

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141863.5 - phase: 0
(139 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
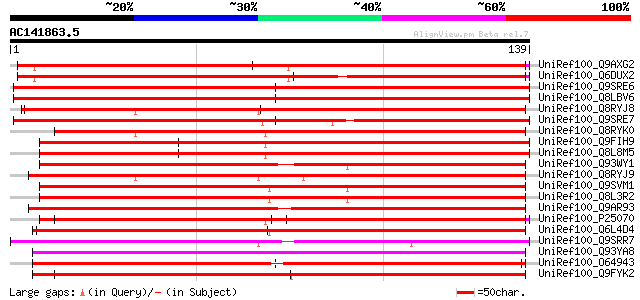
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9AXG2 Regulator of gene silencing [Nicotiana tabacum] 152 1e-36
UniRef100_Q6DUX2 Regulator of gene silencing [Lycopersicon escul... 145 2e-34
UniRef100_Q9SRE6 Putative calmodulin; 4214-3681 [Arabidopsis tha... 129 2e-29
UniRef100_Q8LBV6 Putative calmodulin [Arabidopsis thaliana] 129 2e-29
UniRef100_Q8RYJ8 B1139B11.11 protein [Oryza sativa] 126 1e-28
UniRef100_Q9SRE7 Putative calmodulin; 2575-2096 [Arabidopsis tha... 124 4e-28
UniRef100_Q8RYK0 B1139B11.9 protein [Oryza sativa] 121 4e-27
UniRef100_Q9FIH9 Similarity to calmodulin [Arabidopsis thaliana] 120 8e-27
UniRef100_Q8L8M5 Putative calmodulin [Arabidopsis thaliana] 119 1e-26
UniRef100_Q93WY1 Calmodulin-like protein [Musa acuminata] 119 2e-26
UniRef100_Q8RYJ9 B1139B11.10 protein [Oryza sativa] 116 9e-26
UniRef100_Q9SVM1 Calmodulin-like protein [Arabidopsis thaliana] 115 3e-25
UniRef100_Q8L3R2 Calmodulin-like protein [Arabidopsis thaliana] 115 3e-25
UniRef100_Q9AR93 Putative calmodulin-related protein [Medicago s... 105 3e-22
UniRef100_P25070 Calmodulin-related protein 2, touch-induced [Ar... 105 3e-22
UniRef100_Q6L4D4 Hypothetical protein OSJNBa0088M05.6 [Oryza sat... 104 5e-22
UniRef100_Q9SRR7 Putative calmodulin [Arabidopsis thaliana] 102 1e-21
UniRef100_Q93YA8 Calcium binding protein [Sesbania rostrata] 102 1e-21
UniRef100_O64943 Polcalcin Jun o 2 [Juniperus oxycedrus] 102 2e-21
UniRef100_Q9FYK2 F21J9.28 [Arabidopsis thaliana] 101 3e-21
>UniRef100_Q9AXG2 Regulator of gene silencing [Nicotiana tabacum]
Length = 190
Score = 152 bits (385), Expect = 1e-36
Identities = 80/138 (57%), Positives = 102/138 (72%), Gaps = 2/138 (1%)
Query: 3 NAG-FEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEE 61
N+G E V YFDE+GDGKVSPAELR+ ++ +G E+ ++EAEMA+ DSDGDG L LE+
Sbjct: 51 NSGELERVFTYFDENGDGKVSPAELRRCVKAVGGELTVEEAEMAVRLSDSDGDGLLGLED 110
Query: 62 LIALMEEGGEEQ-KLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKH 120
LME EE+ K +L AF MY+ E G+ITPKSLK ML ++GES SID CKAMI+
Sbjct: 111 FTKLMEGMEEERNKESELIGAFGMYEMEGSGYITPKSLKMMLSRLGESTSIDNCKAMIQR 170
Query: 121 FDLDGDGLLSFDEFITMM 138
FD++GDG+L+FDEF MM
Sbjct: 171 FDINGDGVLNFDEFKAMM 188
Score = 47.0 bits (110), Expect = 9e-05
Identities = 20/74 (27%), Positives = 38/74 (51%)
Query: 66 MEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDG 125
M + +L F +D G ++P L+R +K +G +++E + ++ D DG
Sbjct: 43 MSSSSKSNNSGELERVFTYFDENGDGKVSPAELRRCVKAVGGELTVEEAEMAVRLSDSDG 102
Query: 126 DGLLSFDEFITMMQ 139
DGLL ++F +M+
Sbjct: 103 DGLLGLEDFTKLME 116
>UniRef100_Q6DUX2 Regulator of gene silencing [Lycopersicon esculentum]
Length = 198
Score = 145 bits (365), Expect = 2e-34
Identities = 78/138 (56%), Positives = 98/138 (70%), Gaps = 4/138 (2%)
Query: 3 NAG-FEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEE 61
N+G E V YFDE+GDGKVSP ELR+ ++ +G EI ++EAEM + DSDGDG L E+
Sbjct: 61 NSGELERVFTYFDENGDGKVSPMELRRCMKAVGGEITVEEAEMVVRLSDSDGDGLLGFED 120
Query: 62 LIALMEEGGEEQ-KLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKH 120
LME EE+ K +L AF MY+ E G+ITPKSLK ML ++GES SID+CK MI+
Sbjct: 121 FTKLMEGMEEERNKESELMGAFGMYEME--GYITPKSLKMMLSRLGESTSIDKCKVMIRR 178
Query: 121 FDLDGDGLLSFDEFITMM 138
FD +GDG+LSFDEF MM
Sbjct: 179 FDTNGDGVLSFDEFKVMM 196
Score = 49.3 bits (116), Expect = 2e-05
Identities = 20/63 (31%), Positives = 38/63 (59%)
Query: 77 DLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSFDEFIT 136
+L F +D G ++P L+R +K +G +++E + +++ D DGDGLL F++F
Sbjct: 64 ELERVFTYFDENGDGKVSPMELRRCMKAVGGEITVEEAEMVVRLSDSDGDGLLGFEDFTK 123
Query: 137 MMQ 139
+M+
Sbjct: 124 LME 126
>UniRef100_Q9SRE6 Putative calmodulin; 4214-3681 [Arabidopsis thaliana]
Length = 177
Score = 129 bits (323), Expect = 2e-29
Identities = 64/138 (46%), Positives = 91/138 (65%)
Query: 2 KNAGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEE 61
KN E V Y D + DG++SP EL++ +GE++ +EA A+ D+DGDG L EE
Sbjct: 40 KNRELEAVFSYMDANRDGRISPEELQKSFMTLGEQLSDEEAVAAVRLSDTDGDGMLDFEE 99
Query: 62 LIALMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHF 121
L++ EE+K +L+ AF +Y +E ITP+SLK MLKK+GES++ D+C+ MI F
Sbjct: 100 FSQLIKVDDEEEKKMELKGAFRLYIAEGEDCITPRSLKMMLKKLGESRTTDDCRVMISAF 159
Query: 122 DLDGDGLLSFDEFITMMQ 139
DL+ DG+LSFDEF MM+
Sbjct: 160 DLNADGVLSFDEFALMMR 177
Score = 50.4 bits (119), Expect = 8e-06
Identities = 23/68 (33%), Positives = 40/68 (58%)
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E K ++L F D+ + G I+P+ L++ +GE S +E A ++ D DGDG+L F
Sbjct: 38 EDKNRELEAVFSYMDANRDGRISPEELQKSFMTLGEQLSDEEAVAAVRLSDTDGDGMLDF 97
Query: 132 DEFITMMQ 139
+EF +++
Sbjct: 98 EEFSQLIK 105
>UniRef100_Q8LBV6 Putative calmodulin [Arabidopsis thaliana]
Length = 177
Score = 129 bits (323), Expect = 2e-29
Identities = 64/138 (46%), Positives = 91/138 (65%)
Query: 2 KNAGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEE 61
KN E V Y D + DG++SP EL++ +GE++ +EA A+ D+DGDG L EE
Sbjct: 40 KNRELEAVFSYMDANRDGRISPEELQKSFMTLGEQLSDEEAVAAVRLSDTDGDGMLDFEE 99
Query: 62 LIALMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHF 121
L++ EE+K +L+ AF +Y +E ITP+SLK MLKK+GES++ D+C+ MI F
Sbjct: 100 FSQLIKVDDEEEKKMELKGAFRLYITEGEDCITPRSLKMMLKKLGESRTTDDCRVMISAF 159
Query: 122 DLDGDGLLSFDEFITMMQ 139
DL+ DG+LSFDEF MM+
Sbjct: 160 DLNADGVLSFDEFALMMR 177
Score = 50.4 bits (119), Expect = 8e-06
Identities = 23/68 (33%), Positives = 40/68 (58%)
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E K ++L F D+ + G I+P+ L++ +GE S +E A ++ D DGDG+L F
Sbjct: 38 EDKNRELEAVFSYMDANRDGRISPEELQKSFMTLGEQLSDEEAVAAVRLSDTDGDGMLDF 97
Query: 132 DEFITMMQ 139
+EF +++
Sbjct: 98 EEFSQLIK 105
>UniRef100_Q8RYJ8 B1139B11.11 protein [Oryza sativa]
Length = 146
Score = 126 bits (316), Expect = 1e-28
Identities = 68/139 (48%), Positives = 90/139 (63%), Gaps = 4/139 (2%)
Query: 4 AGFEHVLRYFDEDGDGKVSPAELRQRLRI-MGEEILLKEAEMAIEAMDSDGDGYLSLEEL 62
A F V FD D DGK+S AELR ++ +GE++ +EAE + + D+D DG L EE
Sbjct: 6 AEFRRVFSAFDRDADGKISAAELRLCMKAALGEDMSAEEAEALVSSADTDDDGLLDEEEF 65
Query: 63 IAL---MEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIK 119
L +E G EE++ + L EAF MY+ E G ITP SLKRML K+G + I+EC+ MI
Sbjct: 66 TKLAVQLEMGDEEERCRGLMEAFRMYEMEGEGRITPASLKRMLSKLGSHQGIEECQTMIC 125
Query: 120 HFDLDGDGLLSFDEFITMM 138
FDLDGDG++SF+EF MM
Sbjct: 126 RFDLDGDGVISFEEFKIMM 144
Score = 45.4 bits (106), Expect = 3e-04
Identities = 20/63 (31%), Positives = 36/63 (56%)
Query: 5 GFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIA 64
G R ++ +G+G+++PA L++ L +G ++E + I D DGDG +S EE
Sbjct: 83 GLMEAFRMYEMEGEGRITPASLKRMLSKLGSHQGIEECQTMICRFDLDGDGVISFEEFKI 142
Query: 65 LME 67
+M+
Sbjct: 143 MMD 145
>UniRef100_Q9SRE7 Putative calmodulin; 2575-2096 [Arabidopsis thaliana]
Length = 159
Score = 124 bits (311), Expect = 4e-28
Identities = 67/143 (46%), Positives = 93/143 (64%), Gaps = 7/143 (4%)
Query: 2 KNAGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEE 61
KN E V Y D + DG++S EL++ + +GE++ +EAE A++ D DGDG L + E
Sbjct: 19 KNRDLEAVFAYMDANRDGRISAEELKKSFKTLGEQMSDEEAEAAVKLSDIDGDGMLDINE 78
Query: 62 LIALM---EEGGEEQKLKDLREAFEMY--DSEKCGFITPKSLKRMLKKMGESKSIDECKA 116
L+ +E EE+K + + EAF MY D E C ITP SLK ML K+GES++ D+CK
Sbjct: 79 FALLIKGNDEFTEEEKKRKIMEAFRMYIADGEDC--ITPGSLKMMLMKLGESRTTDDCKV 136
Query: 117 MIKHFDLDGDGLLSFDEFITMMQ 139
MI+ FDL+ DG+LSFDEF MM+
Sbjct: 137 MIQAFDLNADGVLSFDEFALMMR 159
Score = 54.7 bits (130), Expect = 4e-07
Identities = 25/68 (36%), Positives = 42/68 (61%)
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E+K +DL F D+ + G I+ + LK+ K +GE S +E +A +K D+DGDG+L
Sbjct: 17 EEKNRDLEAVFAYMDANRDGRISAEELKKSFKTLGEQMSDEEAEAAVKLSDIDGDGMLDI 76
Query: 132 DEFITMMQ 139
+EF +++
Sbjct: 77 NEFALLIK 84
>UniRef100_Q8RYK0 B1139B11.9 protein [Oryza sativa]
Length = 151
Score = 121 bits (303), Expect = 4e-27
Identities = 64/131 (48%), Positives = 91/131 (68%), Gaps = 5/131 (3%)
Query: 13 FDEDGDGKVSPAELRQRLRI-MGEEILLKEAEMAIEAMDSDGDGYLSLEELIALME---- 67
FD DGDG++S AELR ++ +GEE+ +EA + ++D+DGDG L E + L++
Sbjct: 19 FDHDGDGRISAAELRLCMKTTLGEEVSDEEAGQLVASVDADGDGLLCEAEFVRLVQAAEV 78
Query: 68 EGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDG 127
E +E++ LREAF MY+ E G ITP SL+RML+++G + ID+C+AMI FDL+GDG
Sbjct: 79 EEEDERRGTGLREAFGMYEMEGEGCITPTSLRRMLRRLGSDQDIDDCRAMICRFDLNGDG 138
Query: 128 LLSFDEFITMM 138
+LSFDEF MM
Sbjct: 139 VLSFDEFKIMM 149
Score = 39.3 bits (90), Expect = 0.019
Identities = 18/65 (27%), Positives = 33/65 (50%)
Query: 2 KNAGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEE 61
+ G ++ +G+G ++P LR+ LR +G + + + I D +GDG LS +E
Sbjct: 85 RGTGLREAFGMYEMEGEGCITPTSLRRMLRRLGSDQDIDDCRAMICRFDLNGDGVLSFDE 144
Query: 62 LIALM 66
+M
Sbjct: 145 FKIMM 149
Score = 36.2 bits (82), Expect = 0.16
Identities = 20/59 (33%), Positives = 32/59 (53%), Gaps = 1/59 (1%)
Query: 82 FEMYDSEKCGFITPKSLKRMLKK-MGESKSIDECKAMIKHFDLDGDGLLSFDEFITMMQ 139
F +D + G I+ L+ +K +GE S +E ++ D DGDGLL EF+ ++Q
Sbjct: 16 FATFDHDGDGRISAAELRLCMKTTLGEEVSDEEAGQLVASVDADGDGLLCEAEFVRLVQ 74
>UniRef100_Q9FIH9 Similarity to calmodulin [Arabidopsis thaliana]
Length = 185
Score = 120 bits (300), Expect = 8e-27
Identities = 59/133 (44%), Positives = 88/133 (65%), Gaps = 2/133 (1%)
Query: 9 VLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALME- 67
V Y D + DGK+S EL+ + ++G + +E E ++ D DGDG++ EE + LME
Sbjct: 53 VFDYMDANSDGKISGEELQSCVSLLGGALSSREVEEVVKTSDVDGDGFIDFEEFLKLMEG 112
Query: 68 -EGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGD 126
+G +E++ K+L+EAF MY E FIT SL+R L ++GES ++D CK MI+ FD + D
Sbjct: 113 EDGSDEERRKELKEAFGMYVMEGEEFITAASLRRTLSRLGESCTVDACKVMIRGFDQNDD 172
Query: 127 GLLSFDEFITMMQ 139
G+LSFDEF+ MM+
Sbjct: 173 GVLSFDEFVLMMR 185
Score = 47.8 bits (112), Expect = 5e-05
Identities = 24/94 (25%), Positives = 47/94 (49%)
Query: 46 IEAMDSDGDGYLSLEELIALMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKM 105
+ + S+ L E + G + +LR F+ D+ G I+ + L+ + +
Sbjct: 18 VSSKRSESSRNLEDESRTSSNSSGSSSLNVNELRTVFDYMDANSDGKISGEELQSCVSLL 77
Query: 106 GESKSIDECKAMIKHFDLDGDGLLSFDEFITMMQ 139
G + S E + ++K D+DGDG + F+EF+ +M+
Sbjct: 78 GGALSSREVEEVVKTSDVDGDGFIDFEEFLKLME 111
>UniRef100_Q8L8M5 Putative calmodulin [Arabidopsis thaliana]
Length = 185
Score = 119 bits (299), Expect = 1e-26
Identities = 59/133 (44%), Positives = 88/133 (65%), Gaps = 2/133 (1%)
Query: 9 VLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALME- 67
V Y D + DGK+S EL+ + ++G + +E E ++ D DGDG++ EE + LME
Sbjct: 53 VFDYMDANSDGKISGEELQSCVSLLGGALSSREVEEVVKTSDVDGDGFIDFEEFLKLMEG 112
Query: 68 -EGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGD 126
+G +E++ K+L+EAF MY E FIT SL+R L ++GES ++D CK MI+ FD + D
Sbjct: 113 EDGSDEERRKELKEAFGMYLMEGEEFITAASLRRTLSRLGESCTVDACKVMIRGFDQNDD 172
Query: 127 GLLSFDEFITMMQ 139
G+LSFDEF+ MM+
Sbjct: 173 GVLSFDEFVLMMR 185
Score = 47.8 bits (112), Expect = 5e-05
Identities = 24/94 (25%), Positives = 47/94 (49%)
Query: 46 IEAMDSDGDGYLSLEELIALMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKM 105
+ + S+ L E + G + +LR F+ D+ G I+ + L+ + +
Sbjct: 18 VSSKRSESSRNLEDEARTSSSSSGSSSLNVNELRTVFDYMDANSDGKISGEELQSCVSLL 77
Query: 106 GESKSIDECKAMIKHFDLDGDGLLSFDEFITMMQ 139
G + S E + ++K D+DGDG + F+EF+ +M+
Sbjct: 78 GGALSSREVEEVVKTSDVDGDGFIDFEEFLKLME 111
>UniRef100_Q93WY1 Calmodulin-like protein [Musa acuminata]
Length = 173
Score = 119 bits (297), Expect = 2e-26
Identities = 63/131 (48%), Positives = 87/131 (66%), Gaps = 5/131 (3%)
Query: 9 VLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEE 68
V R+ D+D DGK+S EL +GEE+ ++EAE AI +DSDGD L + + +ME
Sbjct: 46 VFRHIDQDRDGKISGVELLGFFGSIGEEMPMEEAEAAIALLDSDGDRLLDFGDFLRMMER 105
Query: 69 GGEEQKLKDLREAFEMYDSEK-CGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDG 127
E+ DLR AFEM++ K G ITPK L+RM+ ++GE +S+++CKAMI+ +DLDGDG
Sbjct: 106 EEED----DLRRAFEMFEVVKGSGRITPKGLQRMMSRLGEERSVEDCKAMIRAYDLDGDG 161
Query: 128 LLSFDEFITMM 138
L F EF MM
Sbjct: 162 ELDFQEFHQMM 172
Score = 41.6 bits (96), Expect = 0.004
Identities = 21/66 (31%), Positives = 35/66 (52%)
Query: 74 KLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSFDE 133
++ +L + F D ++ G I+ L +GE ++E +A I D DGD LL F +
Sbjct: 39 RVDELLQVFRHIDQDRDGKISGVELLGFFGSIGEEMPMEEAEAAIALLDSDGDRLLDFGD 98
Query: 134 FITMMQ 139
F+ MM+
Sbjct: 99 FLRMME 104
>UniRef100_Q8RYJ9 B1139B11.10 protein [Oryza sativa]
Length = 151
Score = 116 bits (291), Expect = 9e-26
Identities = 67/143 (46%), Positives = 90/143 (62%), Gaps = 10/143 (6%)
Query: 6 FEHVLRYFDEDGDGKVSPAELRQRLRI-MGEEILLKEAEMAIEAMDSDGDGYLSLEELIA 64
F V FD+DGDGK+S ELR ++ +GE++ +E + + D+DGDG L EE +
Sbjct: 7 FRRVFGSFDQDGDGKISATELRLCVKASLGEDMPDEEVQALMALADTDGDGLLDEEEFVR 66
Query: 65 L---MEEGGEEQKLKD------LREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECK 115
L ME G+E++ D LREAF MY+ E G ITP SLK ML K+G + EC+
Sbjct: 67 LVTEMEADGDEEEDDDDETCRCLREAFAMYEMEGRGCITPLSLKLMLSKLGTHLDVAECQ 126
Query: 116 AMIKHFDLDGDGLLSFDEFITMM 138
AMI FD++GDG+L+FDEF TMM
Sbjct: 127 AMICRFDMNGDGVLTFDEFKTMM 149
>UniRef100_Q9SVM1 Calmodulin-like protein [Arabidopsis thaliana]
Length = 205
Score = 115 bits (287), Expect = 3e-25
Identities = 63/136 (46%), Positives = 86/136 (62%), Gaps = 6/136 (4%)
Query: 9 VLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEE 68
V +FD DGDGK+S ELR +GE I + A+ AI +D+D DG L E+ + LM
Sbjct: 68 VFSHFDSDGDGKISAFELRHYFGSVGEYISHEAAQEAINEVDTDADGSLGFEDFVGLMTR 127
Query: 69 -----GGEEQKLKDLREAFEMYDSEK-CGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
GE +L+ AFEM++ EK G ITPK L++ML K+GES++ EC+AMIK +D
Sbjct: 128 RDLYGDGEVDGDGELKTAFEMFEVEKGSGCITPKGLQKMLVKLGESRTYGECEAMIKFYD 187
Query: 123 LDGDGLLSFDEFITMM 138
+DG+G+L F EF MM
Sbjct: 188 IDGNGILDFHEFRQMM 203
Score = 41.2 bits (95), Expect = 0.005
Identities = 20/63 (31%), Positives = 34/63 (53%)
Query: 76 KDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSFDEFI 135
++LR+ F +DS+ G I+ L+ +GE S + + I D D DG L F++F+
Sbjct: 63 EELRQVFSHFDSDGDGKISAFELRHYFGSVGEYISHEAAQEAINEVDTDADGSLGFEDFV 122
Query: 136 TMM 138
+M
Sbjct: 123 GLM 125
>UniRef100_Q8L3R2 Calmodulin-like protein [Arabidopsis thaliana]
Length = 205
Score = 115 bits (287), Expect = 3e-25
Identities = 63/136 (46%), Positives = 86/136 (62%), Gaps = 6/136 (4%)
Query: 9 VLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEE 68
V +FD DGDGK+S ELR +GE I + A+ AI +D+D DG L E+ + LM
Sbjct: 68 VFSHFDSDGDGKISAFELRHYFGSVGEYISHEAAQEAINEVDTDADGSLGFEDFVGLMTR 127
Query: 69 -----GGEEQKLKDLREAFEMYDSEK-CGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
GE +L+ AFEM++ EK G ITPK L++ML K+GES++ EC+AMIK +D
Sbjct: 128 RDLYGDGEVDGDGELKTAFEMFEVEKGSGCITPKGLQKMLVKLGESRTYGECEAMIKFYD 187
Query: 123 LDGDGLLSFDEFITMM 138
+DG+G+L F EF MM
Sbjct: 188 IDGNGILDFHEFRQMM 203
Score = 41.2 bits (95), Expect = 0.005
Identities = 20/63 (31%), Positives = 34/63 (53%)
Query: 76 KDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSFDEFI 135
++LR+ F +DS+ G I+ L+ +GE S + + I D D DG L F++F+
Sbjct: 63 EELRQVFSHFDSDGDGKISAFELRHYFGSVGEYISHEAAQEAINEVDTDADGSLGFEDFV 122
Query: 136 TMM 138
+M
Sbjct: 123 GLM 125
>UniRef100_Q9AR93 Putative calmodulin-related protein [Medicago sativa]
Length = 167
Score = 105 bits (261), Expect = 3e-22
Identities = 52/133 (39%), Positives = 82/133 (61%), Gaps = 3/133 (2%)
Query: 6 FEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIAL 65
F + FD++GDGK+S EL++ + +G + +E +E +D +GDGY+ L+E L
Sbjct: 5 FARIFNKFDKNGDGKISRTELKEMMTALGCKTTTEEVTRMMEELDRNGDGYIDLKEFGEL 64
Query: 66 MEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDG 125
GG+ K+LREAFEMYD K G + K L +++++GE S+ +C+ MI + D D
Sbjct: 65 HNGGGDT---KELREAFEMYDLGKNGLTSAKELHAVMRRLGEKCSLGDCRRMIGNVDADS 121
Query: 126 DGLLSFDEFITMM 138
DG ++F+EF MM
Sbjct: 122 DGNVNFEEFKKMM 134
>UniRef100_P25070 Calmodulin-related protein 2, touch-induced [Arabidopsis thaliana]
Length = 161
Score = 105 bits (261), Expect = 3e-22
Identities = 52/135 (38%), Positives = 82/135 (60%), Gaps = 5/135 (3%)
Query: 9 VLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALME- 67
V + FD++GDGK+S EL++ +R + +E ++ D DG+G++ L+E +AL +
Sbjct: 21 VFQRFDKNGDGKISVDELKEVIRALSPTASPEETVTMMKQFDLDGNGFIDLDEFVALFQI 80
Query: 68 ----EGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDL 123
G + DL+EAFE+YD + G I+ K L ++K +GE S+ +CK MI D+
Sbjct: 81 GIGGGGNNRNDVSDLKEAFELYDLDGNGRISAKELHSVMKNLGEKCSVQDCKKMISKVDI 140
Query: 124 DGDGLLSFDEFITMM 138
DGDG ++FDEF MM
Sbjct: 141 DGDGCVNFDEFKKMM 155
Score = 51.6 bits (122), Expect = 4e-06
Identities = 21/65 (32%), Positives = 39/65 (59%)
Query: 75 LKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSFDEF 134
+ D+++ F+ +D G I+ LK +++ + + S +E M+K FDLDG+G + DEF
Sbjct: 15 MDDIKKVFQRFDKNGDGKISVDELKEVIRALSPTASPEETVTMMKQFDLDGNGFIDLDEF 74
Query: 135 ITMMQ 139
+ + Q
Sbjct: 75 VALFQ 79
Score = 45.8 bits (107), Expect = 2e-04
Identities = 19/58 (32%), Positives = 36/58 (61%)
Query: 13 FDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEEGG 70
+D DG+G++S EL ++ +GE+ +++ + I +D DGDG ++ +E +M GG
Sbjct: 102 YDLDGNGRISAKELHSVMKNLGEKCSVQDCKKMISKVDIDGDGCVNFDEFKKMMSNGG 159
>UniRef100_Q6L4D4 Hypothetical protein OSJNBa0088M05.6 [Oryza sativa]
Length = 201
Score = 104 bits (259), Expect = 5e-22
Identities = 55/134 (41%), Positives = 81/134 (60%), Gaps = 2/134 (1%)
Query: 7 EHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALM 66
E V R FD +GDG++S AEL R +G + E ++ DSDGDGY+SL E A+
Sbjct: 57 ERVFRKFDANGDGRISRAELAALFRSVGHAVTDDEVARMMQEADSDGDGYISLGEFAAIS 116
Query: 67 EE--GGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLD 124
G +DLR AF ++D++ G ITP L R+L+ +GE+ ++ +C+ MI D +
Sbjct: 117 APPPGDAAAAEEDLRHAFGVFDADGNGVITPAELARVLRGIGEAATVAQCRRMIDGVDRN 176
Query: 125 GDGLLSFDEFITMM 138
GDGL++F+EF MM
Sbjct: 177 GDGLINFEEFKLMM 190
Score = 50.1 bits (118), Expect = 1e-05
Identities = 23/62 (37%), Positives = 36/62 (57%)
Query: 8 HVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALME 67
H FD DG+G ++PAEL + LR +GE + + I+ +D +GDG ++ EE +M
Sbjct: 132 HAFGVFDADGNGVITPAELARVLRGIGEAATVAQCRRMIDGVDRNGDGLINFEEFKLMMA 191
Query: 68 EG 69
G
Sbjct: 192 AG 193
>UniRef100_Q9SRR7 Putative calmodulin [Arabidopsis thaliana]
Length = 153
Score = 102 bits (255), Expect = 1e-21
Identities = 56/145 (38%), Positives = 87/145 (59%), Gaps = 9/145 (6%)
Query: 1 MKNAGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLE 60
M A + + FD +GDGK++ EL L +G I K+ IE +D +GDGY+ +E
Sbjct: 1 MDQAELARIFQMFDRNGDGKITKQELNDSLENLGIYIPDKDLVQMIEKIDLNGDGYVDIE 60
Query: 61 ELIAL----MEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMG--ESKSIDEC 114
E L MEE EE+ D+REAF ++D + GFIT + L+ +L +G + +++++C
Sbjct: 61 EFGGLYQTIMEERDEEE---DMREAFNVFDQNRDGFITVEELRSVLASLGLKQGRTLEDC 117
Query: 115 KAMIKHFDLDGDGLLSFDEFITMMQ 139
K MI D+DGDG+++F EF MM+
Sbjct: 118 KRMISKVDVDGDGMVNFKEFKQMMK 142
>UniRef100_Q93YA8 Calcium binding protein [Sesbania rostrata]
Length = 172
Score = 102 bits (255), Expect = 1e-21
Identities = 52/132 (39%), Positives = 77/132 (57%)
Query: 7 EHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALM 66
+ V FD +GDGK+S EL LR +G + E E ++ +D+D DG+++L E A
Sbjct: 34 KRVFSRFDANGDGKISVNELDNVLRALGSTVPSDELERVMKDLDTDNDGFINLTEFAAFC 93
Query: 67 EEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGD 126
+ +LREAF++YD +K G I+ L +L ++G S++EC MIK D DGD
Sbjct: 94 RSDAADGGASELREAFDLYDQDKNGLISAAELCLVLNRLGMKCSVEECHNMIKSVDSDGD 153
Query: 127 GLLSFDEFITMM 138
G ++FDEF MM
Sbjct: 154 GNVNFDEFKQMM 165
Score = 39.7 bits (91), Expect = 0.015
Identities = 17/62 (27%), Positives = 33/62 (52%)
Query: 73 QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSFD 132
+ + +L+ F +D+ G I+ L +L+ +G + DE + ++K D D DG ++
Sbjct: 28 EDMDELKRVFSRFDANGDGKISVNELDNVLRALGSTVPSDELERVMKDLDTDNDGFINLT 87
Query: 133 EF 134
EF
Sbjct: 88 EF 89
>UniRef100_O64943 Polcalcin Jun o 2 [Juniperus oxycedrus]
Length = 165
Score = 102 bits (254), Expect = 2e-21
Identities = 54/132 (40%), Positives = 84/132 (62%), Gaps = 3/132 (2%)
Query: 7 EHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALM 66
E V + FD +GDGK+S +EL LR +G ++ E + +E D+DGDGY+SL+E + L
Sbjct: 28 EEVFKKFDANGDGKISGSELADILRSLGSDVGEAEVKAMMEEADADGDGYVSLQEFVDLN 87
Query: 67 EEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGD 126
+G +KDL+ AF+++D + G I+ L L+ +GE +I+E K +I + D +GD
Sbjct: 88 NKGA---SVKDLKNAFKVFDRDCNGSISAAELCHTLESVGEPCTIEESKNIIHNVDKNGD 144
Query: 127 GLLSFDEFITMM 138
GL+S +EF TMM
Sbjct: 145 GLISVEEFQTMM 156
Score = 48.1 bits (113), Expect = 4e-05
Identities = 23/66 (34%), Positives = 37/66 (55%)
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
EQ + +L E F+ +D+ G I+ L +L+ +G E KAM++ D DGDG +S
Sbjct: 21 EQSVHELEEVFKKFDANGDGKISGSELADILRSLGSDVGEAEVKAMMEEADADGDGYVSL 80
Query: 132 DEFITM 137
EF+ +
Sbjct: 81 QEFVDL 86
>UniRef100_Q9FYK2 F21J9.28 [Arabidopsis thaliana]
Length = 186
Score = 101 bits (252), Expect = 3e-21
Identities = 53/133 (39%), Positives = 82/133 (60%), Gaps = 1/133 (0%)
Query: 7 EHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALM 66
E V + FD +GDGK+S EL + +G E+ +E E AI +D GDGY++ EE + L
Sbjct: 39 EAVFKKFDVNGDGKISSKELGAIMTSLGHEVPEEELEKAITEIDRKGDGYINFEEFVELN 98
Query: 67 EEGGEEQK-LKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDG 125
+G ++ L++L++AF +YD + G I+ + L +L+ +G+ SI EC+ MI D DG
Sbjct: 99 TKGMDQNDVLENLKDAFSVYDIDGNGSISAEELHEVLRSLGDECSIAECRKMIGGVDKDG 158
Query: 126 DGLLSFDEFITMM 138
DG + F+EF MM
Sbjct: 159 DGTIDFEEFKIMM 171
Score = 47.4 bits (111), Expect = 7e-05
Identities = 22/63 (34%), Positives = 33/63 (51%)
Query: 13 FDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEEGGEE 72
+D DG+G +S EL + LR +G+E + E I +D DGDG + EE +M G
Sbjct: 118 YDIDGNGSISAEELHEVLRSLGDECSIAECRKMIGGVDKDGDGTIDFEEFKIMMTMGSRR 177
Query: 73 QKL 75
+
Sbjct: 178 DNV 180
Score = 38.9 bits (89), Expect = 0.025
Identities = 17/64 (26%), Positives = 34/64 (52%)
Query: 74 KLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSFDE 133
++++L F+ +D G I+ K L ++ +G +E + I D GDG ++F+E
Sbjct: 34 EIRELEAVFKKFDVNGDGKISSKELGAIMTSLGHEVPEEELEKAITEIDRKGDGYINFEE 93
Query: 134 FITM 137
F+ +
Sbjct: 94 FVEL 97
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.138 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 234,952,680
Number of Sequences: 2790947
Number of extensions: 9416438
Number of successful extensions: 46560
Number of sequences better than 10.0: 3034
Number of HSP's better than 10.0 without gapping: 1852
Number of HSP's successfully gapped in prelim test: 1204
Number of HSP's that attempted gapping in prelim test: 35522
Number of HSP's gapped (non-prelim): 8420
length of query: 139
length of database: 848,049,833
effective HSP length: 115
effective length of query: 24
effective length of database: 527,090,928
effective search space: 12650182272
effective search space used: 12650182272
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 67 (30.4 bits)
Medicago: description of AC141863.5


