMGPRRFPYHELVDATKS 358
Query: 367 FDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERV-FINEVRIISRLIHRNLV 425
F + LG+GG+G VY+G L LG VA+KR + R + +E+++ISRL HRNLV
Sbjct: 359 FAPEEKLGQGGFGAVYRGYLRELGLAVAIKRFAKNSSKQGRKEYKSEIKVISRLRHRNLV 418
Query: 426 QFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKSLAWEVRYKVALGVANALRYLHDDAEQ 485
Q IGWCH + ELLLV+E PN SLD HL G+ L W +R + G+ +AL YLH++ +Q
Sbjct: 419 QLIGWCHGRTELLLVYELFPNRSLDVHLHGNGTFLTWPMRINIVHGLGSALLYLHEEWDQ 478
Query: 486 CVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQ-RTDVVGTYGYLAPEYINGGRA 544
CV+HRDIK +NV+LD F+ KLGDFG+A+L+D + Q T GT GYL PE + G+A
Sbjct: 479 CVVHRDIKPSNVMLDESFNAKLGDFGLARLIDHAVGIQTMTHPSGTPGYLDPECVITGKA 538
Query: 545 SKESDMYSFGIVALELATGRR--FFQD--GDFHVPLMNWVWGLYVEGNLMCAADEKLNME 600
S ESD+YSFGIV LE+A GRR QD + L+ WVW LY +G ++ AADE+LN E
Sbjct: 539 SAESDVYSFGIVLLEVACGRRPISLQDTQNNCLFRLVEWVWDLYGQGAVLNAADERLNNE 598
Query: 601 FDVSEMKSLLIVGL 614
+D + M+ ++ VGL
Sbjct: 599 YDTTSMECVMAVGL 612
>UniRef100_Q6ZFS1 Putative receptor-type protein kinase LRK1 [Oryza sativa]
Length = 543
Score = 362 bits (929), Expect = 2e-98
Identities = 226/557 (40%), Positives = 319/557 (56%), Gaps = 52/557 (9%)
Query: 131 IPPNSAGGVYGLFNATTNSNFVMNYVVGVEFDT----FVGPTDPPMKHVGIDDNALTSVA 186
+P GG GL + T ++ + V VEFDT F+ P D H+GID NAL SV
Sbjct: 1 MPYMGYGGALGLTSQTFDNATAGDRFVAVEFDTYNNSFLDP-DATYDHIGIDVNALRSVK 59
Query: 187 FGKFDIDKNLGRVCYVLIDYNSDEKMLEV-FWSFKGRFVKGDGSYGNSSISYQIDLMKKL 245
+G + ++DYNS+ ++ V W+ +GS ++S +IDL L
Sbjct: 60 TESLPSFILIGNMT-AIVDYNSNSSIMSVKLWA--------NGSTTPYNLSSKIDLKSAL 110
Query: 246 PEFVNIGFSASTGLSTESNVIHSWEFSSNLEDSNSTTSLVEGNDGKGSLKTVIVVVAVIV 305
PE V +GFSA+TG S E + + SW F+ LE T G +G +V A +
Sbjct: 111 PEKVAVGFSAATGSSFEQHQLRSWYFNLTLEQKQPT-----GQHSRGG----VVAGATVG 161
Query: 306 PVILVFLIASIVGWVIVKRKRKNC--DEGLDEYGIPFPKKFDLDKATIPRRFEYSELVAA 363
++ + L+ ++V ++ +R+RK +E D G P +++ T PRRF Y LV A
Sbjct: 162 AILFIVLLFTMVAILVRRRQRKKMREEEEDDSEGDPI---VEIEMGTGPRRFPYHILVNA 218
Query: 364 TNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERV-FINEVRIISRLIHR 422
T F + LG+GG+G VY+G L LG VA+KR D R + +E+++ISRL HR
Sbjct: 219 TKSFAAEEKLGQGGFGAVYRGNLRELGIDVAIKRFAKDSSKQGRKEYKSEIKVISRLRHR 278
Query: 423 NLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKSLAWEVRYKVALGVANALRYLHDD 482
NLVQ IGWCH ++ELLLV+E +PN SLD HL G+ L W +R + LG+ NAL YLH++
Sbjct: 279 NLVQLIGWCHGRNELLLVYELVPNRSLDVHLHGNGTFLTWPMRINIVLGLGNALLYLHEE 338
Query: 483 AEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQ-RTDVVGTYGYLAPEYING 541
EQCV+HRDIK +NV+LD F+TKLGDFG+A+L+D + Q T GT GY+ PE +
Sbjct: 339 WEQCVVHRDIKPSNVMLDESFNTKLGDFGLARLIDHAIGAQTMTHPSGTPGYVDPECVIT 398
Query: 542 GRASKESDMYSFGIVALELATGRRFF------QDGDFHVPLMNWVWGLYVEGNLMCAADE 595
G+AS ESD+YSFGIV LE+A GRR +G F L+ WVW LY +G ++ AADE
Sbjct: 399 GKASAESDVYSFGIVLLEVACGRRPMNLLDDQNNGLFR--LVEWVWDLYGQGAVLKAADE 456
Query: 596 KLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPELPLDMHDRAPPIV 655
+LN ++D ++M+ +L+VGLWC H + RP + VLQ LP LP M P+
Sbjct: 457 RLNGDYDATDMECVLVVGLWCAHPDRCARPSIRVAMAVLQSNGPLPMLPTKM-----PV- 510
Query: 656 PFRQSNGPSMAPPMTNS 672
P+ PP+ +S
Sbjct: 511 -------PTYGPPVASS 520
>UniRef100_Q7X756 OSJNBb0096E05.1 protein [Oryza sativa]
Length = 746
Score = 351 bits (901), Expect = 4e-95
Identities = 250/693 (36%), Positives = 360/693 (51%), Gaps = 63/693 (9%)
Query: 1 MLNSPSSCSSNII----NQSKNILVLFSLLLVLSYLPKTNSLLFNITNFDDPTVASNMSY 56
+LN SS SS Q + +L+L LLLV +P T SL F+ D V+ +
Sbjct: 4 LLNPSSSSSSKATPRPPQQQQLLLLLLHLLLVA--VPAT-SLTFSYDA--DSFVSEDFRQ 58
Query: 57 QGDGKSTNGSIDLNKVSYLFRV-GRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKL---N 112
+ D T G I+L + R GRA Y +P+ LWD T SF F+FTI +
Sbjct: 59 EDDAMVTAGRIELLGEEFAARARGRALYKRPVQLWDGATGEEASFAASFNFTIRSVAGRG 118
Query: 113 DTTYGDGFVFYLAPLGYQIPPNSAGGVYGLFNATTNSNFVMNYV---------VGVEFDT 163
+ G G F+LAP +P G GLF+ + N + V VEFDT
Sbjct: 119 NALAGHGMTFFLAPFMPDMPQECYEGCLGLFDQSLTRNTASATMGNASGAASFVAVEFDT 178
Query: 164 FVGPTDPPMKHVGIDDNALTSVAFGKFDIDKNL---GRVCYVLIDYNSDEKMLEVFWSFK 220
+ DP +HVG+D N + S + ++ V + Y+S + L+V +
Sbjct: 179 HMDGWDPSGRHVGVDINNVDSRRGNYVVLPEDSLVDAGVMSATVSYDSGARRLDVALA-- 236
Query: 221 GRFVKGDGSYGNSSISYQIDLMKKLPEFVNIGFSASTGLSTESN-VIHSWEFSSNLEDSN 279
+ G + ++S + L LPE V +GFSA+TG SN + S+ FSS L
Sbjct: 237 ---IGGGAATATYNLSAAVHLRSVLPEQVAVGFSAATGDQFASNHTVLSFTFSSTLPTRT 293
Query: 280 STTSLVEGNDGKGSLKTVIVVVAVIVPVILVFLIASIVGWVIVKRKRKNCDEGLD-EYGI 338
+ + K + + V A I ++LV I ++ RKR+ D+G+ + +
Sbjct: 294 TNPPPPSTSSAKTAHLSAAVAAAGIALLLLVLAITILIRRA---RKRRRRDDGVSYDDSL 350
Query: 339 PFPKKFDLDKATIPRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRI 398
+ D++ T PRR Y++L AAT GF + LG GG G VY G + LG+ VA+K +
Sbjct: 351 DDDDEEDMESGTGPRRIPYAQLAAATGGFAEIGKLGEGGSGSVYGGHVRELGRDVAIK-V 409
Query: 399 FADFENSE--RVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGD 456
F + E + + +EV +ISRL HRNLVQ +GWCH + LLLV+E + NGSLD HL+ +
Sbjct: 410 FTRGASMEGRKEYRSEVTVISRLRHRNLVQLMGWCHGRRRLLLVYELVRNGSLDGHLYSN 469
Query: 457 KKSLAWEVRYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLV 516
K++L W +RY++ G+A+A+ YLH + +QCV+H DIK +N++LD F+ KLGDFG+A+L+
Sbjct: 470 KETLTWPLRYQIINGLASAVLYLHQEWDQCVVHGDIKPSNIMLDESFNAKLGDFGLARLI 529
Query: 517 DPMLRTQ-RTDVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFF-------- 567
D + Q T V GT GYL PE + G+AS ESDMYSFGIV LE+A+GRR
Sbjct: 530 DHGMSLQTMTAVAGTPGYLDPECVITGKASTESDMYSFGIVLLEVASGRRPMVVTPRAAA 589
Query: 568 ---------QDGDFHV-PLMNWVWGLYVEG-----NLMCAADEKLNMEFDVSEMKSLLIV 612
DG V L+ W W LY G +L AD +L FD EM+ ++ V
Sbjct: 590 ATAGGGKDDDDGGGQVFRLVEWAWELYGRGDDDQSSLDAIADTRLGGAFDRWEMERVVGV 649
Query: 613 GLWCTHSNDKERPKAYEVIKVLQ-LEMALPELP 644
GLWC H + K RP + + LQ + +P LP
Sbjct: 650 GLWCAHPDPKARPAIRQAAEALQSRKFRMPVLP 682
>UniRef100_Q7XL08 OSJNBa0065J03.1 protein [Oryza sativa]
Length = 746
Score = 350 bits (899), Expect = 6e-95
Identities = 251/693 (36%), Positives = 358/693 (51%), Gaps = 63/693 (9%)
Query: 1 MLNSPSSCSSNII----NQSKNILVLFSLLLVLSYLPKTNSLLFNITNFDDPTVASNMSY 56
+LN SS SS Q + +L+L LLLV +P T SL F+ D V+ +
Sbjct: 4 LLNPSSSSSSKATPRPPQQQQLLLLLLHLLLVA--VPAT-SLTFSYDA--DSFVSEDFRQ 58
Query: 57 QGDGKSTNGSIDLNKVSYLFRV-GRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKL---N 112
+ D T G I+L + R GRA Y +P+ LWD T SF F+FTI +
Sbjct: 59 EDDAMVTAGRIELLGEEFAARARGRALYKRPVQLWDGATGEEASFAASFNFTIRSVAGRG 118
Query: 113 DTTYGDGFVFYLAPLGYQIPPNSAGGVYGLFNATTNSNFVMNYV---------VGVEFDT 163
+ G G F+LAP +P G GLF+ + N + V VEFDT
Sbjct: 119 NALAGHGMTFFLAPFMPDMPQECYEGCLGLFDQSLTRNTASATMGNASGAASFVAVEFDT 178
Query: 164 FVGPTDPPMKHVGIDDNALTSVAFGKFDIDKNL---GRVCYVLIDYNSDEKMLEVFWSFK 220
+ DP +HVG+D N + S + ++ V + Y+S + L+V +
Sbjct: 179 HMDGWDPSGRHVGVDINNVDSRRGNYVVLPEDSLVDAGVMSATVSYDSGARRLDVALA-- 236
Query: 221 GRFVKGDGSYGNSSISYQIDLMKKLPEFVNIGFSASTGLSTESN-VIHSWEFSSNLEDSN 279
+ G + ++S + L LPE V +GFSA+TG SN + S+ FSS L
Sbjct: 237 ---IGGGAATATYNLSAAVHLRSVLPEQVAVGFSAATGDQFASNHTVLSFTFSSTLPTRT 293
Query: 280 STTSLVEGNDGKGSLKTVIVVVAVIVPVILVFLIASIVGWVIVKRKRKNCDEGLD-EYGI 338
+ + K + + V A I +LV I ++ RKR+ D+G+ + I
Sbjct: 294 TNPPQPSTSSAKTAHLSAAVAAAGIALQLLVLAITILIRRA---RKRRRRDDGVSYDDSI 350
Query: 339 PFPKKFDLDKATIPRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRI 398
+ D++ T PRR Y+ L AAT GF + LG GG G VY G + LG+ VA+K +
Sbjct: 351 DDDDEEDMESGTGPRRIPYAHLAAATGGFAEIGKLGEGGSGSVYGGHVRELGRNVAIK-V 409
Query: 399 FADFENSE--RVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGD 456
F + E + + +EV +ISRL HRNLVQ +GWCH + LLLV+E + NGSLD HL+ +
Sbjct: 410 FTRGASMEGRKEYRSEVTVISRLRHRNLVQLMGWCHGRRRLLLVYELVRNGSLDGHLYSN 469
Query: 457 KKSLAWEVRYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLV 516
K++L W +RY++ G+A+A+ YLH + +QCV+H DIK +N++LD F+ KLGDFG+A+L+
Sbjct: 470 KETLTWPLRYQIINGLASAVLYLHQEWDQCVVHGDIKPSNIMLDESFNAKLGDFGLARLI 529
Query: 517 DPMLRTQ-RTDVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFF-------- 567
D + Q T V GT GYL PE + G+AS ESDMYSFGIV LE+A+GRR
Sbjct: 530 DHGMSLQTMTAVAGTPGYLDPECVITGKASTESDMYSFGIVLLEVASGRRPMVVTPRAAA 589
Query: 568 ---------QDGDFHV-PLMNWVWGLYVEG-----NLMCAADEKLNMEFDVSEMKSLLIV 612
DG V L+ W W LY G +L AD +L FD EM+ ++ V
Sbjct: 590 ATAGGGKDDDDGGGQVFRLVEWAWELYGRGDDDQSSLDAIADTRLGGAFDRWEMERVVGV 649
Query: 613 GLWCTHSNDKERPKAYEVIKVLQ-LEMALPELP 644
GLWC H + K RP + + LQ + +P LP
Sbjct: 650 GLWCAHPDPKARPAIRQAAEALQSRKFRMPVLP 682
>UniRef100_Q9FHG4 Serine/threonine-specific kinase like protein [Arabidopsis
thaliana]
Length = 681
Score = 346 bits (888), Expect = 1e-93
Identities = 228/663 (34%), Positives = 349/663 (52%), Gaps = 51/663 (7%)
Query: 16 SKNILVLFSLLLVLSYLPKTNSLLFNITNFDDPTVA-SNMSYQGDGKSTNGSIDLNKVSY 74
S+ +LV+F + + K + + NF + N+++ GD NG + L +
Sbjct: 4 SRKLLVIFFTWITALSMSKPIFVSSDNMNFTFKSFTIRNLTFLGDSHLRNGVVGLTRELG 63
Query: 75 L--FRVGRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLND--TTYGDGFVFYLAPLGYQ 130
+ G Y+ P+ +D +NT SF+T FSFT+ LN T+ GDG F+L+
Sbjct: 64 VPDTSSGTVIYNNPIRFYDPDSNTTASFSTHFSFTVQNLNPDPTSAGDGLAFFLSHDNDT 123
Query: 131 IPPNSAGGVYGLFNATTNSNFVMNYVVGVEFDTFVGP--TDPPMKHVGIDDNALTSVAFG 188
+ S GG GL N+ S + N V +EFDT + P DP H+G+D ++L S++
Sbjct: 124 L--GSPGGYLGLVNS---SQPMKNRFVAIEFDTKLDPHFNDPNGNHIGLDVDSLNSISTS 178
Query: 189 ----KFDIDKNLGRVCYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSSISYQIDLMKK 244
ID G+ IDY +D ++L VF S+ V +S IDL
Sbjct: 179 DPLLSSQIDLKSGKSITSWIDYKNDLRLLNVFLSYTDP-VTTTKKPEKPLLSVNIDLSPF 237
Query: 245 LPEFVNIGFSASTGLSTESNVIHSWEFSS--------------NLEDSNSTTS--LVEGN 288
L + +GFS ST STE ++I +W F + N+ DS+ +V +
Sbjct: 238 LNGEMYVGFSGSTEGSTEIHLIENWSFKTSGFLPVRSKSNHLHNVSDSSVVNDDPVVIPS 297
Query: 289 DGKGSLKTVIVVVAVIVPVILVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLDK 348
+ + + + + PV L+ L + G+ +K+ + + K+ +
Sbjct: 298 KKRRHRHNLAIGLGISCPV-LICLALFVFGYFTLKKWKS----------VKAEKELKTEL 346
Query: 349 ATIPRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERV 408
T R F Y EL AT GF R++GRG +G VY+ G I AVKR + +
Sbjct: 347 ITGLREFSYKELYTATKGFHSSRVIGRGAFGNVYRAMFVSSGTISAVKRSRHNSTEGKTE 406
Query: 409 FINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS----LAWEV 464
F+ E+ II+ L H+NLVQ GWC+E+ ELLLV+E+MPNGSLD L+ + ++ L W
Sbjct: 407 FLAELSIIACLRHKNLVQLQGWCNEKGELLLVYEFMPNGSLDKILYQESQTGAVALDWSH 466
Query: 465 RYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQR 524
R +A+G+A+AL YLH + EQ V+HRDIK++N++LD +F+ +LGDFG+A+L +
Sbjct: 467 RLNIAIGLASALSYLHHECEQQVVHRDIKTSNIMLDINFNARLGDFGLARLTEHDKSPVS 526
Query: 525 TDVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQ---DGDFHVPLMNWVW 581
T GT GYLAPEY+ G A++++D +S+G+V LE+A GRR + V L++WVW
Sbjct: 527 TLTAGTMGYLAPEYLQYGTATEKTDAFSYGVVILEVACGRRPIDKEPESQKTVNLVDWVW 586
Query: 582 GLYVEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALP 641
L+ EG ++ A DE+L EFD MK LL+VGL C H + ERP V+++L E+
Sbjct: 587 RLHSEGRVLEAVDERLKGEFDEEMMKKLLLVGLKCAHPDSNERPSMRRVLQILNNEIEPS 646
Query: 642 ELP 644
+P
Sbjct: 647 PVP 649
>UniRef100_Q84VX6 At3g53810 [Arabidopsis thaliana]
Length = 677
Score = 342 bits (878), Expect = 2e-92
Identities = 240/656 (36%), Positives = 356/656 (53%), Gaps = 57/656 (8%)
Query: 20 LVLFSLLLVLSYLPKTNSLLFNITNFDDPTVASNMSYQGDGKST-NGSIDLNKVSYLFRV 78
L+ F LL + + +L F F P +++S QG T NG + L S + +
Sbjct: 7 LIFFFFLLCQIMISSSQNLNFTYNGFHPPL--TDISLQGLATVTPNGLLKLTNTS-VQKT 63
Query: 79 GRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGDGFVFYLAP---LGYQIPPNS 135
G AF ++ + D ++ ++SF+T F F I T G G F +AP L + +P
Sbjct: 64 GHAFCTERIRFKDSQSGNVSSFSTTFVFAIHSQIPTLSGHGIAFVVAPTLGLPFALPSQ- 122
Query: 136 AGGVYGLFNATTNSNFVMNYVVGVEFDTFVGPT--DPPMKHVGIDDNALTSVAFG----K 189
GLFN + N N N++ VEFDT DP HVGID N L S + +
Sbjct: 123 ---YIGLFNISNNGNDT-NHIFAVEFDTIQSSEFGDPNDNHVGIDLNGLRSANYSTAGYR 178
Query: 190 FDIDK--NLGRVC----YVLIDYNSDEKMLEV----FWSFKGRFVKGDGSYGNSSISYQI 239
D DK NL + V IDY++ ++V F S K R +SY
Sbjct: 179 DDHDKFQNLSLISRKRIQVWIDYDNRSHRIDVTVAPFDSDKPR---------KPLVSYVR 229
Query: 240 DLMKKLPEFVNIGFSASTGLSTESNVIHSWEFSSNLEDSNSTTSLVEGNDGKGSLKTVIV 299
DL L E + +GFS++TG + + W F N E + S + + + +
Sbjct: 230 DLSSILLEDMYVGFSSATGSVLSEHFLVGWSFRLNGEAPMLSLSKLPKLP-RFEPRRISE 288
Query: 300 VVAVIVPVILVFLIASIV--GWVIVKRKRKNCDEGLDEYGIPFPKKFDLDKATIPRRFEY 357
+ +P+I + LI SI+ + IV+RK+K +E LD++ F K RF +
Sbjct: 289 FYKIGMPLISLSLIFSIIFLAFYIVRRKKKY-EEELDDWETEFGKN----------RFRF 337
Query: 358 SELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIIS 417
EL AT GF + +LG GG+G+VY+G L VAVKR+ D + + F+ E+ I
Sbjct: 338 KELYHATKGFKEKDLLGSGGFGRVYRGILPTTKLEVAVKRVSHDSKQGMKEFVAEIVSIG 397
Query: 418 RLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS-LAWEVRYKVALGVANAL 476
R+ HRNLV +G+C + ELLLV++YMPNGSLD +L+ + ++ L W+ R + GVA+ L
Sbjct: 398 RMSHRNLVPLLGYCRRRGELLLVYDYMPNGSLDKYLYNNPETTLDWKQRSTIIKGVASGL 457
Query: 477 RYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGYLAP 536
YLH++ EQ V+HRD+K++NVLLD DF+ +LGDFG+A+L D Q T VVGT GYLAP
Sbjct: 458 FYLHEEWEQVVIHRDVKASNVLLDADFNGRLGDFGLARLYDHGSDPQTTHVVGTLGYLAP 517
Query: 537 EYINGGRASKESDMYSFGIVALELATGRR---FFQDGDFHVPLMNWVWGLYVEGNLMCAA 593
E+ GRA+ +D+Y+FG LE+ +GRR F D L+ WV+ L++ GN+M A
Sbjct: 518 EHSRTGRATTTTDVYAFGAFLLEVVSGRRPIEFHSASDDTFLLVEWVFSLWLRGNIMEAK 577
Query: 594 DEKLNME-FDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPEL-PLDM 647
D KL +D+ E++ +L +GL C+HS+ + RP +V++ L+ +MALPEL PLD+
Sbjct: 578 DPKLGSSGYDLEEVEMVLKLGLLCSHSDPRARPSMRQVLQYLRGDMALPELTPLDL 633
>UniRef100_Q9M345 Serine/threonine-specific kinase like protein [Arabidopsis
thaliana]
Length = 677
Score = 342 bits (877), Expect = 2e-92
Identities = 240/656 (36%), Positives = 355/656 (53%), Gaps = 57/656 (8%)
Query: 20 LVLFSLLLVLSYLPKTNSLLFNITNFDDPTVASNMSYQGDGKST-NGSIDLNKVSYLFRV 78
L+ F LL + + +L F F P +++S QG T NG + L S + +
Sbjct: 7 LIFFFFLLCQIMISSSQNLNFTYNGFHPPL--TDISLQGLATVTPNGLLKLTNTS-VQKT 63
Query: 79 GRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGDGFVFYLAP---LGYQIPPNS 135
G AF ++ + D + ++SF+T F F I T G G F +AP L + +P
Sbjct: 64 GHAFCTERIRFKDSQNGNVSSFSTTFVFAIHSQIPTLSGHGIAFVVAPTLGLPFALPSQ- 122
Query: 136 AGGVYGLFNATTNSNFVMNYVVGVEFDTFVGPT--DPPMKHVGIDDNALTSVAFG----K 189
GLFN + N N N++ VEFDT DP HVGID N L S + +
Sbjct: 123 ---YIGLFNISNNGNDT-NHIFAVEFDTIQSSEFGDPNDNHVGIDLNGLRSANYSTAGYR 178
Query: 190 FDIDK--NLGRVC----YVLIDYNSDEKMLEV----FWSFKGRFVKGDGSYGNSSISYQI 239
D DK NL + V IDY++ ++V F S K R +SY
Sbjct: 179 DDHDKFQNLSLISRKRIQVWIDYDNRSHRIDVTVAPFDSDKPR---------KPLVSYVR 229
Query: 240 DLMKKLPEFVNIGFSASTGLSTESNVIHSWEFSSNLEDSNSTTSLVEGNDGKGSLKTVIV 299
DL L E + +GFS++TG + + W F N E + S + + + +
Sbjct: 230 DLSSILLEDMYVGFSSATGSVLSEHFLVGWSFRLNGEAPMLSLSKLPKLP-RFEPRRISE 288
Query: 300 VVAVIVPVILVFLIASIV--GWVIVKRKRKNCDEGLDEYGIPFPKKFDLDKATIPRRFEY 357
+ +P+I + LI SI+ + IV+RK+K +E LD++ F K RF +
Sbjct: 289 FYKIGMPLISLSLIFSIIFLAFYIVRRKKKY-EEELDDWETEFGKN----------RFRF 337
Query: 358 SELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIIS 417
EL AT GF + +LG GG+G+VY+G L VAVKR+ D + + F+ E+ I
Sbjct: 338 KELYHATKGFKEKDLLGSGGFGRVYRGILPTTKLEVAVKRVSHDSKQGMKEFVAEIVSIG 397
Query: 418 RLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS-LAWEVRYKVALGVANAL 476
R+ HRNLV +G+C + ELLLV++YMPNGSLD +L+ + ++ L W+ R + GVA+ L
Sbjct: 398 RMSHRNLVPLLGYCRRRGELLLVYDYMPNGSLDKYLYNNPETTLDWKQRSTIIKGVASGL 457
Query: 477 RYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGYLAP 536
YLH++ EQ V+HRD+K++NVLLD DF+ +LGDFG+A+L D Q T VVGT GYLAP
Sbjct: 458 FYLHEEWEQVVIHRDVKASNVLLDADFNGRLGDFGLARLYDHGSDPQTTHVVGTLGYLAP 517
Query: 537 EYINGGRASKESDMYSFGIVALELATGRR---FFQDGDFHVPLMNWVWGLYVEGNLMCAA 593
E+ GRA+ +D+Y+FG LE+ +GRR F D L+ WV+ L++ GN+M A
Sbjct: 518 EHSRTGRATTTTDVYAFGAFLLEVVSGRRPIEFHSASDDTFLLVEWVFSLWLRGNIMEAK 577
Query: 594 DEKLNME-FDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPEL-PLDM 647
D KL +D+ E++ +L +GL C+HS+ + RP +V++ L+ +MALPEL PLD+
Sbjct: 578 DPKLGSSGYDLEEVEMVLKLGLLCSHSDPRARPSMRQVLQYLRGDMALPELTPLDL 633
>UniRef100_O81292 T14P8.3 protein [Arabidopsis thaliana]
Length = 674
Score = 335 bits (859), Expect = 3e-90
Identities = 227/693 (32%), Positives = 364/693 (51%), Gaps = 59/693 (8%)
Query: 19 ILVLFSLLLVLSYLPKTNSLLFNITNFDDPTVASNMSYQGDGKSTNGSIDLNKVSYLFRV 78
I F +LL + SL F +F P +N+S QG T+ I +
Sbjct: 8 IFFFFIILLSKPLNSSSQSLNFTYNSFHRPP--TNISIQGIATVTSNGILKLTDKTVIST 65
Query: 79 GRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLNDTTYGDGFVFYLAPLGYQIPPNSAGG 138
G AFY++P+ D +T++SF+T F I T G G F++AP + A
Sbjct: 66 GHAFYTEPIRFKDSPNDTVSSFSTTFVIGIYSGIPTISGHGMAFFIAP-NPVLSSAMASQ 124
Query: 139 VYGLFNATTNSNFVMNYVVGVEFDTFVGP--TDPPMKHVGIDDNALTSV---AFGKFDID 193
GLF++T N N N+++ VEFDT + P D HVGI+ N+LTSV G +D
Sbjct: 125 YLGLFSSTNNGNDT-NHILAVEFDTIMNPEFDDTNDNHVGININSLTSVKSSLVGYWDEI 183
Query: 194 KNLGRV-------CYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSSISYQIDLMKKLP 246
+ V +DY+ ++V + G+ + +S DL
Sbjct: 184 NQFNNLTLISRKRMQVWVDYDDRTNQIDVTMA-----PFGEVKPRKALVSVVRDLSSVFL 238
Query: 247 EFVNIGFSASTGLSTESNVIHSWEFSSNLEDSN-------------STTSLVEGNDGKGS 293
+ + +GFSA+TG + + W F + + TSL +
Sbjct: 239 QDMYLGFSAATGYVLSEHFVFGWSFMVKGKTAPPLTLSKVPKFPRVGPTSLQRFYKNRMP 298
Query: 294 LKTVIVVVAVIVPVILVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLDKATIPR 353
L +++++ + V V L+FL+ IV R+R+ E +++ F K
Sbjct: 299 LFSLLLIPVLFV-VSLIFLVRFIV------RRRRKFAEEFEDWETEFGK----------N 341
Query: 354 RFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEV 413
R + +L AT GF D +LG GG+G+VY+G + K +AVKR+ + + F+ E+
Sbjct: 342 RLRFKDLYYATKGFKDKDLLGSGGFGRVYRGVMPTTKKEIAVKRVSNESRQGLKEFVAEI 401
Query: 414 RIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFG-DKKSLAWEVRYKVALGV 472
I R+ HRNLV +G+C +DELLLV++YMPNGSLD +L+ + +L W+ R+ V +GV
Sbjct: 402 VSIGRMSHRNLVPLLGYCRRRDELLLVYDYMPNGSLDKYLYDCPEVTLDWKQRFNVIIGV 461
Query: 473 ANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYG 532
A+ L YLH++ EQ V+HRDIK++NVLLD +++ +LGDFG+A+L D Q T VVGT+G
Sbjct: 462 ASGLFYLHEEWEQVVIHRDIKASNVLLDAEYNGRLGDFGLARLCDHGSDPQTTRVVGTWG 521
Query: 533 YLAPEYINGGRASKESDMYSFGIVALELATGRRFFQ---DGDFHVPLMNWVWGLYVEGNL 589
YLAP+++ GRA+ +D+++FG++ LE+A GRR + + D V L++ V+G ++EGN+
Sbjct: 522 YLAPDHVRTGRATTATDVFAFGVLLLEVACGRRPIEIEIESDESVLLVDSVFGFWIEGNI 581
Query: 590 MCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPEL-PLDMH 648
+ A D L +D E++++L +GL C+HS+ + RP +V++ L+ + LP+L PLD
Sbjct: 582 LDATDPNLGSVYDQREVETVLKLGLLCSHSDPQVRPTMRQVLQYLRGDATLPDLSPLDFR 641
Query: 649 DRAPPI---VPFRQSNGPSMAPPMTNSLITSGR 678
+ F +S S + S+++ GR
Sbjct: 642 GSGKMLGMNHRFSESCTFSSGSSIAYSIVSGGR 674
>UniRef100_Q9XIS3 Lectin-like protein kinase [Populus nigra]
Length = 676
Score = 333 bits (855), Expect = 8e-90
Identities = 207/629 (32%), Positives = 340/629 (53%), Gaps = 50/629 (7%)
Query: 36 NSLLFNITNFDDPTVASNMSYQGDGKSTNGSIDLNKVSYLFRVGRAFYSQPLHLWDKKTN 95
N ++I + P SN + Q S NG+ L R GR ++ LW+
Sbjct: 35 NETYYDIFQVERPATISNNALQITPDSINGNFTLAN-----RSGRVLLNKSFILWEDDGA 89
Query: 96 ---TLTSFTTRFSFTIDKLNDTTYGDGFVFYLAPLGYQIPPNSAGGVYGLFNATTNSNFV 152
+ SF + F I ++N++T G+G F +AP +P NS G GL N+TT+ N
Sbjct: 90 GGVRVASFNSSFVINIFRVNNSTPGEGLAFLIAP-DLALPENSDGQYLGLTNSTTDRN-P 147
Query: 153 MNYVVGVEFDTFVGPTDPPMKHVGIDDNALTSVAFGKFDIDKNLG--------RVCYVLI 204
N +V +E DT DP H+G++ +++ S+ D +LG R V +
Sbjct: 148 ENGIVAIELDTVKQEFDPDDNHMGLNIHSVISLKTVSLD---DLGIEIAPVGARNHMVWV 204
Query: 205 DYNSDEKMLEVFWSFKGRFVKGDGSYGNSSISYQIDLMKKLPEFVNIGFSASTGLSTESN 264
Y+ + K +EV+ + +GR +++ +++L + E GF+ASTG + + N
Sbjct: 205 HYDGNSKKMEVYMAEEGR-----AKPATPALAAELNLKDLVREKSYFGFAASTGRNFQLN 259
Query: 265 VIHSWEFSSNLEDSNSTTSLVEGNDGKGSLKTVIVVVAVIVPVILVFLIASIVGWVIVKR 324
+ W + + +S + GND K +K + + + ++L+ + W+
Sbjct: 260 CVLKWNLTVEMLSDDSVEN--GGNDNKKLIKICVGIGVGLFSLLLI----GVGTWLYYLH 313
Query: 325 KRKNCDEGLDEYGIPFPKKFDLDKAT--IPRRFEYSELVAATNGFDDDRMLGRGGYGQVY 382
K+K + P K+ PR F + +L ATN FD+ LG+GG+G VY
Sbjct: 314 KKKAASD---------PNLLGALKSLPGTPREFPFKDLKKATNNFDERHKLGQGGFGVVY 364
Query: 383 KGALSYLGKIVAVKRIFADFENSERVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFE 442
KG L+ +AVK+ + + F++E+ II+RL H++LV+ +GWCH LLLV++
Sbjct: 365 KGLLTKENIQIAVKKFSRENIKGQDDFLSELTIINRLRHKHLVRLLGWCHRNGMLLLVYD 424
Query: 443 YMPNGSLDTHLFGDKKS---LAWEVRYKVALGVANALRYLHDDAEQCVLHRDIKSANVLL 499
YMPNGSLD HLF + + L W +RYK+ GVA+AL YLH++ +Q V+HRD+K++N++L
Sbjct: 425 YMPNGSLDNHLFHELEGNVILEWNLRYKIISGVASALHYLHNEYDQTVVHRDLKASNIML 484
Query: 500 DTDFSTKLGDFGMAKLV--DPMLRTQRTDVVGTYGYLAPEYINGGRASKESDMYSFGIVA 557
D++F+ +LGDFG+A+ + + + V GT GY+APE + G+A++ESD+Y FG V
Sbjct: 485 DSEFNARLGDFGLARALENEKTSYAELEGVPGTMGYIAPECFHTGKATRESDVYGFGAVV 544
Query: 558 LELATGRR-FFQDGDFHVPLMNWVWGLYVEGNLMCAADEKLNMEFDVSEMKSLLIVGLWC 616
LE+ G+R + + G F L++WVW L+ E ++ A DE+LN ++ E K LL++GL C
Sbjct: 545 LEVVCGQRPWTKIGGFQF-LVDWVWSLHREERILEAVDERLNSDYVAEEAKRLLLLGLAC 603
Query: 617 THSNDKERPKAYEVIKVLQLEMALPELPL 645
+H ERPK + +++ + P +PL
Sbjct: 604 SHPIASERPKTQAIFQIISGSVPPPHVPL 632
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.139 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,193,142,722
Number of Sequences: 2790947
Number of extensions: 53807205
Number of successful extensions: 196080
Number of sequences better than 10.0: 18299
Number of HSP's better than 10.0 without gapping: 8398
Number of HSP's successfully gapped in prelim test: 9902
Number of HSP's that attempted gapping in prelim test: 155402
Number of HSP's gapped (non-prelim): 21112
length of query: 678
length of database: 848,049,833
effective HSP length: 134
effective length of query: 544
effective length of database: 474,062,935
effective search space: 257890236640
effective search space used: 257890236640
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 78 (34.7 bits)
Medicago: description of AC143339.11


