

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142096.4 - phase: 0 /pseudo
(213 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
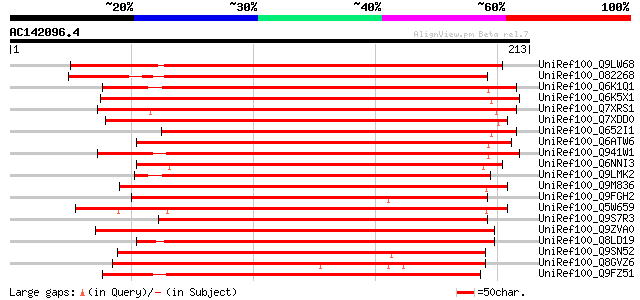
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LW68 Gb|AAC63835.1 [Arabidopsis thaliana] 295 5e-79
UniRef100_O82268 Expressed protein [Arabidopsis thaliana] 280 1e-74
UniRef100_Q6K1Q1 Hypothetical protein OSJNOa148N02.6 [Oryza sativa] 277 1e-73
UniRef100_Q6K5X1 Hypothetical protein P0016F11.6 [Oryza sativa] 277 2e-73
UniRef100_Q7XRS1 OSJNBb0072M01.12 protein [Oryza sativa] 274 1e-72
UniRef100_Q7XDD0 Hypothetical protein [Oryza sativa] 270 2e-71
UniRef100_Q652I1 Hypothetical protein OSJNBa0032M14.22 [Oryza sa... 268 5e-71
UniRef100_Q6ATW6 Hypothetical protein P0015C02.10 [Oryza sativa] 254 1e-66
UniRef100_Q941W1 Hypothetical protein B1088C09.26 [Oryza sativa] 249 3e-65
UniRef100_Q6NNI3 At5g28490 [Arabidopsis thaliana] 249 3e-65
UniRef100_Q9LMK2 F10K1.20 protein [Arabidopsis thaliana] 249 3e-65
UniRef100_Q9M836 T27C4.16 protein [Arabidopsis thaliana] 243 2e-63
UniRef100_Q9FGH2 Gb|AAC63835.1 [Arabidopsis thaliana] 242 4e-63
UniRef100_Q5W659 Hypothetical protein OSJNBb0052F16.2 [Oryza sat... 235 7e-61
UniRef100_Q9S7R3 177 protein [Arabidopsis thaliana] 226 3e-58
UniRef100_Q9ZVA0 F9K20.14 protein [Arabidopsis thaliana] 204 2e-51
UniRef100_Q8LD19 Hypothetical protein [Arabidopsis thaliana] 202 5e-51
UniRef100_Q9SN52 Hypothetical protein F28A21.20 [Arabidopsis tha... 196 3e-49
UniRef100_Q8GVZ6 Hypothetical protein OJ1417_E01.118 [Oryza sativa] 194 1e-48
UniRef100_Q9FZ51 F6I1.8 protein [Arabidopsis thaliana] 183 3e-45
>UniRef100_Q9LW68 Gb|AAC63835.1 [Arabidopsis thaliana]
Length = 195
Score = 295 bits (755), Expect = 5e-79
Identities = 142/177 (80%), Positives = 156/177 (87%), Gaps = 2/177 (1%)
Query: 26 TNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPSRYENQKRRDWNTFGQYLRNHR 85
TN S + + I +++TT+ +SSS++++S S T+ SRYENQKRRDWNTFGQYLRNHR
Sbjct: 11 TNMSHNTNLMIAAAATTTTTSSSSSSSSGGSGTNQL--SRYENQKRRDWNTFGQYLRNHR 68
Query: 86 PPLSLSRCSGAHVLEFLRYLDQFGKTKVHTLICPFYGHPNPPASCPCPLRQAWGSLDALI 145
PPLSLSRCSGAHVLEFLRYLDQFGKTKVHT +CPF+GHPNPPA C CPLRQAWGSLDALI
Sbjct: 69 PPLSLSRCSGAHVLEFLRYLDQFGKTKVHTHLCPFFGHPNPPAPCACPLRQAWGSLDALI 128
Query: 146 GRLRAAFEENGGKPEANPFGARAVRLYLREVRDSQAKARGISYEKKKRKRPPQPPPP 202
GRLRAAFEENGG PE NPFGARAVRLYLREVRDSQAKARGISYEKKKRKRPP P PP
Sbjct: 129 GRLRAAFEENGGSPETNPFGARAVRLYLREVRDSQAKARGISYEKKKRKRPPPPLPP 185
>UniRef100_O82268 Expressed protein [Arabidopsis thaliana]
Length = 219
Score = 280 bits (717), Expect = 1e-74
Identities = 140/172 (81%), Positives = 146/172 (84%), Gaps = 9/172 (5%)
Query: 25 ITNPSSSMTMTIPSSSTTSASSSSTATTSPPSTTSTTTPSRYENQKRRDWNTFGQYLRNH 84
I+N SSS+T +T SSSS+ PS S SRYENQKRRDWNTFGQYLRNH
Sbjct: 23 ISNNSSSVTGATGGEATQPLSSSSS-----PSANS----SRYENQKRRDWNTFGQYLRNH 73
Query: 85 RPPLSLSRCSGAHVLEFLRYLDQFGKTKVHTLICPFYGHPNPPASCPCPLRQAWGSLDAL 144
RPPLSLSRCSGAHVLEFLRYLDQFGKTKVHT IC FYGHPNPPA CPCPLRQAWGSLDAL
Sbjct: 74 RPPLSLSRCSGAHVLEFLRYLDQFGKTKVHTNICHFYGHPNPPAPCPCPLRQAWGSLDAL 133
Query: 145 IGRLRAAFEENGGKPEANPFGARAVRLYLREVRDSQAKARGISYEKKKRKRP 196
IGRLRAAFEENGGKPE NPFGARAVRLYLREVRD Q+KARG+SYEKKKRKRP
Sbjct: 134 IGRLRAAFEENGGKPETNPFGARAVRLYLREVRDMQSKARGVSYEKKKRKRP 185
>UniRef100_Q6K1Q1 Hypothetical protein OSJNOa148N02.6 [Oryza sativa]
Length = 253
Score = 277 bits (709), Expect = 1e-73
Identities = 133/184 (72%), Positives = 146/184 (79%), Gaps = 19/184 (10%)
Query: 39 SSTTSASSSSTATTSPPSTTSTTTPSRYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHV 98
SS+ ASSS+ + P T PSRYE QKRRDWNTFGQYLRNHRPPL L++CSGAHV
Sbjct: 73 SSSAGASSSAGGGAATPQT-----PSRYEAQKRRDWNTFGQYLRNHRPPLGLAQCSGAHV 127
Query: 99 LEFLRYLDQFGKTKVHTLICPFYGHPNPPASCPCPLRQAWGSLDALIGRLRAAFEENGGK 158
LEFLRYLDQFGKTKVHT CPF+GHPNPPA CPCPLRQAWGSLDAL+GRLRAAFEENGG+
Sbjct: 128 LEFLRYLDQFGKTKVHTAACPFFGHPNPPAPCPCPLRQAWGSLDALVGRLRAAFEENGGR 187
Query: 159 PEANPFGARAVRLYLREVRDSQAKARGISYEKKKRKR--------------PPQPPPPPP 204
PE+NPF RAVRLYLREVR+ QA+ARG+SYEKKKRK+ PP PPPPPP
Sbjct: 188 PESNPFAVRAVRLYLREVREHQARARGVSYEKKKRKKPQPADTSGGGGHPHPPPPPPPPP 247
Query: 205 SNNA 208
S A
Sbjct: 248 SAGA 251
>UniRef100_Q6K5X1 Hypothetical protein P0016F11.6 [Oryza sativa]
Length = 248
Score = 277 bits (708), Expect = 2e-73
Identities = 131/182 (71%), Positives = 151/182 (81%), Gaps = 10/182 (5%)
Query: 38 SSSTTSASSSSTATTSPPSTTSTTTPSRYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAH 97
SSS+ + +S+ P ++S+ + SRYE+QKRRDWNTFGQYLRNHRPPLSLSRCSGAH
Sbjct: 53 SSSSGAGTSAGGGGGGPSPSSSSPSLSRYESQKRRDWNTFGQYLRNHRPPLSLSRCSGAH 112
Query: 98 VLEFLRYLDQFGKTKVHTLICPFYGHPNPPASCPCPLRQAWGSLDALIGRLRAAFEENGG 157
VLEFL+Y+DQFGKTKVHT +CPFYGHPNPPA CPCPLRQAWGSLDALIGRLRAA+EENGG
Sbjct: 113 VLEFLKYMDQFGKTKVHTPVCPFYGHPNPPAPCPCPLRQAWGSLDALIGRLRAAYEENGG 172
Query: 158 KPEANPFGARAVRLYLREVRDSQAKARGISYEKKKRKRP----------PQPPPPPPSNN 207
PE NPFGARAVRLYLREVR++QA+ARGISYEKKKRK+P + PPPP +
Sbjct: 173 TPEMNPFGARAVRLYLREVRETQARARGISYEKKKRKKPSSAGAGAGPSSEGSPPPPGGS 232
Query: 208 AT 209
A+
Sbjct: 233 AS 234
>UniRef100_Q7XRS1 OSJNBb0072M01.12 protein [Oryza sativa]
Length = 202
Score = 274 bits (701), Expect = 1e-72
Identities = 131/191 (68%), Positives = 148/191 (76%), Gaps = 19/191 (9%)
Query: 37 PSSSTTSASSSSTATTSPPS--TTSTTTPSRYENQKRRDWNTFGQYLRNHRPPLSLSRCS 94
PS S+++ S PS + +PSRYE QKRRDWNTFGQYLRNHRPPLSL++CS
Sbjct: 11 PSGGGNGGGGGSSSSNSSPSMGAGAPQSPSRYEAQKRRDWNTFGQYLRNHRPPLSLAQCS 70
Query: 95 GAHVLEFLRYLDQFGKTKVHTLICPFYGHPNPPASCPCPLRQAWGSLDALIGRLRAAFEE 154
GAHVLEFLRYLDQFGKTKVHT CPF+GHP+PPA CPCPLRQAWGSLDAL+GRLRAAFEE
Sbjct: 71 GAHVLEFLRYLDQFGKTKVHTAACPFFGHPSPPAPCPCPLRQAWGSLDALVGRLRAAFEE 130
Query: 155 NGGKPEANPFGARAVRLYLREVRDSQAKARGISYEKKKRKRPPQ---------------- 198
NGG+PE+NPF ARAVRLYLREVR+ QA+ARG+SYEKKKRK+P Q
Sbjct: 131 NGGRPESNPFAARAVRLYLREVREHQARARGVSYEKKKRKKPQQQQLQGGDSSGLHGHQH 190
Query: 199 -PPPPPPSNNA 208
PPPPPP+ A
Sbjct: 191 HPPPPPPAGAA 201
>UniRef100_Q7XDD0 Hypothetical protein [Oryza sativa]
Length = 204
Score = 270 bits (690), Expect = 2e-71
Identities = 127/186 (68%), Positives = 143/186 (76%), Gaps = 21/186 (11%)
Query: 40 STTSASSSSTATTSPPSTTSTTTPSRYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHVL 99
S + AT + ++ + +PSRYE+QKRRDWNTFGQYLRNHRPPLSL+RCSGAHVL
Sbjct: 15 SDSGGGGGGMATGATSASAAGASPSRYESQKRRDWNTFGQYLRNHRPPLSLARCSGAHVL 74
Query: 100 EFLRYLDQFGKTKVHTLICPFYGHPNPPASCPCPLRQAWGSLDALIGRLRAAFEENGGKP 159
EFLRYLDQFGKTKVH CPF+GHP PPA CPCPLRQAWGSLDAL+GRLRAA+EENGG+P
Sbjct: 75 EFLRYLDQFGKTKVHAPACPFFGHPAPPAPCPCPLRQAWGSLDALVGRLRAAYEENGGRP 134
Query: 160 EANPFGARAVRLYLREVRDSQAKARGISYEKKKRKRPPQP-------------------- 199
E NPFGARAVRLYLREVR+ QA+ARG+SYEKKKRK+PP P
Sbjct: 135 ENNPFGARAVRLYLREVREHQARARGVSYEKKKRKKPPHPSSAAAAHDDAANGALHHHHH 194
Query: 200 -PPPPP 204
PPPPP
Sbjct: 195 MPPPPP 200
>UniRef100_Q652I1 Hypothetical protein OSJNBa0032M14.22 [Oryza sativa]
Length = 277
Score = 268 bits (686), Expect = 5e-71
Identities = 127/171 (74%), Positives = 138/171 (80%), Gaps = 25/171 (14%)
Query: 63 PSRYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHVLEFLRYLDQFGKTKVHTLICPFYG 122
PSRYE+QKRRDW+TFGQYLRNHRPPL LSRCSGAHVLEFLRYLDQFGKTKVH CPF+G
Sbjct: 20 PSRYESQKRRDWHTFGQYLRNHRPPLELSRCSGAHVLEFLRYLDQFGKTKVHAAGCPFFG 79
Query: 123 HPNPPASCPCPLRQAWGSLDALIGRLRAAFEENGGKPEANPFGARAVRLYLREVRDSQAK 182
HP+PPA CPCPLRQAWGSLDAL+GRLRAAFEE+GG+PEANPFGARAVRLYLREVRDSQAK
Sbjct: 80 HPSPPAPCPCPLRQAWGSLDALVGRLRAAFEEHGGRPEANPFGARAVRLYLREVRDSQAK 139
Query: 183 ARGISYEKKKRKRP-------------------------PQPPPPPPSNNA 208
ARGI+YEKK+RKRP P PPPPPP+ +
Sbjct: 140 ARGIAYEKKRRKRPPTSSSSSQAAAAAAAATSPASPAASPTPPPPPPTERS 190
>UniRef100_Q6ATW6 Hypothetical protein P0015C02.10 [Oryza sativa]
Length = 238
Score = 254 bits (648), Expect = 1e-66
Identities = 118/159 (74%), Positives = 134/159 (84%), Gaps = 5/159 (3%)
Query: 53 SPPSTTSTTTPSRYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHVLEFLRYLDQFGKTK 112
+PP+ SRYE+QKRRDWNTF QYL+NHRPPL+L+RCSGAHV+EFL+YLDQFGKTK
Sbjct: 18 APPAMQPMQQLSRYESQKRRDWNTFLQYLKNHRPPLTLARCSGAHVIEFLKYLDQFGKTK 77
Query: 113 VHTLICPFYGHPNPPASCPCPLRQAWGSLDALIGRLRAAFEENGGKPEANPFGARAVRLY 172
VH C +YG P+PPA CPCPLRQAWGSLDALIGRLRAA+EE+G PE+NPF ARAVR+Y
Sbjct: 78 VHASGCAYYGQPSPPAPCPCPLRQAWGSLDALIGRLRAAYEESGHAPESNPFAARAVRIY 137
Query: 173 LREVRDSQAKARGISYEKKKRKR-----PPQPPPPPPSN 206
LREVRD+QAKARGI YEKKKRKR PP PPPPPP +
Sbjct: 138 LREVRDAQAKARGIPYEKKKRKRTQQQQPPPPPPPPPQH 176
>UniRef100_Q941W1 Hypothetical protein B1088C09.26 [Oryza sativa]
Length = 212
Score = 249 bits (637), Expect = 3e-65
Identities = 122/175 (69%), Positives = 138/175 (78%), Gaps = 7/175 (4%)
Query: 37 PSSSTTSASSSSTATTSPPSTTSTTTPSRYENQKRRDWNTFGQYLRNHRPPLSLSRCSGA 96
PSS+ + + A PP+ S RYE+QKRRDWNTF QYLRNHRPPL+L+RCSGA
Sbjct: 8 PSSAAAGGAPAVAAAPQPPAQLS-----RYESQKRRDWNTFLQYLRNHRPPLTLARCSGA 62
Query: 97 HVLEFLRYLDQFGKTKVHTLICPFYGHPNPPASCPCPLRQAWGSLDALIGRLRAAFEENG 156
HV+EFLRYLDQFGKTKVH C FYG P+PP CPCPLRQAWGSLDALIGRLRAA+EE+G
Sbjct: 63 HVIEFLRYLDQFGKTKVHASGCAFYGQPSPPGPCPCPLRQAWGSLDALIGRLRAAYEESG 122
Query: 157 GKPEANPFGARAVRLYLREVRDSQAKARGISYEKKKRKR--PPQPPPPPPSNNAT 209
G PE+NPF ARAVR+YLREVRDSQAKARGI YEKKKRKR QP PS +++
Sbjct: 123 GTPESNPFAARAVRIYLREVRDSQAKARGIPYEKKKRKRSQAAQPAGVEPSGSSS 177
>UniRef100_Q6NNI3 At5g28490 [Arabidopsis thaliana]
Length = 190
Score = 249 bits (637), Expect = 3e-65
Identities = 121/155 (78%), Positives = 132/155 (85%), Gaps = 5/155 (3%)
Query: 53 SPPSTTSTTTPS--RYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHVLEFLRYLDQFGK 110
+P S+T T PS RYENQKRRDWNTF QYLRNHRPPLSL CSGAHVLEFLRYLDQFGK
Sbjct: 11 NPNSSTQLTPPSSSRYENQKRRDWNTFCQYLRNHRPPLSLPSCSGAHVLEFLRYLDQFGK 70
Query: 111 TKVHTLICPFYGHPNPPASCPCPLRQAWGSLDALIGRLRAAFEENGGKPEANPFGARAVR 170
TKVH C F+G PNPPA CPCPLRQAWGSLDALIGRLRAA+EENGG PEANPFG+RAVR
Sbjct: 71 TKVHHQNCAFFGLPNPPAPCPCPLRQAWGSLDALIGRLRAAYEENGGPPEANPFGSRAVR 130
Query: 171 LYLREVRDSQAKARGISYEKKKR---KRPPQPPPP 202
L+LREVRD QAKARG+SYEKK++ ++ PQ PP
Sbjct: 131 LFLREVRDFQAKARGVSYEKKRKRVNRQKPQTQPP 165
>UniRef100_Q9LMK2 F10K1.20 protein [Arabidopsis thaliana]
Length = 196
Score = 249 bits (636), Expect = 3e-65
Identities = 113/146 (77%), Positives = 131/146 (89%), Gaps = 5/146 (3%)
Query: 52 TSPPSTTSTTTPSRYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHVLEFLRYLDQFGKT 111
+SPP+T PSRYE+QKRRDWNTF QYL+NH+PPL+LSRCSGAHV+EFL+YLDQFGKT
Sbjct: 23 SSPPAT-----PSRYESQKRRDWNTFLQYLKNHKPPLALSRCSGAHVIEFLKYLDQFGKT 77
Query: 112 KVHTLICPFYGHPNPPASCPCPLRQAWGSLDALIGRLRAAFEENGGKPEANPFGARAVRL 171
KVH CP++GH PP+ C CPL+QAWGSLDALIGRLRAA+EENGG+P++NPF ARAVR+
Sbjct: 78 KVHVAACPYFGHQQPPSPCSCPLKQAWGSLDALIGRLRAAYEENGGRPDSNPFAARAVRI 137
Query: 172 YLREVRDSQAKARGISYEKKKRKRPP 197
YLREVR+SQAKARGI YEKKKRKRPP
Sbjct: 138 YLREVRESQAKARGIPYEKKKRKRPP 163
>UniRef100_Q9M836 T27C4.16 protein [Arabidopsis thaliana]
Length = 201
Score = 243 bits (620), Expect = 2e-63
Identities = 115/167 (68%), Positives = 135/167 (79%), Gaps = 8/167 (4%)
Query: 46 SSSTATTSPPSTTSTTTPSRYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHVLEFLRYL 105
++S +T +P S +S + SRYENQKRRDWNTF QYLRNH PPLSL+ CSGAHVL+FLRYL
Sbjct: 14 NTSLSTQTPSSFSSPPSSSRYENQKRRDWNTFCQYLRNHHPPLSLASCSGAHVLDFLRYL 73
Query: 106 DQFGKTKVHTLICPFYGHPNPPASCPCPLRQAWGSLDALIGRLRAAFEENGGKPEANPFG 165
DQFGKTKVH C F+G PNPPA CPCPLRQAWGSLDALIGRLRAA+EENGG PE +PFG
Sbjct: 74 DQFGKTKVHHQNCAFFGLPNPPAPCPCPLRQAWGSLDALIGRLRAAYEENGGAPETSPFG 133
Query: 166 ARAVRLYLREVRDSQAKARGISYEKKKRK--------RPPQPPPPPP 204
+R+VR++LREVRD QAK+RG+SYEKK+++ PQ PP P
Sbjct: 134 SRSVRIFLREVRDFQAKSRGVSYEKKRKRVNNKQITQSQPQSQPPLP 180
>UniRef100_Q9FGH2 Gb|AAC63835.1 [Arabidopsis thaliana]
Length = 182
Score = 242 bits (618), Expect = 4e-63
Identities = 111/147 (75%), Positives = 130/147 (87%), Gaps = 1/147 (0%)
Query: 51 TTSPPSTTSTTTPSRYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHVLEFLRYLDQFGK 110
T + + +S+++PSRYE+QKRRDWNTF QYLRNH+PPL+LSRCSGAHVLEFL+YLDQFGK
Sbjct: 5 TAAKAAASSSSSPSRYESQKRRDWNTFLQYLRNHKPPLNLSRCSGAHVLEFLKYLDQFGK 64
Query: 111 TKVHTLICPFYGHPNPPASCPCPLRQAWGSLDALIGRLRAAFEE-NGGKPEANPFGARAV 169
TKVH CPF+G PNPP+ C CPL+QAWGSLDALIGRLRAAFEE GG PE+NPF A+AV
Sbjct: 65 TKVHATACPFFGQPNPPSQCTCPLKQAWGSLDALIGRLRAAFEEIGGGLPESNPFAAKAV 124
Query: 170 RLYLREVRDSQAKARGISYEKKKRKRP 196
R+YL+EVR +QAKARGI Y+KKKRKRP
Sbjct: 125 RIYLKEVRQTQAKARGIPYDKKKRKRP 151
>UniRef100_Q5W659 Hypothetical protein OSJNBb0052F16.2 [Oryza sativa]
Length = 284
Score = 235 bits (599), Expect = 7e-61
Identities = 116/195 (59%), Positives = 141/195 (71%), Gaps = 18/195 (9%)
Query: 28 PSSSMTMTIPSSSTTS------ASSSSTATTSPPSTTSTTTP-----SRYENQKRRDWNT 76
PSSS + P++S+ + + + P + P SRYE+QKRRDWNT
Sbjct: 19 PSSSASAPAPAASSNEEEGRHQSQAQQQVQEAQPQPLAQQAPAAAGLSRYESQKRRDWNT 78
Query: 77 FGQYLRNHRPPLSLSRCSGAHVLEFLRYLDQFGKTKVHTLICPFYGHPNPPASCPCPLRQ 136
F QYLRNH+PPL+L RCSGAHV+EFL+YLDQFGKTKVH C ++G PNPPA C CPLRQ
Sbjct: 79 FLQYLRNHKPPLTLPRCSGAHVIEFLKYLDQFGKTKVHADGCAYFGEPNPPAPCACPLRQ 138
Query: 137 AWGSLDALIGRLRAAFEENGGKPEANPFGARAVRLYLREVRDSQAKARGISYEKKKRK-- 194
AWGSLDALIGRLRAA+EE+GG+PE+NPF ARAVR+YLREVR++QAKARGI YEKK+++
Sbjct: 139 AWGSLDALIGRLRAAYEESGGRPESNPFAARAVRIYLREVREAQAKARGIPYEKKRKRGA 198
Query: 195 -----RPPQPPPPPP 204
PP PPP
Sbjct: 199 AAAAAAPPVVVAPPP 213
>UniRef100_Q9S7R3 177 protein [Arabidopsis thaliana]
Length = 177
Score = 226 bits (576), Expect = 3e-58
Identities = 103/135 (76%), Positives = 116/135 (85%)
Query: 62 TPSRYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHVLEFLRYLDQFGKTKVHTLICPFY 121
TPSRYE+QKRRDWNTFGQYL+N RPP+ +S CS HVL+FLRYLDQFGKTKVH C FY
Sbjct: 22 TPSRYESQKRRDWNTFGQYLKNQRPPVPMSHCSCNHVLDFLRYLDQFGKTKVHVPGCMFY 81
Query: 122 GHPNPPASCPCPLRQAWGSLDALIGRLRAAFEENGGKPEANPFGARAVRLYLREVRDSQA 181
G P PPA C CPLRQAWGSLDALIGRLRAA+EENGG PE NPF + A+R+YLREVR+ QA
Sbjct: 82 GQPEPPAPCTCPLRQAWGSLDALIGRLRAAYEENGGPPETNPFASGAIRVYLREVRECQA 141
Query: 182 KARGISYEKKKRKRP 196
KARGI Y+KKK+K+P
Sbjct: 142 KARGIPYKKKKKKKP 156
>UniRef100_Q9ZVA0 F9K20.14 protein [Arabidopsis thaliana]
Length = 195
Score = 204 bits (518), Expect = 2e-51
Identities = 95/164 (57%), Positives = 116/164 (69%)
Query: 36 IPSSSTTSASSSSTATTSPPSTTSTTTPSRYENQKRRDWNTFGQYLRNHRPPLSLSRCSG 95
I S+ S P S + SRYE+QKRRDWNTF QYLRN +PP+ +S+C
Sbjct: 11 IAEGSSQPQSQPQPQPHQPQSPPNPPALSRYESQKRRDWNTFCQYLRNQQPPVHISQCGS 70
Query: 96 AHVLEFLRYLDQFGKTKVHTLICPFYGHPNPPASCPCPLRQAWGSLDALIGRLRAAFEEN 155
H+L+FL+YLDQFGKTKVH C F+G P C CPL+QAWGSLDALIGRLRAAFEEN
Sbjct: 71 NHILDFLQYLDQFGKTKVHIHGCVFFGQVEPAGQCNCPLKQAWGSLDALIGRLRAAFEEN 130
Query: 156 GGKPEANPFGARAVRLYLREVRDSQAKARGISYEKKKRKRPPQP 199
GG PE NPF +R++LREVRDSQAKARG+ Y+K+K+++ P
Sbjct: 131 GGLPERNPFAGGGIRVFLREVRDSQAKARGVPYKKRKKRKKRNP 174
>UniRef100_Q8LD19 Hypothetical protein [Arabidopsis thaliana]
Length = 195
Score = 202 bits (514), Expect = 5e-51
Identities = 93/147 (63%), Positives = 114/147 (77%), Gaps = 3/147 (2%)
Query: 53 SPPSTTSTTTPSRYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHVLEFLRYLDQFGKTK 112
SPP+ + SRYE+QKRRDWNTF QYLRN +PP+ +S+C H+L+FL+YLDQFGKTK
Sbjct: 31 SPPNPPAL---SRYESQKRRDWNTFCQYLRNQQPPVHISQCGSNHILDFLQYLDQFGKTK 87
Query: 113 VHTLICPFYGHPNPPASCPCPLRQAWGSLDALIGRLRAAFEENGGKPEANPFGARAVRLY 172
VH C F+G P C CPL+QAWGSLDALIGRLRAAFEENGG PE NPF +R++
Sbjct: 88 VHIHGCVFFGQVEPAGQCNCPLKQAWGSLDALIGRLRAAFEENGGLPERNPFAGGGIRVF 147
Query: 173 LREVRDSQAKARGISYEKKKRKRPPQP 199
LREVRDSQAKARG+ Y+K+K+++ P
Sbjct: 148 LREVRDSQAKARGVPYKKRKKRKKRNP 174
>UniRef100_Q9SN52 Hypothetical protein F28A21.20 [Arabidopsis thaliana]
Length = 191
Score = 196 bits (498), Expect = 3e-49
Identities = 95/152 (62%), Positives = 113/152 (73%), Gaps = 1/152 (0%)
Query: 45 SSSSTATTSPPSTTSTTTPSRYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHVLEFLRY 104
SSSS P SRYE+QKRRDWNTF QYL++ PPL +S+ HVL FLRY
Sbjct: 17 SSSSNHHKQPLPPQPQQPLSRYESQKRRDWNTFVQYLKSQNPPLMMSQFDYTHVLSFLRY 76
Query: 105 LDQFGKTKVHTLICPFYGHPNPPASCPCPLRQAWGSLDALIGRLRAAFEEN-GGKPEANP 163
LDQFGKTKVH C F+G P+PP C CPL+QAWGSLDALIGRLRAA+EE+ GG P+ NP
Sbjct: 77 LDQFGKTKVHHQACVFFGQPDPPGPCTCPLKQAWGSLDALIGRLRAAYEEHGGGSPDTNP 136
Query: 164 FGARAVRLYLREVRDSQAKARGISYEKKKRKR 195
F ++R++LREVR+SQAKARGI Y KKKR++
Sbjct: 137 FANGSIRVHLREVRESQAKARGIPYRKKKRRK 168
>UniRef100_Q8GVZ6 Hypothetical protein OJ1417_E01.118 [Oryza sativa]
Length = 276
Score = 194 bits (493), Expect = 1e-48
Identities = 103/185 (55%), Positives = 123/185 (65%), Gaps = 32/185 (17%)
Query: 43 SASSSSTATTSPPSTTSTTTPSRYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHVLEFL 102
S+SS++ + + + PSRYE+QKRRDW TF QYL HRPPL L RCSGAHVLEFL
Sbjct: 2 SSSSAAALGSDDGCSPAELRPSRYESQKRRDWQTFTQYLAAHRPPLELRRCSGAHVLEFL 61
Query: 103 RYLDQFGKTKVHTLICPFYGHPNP---------PASCPCPLRQAWGSLDALIGRLRAAFE 153
RYLD+FGKT+VH CP YG +P A+C CPLRQAWGSLDAL+GRLRAA++
Sbjct: 62 RYLDRFGKTRVHEPPCPSYGGRSPSAAGPVAAAAAACQCPLRQAWGSLDALVGRLRAAYD 121
Query: 154 E---NGGKPE--------------------ANPFGARAVRLYLREVRDSQAKARGISYEK 190
E G+P+ ANPF ARAVRLYLR+VRD+QA ARGISY K
Sbjct: 122 ERHGRAGEPDAVAGAGAVATDSTSSSSAAAANPFAARAVRLYLRDVRDAQAMARGISYHK 181
Query: 191 KKRKR 195
KK++R
Sbjct: 182 KKKRR 186
>UniRef100_Q9FZ51 F6I1.8 protein [Arabidopsis thaliana]
Length = 164
Score = 183 bits (464), Expect = 3e-45
Identities = 88/155 (56%), Positives = 105/155 (66%), Gaps = 5/155 (3%)
Query: 39 SSTTSASSSSTATTSPPSTTSTTTPSRYENQKRRDWNTFGQYLRNHRPPLSLSRCSGAHV 98
+ST + + PP T S RYE+QK RDWNTF QYL PP+ + C H+
Sbjct: 2 TSTNTRNKGKCIVEGPPPTLS-----RYESQKSRDWNTFCQYLMTKMPPVHVWECESNHI 56
Query: 99 LEFLRYLDQFGKTKVHTLICPFYGHPNPPASCPCPLRQAWGSLDALIGRLRAAFEENGGK 158
L+FL+ DQFGKTKVH C F+G PP C CPL+QAWGSLDALIGRLRAA+EENGG
Sbjct: 57 LDFLQSRDQFGKTKVHIQGCVFFGQKEPPGECNCPLKQAWGSLDALIGRLRAAYEENGGL 116
Query: 159 PEANPFGARAVRLYLREVRDSQAKARGISYEKKKR 193
E NPF +R++LREVR SQAKARG+ Y+KKKR
Sbjct: 117 TEKNPFARGGIRIFLREVRGSQAKARGVLYKKKKR 151
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.313 0.128 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 405,958,602
Number of Sequences: 2790947
Number of extensions: 17908187
Number of successful extensions: 313257
Number of sequences better than 10.0: 2366
Number of HSP's better than 10.0 without gapping: 1577
Number of HSP's successfully gapped in prelim test: 842
Number of HSP's that attempted gapping in prelim test: 242078
Number of HSP's gapped (non-prelim): 33570
length of query: 213
length of database: 848,049,833
effective HSP length: 122
effective length of query: 91
effective length of database: 507,554,299
effective search space: 46187441209
effective search space used: 46187441209
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 72 (32.3 bits)
Medicago: description of AC142096.4


