

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142143.3 + phase: 0 /pseudo
(212 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
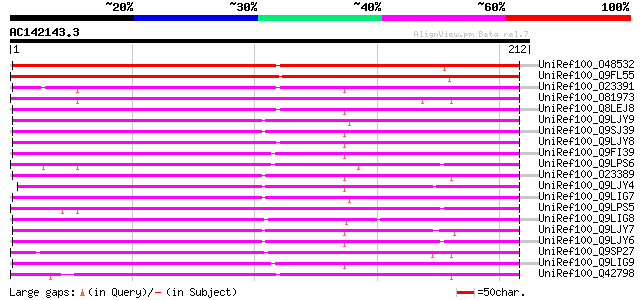
Sequences producing significant alignments: (bits) Value
UniRef100_O48532 Putative cytochrome P450 [Arabidopsis thaliana] 186 5e-46
UniRef100_Q9FL55 Cytochrome P450-like protein [Arabidopsis thali... 175 6e-43
UniRef100_O23391 Cytochrome P450 like protein [Arabidopsis thali... 139 4e-32
UniRef100_O81973 Cytochrome P450 93A3 [Glycine max] 137 2e-31
UniRef100_Q8LEJ8 Cytochrome P450, putative [Arabidopsis thaliana] 136 4e-31
UniRef100_Q9LJY9 Cytochrome P450-like protein [Arabidopsis thali... 135 5e-31
UniRef100_Q9SJ39 Putative cytochrome P450 [Arabidopsis thaliana] 135 7e-31
UniRef100_Q9LJY8 Cytochrome P450-like protein [Arabidopsis thali... 134 2e-30
UniRef100_Q9FI39 Cytochrome P450 [Arabidopsis thaliana] 133 3e-30
UniRef100_Q9LPS6 Cytochrome P450, putative [Arabidopsis thaliana] 133 3e-30
UniRef100_O23389 Cytochrome P450 like protein [Arabidopsis thali... 133 3e-30
UniRef100_Q9LJY4 Cytochrome P450-like protein [Arabidopsis thali... 132 5e-30
UniRef100_Q9LIG7 Cytochrome P450-like protein [Arabidopsis thali... 132 5e-30
UniRef100_Q9LPS5 Cytochrome P450, putative [Arabidopsis thaliana] 131 1e-29
UniRef100_Q9LIG8 Cytochrome P450-like protein [Arabidopsis thali... 131 1e-29
UniRef100_Q9LJY7 Putative cytochrome P450 [Arabidopsis thaliana] 130 2e-29
UniRef100_Q9LJY6 Cytochrome P450-like protein [Arabidopsis thali... 130 2e-29
UniRef100_Q9SP27 Flavone synthase II [Callistephus chinensis] 129 4e-29
UniRef100_Q9LIG9 Cytochrome P450-like protein [Arabidopsis thali... 129 4e-29
UniRef100_Q42798 Cytochrome P450 93A1 [Glycine max] 129 5e-29
>UniRef100_O48532 Putative cytochrome P450 [Arabidopsis thaliana]
Length = 514
Score = 186 bits (471), Expect = 5e-46
Identities = 94/208 (45%), Positives = 138/208 (66%), Gaps = 2/208 (0%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
K +LNFS RP+FGS+ + Y+GS F+ A YG YWRFMKK+C+TKLL+ QL +F IRE
Sbjct: 99 KEQELNFSSRPEFGSAEYFKYRGSRFVLAQYGDYWRFMKKLCMTKLLAVPQLEKFADIRE 158
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
+E KL+ S+ C E DL F TNN++CRM+M T C N + E I+ LV++
Sbjct: 159 EEKLKLVDSVAKCCREGLPCDLSSQFIKYTNNVICRMAMSTRCSGTDN-EAEEIRELVKK 217
Query: 122 FMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTED-CQGD 180
+ + K+S+G+VLGPL D G GKKL ++ ++D ++E +KE E + +D + D
Sbjct: 218 SLELAGKISVGDVLGPLKVMDFSGNGKKLVAVMEKYDLLVERIMKEREAKAKKKDGTRKD 277
Query: 181 MMDILLQVYRNENAEVRLTRNDIKAFFL 208
++DILL+ YR+ AE+++TRND+K+F L
Sbjct: 278 ILDILLETYRDPTAEMKITRNDMKSFLL 305
>UniRef100_Q9FL55 Cytochrome P450-like protein [Arabidopsis thaliana]
Length = 528
Score = 175 bits (444), Expect = 6e-43
Identities = 95/210 (45%), Positives = 130/210 (61%), Gaps = 3/210 (1%)
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTHD +F+ + FG +YKGS F APYG YWRFMKK+C+TKL + QL RF+ IR
Sbjct: 95 LKTHDPDFASKFVFGPRQFNVYKGSEFFNAPYGSYWRFMKKLCMTKLFAGYQLDRFVDIR 154
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
E+E LL +L+ S + DLGL+FT LT IL +M MG C + NI P+ I+ +V
Sbjct: 155 EEETLALLSTLVERSRNGEACDLGLEFTALTTKILSKMVMGKRCRQNSNI-PKEIRKIVS 213
Query: 121 EFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQ-- 178
+ M + E+ GPL DLFG GKKLR + +D+++E LKE+E + E+ +
Sbjct: 214 DIMACATRFGFMELFGPLRDLDLFGNGKKLRSSIWRYDELVEKILKEYENDKSNEEEEKD 273
Query: 179 GDMMDILLQVYRNENAEVRLTRNDIKAFFL 208
D++DILL Y + AE+RLT N IK F L
Sbjct: 274 KDIVDILLDTYNDPKAELRLTMNQIKFFIL 303
>UniRef100_O23391 Cytochrome P450 like protein [Arabidopsis thaliana]
Length = 509
Score = 139 bits (351), Expect = 4e-32
Identities = 79/209 (37%), Positives = 125/209 (59%), Gaps = 4/209 (1%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSY-FITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+ D+N S+R + L+ GSY FI+APYG YW+FM+K+ VTK+L L R R
Sbjct: 94 RDQDVNVSFRHS-PPIEESLFLGSYSFISAPYGDYWKFMRKLMVTKILGPQALERSRRFR 152
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
E EL++ K+LL + + S ++ + L NN +C+M MG +C ++ + E I+ LV
Sbjct: 153 EDELDRFYKTLLDKAMKKESVEIVEEAAKLNNNTICKMIMGRSCSEETG-EAERIRGLVT 211
Query: 121 EFMHVGAKLSMGEVL-GPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQG 179
E M + K+ + + PL K + + K++ + +FD++LE L EHEE++
Sbjct: 212 ESMALTKKIFLATIFHKPLKKLGISLFKKEIMSVSRKFDELLEKILVEHEEKMEEHHQGT 271
Query: 180 DMMDILLQVYRNENAEVRLTRNDIKAFFL 208
DMMD+LL+ YR+ENAE ++TRN IK+ F+
Sbjct: 272 DMMDVLLEAYRDENAEYKITRNHIKSMFV 300
>UniRef100_O81973 Cytochrome P450 93A3 [Glycine max]
Length = 510
Score = 137 bits (345), Expect = 2e-31
Identities = 76/215 (35%), Positives = 127/215 (58%), Gaps = 7/215 (3%)
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSY-FITAPYGPYWRFMKKICVTKLLSSSQLGRFMHI 59
+KTH+ FS RP + + L G F+ APYGPYW+FMKK+C+++LL L +F+ +
Sbjct: 88 LKTHEPAFSNRPANTVAVETLTYGFQDFLFAPYGPYWKFMKKLCMSELLGGHMLDQFLPV 147
Query: 60 REQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLV 119
R+ E +K +K +L + D G +F TL+NNI+ RM + T + + E ++ LV
Sbjct: 148 RQXETKKFIKRVLQKGISGEAVDFGGEFITLSNNIVSRMIVSQTSTTEDENEVEEMRKLV 207
Query: 120 REFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKE-HEERINTEDCQ 178
++ + K ++ + + L +FDL G+ K+L KI FD +L+ +K+ EER N +
Sbjct: 208 KDAAELSGKFNISDFVSFLKRFDLQGFNKRLEKIRDCFDTVLDRIIKQREEERRNKNETV 267
Query: 179 G-----DMMDILLQVYRNENAEVRLTRNDIKAFFL 208
G DM+D+L + +E++E++L + +IKAF L
Sbjct: 268 GKREFKDMLDVLFDISEDESSEIKLNKENIKAFIL 302
>UniRef100_Q8LEJ8 Cytochrome P450, putative [Arabidopsis thaliana]
Length = 513
Score = 136 bits (342), Expect = 4e-31
Identities = 79/208 (37%), Positives = 120/208 (56%), Gaps = 2/208 (0%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
+ HDLN S R L+ + FI+APYG Y++FMKK VTKLL L R IR
Sbjct: 100 RVHDLNVSSRGSPPFEESLLFGSTGFISAPYGDYFKFMKKHLVTKLLGPQALERSRLIRT 159
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
ELE+ +LL + + S ++G + L+NN +C+M MGT+C ++ + E ++ L+ E
Sbjct: 160 NELERFYINLLDKATKKESVEIGKEAMKLSNNSICKMIMGTSCLEEKG-EAERVRGLIIE 218
Query: 122 FMHVGAKLSMGEVL-GPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQGD 180
++ K + L G L K + + K++ + FD +LE +L+EHEE+ + E D
Sbjct: 219 SFYLTKKFFLAFTLRGLLEKLGISLFKKEIMGVSRRFDDLLERYLREHEEKPDNEHQDTD 278
Query: 181 MMDILLQVYRNENAEVRLTRNDIKAFFL 208
M+D LL YR+E AE ++TRN IKAF +
Sbjct: 279 MIDALLAAYRDEKAEYKITRNQIKAFLV 306
>UniRef100_Q9LJY9 Cytochrome P450-like protein [Arabidopsis thaliana]
Length = 524
Score = 135 bits (341), Expect = 5e-31
Identities = 78/208 (37%), Positives = 119/208 (56%), Gaps = 2/208 (0%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
K HD+N S R L+ S FI APYG YW+FMKKI TKLL L +R
Sbjct: 102 KAHDVNVSSRGAAAIDESLLFGSSSFIYAPYGDYWKFMKKIFATKLLRPQVLENSRGVRA 161
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
+EL++ + +L + +N + ++ + L NNILCRMSMG + F + N + E ++ LV E
Sbjct: 162 EELKRFYRRILDKARKNENIEISKESAMLMNNILCRMSMGRS-FSEENGEAERVRGLVGE 220
Query: 122 FMHVGAKLSMGEVLGP-LGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQGD 180
+ K+ L L K + + K++ + FD++LE + EH++++ E D
Sbjct: 221 SYALVKKIFFAATLRRLLEKLRIPLFRKEIMGVSDRFDELLERIIVEHKDKLEKEHQVMD 280
Query: 181 MMDILLQVYRNENAEVRLTRNDIKAFFL 208
MMD+LL YR++NAE ++TRN IK+FF+
Sbjct: 281 MMDVLLAAYRDKNAEYKITRNHIKSFFV 308
>UniRef100_Q9SJ39 Putative cytochrome P450 [Arabidopsis thaliana]
Length = 442
Score = 135 bits (340), Expect = 7e-31
Identities = 81/208 (38%), Positives = 115/208 (54%), Gaps = 2/208 (0%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
+ HD+N S R L+ S ITAPYG YW+FMKK+ TKLL L IR
Sbjct: 100 RAHDVNVSSRGVAAIDESLLFGSSGIITAPYGDYWKFMKKLIATKLLRPQSLESSRGIRG 159
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
+EL + SLL + S ++ + L NN LCRMSMG + F + N + E I+ LV E
Sbjct: 160 EELTRFYNSLLDKARMTESVEISTEAMKLVNNTLCRMSMGRS-FSEENGEAEKIRGLVGE 218
Query: 122 FMHVGAKLSMGEVL-GPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQGD 180
+ K+ + +L PL K + + K++ + D++LE L EHEE+++ E D
Sbjct: 219 SYALTKKMFLASLLRKPLKKLRISLFEKEIMGVSDRLDELLERILVEHEEKLHEEHQGTD 278
Query: 181 MMDILLQVYRNENAEVRLTRNDIKAFFL 208
MMD+LL +ENAE +TRN IK+FF+
Sbjct: 279 MMDVLLAASGDENAEYNITRNHIKSFFV 306
>UniRef100_Q9LJY8 Cytochrome P450-like protein [Arabidopsis thaliana]
Length = 513
Score = 134 bits (336), Expect = 2e-30
Identities = 78/208 (37%), Positives = 119/208 (56%), Gaps = 2/208 (0%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
+ HDLN S R L+ + FI+APYG Y++FMKK VTKLL L R IR
Sbjct: 100 RVHDLNVSSRGSPPFEESLLFGSTGFISAPYGDYFKFMKKHLVTKLLGPQALERSRLIRT 159
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
ELE+ +LL + + S ++G + L+NN +C+M MG +C ++ + E ++ L+ E
Sbjct: 160 NELERFYINLLDKATKKESVEIGKEAMKLSNNSICKMIMGRSCLEEKG-EAERVRGLIIE 218
Query: 122 FMHVGAKLSMGEVL-GPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQGD 180
++ K + L G L K + + K++ + FD +LE +L+EHEE+ + E D
Sbjct: 219 SFYLTKKFFLAFTLRGLLEKLGISLFKKEIMGVSRRFDDLLERYLREHEEKPDNEHQDTD 278
Query: 181 MMDILLQVYRNENAEVRLTRNDIKAFFL 208
M+D LL YR+E AE ++TRN IKAF +
Sbjct: 279 MIDALLAAYRDEKAEYKITRNQIKAFLV 306
>UniRef100_Q9FI39 Cytochrome P450 [Arabidopsis thaliana]
Length = 511
Score = 133 bits (335), Expect = 3e-30
Identities = 76/208 (36%), Positives = 118/208 (56%), Gaps = 2/208 (0%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
K D+N S RP + S FI PYG Y +FMKK V KLL L R +IR
Sbjct: 101 KAQDVNVSSRPPPPIEESLILGSSSFINTPYGDYSKFMKKFMVQKLLGPQALQRSRNIRA 160
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
ELE+ K+LL + + ++ ++ + LTNN +C+M MG +C ++ N + E ++ LV E
Sbjct: 161 DELERFYKTLLDKAMKKQTVEIRNEAMKLTNNTICKMIMGRSCSEE-NGEAETVRGLVTE 219
Query: 122 FMHVGAKLSMGEVL-GPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQGD 180
+ + K +G + PL K + + K+L + FD++LE L EHEE++ D
Sbjct: 220 SIFLTKKHFLGAMFHKPLKKLGISLFAKELMNVSNRFDELLEKILVEHEEKLQEHHQTSD 279
Query: 181 MMDILLQVYRNENAEVRLTRNDIKAFFL 208
M+D+LL+ Y +ENAE ++TR+ IK+ F+
Sbjct: 280 MLDMLLEAYGDENAEYKITRDQIKSLFV 307
>UniRef100_Q9LPS6 Cytochrome P450, putative [Arabidopsis thaliana]
Length = 519
Score = 133 bits (334), Expect = 3e-30
Identities = 77/212 (36%), Positives = 126/212 (59%), Gaps = 6/212 (2%)
Query: 1 MKTHDLNFSYRP--QFGSSHDYLYKGSY-FITAPYGPYWRFMKKICVTKLLSSSQLGRFM 57
++ DLNF+ R Q L GS+ F++ PYG YWRFMKK+ V KLL S L +
Sbjct: 101 LRIQDLNFASRDSGQTPIMEKSLLFGSFGFVSVPYGDYWRFMKKLLVKKLLGSHSLEQTR 160
Query: 58 HIREQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQC 117
+R +EL+ L + +N + D+G + LTNN +CRM+MG +C ++ N + E ++
Sbjct: 161 LLRGKELQTFRAMLFDKAAKNETVDVGKEMMKLTNNSICRMTMGRSCSEE-NGEAEQVRG 219
Query: 118 LVREFMHVGAKLSMGEVLGPLGKF-DLFGYGKKLRKIVGEFDKILEGFLKEHEERINTED 176
LV + + + K + ++G K + +GK++ ++ +D++LE +KEHEE N +
Sbjct: 220 LVTKSLSLTKKFLIASIVGQFSKLVGISLFGKEIMEVSQRYDELLEKIIKEHEENPNNGE 279
Query: 177 CQGDMMDILLQVYRNENAEVRLTRNDIKAFFL 208
DMMD+LL+V ++NAE +++RN IKA F+
Sbjct: 280 -DRDMMDVLLEVCADDNAEFKISRNQIKALFV 310
>UniRef100_O23389 Cytochrome P450 like protein [Arabidopsis thaliana]
Length = 527
Score = 133 bits (334), Expect = 3e-30
Identities = 74/209 (35%), Positives = 121/209 (57%), Gaps = 3/209 (1%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
K HD+N S R ++ S + APYG YW+FMKK+ TKLL L R +R
Sbjct: 99 KAHDVNVSSRGIIALDESLMFGASGILNAPYGDYWKFMKKLMATKLLRPQVLERSRGVRV 158
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
+EL + +S+L + +N S ++G + L NN LC++ MG + F + N + ++ LV E
Sbjct: 159 EELHRFYRSILDKATKNESVEIGKEAMKLMNNTLCKLIMGRS-FSEDNGESNRVRGLVDE 217
Query: 122 FMHVGAKLSMGEVL-GPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQG- 179
+ K+ + +L PL K + + K++ + +FD++LE L+E +E + ++ +G
Sbjct: 218 TYALSEKIFLAAILRRPLAKLRISLFKKEIMGVSNKFDELLERILQERKENLEEKNNEGM 277
Query: 180 DMMDILLQVYRNENAEVRLTRNDIKAFFL 208
DMMD+LL+ Y +ENAE ++T IKAFF+
Sbjct: 278 DMMDVLLEAYGDENAEYKITWKHIKAFFV 306
>UniRef100_Q9LJY4 Cytochrome P450-like protein [Arabidopsis thaliana]
Length = 510
Score = 132 bits (333), Expect = 5e-30
Identities = 80/206 (38%), Positives = 116/206 (55%), Gaps = 3/206 (1%)
Query: 4 HDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIREQE 63
HD+N S+R L F TAPYG YW+FMKK+ VTKLL L R IR
Sbjct: 103 HDVNISFRGNPPIEESLLVGSFGFFTAPYGDYWKFMKKVMVTKLLGPQALQRSRGIRADA 162
Query: 64 LEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVREFM 123
LE+ +LL + + S ++G + L + +C+M MG F + N + E ++ LV E
Sbjct: 163 LERFYMNLLDKAMKKESVEIGKETMKLIYDSICKMIMGRN-FSEENGEAERVRGLVTEST 221
Query: 124 HVGAKLSMGEVL-GPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQGDMM 182
+ K+ M VL PL K + + K++ + FD++LE FL EHEE++N ED DMM
Sbjct: 222 ALTKKIFMANVLHKPLKKLGISLFKKEIMDVSNSFDELLERFLVEHEEKLN-EDQDMDMM 280
Query: 183 DILLQVYRNENAEVRLTRNDIKAFFL 208
+LL R++NAE ++TRN IK+ F+
Sbjct: 281 GVLLAACRDKNAECKITRNHIKSLFV 306
>UniRef100_Q9LIG7 Cytochrome P450-like protein [Arabidopsis thaliana]
Length = 521
Score = 132 bits (333), Expect = 5e-30
Identities = 77/208 (37%), Positives = 117/208 (56%), Gaps = 2/208 (0%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
+ HD+N S R ++ S + APYG Y +F+KKI TKLL L R +R
Sbjct: 100 RIHDMNISSRGAAAIDESLVFGSSGVVYAPYGDYLKFVKKIIATKLLRPQVLERSRGLRA 159
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
+EL++L +L + N + ++G + T L NNI CRMSMG + F + N + E ++ LV E
Sbjct: 160 EELQRLYNRILDKAKMNENVEIGKEATMLMNNIFCRMSMGRS-FSEENGEAERVRGLVAE 218
Query: 122 FMHVGAKLSMGEVLGP-LGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQGD 180
+ K+ VL L K + + K + + F+++LE L EH E+++ E D
Sbjct: 219 SYALAKKIFFASVLRRLLKKLRIPLFKKDIMDVSNRFNELLEKILVEHNEKLDEEHKDTD 278
Query: 181 MMDILLQVYRNENAEVRLTRNDIKAFFL 208
MMD+LL Y +ENAE ++TRN IK+FF+
Sbjct: 279 MMDVLLAAYADENAEYKITRNHIKSFFV 306
>UniRef100_Q9LPS5 Cytochrome P450, putative [Arabidopsis thaliana]
Length = 533
Score = 131 bits (329), Expect = 1e-29
Identities = 79/210 (37%), Positives = 123/210 (57%), Gaps = 3/210 (1%)
Query: 1 MKTHDLNFSYRPQFGSSHDY-LYKGSY-FITAPYGPYWRFMKKICVTKLLSSSQLGRFMH 58
++T DLNF+ R + S + L GS+ F++APYG YWRFMKK+ VT L S L +
Sbjct: 101 LRTQDLNFATRQREVSIMEKSLLFGSFGFVSAPYGDYWRFMKKLLVTNLFGSHSLEQTRL 160
Query: 59 IREQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCL 118
IRE+EL+ L + + + D+G + LTNN +CRM MG C ++ + +V +
Sbjct: 161 IREKELKTFRTMLFDKAAKKGTVDVGKEMMKLTNNSICRMIMGRRCSEENSEAEKVEDLV 220
Query: 119 VREFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQ 178
++ F V L V L KF + + K++ ++ +D++LE +KEHEE N ++
Sbjct: 221 IKSFSLVKKVLIANTVGRLLKKFGISLFEKEIMEVSQRYDELLEKIIKEHEEDPNKKE-D 279
Query: 179 GDMMDILLQVYRNENAEVRLTRNDIKAFFL 208
DMMD+LL+V ++ AEV++TRN IKA +
Sbjct: 280 RDMMDVLLEVCADDKAEVKITRNQIKALIV 309
>UniRef100_Q9LIG8 Cytochrome P450-like protein [Arabidopsis thaliana]
Length = 526
Score = 131 bits (329), Expect = 1e-29
Identities = 78/208 (37%), Positives = 118/208 (56%), Gaps = 3/208 (1%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
K D+N S R + L+ S F APYG Y++FM+K+ TKLL L R IR
Sbjct: 105 KAQDVNVSSRGHAPAGESLLFGSSSFFFAPYGDYFKFMRKLIATKLLGPQALERSRKIRA 164
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
EL++ ++LL + + S D+ + L NNI+C+M MG +C + N + E ++ LV E
Sbjct: 165 DELDRFYRNLLDKAMKKESVDIVEEAAKLNNNIICKMIMGRSC-SEDNGEAERVRGLVIE 223
Query: 122 FMHVGAKLSMGEVLG-PLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQGD 180
+ ++ +G + PL K + + K + K V FD++LE L EHEER+ D
Sbjct: 224 STALTKQIFLGMIFDKPLKKLGISLFQKDI-KSVSRFDELLEKILVEHEERMGKHYKAND 282
Query: 181 MMDILLQVYRNENAEVRLTRNDIKAFFL 208
MMD+LL+ Y +ENAE ++TRN IK+ F+
Sbjct: 283 MMDLLLEAYGDENAEYKITRNHIKSLFV 310
>UniRef100_Q9LJY7 Putative cytochrome P450 [Arabidopsis thaliana]
Length = 510
Score = 130 bits (328), Expect = 2e-29
Identities = 80/209 (38%), Positives = 122/209 (58%), Gaps = 5/209 (2%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
K HDLN S R + L S F+ APYG YW+FMKK+ VTKLL L R IR
Sbjct: 98 KAHDLNISSRDNPPINESLLVGSSVFVGAPYGDYWKFMKKLLVTKLLGPQALERSRSIRA 157
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
ELE+ +SLL + + S ++G + T L+ N +CRMSMG + F + + + E ++ LV E
Sbjct: 158 DELERFYRSLLDKAMKKESVEIGKEATKLSINSICRMSMGRS-FSEESGEAERVRGLVTE 216
Query: 122 FMHVGAKLSMGEVL-GPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQGD 180
+ K+ + +L PL K + + K++ + FD++LE + E E++ N + QG
Sbjct: 217 LDGLTKKVLLVNILRWPLEKLRISLFKKEIMYVSNSFDELLERIIVEREKKPN--EHQGT 274
Query: 181 -MMDILLQVYRNENAEVRLTRNDIKAFFL 208
+MD+LL+ Y +E AE ++TRN IK+ F+
Sbjct: 275 YLMDVLLEAYEDEKAEHKITRNHIKSLFV 303
>UniRef100_Q9LJY6 Cytochrome P450-like protein [Arabidopsis thaliana]
Length = 514
Score = 130 bits (327), Expect = 2e-29
Identities = 82/208 (39%), Positives = 119/208 (56%), Gaps = 3/208 (1%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
K HDLN S R L S F +APYG Y++FMKK+ VTKLL L R IR
Sbjct: 100 KAHDLNISSRDNPSIDDSLLIGSSVFGSAPYGDYFKFMKKLLVTKLLGPQALERSRGIRA 159
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
ELE+ SLL + + ++G + T L+NN L RMS+G + F + N + E ++ LV E
Sbjct: 160 DELERFHSSLLDKAVKKEIVEIGKEATKLSNNSLWRMSIGRS-FSEKNGEAERVRGLVTE 218
Query: 122 FMHVGAKLSMGEVL-GPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQGD 180
+ K+ +L PL K + + K++ + FD++LE L EHE++++ D
Sbjct: 219 LDGLTKKVLFATLLHRPLEKLGISLFKKEIMCVSDSFDEVLERVLVEHEQKLDDHQ-DRD 277
Query: 181 MMDILLQVYRNENAEVRLTRNDIKAFFL 208
MMD+LL Y +ENAE +++RN IKAFF+
Sbjct: 278 MMDVLLAAYGDENAEHKISRNHIKAFFV 305
>UniRef100_Q9SP27 Flavone synthase II [Callistephus chinensis]
Length = 514
Score = 129 bits (325), Expect = 4e-29
Identities = 75/212 (35%), Positives = 124/212 (58%), Gaps = 6/212 (2%)
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KT++L FS R + + D++ G F APYGPYW+F+KK + +LL + L F+ IR
Sbjct: 95 LKTNELAFSSR-KHSLAIDHVTYGVSFAFAPYGPYWKFIKKTSIVELLGNQNLSNFLPIR 153
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
QE+ +LL++L++ S +N S +L + LTNN++C+M M C N + + + LVR
Sbjct: 154 TQEVHELLQTLMVKSKKNESVNLSEELLKLTNNVICQMMMSIRC-SGTNNEADEAKNLVR 212
Query: 121 EFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEER---INTEDC 177
E + + ++ + + DL G+ K+ ++ +LE + E EE+ +ED
Sbjct: 213 EVTKIFGEFNISDFICLFKNIDLQGFKKRYVDTHTRYNALLEKMIFEREEKRKQKKSEDG 272
Query: 178 QG-DMMDILLQVYRNENAEVRLTRNDIKAFFL 208
+G D +DILL V +ENAE+++TR+ IKA L
Sbjct: 273 KGKDFLDILLDVLEDENAEIKITRDHIKALIL 304
>UniRef100_Q9LIG9 Cytochrome P450-like protein [Arabidopsis thaliana]
Length = 523
Score = 129 bits (325), Expect = 4e-29
Identities = 76/208 (36%), Positives = 116/208 (55%), Gaps = 2/208 (0%)
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
K D+N S R + S F APYG Y++FM+K+ TKLL L R IR
Sbjct: 101 KAQDVNVSSRGHAPVGESLWFGSSSFFFAPYGDYFKFMRKLIATKLLGPQALERSRKIRA 160
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
EL++ K+LL + + S ++G + L NNI+C+M MG +C ++ N + E + LV E
Sbjct: 161 DELDRFYKTLLDKAMKKESVEIGEEAAKLNNNIICKMIMGRSCSEE-NGEAEKFRHLVIE 219
Query: 122 FMHVGAKLSMGEVL-GPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQGD 180
M + ++ G + PL K + + K + + +FD++LE L EHEE+ + D
Sbjct: 220 SMALTKQIFFGMIFHKPLKKLGISLFQKDILSLSRKFDELLEKILFEHEEKKAEHNQAND 279
Query: 181 MMDILLQVYRNENAEVRLTRNDIKAFFL 208
MMD LL+ Y +ENAE ++TRN IK+ F+
Sbjct: 280 MMDFLLEAYGDENAEYKITRNHIKSLFV 307
>UniRef100_Q42798 Cytochrome P450 93A1 [Glycine max]
Length = 509
Score = 129 bits (324), Expect = 5e-29
Identities = 70/222 (31%), Positives = 124/222 (55%), Gaps = 20/222 (9%)
Query: 1 MKTHDLNFSYRPQFG--------SSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQ 52
+KTH++NFS RP S D+L F AP+GPYW+FMKK+C+++LLS
Sbjct: 86 LKTHEINFSNRPGQNVAVKGLAYDSQDFL-----FAFAPFGPYWKFMKKLCMSELLSGRM 140
Query: 53 LGRFMHIREQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDP 112
+ +F+ +R+QE ++ + + + D G + TL+NNI+ RM++ + N
Sbjct: 141 MDQFLPVRQQETKRFISRVFRKGVAGEAVDFGDELMTLSNNIVSRMTLSQKTSENDN-QA 199
Query: 113 EVIQCLVREFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERI 172
E ++ LV + K ++ + + L FDL G+ +K+++ FD +++G +K+ +E
Sbjct: 200 EEMKKLVSNIAELMGKFNVSDFIWYLKPFDLQGFNRKIKETRDRFDVVVDGIIKQRQEER 259
Query: 173 NTEDCQG------DMMDILLQVYRNENAEVRLTRNDIKAFFL 208
G DM+D+LL ++ +ENAE++L + +IKAF +
Sbjct: 260 RKNKETGTAKQFKDMLDVLLDMHEDENAEIKLDKKNIKAFIM 301
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.325 0.142 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 355,336,812
Number of Sequences: 2790947
Number of extensions: 14392222
Number of successful extensions: 44916
Number of sequences better than 10.0: 608
Number of HSP's better than 10.0 without gapping: 506
Number of HSP's successfully gapped in prelim test: 102
Number of HSP's that attempted gapping in prelim test: 44009
Number of HSP's gapped (non-prelim): 637
length of query: 212
length of database: 848,049,833
effective HSP length: 122
effective length of query: 90
effective length of database: 507,554,299
effective search space: 45679886910
effective search space used: 45679886910
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 72 (32.3 bits)
Medicago: description of AC142143.3


