

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142096.6 - phase: 0 /pseudo
(930 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
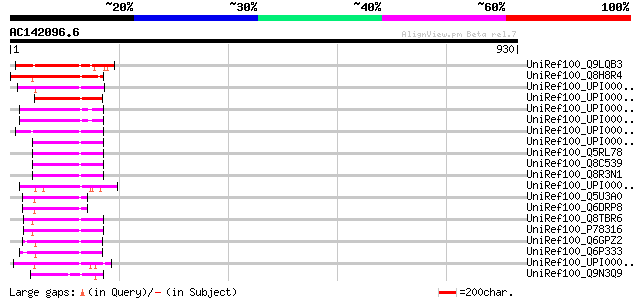
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LQB3 F4N2.4 [Arabidopsis thaliana] 155 7e-36
UniRef100_Q8H8R4 Hypothetical protein OJ1523_A02.5 [Oryza sativa] 152 3e-35
UniRef100_UPI00003C14CD UPI00003C14CD UniRef100 entry 87 2e-15
UniRef100_UPI0000364504 UPI0000364504 UniRef100 entry 87 3e-15
UniRef100_UPI00003ADA76 UPI00003ADA76 UniRef100 entry 83 3e-14
UniRef100_UPI00003ADA75 UPI00003ADA75 UniRef100 entry 83 3e-14
UniRef100_UPI000036CBCC UPI000036CBCC UniRef100 entry 80 4e-13
UniRef100_UPI000017E2CF UPI000017E2CF UniRef100 entry 79 6e-13
UniRef100_Q5RL78 RIKEN cDNA 2610033H07 [Mus musculus] 79 8e-13
UniRef100_Q8C539 Mus musculus adult male spinal cord cDNA, RIKEN... 79 8e-13
UniRef100_Q8R3N1 Probable nucleolar complex protein 14 [Mus musc... 79 8e-13
UniRef100_UPI0000235549 UPI0000235549 UniRef100 entry 78 1e-12
UniRef100_Q5U3A0 Nop14-like [Brachydanio rerio] 78 1e-12
UniRef100_Q6DRP8 Nop14-like [Brachydanio rerio] 78 1e-12
UniRef100_Q8TBR6 C4orf9 protein [Homo sapiens] 78 1e-12
UniRef100_P78316 Probable nucleolar complex protein 14 [Homo sap... 78 1e-12
UniRef100_Q6GPZ2 MGC82495 protein [Xenopus laevis] 76 5e-12
UniRef100_Q6P333 LOC394902 protein [Xenopus tropicalis] 75 1e-11
UniRef100_UPI000042FEFA UPI000042FEFA UniRef100 entry 74 2e-11
UniRef100_Q9N3Q9 Hypothetical protein Y48G1A.4 [Caenorhabditis e... 73 3e-11
>UniRef100_Q9LQB3 F4N2.4 [Arabidopsis thaliana]
Length = 891
Score = 155 bits (391), Expect = 7e-36
Identities = 93/201 (46%), Positives = 133/201 (65%), Gaps = 26/201 (12%)
Query: 12 GTNTSNKSTSKKKKNNKMGPEAVAMKAKAQKTNTNPFESIWSRRKFEVMGQKRKGDTKRM 71
G + K KK +MGP+AVAMKAK QK + NPFESI SRRKF+++G+KRKG+ + +
Sbjct: 2 GKDNKMKKKPNKKGERRMGPDAVAMKAKTQKVD-NPFESIRSRRKFDILGKKRKGEERFV 60
Query: 72 GLARSLAVEKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRERQLNVKS 131
++R+ AV+KRK TL KEYEQS KSS F+D+RIGE ++ L +F K I+RSQR+RQL K
Sbjct: 61 SVSRTRAVDKRKNTLEKEYEQSLKSSVFLDKRIGEQNDELGEFDKGIIRSQRQRQL--KL 118
Query: 132 SKKSKYHLSEEDDDEFEGIDG-LG----RDDFEDEMLGEDDDETDE------------T* 174
+KKS Y+LS+ ++D +E DG LG +DDF+ +L ++D + D+
Sbjct: 119 AKKSMYNLSDGEEDVYE--DGALGGSSVKDDFDSGLLSDEDLQDDDLEASASKRLKHLNR 176
Query: 175 NQE----GSDERSYCKKQVLQ 191
N+E G +ER KK+V++
Sbjct: 177 NREVDASGEEERRKSKKEVME 197
>UniRef100_Q8H8R4 Hypothetical protein OJ1523_A02.5 [Oryza sativa]
Length = 952
Score = 152 bits (385), Expect = 3e-35
Identities = 90/186 (48%), Positives = 121/186 (64%), Gaps = 18/186 (9%)
Query: 1 MAKSSKRSIANGTNTSNKSTSK--KKKNNKMGPEAVAMKAK---------AQKTNTNPFE 49
MAK+ + A ++K K KKK K GP AVAMKA+ A + NPFE
Sbjct: 1 MAKTKPMAAAAAAAAADKKKGKGKKKKQGKNGPAAVAMKARGAAAAAAAAAASGSNNPFE 60
Query: 50 SIWSRRKFEVMGQKRKGDTKRMGLARSLAVEKRKKTLLKEYEQSTKSSEFIDRRIGENDE 109
+IWSRRKF+V+G+KRKG+ +R+G ARS A+ KR+ TLLKE+EQS KSS F DRRIGE DE
Sbjct: 61 AIWSRRKFDVLGKKRKGEERRIGRARSEAIHKRENTLLKEFEQSAKSSVFQDRRIGERDE 120
Query: 110 GLDDFGKAILRSQRERQLNVKSSKKSKYHLSEEDDDEFE---GIDGLGRDDFEDEMLGED 166
L +F K ILR QRE +K ++SKY+LS++++DE + G+DDF++E+
Sbjct: 121 TLPEFDKVILRQQREHMAKLK--RESKYNLSDDEEDEVDVHLPHSLSGKDDFDEEV--PL 176
Query: 167 DDETDE 172
DD +DE
Sbjct: 177 DDYSDE 182
>UniRef100_UPI00003C14CD UPI00003C14CD UniRef100 entry
Length = 981
Score = 87.0 bits (214), Expect = 2e-15
Identities = 56/166 (33%), Positives = 92/166 (54%), Gaps = 9/166 (5%)
Query: 15 TSNKSTSKKKKNNKMGP-EAVAMKAKAQKTNT-----NPFESIWSRRKFEVMGQKRKGDT 68
T + SKK ++ K G E A +A+ K N NPFE ++ K EV+G+K KG
Sbjct: 22 TDRRQLSKKSRSRKGGQDEKDAAEARRNKINAIVADLNPFEQKVTKPKHEVLGRKIKGAV 81
Query: 69 KRMGLARSLAVEKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRERQLN 128
R G A++ + +R++TLL E++ ++ F+DRR GEND + K + R E+Q
Sbjct: 82 GRPGAAKASGLAQRRETLLPEWQARDRTGTFVDRRFGENDANMTPEEKMLERFATEKQR- 140
Query: 129 VKSSKKSKYHLSEEDD-DEFEGIDGLGRDDFEDEMLGEDDDETDET 173
+++K + ++L+++DD G G DD D L EDD++ D++
Sbjct: 141 -RAAKGAAFNLNDDDDLLTHYGQSLSGLDDLADIRLPEDDEDEDDS 185
Score = 37.0 bits (84), Expect = 2.8
Identities = 36/134 (26%), Positives = 60/134 (43%), Gaps = 16/134 (11%)
Query: 58 EVMGQKRKGDTKRMGLARSLAVEKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKA 117
E+ ++R R+ LA E+ ++ L++ E++ D E DE DD GK
Sbjct: 310 ELAFERRAKPQDRLKSEEELAAEQAER--LRKAEKARLKRMRGDESDDEADED-DDGGK- 365
Query: 118 ILRSQRERQLNVKSSKKSKYHLSEEDDD-EFEGID---------GLGRDDFEDEMLGEDD 167
R +R + N S+ K + DDD E +G+ GL D E+D
Sbjct: 366 --RRKRSKGANSSSAPKRAAQADDLDDDFELDGMTAGEVYGLGAGLAESGTVDASDAEED 423
Query: 168 DETDET*NQEGSDE 181
D+T++ ++EG D+
Sbjct: 424 DDTEDDNDEEGQDD 437
>UniRef100_UPI0000364504 UPI0000364504 UniRef100 entry
Length = 846
Score = 86.7 bits (213), Expect = 3e-15
Identities = 50/124 (40%), Positives = 75/124 (60%), Gaps = 2/124 (1%)
Query: 46 NPFESIWSRRKFEVMGQKRKGDTKRMGLARSLAVEKRKKTLLKEYEQSTKSSEFIDRRIG 105
NPFE +R+KF+V+G+K K D G++RS A+ KRK+TLLKEY+Q KSS+FIDRR G
Sbjct: 2 NPFEVKINRKKFDVLGRKVKHDVGLPGVSRSKAITKRKETLLKEYKQKNKSSKFIDRRFG 61
Query: 106 ENDEGLDDFGKAILRSQRERQLNVKSSKKSKYHLSEEDDDEFEGIDGLGRDDFEDEMLGE 165
E D + K + R ER+ + KK ++L+E+++ G + F D + +
Sbjct: 62 EYDTKMAPEDKILQRFAMERKRSY--DKKDVFNLNEDEELTHFGQSLAKMETFNDFVNSD 119
Query: 166 DDDE 169
D+ E
Sbjct: 120 DESE 123
>UniRef100_UPI00003ADA76 UPI00003ADA76 UniRef100 entry
Length = 837
Score = 83.2 bits (204), Expect = 3e-14
Identities = 53/158 (33%), Positives = 86/158 (53%), Gaps = 10/158 (6%)
Query: 18 KSTSKKKKNNKMGPEAVAMKAKAQKTNTNPFESIWSRRKFEVMGQKRKGDTKRMGLARSL 77
K+ +KKK + + + +NPFE +R+KF ++G+K K D G++RS
Sbjct: 3 KAAKRKKKASTSSKAKGSAEPTIAMVKSNPFEVKVNRQKFNILGRKTKNDVGLPGVSRSK 62
Query: 78 AVEKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRERQLNVKSSKKSKY 137
A++KR +TLLKEY++ K++ F D+R GE + + K I R ERQ N KK+ Y
Sbjct: 63 AIKKRTQTLLKEYKEKEKTNVFRDKRFGEYNTKISPEEKMIKRFTLERQQNY--GKKNIY 120
Query: 138 HLSEEDDDEFEGIDGLGRDDFEDEMLG---EDDDETDE 172
+L+E+++ + G+ E E L + D +TDE
Sbjct: 121 NLNEDEE-----LTHFGQSLAEIEKLNDVIDSDSDTDE 153
>UniRef100_UPI00003ADA75 UPI00003ADA75 UniRef100 entry
Length = 779
Score = 83.2 bits (204), Expect = 3e-14
Identities = 53/158 (33%), Positives = 86/158 (53%), Gaps = 10/158 (6%)
Query: 18 KSTSKKKKNNKMGPEAVAMKAKAQKTNTNPFESIWSRRKFEVMGQKRKGDTKRMGLARSL 77
K+ +KKK + + + +NPFE +R+KF ++G+K K D G++RS
Sbjct: 1 KAAKRKKKASTSSKAKGSAEPTIAMVKSNPFEVKVNRQKFNILGRKTKNDVGLPGVSRSK 60
Query: 78 AVEKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRERQLNVKSSKKSKY 137
A++KR +TLLKEY++ K++ F D+R GE + + K I R ERQ N KK+ Y
Sbjct: 61 AIKKRTQTLLKEYKEKEKTNVFRDKRFGEYNTKISPEEKMIKRFTLERQQNY--GKKNIY 118
Query: 138 HLSEEDDDEFEGIDGLGRDDFEDEMLG---EDDDETDE 172
+L+E+++ + G+ E E L + D +TDE
Sbjct: 119 NLNEDEE-----LTHFGQSLAEIEKLNDVIDSDSDTDE 151
>UniRef100_UPI000036CBCC UPI000036CBCC UniRef100 entry
Length = 873
Score = 79.7 bits (195), Expect = 4e-13
Identities = 51/161 (31%), Positives = 87/161 (53%), Gaps = 4/161 (2%)
Query: 11 NGTNTSNKSTSKKKKNNKMGPEAVAMKAKAQKTNTNPFESIWSRRKFEVMGQKRKGDTKR 70
+G K+ + + G A A A K N+NPFE +R+KF+++G+K + D
Sbjct: 12 SGARAMAKAKKVRARRKASGAPAGARGGPA-KANSNPFEVKVNRQKFQILGRKTRHDVGL 70
Query: 71 MGLARSLAVEKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRERQLNVK 130
G++R+ A+ KR +TLLKEY++ KS+ F D+R GE + + K + R E+Q +
Sbjct: 71 PGVSRARALRKRTQTLLKEYKERDKSNVFRDKRFGEYNSNMSPEEKMMKRFALEQQRH-- 128
Query: 131 SSKKSKYHLSEEDDDEFEGIDGLGRDDFEDEMLGEDDDETD 171
KKS Y+L+E+++ G L + ++++ D D D
Sbjct: 129 HEKKSIYNLNEDEELTHYG-QSLADIEKHNDIVDSDSDAED 168
>UniRef100_UPI000017E2CF UPI000017E2CF UniRef100 entry
Length = 862
Score = 79.0 bits (193), Expect = 6e-13
Identities = 45/130 (34%), Positives = 77/130 (58%), Gaps = 3/130 (2%)
Query: 42 KTNTNPFESIWSRRKFEVMGQKRKGDTKRMGLARSLAVEKRKKTLLKEYEQSTKSSEFID 101
KTN NPFE +R+KF+++G+K + D G++R+ A++KR +TLLKEY++ KS+ F D
Sbjct: 26 KTNPNPFEVKVNRQKFQILGRKTRHDVGLPGVSRARAIKKRTQTLLKEYKERNKSNVFTD 85
Query: 102 RRIGENDEGLDDFGKAILRSQRERQLNVKSSKKSKYHLSEEDDDEFEGIDGLGRDDFEDE 161
+R GE + + K + R E+Q KK+ Y+L+E+++ G L + ++
Sbjct: 86 KRFGEYNSNISPEEKMMKRFALEQQR--YHEKKNIYNLNEDEELTHYG-QSLADIEKHND 142
Query: 162 MLGEDDDETD 171
++ D D D
Sbjct: 143 IVDSDSDTED 152
Score = 37.0 bits (84), Expect = 2.8
Identities = 40/186 (21%), Positives = 82/186 (43%), Gaps = 28/186 (15%)
Query: 37 KAKAQKTNTNPFESIWSRRKFEVMGQKRKGDTKRMGLARSLAVEKRKKTLLKEYEQSTKS 96
K K +K + ++ + FE+ Q + RM LA E+++K L++ E
Sbjct: 240 KEKKEKPQPDAYDMMVRELGFEMKAQP----SNRMKTEEELAKEEQEK--LRKLE----- 288
Query: 97 SEFIDRRIGENDE---------GLDDFGKAILRSQRERQLNVKSSKKSKYHLSEEDDDEF 147
+E + R +G+++ DD + + +R+L S K K ++ + +++
Sbjct: 289 AERLRRMLGKDEHENKKKPKHTSADDLNDGFILDKDDRRL--LSYKDGKMNIEDAQEEQS 346
Query: 148 EGIDGLGRD----DFEDEMLGEDDDETDET*NQEGSDERSYCKK--QVLQG*KSKRKGKR 201
+G DG D + E E GED + ++ +G D S + + + ++ +K +R
Sbjct: 347 KGADGQENDQKEEEDESEEEGEDGESCEDPDESDGPDSHSDLESSAETEEEDETPKKEQR 406
Query: 202 *RPAGR 207
P G+
Sbjct: 407 QTPGGK 412
>UniRef100_Q5RL78 RIKEN cDNA 2610033H07 [Mus musculus]
Length = 860
Score = 78.6 bits (192), Expect = 8e-13
Identities = 45/130 (34%), Positives = 76/130 (57%), Gaps = 3/130 (2%)
Query: 42 KTNTNPFESIWSRRKFEVMGQKRKGDTKRMGLARSLAVEKRKKTLLKEYEQSTKSSEFID 101
KTN NPFE +R+KF+++G+K + D G++R+ A+ KR +TLLKEY++ KS+ F D
Sbjct: 26 KTNPNPFEVKVNRQKFQILGRKTRHDVGLPGVSRARAIRKRTQTLLKEYKERNKSNVFAD 85
Query: 102 RRIGENDEGLDDFGKAILRSQRERQLNVKSSKKSKYHLSEEDDDEFEGIDGLGRDDFEDE 161
+R GE + + K + R E+Q KK+ Y+L+E+++ G L + ++
Sbjct: 86 KRFGEYNSNISPEEKMMKRFALEQQR--YHEKKNIYNLNEDEELTHYG-QSLADIEKHND 142
Query: 162 MLGEDDDETD 171
++ D D D
Sbjct: 143 IVDSDSDTED 152
Score = 35.8 bits (81), Expect = 6.1
Identities = 37/163 (22%), Positives = 73/163 (44%), Gaps = 23/163 (14%)
Query: 31 PEAVAMKAKAQKTNTNPFESIWSRRKFEVMGQKRKGDTKRMGLARSLAVEKRKKTLLKEY 90
P+ K K +K + ++ + FE+ Q + RM LA E++++ LK+
Sbjct: 234 PKKSEDKEKKEKPQPDEYDMMVRELGFEMKAQP----SNRMKTEEELAKEEQER--LKKL 287
Query: 91 EQSTKSSEFIDRRIGENDE---------GLDDFGKAILRSQRERQLNVKSSKKSKYHLSE 141
E +E + R +G+++ DD + + +R+L S K K ++ +
Sbjct: 288 E-----AERLRRMLGKDEHENKKKPKHTSADDLNDGFILDKDDRRL--LSYKDGKMNIED 340
Query: 142 EDDDEFEGIDGLGRDDFEDEMLGEDDDETDET*NQEGSDERSY 184
+++ + DG D E E E++DE+ E ++E D S+
Sbjct: 341 VQEEQSKEADGQENDQKEGEDDSEEEDESHED-SEESEDPDSH 382
>UniRef100_Q8C539 Mus musculus adult male spinal cord cDNA, RIKEN full-length
enriched library, clone:A330071M14 product:RES4-25
protein homolog [Mus musculus]
Length = 190
Score = 78.6 bits (192), Expect = 8e-13
Identities = 45/130 (34%), Positives = 76/130 (57%), Gaps = 3/130 (2%)
Query: 42 KTNTNPFESIWSRRKFEVMGQKRKGDTKRMGLARSLAVEKRKKTLLKEYEQSTKSSEFID 101
KTN NPFE +R+KF+++G+K + D G++R+ A+ KR +TLLKEY++ KS+ F D
Sbjct: 26 KTNPNPFEVKVNRQKFQILGRKTRHDVGLPGVSRARAIRKRTQTLLKEYKERNKSNVFAD 85
Query: 102 RRIGENDEGLDDFGKAILRSQRERQLNVKSSKKSKYHLSEEDDDEFEGIDGLGRDDFEDE 161
+R GE + + K + R E+Q KK+ Y+L+E+++ G L + ++
Sbjct: 86 KRFGEYNSNISPEEKMMKRFALEQQR--YHEKKNIYNLNEDEELTHYG-QSLADIEKHND 142
Query: 162 MLGEDDDETD 171
++ D D D
Sbjct: 143 IVDSDSDTED 152
>UniRef100_Q8R3N1 Probable nucleolar complex protein 14 [Mus musculus]
Length = 860
Score = 78.6 bits (192), Expect = 8e-13
Identities = 45/130 (34%), Positives = 76/130 (57%), Gaps = 3/130 (2%)
Query: 42 KTNTNPFESIWSRRKFEVMGQKRKGDTKRMGLARSLAVEKRKKTLLKEYEQSTKSSEFID 101
KTN NPFE +R+KF+++G+K + D G++R+ A+ KR +TLLKEY++ KS+ F D
Sbjct: 26 KTNPNPFEVKVNRQKFQILGRKTRHDVGLPGVSRARAIRKRTQTLLKEYKERNKSNVFAD 85
Query: 102 RRIGENDEGLDDFGKAILRSQRERQLNVKSSKKSKYHLSEEDDDEFEGIDGLGRDDFEDE 161
+R GE + + K + R E+Q KK+ Y+L+E+++ G L + ++
Sbjct: 86 KRFGEYNSNISPEEKMMKRFALEQQR--YHEKKNIYNLNEDEELTHYG-QSLADIEKHND 142
Query: 162 MLGEDDDETD 171
++ D D D
Sbjct: 143 IVDSDSDTED 152
Score = 36.6 bits (83), Expect = 3.6
Identities = 37/163 (22%), Positives = 73/163 (44%), Gaps = 23/163 (14%)
Query: 31 PEAVAMKAKAQKTNTNPFESIWSRRKFEVMGQKRKGDTKRMGLARSLAVEKRKKTLLKEY 90
P+ K K +K + ++ + FE+ Q + RM LA E++++ LK+
Sbjct: 234 PKKSENKEKKEKPQPDEYDMMVRELGFEMKAQP----SNRMKTEEELAKEEQER--LKKL 287
Query: 91 EQSTKSSEFIDRRIGENDE---------GLDDFGKAILRSQRERQLNVKSSKKSKYHLSE 141
E +E + R +G+++ DD + + +R+L S K K ++ +
Sbjct: 288 E-----AERLRRMLGKDEHENKKKPKHTSADDLNDGFILDKDDRRL--LSYKDGKMNIED 340
Query: 142 EDDDEFEGIDGLGRDDFEDEMLGEDDDETDET*NQEGSDERSY 184
+++ + DG D E E E++DE+ E ++E D S+
Sbjct: 341 VQEEQSKEADGQENDQKEGEDESEEEDESHED-SEESEDPDSH 382
>UniRef100_UPI0000235549 UPI0000235549 UniRef100 entry
Length = 913
Score = 78.2 bits (191), Expect = 1e-12
Identities = 70/219 (31%), Positives = 106/219 (47%), Gaps = 41/219 (18%)
Query: 18 KSTSKKKKNNKMGPEAVAMKAKAQKTNT-----NPFE-SIWSRRKFEVM----GQKRKGD 67
KS +K++N K G A + N NPFE SR KF+V G K
Sbjct: 23 KSKKQKRQNAKTGANAQNRAQRESALNAIRERFNPFEIRTVSRGKFDVTTRDGGAKSASG 82
Query: 68 TKRMGLARSLAVEKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRERQL 127
R G+ +SL E+R++TLL+E ++ K +DRR GEND + +A R R Q
Sbjct: 83 NFRPGVTKSLGEERRRQTLLQEMDRRNKVGGIMDRRFGENDPTMTPEERAAERFARASQK 142
Query: 128 NVKSSKKSKYHLSEEDDDEF------EGI-----DGLGRDDFEDEM-LGE---------- 165
++ K S ++L E+D++EF E + DG+ RDDF++++ G+
Sbjct: 143 KLR--KDSMFNLEEDDEEEFTLTHKGESLSFGDEDGI-RDDFQEDLDAGDMSDTEMPRKR 199
Query: 166 ----DDDETDET*NQEGSD--ERSYCKKQVLQG*KSKRK 198
DD E +E +Q+G D ER K +V+Q +K K
Sbjct: 200 KRFIDDPEAEEDGSQDGEDVPERKKSKHEVMQEVIAKSK 238
>UniRef100_Q5U3A0 Nop14-like [Brachydanio rerio]
Length = 860
Score = 78.2 bits (191), Expect = 1e-12
Identities = 48/124 (38%), Positives = 72/124 (57%), Gaps = 6/124 (4%)
Query: 24 KKNNKMGPEAVAMKAKAQKT----NTNPFESIWSRRKFEVMGQKRKGDTKRMGLARSLAV 79
KK + G +A K + K NPFE +R+KF+V+G+K K D G++RS A
Sbjct: 2 KKQQQKGVRGLADKVRRSKAAPEIQRNPFEVKINRKKFDVLGRKSKHDVGLPGVSRSRAN 61
Query: 80 EKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRERQLNVKSSKKSKYHL 139
+KRK+TLLKE + KS++FIDRR GE D L+ K + R ERQ ++ ++L
Sbjct: 62 QKRKETLLKELKVKGKSNKFIDRRFGEYDSKLNPEEKILQRFTLERQR--LHDRRDVFNL 119
Query: 140 SEED 143
+E++
Sbjct: 120 NEDE 123
>UniRef100_Q6DRP8 Nop14-like [Brachydanio rerio]
Length = 859
Score = 78.2 bits (191), Expect = 1e-12
Identities = 48/124 (38%), Positives = 72/124 (57%), Gaps = 6/124 (4%)
Query: 24 KKNNKMGPEAVAMKAKAQKT----NTNPFESIWSRRKFEVMGQKRKGDTKRMGLARSLAV 79
KK + G +A K + K NPFE +R+KF+V+G+K K D G++RS A
Sbjct: 2 KKQQQKGVRGLADKVRRSKAAPEIQRNPFEVKINRKKFDVLGRKSKHDVGLPGVSRSRAN 61
Query: 80 EKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRERQLNVKSSKKSKYHL 139
+KRK+TLLKE + KS++FIDRR GE D L+ K + R ERQ ++ ++L
Sbjct: 62 QKRKETLLKELKVKGKSNKFIDRRFGEYDSKLNPEEKILQRFTLERQR--LHDRRDVFNL 119
Query: 140 SEED 143
+E++
Sbjct: 120 NEDE 123
>UniRef100_Q8TBR6 C4orf9 protein [Homo sapiens]
Length = 806
Score = 78.2 bits (191), Expect = 1e-12
Identities = 51/153 (33%), Positives = 84/153 (54%), Gaps = 9/153 (5%)
Query: 25 KNNKMGPEAVAMKAKA------QKTNTNPFESIWSRRKFEVMGQKRKGDTKRMGLARSLA 78
K K+G A A A K N+NPFE +R+KF+++G+K + D G++R+ A
Sbjct: 3 KAKKVGARRKASGAPAGARGGPAKANSNPFEVKVNRQKFQILGRKTRHDVGLPGVSRARA 62
Query: 79 VEKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRERQLNVKSSKKSKYH 138
+ KR +TLLKEY++ KS+ F D+R GE + + K + R E+Q + KKS Y+
Sbjct: 63 LRKRTQTLLKEYKERDKSNVFRDKRFGEYNSNMSPEEKMMKRFALEQQRH--HEKKSIYN 120
Query: 139 LSEEDDDEFEGIDGLGRDDFEDEMLGEDDDETD 171
L+E+++ G L + ++++ D D D
Sbjct: 121 LNEDEELTHYG-QSLADIEKHNDIVDSDSDAED 152
>UniRef100_P78316 Probable nucleolar complex protein 14 [Homo sapiens]
Length = 857
Score = 78.2 bits (191), Expect = 1e-12
Identities = 51/153 (33%), Positives = 84/153 (54%), Gaps = 9/153 (5%)
Query: 25 KNNKMGPEAVAMKAKA------QKTNTNPFESIWSRRKFEVMGQKRKGDTKRMGLARSLA 78
K K+G A A A K N+NPFE +R+KF+++G+K + D G++R+ A
Sbjct: 3 KAKKVGARRKASGAPAGARGGPAKANSNPFEVKVNRQKFQILGRKTRHDVGLPGVSRARA 62
Query: 79 VEKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRERQLNVKSSKKSKYH 138
+ KR +TLLKEY++ KS+ F D+R GE + + K + R E+Q + KKS Y+
Sbjct: 63 LRKRTQTLLKEYKERDKSNVFRDKRFGEYNSNMSPEEKMMKRFALEQQRH--HEKKSIYN 120
Query: 139 LSEEDDDEFEGIDGLGRDDFEDEMLGEDDDETD 171
L+E+++ G L + ++++ D D D
Sbjct: 121 LNEDEELTHYG-QSLADIEKHNDIVDSDSDAED 152
>UniRef100_Q6GPZ2 MGC82495 protein [Xenopus laevis]
Length = 180
Score = 75.9 bits (185), Expect = 5e-12
Identities = 53/151 (35%), Positives = 83/151 (54%), Gaps = 9/151 (5%)
Query: 24 KKNNKMGPEAVAMKAKAQKTNT----NPFESIWSRRKFEVMGQKR-KGDTKRMGLARSLA 78
KK K+ A A + AQK T NPFE +R+KF V+G+K K D G++++ A
Sbjct: 7 KKKGKV--PAAAARVVAQKARTEIRDNPFEVKINRQKFNVLGRKTGKHDVGLPGVSKTKA 64
Query: 79 VEKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRERQLNVKSSKKSKYH 138
+++RK+T+L EY++ K S F+D+R GE D L K + R ERQ + + KK+ Y+
Sbjct: 65 LKRRKETILVEYQRRGKVSVFMDKRFGEYDTKLTPDEKMMKRFAMERQRS--TEKKNLYN 122
Query: 139 LSEEDDDEFEGIDGLGRDDFEDEMLGEDDDE 169
L+EE++ G + D + + D E
Sbjct: 123 LNEEEELTHYGQSLASLEKLNDAVESDSDSE 153
>UniRef100_Q6P333 LOC394902 protein [Xenopus tropicalis]
Length = 432
Score = 74.7 bits (182), Expect = 1e-11
Identities = 56/159 (35%), Positives = 85/159 (53%), Gaps = 14/159 (8%)
Query: 18 KSTSKKKKNNKMGPEAVAMKAK--AQKTNT----NPFESIWSRRKFEVMGQKR-KGDTKR 70
KS KK K P A A A+ AQK T NPFE +++KF V+G+K K D
Sbjct: 3 KSAGKKGK-----PPAAAAAARVIAQKARTEIRDNPFEVKINKQKFNVLGRKTGKHDVGL 57
Query: 71 MGLARSLAVEKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRERQLNVK 130
G+++S A+++RK T+L EY++ K S F+D+R GE D L K + R ER+ N
Sbjct: 58 PGVSKSKALKRRKDTILVEYQRRGKVSVFMDKRFGEYDTKLTPDEKMMKRFAMERKKN-- 115
Query: 131 SSKKSKYHLSEEDDDEFEGIDGLGRDDFEDEMLGEDDDE 169
+ +K+ Y+L+EE++ G + D + + D E
Sbjct: 116 AERKTLYNLNEEEELTHYGQSLANLEKLNDTVDSDSDSE 154
>UniRef100_UPI000042FEFA UPI000042FEFA UniRef100 entry
Length = 933
Score = 74.3 bits (181), Expect = 2e-11
Identities = 61/205 (29%), Positives = 92/205 (44%), Gaps = 28/205 (13%)
Query: 7 RSIANGTNTSNKSTSKKKKN--NKMGPEAVAMKAKAQKT-----NTNPFESIWSRRKFEV 59
++ N S K+ SKK K K G K +K N N F+ +R K +V
Sbjct: 10 KAALNTAGLSRKTHSKKDKKAYKKGGARETDRAKKVEKLEEIRKNLNKFDERETRVKHDV 69
Query: 60 MGQKRKGDTKRMGLARSLAVEKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAIL 119
G+ KG R ++ +E+RKKTLL E++ F DRR GEND + + +
Sbjct: 70 GGRNLKGVVGRPSASKQAGLEQRKKTLLPEHQLRDHRGTFRDRRFGENDPSMSIEDRMLE 129
Query: 120 RSQRERQLNVKSSKKSKYHLSEEDDDE----------FEGIDGLGR-------DDFEDEM 162
R RERQ KK ++L +ED+DE G+ GR DDF +
Sbjct: 130 RYTRERQRG--QGKKGMFNLEDEDEDEPFGELDDGFALGGLTHGGRSVMDLPGDDFVAQG 187
Query: 163 LGEDDDETDET*NQEGSDERSYCKK 187
L ++D++ +E N+ G ++ K
Sbjct: 188 LADEDEDEEE--NRAGRIDKRVVSK 210
>UniRef100_Q9N3Q9 Hypothetical protein Y48G1A.4 [Caenorhabditis elegans]
Length = 813
Score = 73.2 bits (178), Expect = 3e-11
Identities = 41/138 (29%), Positives = 79/138 (56%), Gaps = 8/138 (5%)
Query: 39 KAQKTNTNPFESIWSRRKFEVMGQKRKGDTKRMGLARSLAVEKRKKTLLKEYEQSTKSSE 98
K ++ TNPFE +++ K +++G+K+ +R A E+R++TL EY++ K S+
Sbjct: 6 KQKQRKTNPFELKFNKSKHDILGRKKGAQVGAPTASRKRAHEQREQTLGVEYDRKNKISK 65
Query: 99 FIDRRIGENDEGLDDFGKAILRSQRERQLNVKSSKKSKYHLSEEDDDEFEGIDGLGR--- 155
+D+R+GE D G + K +R ER N K + SK++L+++ D+E E + G+
Sbjct: 66 IVDKRLGEKD-GKSEEEKGAMRFTEERVKNYK--RASKFNLTDDGDEEEEVLTHKGKALS 122
Query: 156 --DDFEDEMLGEDDDETD 171
+ ++ M+ + DD+ +
Sbjct: 123 DIEKYDKSMISDSDDDEE 140
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.377 0.171 0.654
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,115,474,675
Number of Sequences: 2790947
Number of extensions: 37909461
Number of successful extensions: 443728
Number of sequences better than 10.0: 668
Number of HSP's better than 10.0 without gapping: 77
Number of HSP's successfully gapped in prelim test: 636
Number of HSP's that attempted gapping in prelim test: 434134
Number of HSP's gapped (non-prelim): 4168
length of query: 930
length of database: 848,049,833
effective HSP length: 137
effective length of query: 793
effective length of database: 465,690,094
effective search space: 369292244542
effective search space used: 369292244542
T: 11
A: 40
X1: 13 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 35 (21.6 bits)
S2: 80 (35.4 bits)
Medicago: description of AC142096.6


