

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141865.9 - phase: 0 /pseudo
(547 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
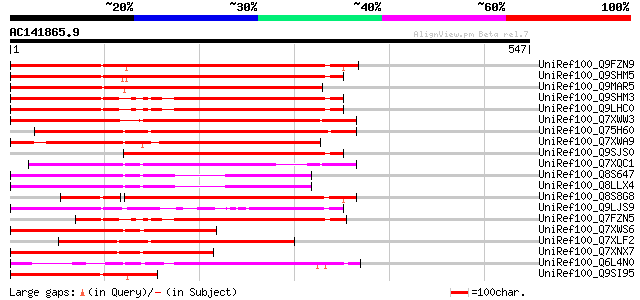
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FZN9 Retroelement pol polyprotein-like [Arabidopsis ... 442 e-122
UniRef100_Q9SHM5 F7F22.15 [Arabidopsis thaliana] 433 e-120
UniRef100_Q9MAR5 F28H19.8 protein [Arabidopsis thaliana] 424 e-117
UniRef100_Q9SHM3 F7F22.17 [Arabidopsis thaliana] 363 7e-99
UniRef100_Q9LHC0 Retroelement pol polyprotein-like [Arabidopsis ... 353 9e-96
UniRef100_Q7XWW3 OSJNBb0058J09.16 protein [Oryza sativa] 330 5e-89
UniRef100_Q75H60 Putative reverse transcriptase [Oryza sativa] 307 6e-82
UniRef100_Q7XWA9 OSJNBa0061A09.17 protein [Oryza sativa] 305 3e-81
UniRef100_Q9SJS0 Putative retroelement pol polyprotein [Arabidop... 279 1e-73
UniRef100_Q7XQC1 OSJNBb0060M15.10 protein [Oryza sativa] 276 9e-73
UniRef100_Q8S647 Putative retroelement [Oryza sativa] 275 2e-72
UniRef100_Q8LLX4 Putative retroelement [Oryza sativa] 275 2e-72
UniRef100_Q8S8G8 Putative retroelement pol polyprotein [Arabidop... 266 2e-69
UniRef100_Q9LJS9 Retroelement pol polyprotein-like [Arabidopsis ... 250 9e-65
UniRef100_Q7FZN5 T14A16.1 protein [Arabidopsis thaliana] 247 7e-64
UniRef100_Q7XWS6 OSJNBa0091C12.6 protein [Oryza sativa] 233 1e-59
UniRef100_Q7XLF2 OSJNBa0013A04.17 protein [Oryza sativa] 226 1e-57
UniRef100_Q7XNX7 OSJNBb0026I12.3 protein [Oryza sativa] 212 3e-53
UniRef100_Q6L4N0 Hypothetical protein P0530H10.3 [Oryza sativa] 201 6e-50
UniRef100_Q9SI95 Putative retroelement pol polyprotein [Arabidop... 188 3e-46
>UniRef100_Q9FZN9 Retroelement pol polyprotein-like [Arabidopsis thaliana]
Length = 1864
Score = 442 bits (1136), Expect = e-122
Identities = 223/389 (57%), Positives = 277/389 (70%), Gaps = 29/389 (7%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
IYYAS+T+D A V YATTEKELLAVV+A +KFR YLVGSK+TVYTDH+ ++++ KKD K
Sbjct: 1299 IYYASRTMDDAQVRYATTEKELLAVVFAFEKFRSYLVGSKVTVYTDHAALRHIYAKKDTK 1358
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLAQTN 120
PRL++ I LLQEFD+EI DKKG+EN VADHLSR+R EDE+P+DDS P++QL + Q N
Sbjct: 1359 PRLLRWILLLQEFDMEIVDKKGIENGVADHLSRMRI--EDEVPIDDSMPEEQLMAIQQLN 1416
Query: 121 A------------------PWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYL 162
PWYAD VN+L +G PP LS +KKKFF D+ H+Y DEPYL
Sbjct: 1417 ESAQIRKSLDQVCTIEEKLPWYADHVNYLVSGEEPPNLSSYEKKKFFKDINHFYWDEPYL 1476
Query: 163 FKRG*DGIFRRCIPENEVSSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLF 222
+ D I+RRC+ E+E+ IL HCH S+YGGH +T K KIL +GFWWPS+FKD F
Sbjct: 1477 YTLCKDKIYRRCVSEDEIEGILLHCHGSAYGGHFATFKTVSKILQAGFWWPSMFKDAQEF 1536
Query: 223 ISKCDKCQRTGNITKRNEMPLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKW 282
ISKCD CQR GNI++RNEMP N ILEVEIFDVW IDF+GPFPSS+ N+YILVAVDYVSKW
Sbjct: 1537 ISKCDSCQRRGNISRRNEMPQNPILEVEIFDVWGIDFMGPFPSSYGNKYILVAVDYVSKW 1596
Query: 283 VEVIASLTNDAQVVIKLFKKNHFSRFGVPRVVISDGGSQNILKNFCKNLE*GTKLQHPIT 342
VE IAS TNDA+VV+KLFK F RFGVPR++ISDGG I K F L+ +H +
Sbjct: 1597 VEAIASPTNDARVVLKLFKTIIFPRFGVPRIMISDGGKHFINKVFENLLK-----KHGVK 1651
Query: 343 HKLVDKWR----SQIDKSKLSLKKLFQPL 367
HK+ + Q++ S +K + + +
Sbjct: 1652 HKVATPYHPQTSGQVEISNREIKAILEKI 1680
>UniRef100_Q9SHM5 F7F22.15 [Arabidopsis thaliana]
Length = 1862
Score = 433 bits (1114), Expect = e-120
Identities = 222/369 (60%), Positives = 268/369 (72%), Gaps = 24/369 (6%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
IYYAS+TLD A YATTEKELLAVV+A +KFR YLVGSK+TVYTDH+ +++L KKD K
Sbjct: 1298 IYYASRTLDDAQGRYATTEKELLAVVFAFEKFRSYLVGSKVTVYTDHAALRHLYAKKDTK 1357
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLA--- 117
PRL++ I LLQEFD+EI DKKG+EN ADHLSR+R E+ L +DDS P++QL ++
Sbjct: 1358 PRLLRWILLLQEFDMEIVDKKGIENGAADHLSRMRI--EEPLLIDDSMPEEQLMVVEFFG 1415
Query: 118 ---------QTNA-----PWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLF 163
Q NA PWYAD VN+LA GV PP L+ ++KKFF D+ HYY DEPYL+
Sbjct: 1416 KSYSGKEFHQLNAVEGESPWYADHVNYLACGVEPPNLTSYERKKFFRDIHHYYWDEPYLY 1475
Query: 164 KRG*DGIFRRCIPENEVSSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFI 223
D I+RRC+ E+EV IL HCH S+YGGH +T K KIL +GFWWP++FKD F+
Sbjct: 1476 TLCKDKIYRRCVSEDEVEGILLHCHGSAYGGHFATFKTVSKILQAGFWWPTMFKDAQEFV 1535
Query: 224 SKCDKCQRTGNITKRNEMPLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWV 283
SKCD CQR GNI++RNEMP N ILEVEIFDVW IDF+GPFPSS+ N+YILVAVDYVSKWV
Sbjct: 1536 SKCDSCQRKGNISRRNEMPQNPILEVEIFDVWGIDFMGPFPSSYGNKYILVAVDYVSKWV 1595
Query: 284 EVIASLTNDAQVVIKLFKKNHFSRFGVPRVVISDGGSQNILKNFCKNLE*GTKLQHPITH 343
E IAS TNDA+VV+KLFK F RFGVPRVVISDGG I K F L+ +H + H
Sbjct: 1596 EAIASPTNDAKVVLKLFKTIIFPRFGVPRVVISDGGKHFINKVFENLLK-----KHGVKH 1650
Query: 344 KLVDKWRSQ 352
K+ + Q
Sbjct: 1651 KVATPYNPQ 1659
>UniRef100_Q9MAR5 F28H19.8 protein [Arabidopsis thaliana]
Length = 640
Score = 424 bits (1089), Expect = e-117
Identities = 209/345 (60%), Positives = 256/345 (73%), Gaps = 18/345 (5%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
IYYAS+TLD YATTEKELL VV+A +KFR YLVGSK+TVYTDH+ +K++ KKD K
Sbjct: 297 IYYASRTLDDTQSRYATTEKELLVVVFAFEKFRSYLVGSKVTVYTDHAALKHIYAKKDTK 356
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLAQT- 119
PRL++ I LQEFD+EI DK+G EN VADHLSR+R T + +P+DDS P++QL + +
Sbjct: 357 PRLLRWILFLQEFDMEIVDKRGSENGVADHLSRMRMT--EAVPIDDSMPEEQLLAVGTSF 414
Query: 120 --------------NAPWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKR 165
PWY D VN+L AG++P ELS QKKKFF D+ HYY DEP L+K+
Sbjct: 415 DKFKIETMCATRSGKLPWYVDLVNYLTAGIVPSELSSYQKKKFFRDINHYYWDEPVLYKK 474
Query: 166 G*DGIFRRCIPENEVSSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISK 225
G DG+FRRCI E EV +L CH+S YGGH +T K K+L +G WWPS+FKD H I++
Sbjct: 475 GVDGLFRRCIAEEEVQGVLELCHNSPYGGHFATFKTVQKVLQAGLWWPSMFKDAHEHITR 534
Query: 226 CDKCQRTGNITKRNEMPLNNILEVEIFDVWVIDFIGPF-PSSFVNQYILVAVDYVSKWVE 284
CD+CQR GNI++RNEMP N ILEVE+FDVW IDF+GPF P+S+ N+YILVAVDYVSKWVE
Sbjct: 535 CDRCQRVGNISRRNEMPQNPILEVEVFDVWGIDFMGPFEPASYGNKYILVAVDYVSKWVE 594
Query: 285 VIASLTNDAQVVIKLFKKNHFSRFGVPRVVISDGGSQNILKNFCK 329
IAS TND++VV+KLFK F RFG+PRVVISDGGS I K F K
Sbjct: 595 AIASPTNDSKVVLKLFKTIIFPRFGIPRVVISDGGSHFINKVFEK 639
>UniRef100_Q9SHM3 F7F22.17 [Arabidopsis thaliana]
Length = 1799
Score = 363 bits (932), Expect = 7e-99
Identities = 196/352 (55%), Positives = 238/352 (66%), Gaps = 39/352 (11%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
IYYAS+TLD A YATTEKELL VV+A +KFR YLVGSK+TVYTDH+ +++L KKD K
Sbjct: 1284 IYYASRTLDDAQGRYATTEKELLVVVFAFEKFRSYLVGSKVTVYTDHAALRHLYAKKDTK 1343
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLAQTN 120
PRL++ I LLQEFD+EI DKKG+EN ADHLSR+R E+ L +DDS P++QL +
Sbjct: 1344 PRLLRWILLLQEFDMEIVDKKGIENGAADHLSRMRI--EEPLLIDDSMPEEQLMV----- 1396
Query: 121 APWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENEV 180
V F SY K+ F+ L + P+ RC+ E+EV
Sbjct: 1397 -------VEFFGK-------SYSGKE--FHQLNAVEGESPW-----------RCVSEDEV 1429
Query: 181 SSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNE 240
IL HCH S+YGGH +T K KIL +GFWWP++FKD F+SKCD CQR GNI++RNE
Sbjct: 1430 EGILLHCHGSAYGGHFATFKTVSKILQAGFWWPTMFKDAQEFVSKCDSCQRKGNISRRNE 1489
Query: 241 MPLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQVVIKLF 300
MP N ILEVEIFDVW IDF+GPFPSS+ N+YILVAVDYVSKWVE IAS TNDA+VV+KLF
Sbjct: 1490 MPQNPILEVEIFDVWGIDFMGPFPSSYGNKYILVAVDYVSKWVEAIASPTNDAKVVLKLF 1549
Query: 301 KKNHFSRFGVPRVVISDGGSQNILKNFCKNLE*GTKLQHPITHKLVDKWRSQ 352
K F RFGVPRVVISDGG I K F L+ +H + HK+ + Q
Sbjct: 1550 KTIIFPRFGVPRVVISDGGKHFINKVFENLLK-----KHGVKHKVATPYNPQ 1596
>UniRef100_Q9LHC0 Retroelement pol polyprotein-like [Arabidopsis thaliana]
Length = 897
Score = 353 bits (905), Expect = 9e-96
Identities = 191/352 (54%), Positives = 235/352 (66%), Gaps = 39/352 (11%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
IYYAS+TLD A YATTEKELLAVV+A +KFR YLVGSK+TVYTDH+ +++L KKD K
Sbjct: 382 IYYASRTLDDAQGRYATTEKELLAVVFAFEKFRSYLVGSKVTVYTDHAALRHLYAKKDTK 441
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLAQTN 120
PRL++ I LLQEFD+EI DKKG+EN ADHL R+R E+ LP+DDS P++QL +
Sbjct: 442 PRLLRWILLLQEFDMEIVDKKGIENGAADHLLRMRI--EEPLPIDDSMPEEQLMV----- 494
Query: 121 APWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENEV 180
V F SY K+ F+ L + P+ RC+ E+EV
Sbjct: 495 -------VEFFGK-------SYSGKE--FHQLNDVEGESPW-----------RCVSEDEV 527
Query: 181 SSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNE 240
IL HCH S+YGGH +T K KIL +GFWWP++FKD F+SKCD CQR NI++ NE
Sbjct: 528 EGILLHCHCSAYGGHFATFKTVSKILQAGFWWPTMFKDAQEFVSKCDSCQRKDNISRINE 587
Query: 241 MPLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQVVIKLF 300
MP N I+EVEIFDVW IDF+G FPSS+ N+YILVA+DYVSKWVE IA TNDA+VV+KLF
Sbjct: 588 MPQNPIVEVEIFDVWGIDFMGLFPSSYGNKYILVAIDYVSKWVEAIAIPTNDAKVVLKLF 647
Query: 301 KKNHFSRFGVPRVVISDGGSQNILKNFCKNLE*GTKLQHPITHKLVDKWRSQ 352
K F RFGVPRVVISDGG I K F L+ +H + HK+ + Q
Sbjct: 648 KTIIFPRFGVPRVVISDGGKHFINKVFENLLK-----KHGVKHKVATPYHPQ 694
>UniRef100_Q7XWW3 OSJNBb0058J09.16 protein [Oryza sativa]
Length = 1045
Score = 330 bits (847), Expect = 5e-89
Identities = 180/365 (49%), Positives = 229/365 (62%), Gaps = 23/365 (6%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
I YASKTL GA +NYATTEKELL VV+AIDKFR YLVG+K+ +YTDH+ +KYLL KKDAK
Sbjct: 602 ICYASKTLTGAQLNYATTEKELLTVVFAIDKFRSYLVGAKVIIYTDHAALKYLLTKKDAK 661
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLAQTN 120
PRL++ I LLQEFD+EIKDKKGVEN VADHLSRL TN E P++D DD L
Sbjct: 662 PRLLRWILLLQEFDIEIKDKKGVENYVADHLSRLHITNMQEQPINDLLRDDMLMTC---- 717
Query: 121 APWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENEV 180
+PP Q+K + +H + DEPYL++ DG+ RRC+P +E
Sbjct: 718 ---------------IPP--GENQRKLKYESHRHIW-DEPYLYRVCSDGLLRRCVPTDEG 759
Query: 181 SSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNE 240
I+ CH+S YGGH + KI SGF+WP++++D FI +C CQR G IT R+
Sbjct: 760 LKIIERCHASPYGGHYGAFRTQAKIWQSGFFWPTMYEDSKEFIRRCTSCQRQGGITARDA 819
Query: 241 MPLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQVVIKLF 300
MPL L+VEIFDVW IDF+GPFP S +YILVAVDYVSKWVE + DA+ K+F
Sbjct: 820 MPLTYNLQVEIFDVWGIDFMGPFPKSRNCEYILVAVDYVSKWVEAMPCSAADARHAKKMF 879
Query: 301 KKNHFSRFGVPRVVISDGGSQNILKNFCKNLE*GTKLQHPITHKLVDKWRSQIDKSKLSL 360
+ F RFG R+VISD G I K F ++L + +H + + Q + S +
Sbjct: 880 TETIFPRFGTRRMVISDRGFHFIDKTF-RDLLREMEAKHNVATPYHPQTSGQAETSNKQI 938
Query: 361 KKLFQ 365
K + Q
Sbjct: 939 KNILQ 943
>UniRef100_Q75H60 Putative reverse transcriptase [Oryza sativa]
Length = 743
Score = 307 bits (786), Expect = 6e-82
Identities = 163/340 (47%), Positives = 216/340 (62%), Gaps = 7/340 (2%)
Query: 27 YAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAKPRLIQLIFLLQEFDLEIKDKKGVENV 86
+A+ FR YLV +K+ +YTDH+ +KY L KKDAKPRL++ I LLQEFDLEIKDKKGVEN
Sbjct: 220 FAVGAFRSYLVRAKVIIYTDHAALKYPLTKKDAKPRLLRWILLLQEFDLEIKDKKGVENS 279
Query: 87 VADHLSRLRETNEDELPLDDSFPDDQLFLLAQTNAPWYADFVNFLAAGVLPPELSYQQKK 146
VADHLSRL+ TN E P++D DD L + +N PWYA+ VN++ +PP + + K
Sbjct: 280 VADHLSRLQITNMPEQPINDFLRDDMLMTVRDSN-PWYANIVNYMVFKYIPPGKNQRNLK 338
Query: 147 KFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENEVSSILTHCHSSSYGGHASTQKASFKIL 206
+ +H + DEPYL++ DG+ RRC+P E I+ CH+S YGGH + KI
Sbjct: 339 --YESRRHIW-DEPYLYRVCSDGLLRRCVPTEEGLKIIERCHASPYGGHYGVFRTQAKIW 395
Query: 207 HSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNEMPLNNILEVEIFDVWVIDFIGPFPSS 266
SGF+WP++++D F+ +C CQR G IT R+ MPL L+VEIFDVW IDF+GPFP S
Sbjct: 396 QSGFFWPTMYEDSKEFVRRCTSCQRQGGITARDAMPLTYNLQVEIFDVWGIDFMGPFPKS 455
Query: 267 FVNQYILVAVDYVSKWVEVIASLTNDAQVVIKLFKKNHFSRFGVPRVVISDGGSQNILKN 326
+YILVAVDYVSKWV+ + DA+ K+F + F +F PR+VISDGGS I K
Sbjct: 456 RNCEYILVAVDYVSKWVKAMPCSATDARHAKKMFIEIIFPKFDTPRMVISDGGSHFIDKT 515
Query: 327 FCKNL-E*GTKLQHPITHKLVDKWRSQIDKSKLSLKKLFQ 365
F L E G K H + + Q + S +K + Q
Sbjct: 516 FRDLLREMGAK--HNVATPYHPQTSGQAETSNKQIKNILQ 553
>UniRef100_Q7XWA9 OSJNBa0061A09.17 protein [Oryza sativa]
Length = 430
Score = 305 bits (780), Expect = 3e-81
Identities = 161/332 (48%), Positives = 212/332 (63%), Gaps = 25/332 (7%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
I YASKTL GA +NY+TTEKELLAVV G+K+ VYTDH+ +KYLL KKDAK
Sbjct: 24 IAYASKTLTGAQLNYSTTEKELLAVV-----------GAKVVVYTDHAALKYLLTKKDAK 72
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLAQTN 120
PRLI+ I +LQEFD+EI +KKG+EN VADHLSR++ TN EL ++D DD L + ++
Sbjct: 73 PRLIRWILILQEFDIEINNKKGIENSVADHLSRMQITNMQELSINDYLRDDMLLKVTDSD 132
Query: 121 APWYADFVNFLAAGVLPP-----ELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCI 175
PWYA VNF+ A +PP LSY+ +K+ + D PYL++ D + RRC+
Sbjct: 133 -PWYAPIVNFMVAWHVPPGENKKRLSYESRKRLW--------DAPYLYRVCSDSLLRRCV 183
Query: 176 PENEVSSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNI 235
P +E I+ C + YG H + KI SGF+WP+++ D FI +C CQ+ G I
Sbjct: 184 PADEGMEIIEKCRVAPYGSHYGAFRTHAKIWQSGFFWPTMYGDTKEFIRRCTSCQKHGGI 243
Query: 236 TKRNEMPLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQV 295
T ++ MPL L+VE+FDVW IDF+GPFP + +YILVAVDYVSKWVE + + +
Sbjct: 244 TAQDAMPLTYNLQVELFDVWGIDFMGPFPKFYDCEYILVAVDYVSKWVEAMPCRAANTKH 303
Query: 296 VIKLFKKNHFSRFGVPRVVISDGGSQNILKNF 327
++F + F FG PR VISDGGS I K F
Sbjct: 304 AHRMFDEIIFPCFGTPRTVISDGGSHFIDKTF 335
>UniRef100_Q9SJS0 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 466
Score = 279 bits (714), Expect = 1e-73
Identities = 136/232 (58%), Positives = 166/232 (70%), Gaps = 5/232 (2%)
Query: 121 APWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENEV 180
+PWYAD VN+LA G+ PP L+ ++K FF D+ HYY DEPYL+ D I+R + E+EV
Sbjct: 37 SPWYADHVNYLACGIEPPNLTSYERKNFFRDIHHYYWDEPYLYTLCKDKIYRSYVSEDEV 96
Query: 181 SSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNE 240
IL HCH S+YGGH +T K KIL +GFWWP++FKD F+SKCD CQR GNI++RNE
Sbjct: 97 EGILLHCHDSAYGGHFATFKTVSKILQAGFWWPTMFKDAQEFVSKCDSCQRKGNISRRNE 156
Query: 241 MPLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQVVIKLF 300
MP N ILEV+IFDVW IDF+GPFPSS+ N+YILV VDYVSKWVE IAS TNDA+VV+KLF
Sbjct: 157 MPQNPILEVDIFDVWGIDFMGPFPSSYGNKYILVVVDYVSKWVEAIASPTNDAKVVLKLF 216
Query: 301 KKNHFSRFGVPRVVISDGGSQNILKNFCKNLE*GTKLQHPITHKLVDKWRSQ 352
K F RFGV VVISDGG I K F L+ +H + HK+ + Q
Sbjct: 217 KTIIFPRFGVSWVVISDGGKHFINKVFENLLK-----KHGVKHKVATPYHPQ 263
>UniRef100_Q7XQC1 OSJNBb0060M15.10 protein [Oryza sativa]
Length = 1028
Score = 276 bits (707), Expect = 9e-73
Identities = 153/346 (44%), Positives = 206/346 (59%), Gaps = 36/346 (10%)
Query: 20 KELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAKPRLIQLIFLLQEFDLEIKD 79
K VV+ IDKFR YLVG+K+ +YTDH+ +KYLL KKDAK RL++ I LLQEFDLEIKD
Sbjct: 513 KRAATVVFDIDKFRSYLVGAKVIIYTDHAALKYLLTKKDAKQRLLRWILLLQEFDLEIKD 572
Query: 80 KKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLAQTNAPWYADFVNFLAAGVLPPE 139
KKGVEN VADHLSRL+ T+ E P++D DD L + +N PWYA+ VN++ + +PP
Sbjct: 573 KKGVENSVADHLSRLQITDMQEQPINDFLRDDMLMTVRDSN-PWYANIVNYMVSKYIPP- 630
Query: 140 LSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENEVSSILTHCHSSSYGGHASTQ 199
Q+K + + +H + DEPYL++ DG+ RRC+P N+ I+ C +S YG H
Sbjct: 631 -GENQRKLKYENRRHIW-DEPYLYRVCSDGLLRRCVPTNKGLRIIERCLASPYGSHYGAF 688
Query: 200 KASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNEMPLNNILEVEIFDVWVIDF 259
+ KI SGF+WP++++D F+ +C CQR G IT R+ MPL L+ EIFDVW IDF
Sbjct: 689 RTQAKIWQSGFFWPTMYEDSTEFVRRCTSCQRQGGITARDAMPLTYNLQGEIFDVWGIDF 748
Query: 260 IGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQVVIKLFKKNHFSRFGVPRVVISDGG 319
+GPFP S +YILVAVDYVSK +VISDGG
Sbjct: 749 MGPFPKSRNCEYILVAVDYVSK-------------------------------MVISDGG 777
Query: 320 SQNILKNFCKNLE*GTKLQHPITHKLVDKWRSQIDKSKLSLKKLFQ 365
S I K F ++L + +H + + Q + S +K + Q
Sbjct: 778 SHFIDKTF-RDLLREMRAKHNVATPYHPQTSGQAETSNKQIKNILQ 822
>UniRef100_Q8S647 Putative retroelement [Oryza sativa]
Length = 1085
Score = 275 bits (704), Expect = 2e-72
Identities = 154/318 (48%), Positives = 192/318 (59%), Gaps = 56/318 (17%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
I YASKTL GA +NYATTEKELLAVV++IDKFR YLVG+++ +YTDH+ +KYLL KKDAK
Sbjct: 549 ICYASKTLTGAQLNYATTEKELLAVVFSIDKFRSYLVGAEVIIYTDHAALKYLLTKKDAK 608
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLAQTN 120
PRL++ I LLQEFDLEIKD+KGVEN VADHLSRL+ TN E P++D DD L ++ +N
Sbjct: 609 PRLLRWILLLQEFDLEIKDRKGVENSVADHLSRLQITNMQEQPINDFLRDDMLMTVSDSN 668
Query: 121 APWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENEV 180
PWY + VN++ + +PP Q+K + +H + DEPYL++ RRC
Sbjct: 669 -PWYVNIVNYMVSMYIPP--GENQRKLKYESRRHIW-DEPYLYREDSREFIRRC------ 718
Query: 181 SSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNE 240
CQR G IT R+
Sbjct: 719 ----------------------------------------------TSCQRQGGITTRDA 732
Query: 241 MPLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQVVIKLF 300
MPL L+VEIFDVW IDF+GPFP S +YILVAVDYVSKWVE + DA+ K+F
Sbjct: 733 MPLTYNLQVEIFDVWGIDFMGPFPKSRSCEYILVAVDYVSKWVEAMPCSAADARHAKKMF 792
Query: 301 KKNHFSRFGVPRVVISDG 318
+ F RFG PR+VISDG
Sbjct: 793 TEIIFPRFGTPRMVISDG 810
>UniRef100_Q8LLX4 Putative retroelement [Oryza sativa]
Length = 1093
Score = 275 bits (704), Expect = 2e-72
Identities = 154/318 (48%), Positives = 192/318 (59%), Gaps = 56/318 (17%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
I YASKTL GA +NYATTEKELLAVV++IDKFR YLVG+++ +YTDH+ +KYLL KKDAK
Sbjct: 557 ICYASKTLTGAQLNYATTEKELLAVVFSIDKFRSYLVGAEVIIYTDHAALKYLLTKKDAK 616
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLAQTN 120
PRL++ I LLQEFDLEIKD+KGVEN VADHLSRL+ TN E P++D DD L ++ +N
Sbjct: 617 PRLLRWILLLQEFDLEIKDRKGVENSVADHLSRLQITNMQEQPINDFLRDDMLMTVSDSN 676
Query: 121 APWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENEV 180
PWY + VN++ + +PP Q+K + +H + DEPYL++ RRC
Sbjct: 677 -PWYVNIVNYMVSMYIPP--GENQRKLKYESRRHIW-DEPYLYREDSREFIRRC------ 726
Query: 181 SSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNE 240
CQR G IT R+
Sbjct: 727 ----------------------------------------------TSCQRQGGITTRDA 740
Query: 241 MPLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQVVIKLF 300
MPL L+VEIFDVW IDF+GPFP S +YILVAVDYVSKWVE + DA+ K+F
Sbjct: 741 MPLTYNLQVEIFDVWGIDFMGPFPKSRSCEYILVAVDYVSKWVEAMPCSAADARHAKKMF 800
Query: 301 KKNHFSRFGVPRVVISDG 318
+ F RFG PR+VISDG
Sbjct: 801 TEIIFPRFGTPRMVISDG 818
>UniRef100_Q8S8G8 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 930
Score = 266 bits (679), Expect = 2e-69
Identities = 134/248 (54%), Positives = 167/248 (67%), Gaps = 9/248 (3%)
Query: 122 PWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENEVS 181
PWYAD VN+LAA V P + KK+F +++ Y DEPYL+K DGI+RRCI EV
Sbjct: 137 PWYADIVNYLAADVEPDNFTDYNKKRFLREIRRYQWDEPYLYKHSYDGIYRRCIAATEVP 196
Query: 182 SILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNEM 241
SIL+HCHSSSYGGH +T K K+L + FWWP++F+D FIS+CD CQR G I+KRNEM
Sbjct: 197 SILSHCHSSSYGGHFATFKTVSKVLQADFWWPTMFRDTQKFISQCDPCQRRGKISKRNEM 256
Query: 242 PLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQVVIKLFK 301
P N LEVE+FD W IDF+GPFP S N YILV VDYVSKWVE IASL ND+ VV+KLFK
Sbjct: 257 PPNFRLEVEVFDRWGIDFMGPFPPSNKNLYILVDVDYVSKWVEAIASLKNDSAVVMKLFK 316
Query: 302 KNHFSRFGVPRVVISDGGSQNILKNFCKNLE*GTKLQHPITHKLVDKWR----SQIDKSK 357
F RFGVPR+VISDG I K K L LQ+ + H++ + Q++ S
Sbjct: 317 SIIFPRFGVPRIVISDGDKHFINKILEKLL-----LQYGVQHRVATPYHPQTSGQVEVSN 371
Query: 358 LSLKKLFQ 365
+K++ +
Sbjct: 372 RQIKEILE 379
Score = 78.2 bits (191), Expect = 6e-13
Identities = 35/63 (55%), Positives = 52/63 (81%), Gaps = 2/63 (3%)
Query: 54 LNKKDAKPRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQL 113
+ KKDAKPRL+ I LLQEFD+E++DKKGVEN VADHLSR+R +D++P++D P++ +
Sbjct: 1 MQKKDAKPRLLIWILLLQEFDIEVRDKKGVENGVADHLSRIR--IDDDVPINDFLPEENI 58
Query: 114 FLL 116
+++
Sbjct: 59 YMI 61
>UniRef100_Q9LJS9 Retroelement pol polyprotein-like [Arabidopsis thaliana]
Length = 1118
Score = 250 bits (638), Expect = 9e-65
Identities = 162/355 (45%), Positives = 208/355 (57%), Gaps = 58/355 (16%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
IYYAS+TLDGA VNYATTEKELL +V+A +KFR YLV SK+ V+TDH+T++YLL+KKDAK
Sbjct: 615 IYYASRTLDGAQVNYATTEKELLVIVFAFEKFRSYLVRSKVIVHTDHATLRYLLSKKDAK 674
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLAQTN 120
PRL++ I LLQ+FDLEIKDKKG+EN VAD+LSRLR E+E+P+ D+ P + L +
Sbjct: 675 PRLLRWILLLQDFDLEIKDKKGIENGVADNLSRLRV--EEEVPMRDTLPGENLASIEDC- 731
Query: 121 APWYADFVNFLAAGVLPPELSY--QQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPEN 178
Y D V L L + ++ D +Y S E +P
Sbjct: 732 ---YMDEVERLRVSTLELMTLHTGDSNLPWYADFANYLSCE---------------LPPP 773
Query: 179 EVSSILTHCHSSSYGGHASTQKASFKILHSGFW-WPSLFKDVHLFISKCDKCQRTGNITK 237
+ + ++K K + +W P L+K KC N+ +
Sbjct: 774 DFTGY--------------SKKKFLKDVQRYYWDEPYLYK----------KC--ADNLFR 807
Query: 238 RNEMPLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQVVI 297
R +P + + EV FDVW IDF+GPFPSS+ N+YILVAVDYVSKWVE IA NDA VV+
Sbjct: 808 RC-IPESEVEEV--FDVWGIDFMGPFPSSYGNEYILVAVDYVSKWVEAIACRKNDAVVVL 864
Query: 298 KLFKKNHFSRFGVPRVVISDGGSQNILKNFCKNLE*GTKLQHPITHKLVDKWRSQ 352
KLFKK F RFGVPRVVISDGG I K F L +H + HK+ + Q
Sbjct: 865 KLFKKIIFPRFGVPRVVISDGGRHFINKVFANLL-----AKHGVNHKVATPYHPQ 914
>UniRef100_Q7FZN5 T14A16.1 protein [Arabidopsis thaliana]
Length = 276
Score = 247 bits (630), Expect = 7e-64
Identities = 140/286 (48%), Positives = 178/286 (61%), Gaps = 39/286 (13%)
Query: 70 LQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLAQTNAPWYADFVN 129
++ FD+EI DKK +EN ADHLSR+R E+ +P+DDS ++QL + V
Sbjct: 5 IRGFDIEIVDKKDIENGAADHLSRMRI--EERIPIDDSMLEEQLMV------------VE 50
Query: 130 FLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENEVSSILTHCHS 189
F SY K+ F+ L P+ R + E+EV IL HCH
Sbjct: 51 FFGG-------SYSGKE--FHQLNVVEGRSPW-----------RYVSEDEVEGILLHCHC 90
Query: 190 SSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNEMPLNNILEV 249
S+YGG+ +T K KIL +GFWWP++FK+ F+ KCD C R GNI+KRNEMP N ILEV
Sbjct: 91 STYGGNFTTFKTVSKILQAGFWWPTMFKEAQEFVLKCDSCHRKGNISKRNEMPQNPILEV 150
Query: 250 EIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQVVIKLFKKNHFSRFG 309
E+FDVW IDFIGPFPSS+ N+YILVAVDYVSKW+E AS TNDA+VV+KLFK FSRFG
Sbjct: 151 ELFDVWGIDFIGPFPSSYGNKYILVAVDYVSKWIEATASPTNDAKVVLKLFKTIIFSRFG 210
Query: 310 VPRVVISDGGSQNILKNFCKNLE*GTKLQHPITHKLVDKWRSQIDK 355
VPRVVISDGG I K F L+ +H + H + +++Q K
Sbjct: 211 VPRVVISDGGKHFINKVFENLLK-----KHGVKHGVYVMFKTQKKK 251
>UniRef100_Q7XWS6 OSJNBa0091C12.6 protein [Oryza sativa]
Length = 781
Score = 233 bits (594), Expect = 1e-59
Identities = 117/218 (53%), Positives = 153/218 (69%), Gaps = 4/218 (1%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
I YASKTL GA +NYATTEKELLAVV+AI+KFR YLVG+K+ VYTDH+ +KYLL KKDAK
Sbjct: 566 ITYASKTLTGAQLNYATTEKELLAVVFAIEKFRSYLVGAKVVVYTDHAALKYLLTKKDAK 625
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLAQTN 120
PR+I+ I LLQEFD+EI DKKGVEN VADHLSR++ TN +L ++D DD + ++
Sbjct: 626 PRVIRWILLLQEFDIEIMDKKGVENYVADHLSRMQITNMQKLLINDYLRDDMRLKVIDSD 685
Query: 121 APWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENEV 180
PWYA VNF+A G +PP + KK+ + + + D PYLF+ DG+ R C+ +E
Sbjct: 686 -PWYATIVNFMATGHVPP---VENKKRLIYESRRHLWDAPYLFRVCSDGLLRICVSTDEG 741
Query: 181 SSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKD 218
I+ CH++ YGGH + KI SGF+WPS++ D
Sbjct: 742 IMIIEKCHAAPYGGHYGAFRTHAKIWQSGFFWPSMYDD 779
>UniRef100_Q7XLF2 OSJNBa0013A04.17 protein [Oryza sativa]
Length = 632
Score = 226 bits (577), Expect = 1e-57
Identities = 119/249 (47%), Positives = 157/249 (62%), Gaps = 4/249 (1%)
Query: 52 YLLNKKDAKPRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDD 111
YLL KKDAKPRL+Q I LLQEFDLEIKDKKGVEN +A HLSRL TN E P+++ DD
Sbjct: 387 YLLTKKDAKPRLLQWILLLQEFDLEIKDKKGVENSIAYHLSRLPITNTQEQPINEFLRDD 446
Query: 112 QLFLLAQTNAPWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIF 171
L + + + PWYA+ VN++ + +PP + + K + +H + DE YL+ DG+
Sbjct: 447 ML-MTVRDSKPWYANIVNYMVSKYIPPRENQRMLK--YESRRHIW-DEAYLYTVCSDGLL 502
Query: 172 RRCIPENEVSSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQR 231
RRC+P E I+ CH+ YG H + KI GF+WP++++D F+ + CQR
Sbjct: 503 RRCVPTEEGFMIIEQCHAPPYGRHYVAFRTQAKIWQCGFFWPTMYEDSKEFVRRYPSCQR 562
Query: 232 TGNITKRNEMPLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTN 291
G IT R+ MPL L+VEIFDVW IDF+ PFP + ILVAVDYVSKWVE +
Sbjct: 563 QGGITARDAMPLTYNLQVEIFDVWGIDFMRPFPKCRNCENILVAVDYVSKWVEAMPCSAV 622
Query: 292 DAQVVIKLF 300
DA+ K+F
Sbjct: 623 DAKHAKKMF 631
>UniRef100_Q7XNX7 OSJNBb0026I12.3 protein [Oryza sativa]
Length = 589
Score = 212 bits (539), Expect = 3e-53
Identities = 113/216 (52%), Positives = 150/216 (69%), Gaps = 5/216 (2%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQY-LVGSKITVYTDHSTIKYLLNKKDA 59
I YAS+TL GA NYATTEKELLAVV+AIDKFR+ LVG+K+ VYT+H+ +KYLL KKDA
Sbjct: 292 ICYASETLAGAQSNYATTEKELLAVVFAIDKFRRSDLVGAKVIVYTEHAALKYLLTKKDA 351
Query: 60 KPRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLAQT 119
KPRL++ I LLQEFDLEIKDKKGVEN VADHLSRL+ TN E P++ DD L + +
Sbjct: 352 KPRLLRWIPLLQEFDLEIKDKKGVENSVADHLSRLQLTNMQEPPINYFLQDDML-MAVRN 410
Query: 120 NAPWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENE 179
+ PWYA+ VN++ + +P Q+K + H + DEPYL++ +G+ RRC+P+ E
Sbjct: 411 SDPWYANIVNYMVSKYVPQ--GENQRKLKYESHCHIW-DEPYLYRVCSNGLLRRCVPKEE 467
Query: 180 VSSILTHCHSSSYGGHASTQKASFKILHSGFWWPSL 215
I+ CH++ Y GH + KI S F+WP++
Sbjct: 468 GFKIIERCHAAPYRGHYGAFRTQAKIWQSRFFWPTM 503
>UniRef100_Q6L4N0 Hypothetical protein P0530H10.3 [Oryza sativa]
Length = 903
Score = 201 bits (510), Expect = 6e-50
Identities = 135/375 (36%), Positives = 181/375 (48%), Gaps = 82/375 (21%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
I YASKTL GA +NYATTEKELLA
Sbjct: 555 IAYASKTLTGAQLNYATTEKELLA------------------------------------ 578
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLAQTN 120
L+F + +F L + K ELP++D D+ L + ++
Sbjct: 579 -----LVFAIDKFRLYLVGAK-------------------ELPINDYLRDNMLLKVTDSD 614
Query: 121 APWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENEV 180
PWYA VNF+ AG +PP + KK+ + + + D PYL+ R ++E
Sbjct: 615 -PWYATIVNFMVAGHVPPG---ENKKRLIYESRRHLWDAPYLY---------RVCSDDEG 661
Query: 181 SSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNE 240
I+ CH++ YGGH + KI SGF+WPS++ D FI +C CQ+ G IT R+
Sbjct: 662 MKIIKKCHAALYGGHYGAFRTHAKIWQSGFFWPSMYDDTKEFIRRCTSCQKHGGITARDA 721
Query: 241 MPLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQVVIKLF 300
MPL L+VE+FDVW IDF+GPFP S+ +YILVAVDYVSKWVE + DA+ +F
Sbjct: 722 MPLTYNLQVELFDVWGIDFMGPFPKSYDCEYILVAVDYVSKWVEAMPCRAADAKHARWMF 781
Query: 301 KKNHFSRFGVPRVVISDGGSQNI---LKNFCKNL---E*GTKLQHPITHKLVDKWRSQID 354
+ F RF R+VISDGGS + +NF K L HP T + QI
Sbjct: 782 HEIIFPRFRTLRMVISDGGSHFVDKTFRNFLKELGARHNSATPYHPQTSGQAETSNKQI- 840
Query: 355 KSKLSLKKLFQPLGK 369
K L+K +GK
Sbjct: 841 --KNILQKTINEMGK 853
>UniRef100_Q9SI95 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 301
Score = 188 bits (478), Expect = 3e-46
Identities = 96/173 (55%), Positives = 119/173 (68%), Gaps = 20/173 (11%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
IYYAS T+D V YATTEKELLAVV+A +KFR YLVGSK+TVYTDH+ ++++ KKD K
Sbjct: 131 IYYASGTMDDTQVRYATTEKELLAVVFAFEKFRSYLVGSKVTVYTDHAALRHIYAKKDTK 190
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLAQTN 120
PRL++ I LLQEFD+EI DKKG+EN VADHLSR+R EDE+P+DDS P++QL + Q N
Sbjct: 191 PRLLRWILLLQEFDMEIVDKKGIENGVADHLSRMR--IEDEVPIDDSMPEEQLMAIQQLN 248
Query: 121 AP------------------WYADFVNFLAAGVLPPELSYQQKKKFFNDLKHY 155
WYAD V +L G PP LS +KKKFF D+ H+
Sbjct: 249 ESAQINKSLDQVCAVEDKLMWYADHVKYLFGGEEPPNLSSYEKKKFFKDINHF 301
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.336 0.148 0.464
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 831,829,329
Number of Sequences: 2790947
Number of extensions: 34183570
Number of successful extensions: 132783
Number of sequences better than 10.0: 2190
Number of HSP's better than 10.0 without gapping: 1586
Number of HSP's successfully gapped in prelim test: 604
Number of HSP's that attempted gapping in prelim test: 126798
Number of HSP's gapped (non-prelim): 3741
length of query: 547
length of database: 848,049,833
effective HSP length: 132
effective length of query: 415
effective length of database: 479,644,829
effective search space: 199052604035
effective search space used: 199052604035
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 77 (34.3 bits)
Medicago: description of AC141865.9


