

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141436.5 + phase: 0 /pseudo
(364 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
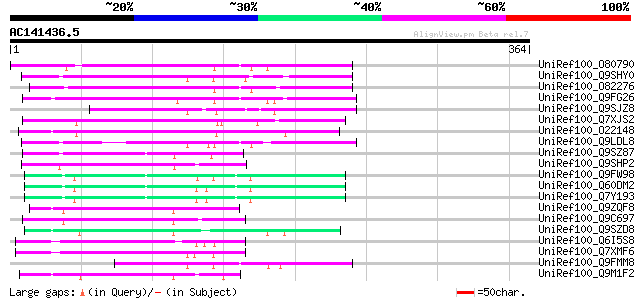
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O80790 Putative non-LTR retroelement reverse transcrip... 109 1e-22
UniRef100_Q9SHY0 F1E22.12 [Arabidopsis thaliana] 109 1e-22
UniRef100_O82276 Putative non-LTR retroelement reverse transcrip... 108 2e-22
UniRef100_Q9FG26 Non-LTR retroelement reverse transcriptase-like... 106 9e-22
UniRef100_Q9SJZ8 Putative non-LTR retroelement reverse transcrip... 96 2e-18
UniRef100_Q7XJS2 Putative non-LTR retroelement reverse transcrip... 92 2e-17
UniRef100_O22148 Putative non-LTR retroelement reverse transcrip... 91 5e-17
UniRef100_Q9LDL8 Hypothetical protein AT4g08830 [Arabidopsis tha... 86 2e-15
UniRef100_Q9SZ87 RNA-directed DNA polymerase-like protein [Arabi... 81 4e-14
UniRef100_Q9SHP2 Putative non-LTR retroelement reverse transcrip... 81 5e-14
UniRef100_Q9FW98 Putative non-LTR retroelement reverse transcrip... 79 2e-13
UniRef100_Q60DM2 F-box domain containing protein [Oryza sativa] 78 3e-13
UniRef100_Q7Y193 Putative reverse transcriptase [Oryza sativa] 78 3e-13
UniRef100_Q9ZQF8 Putative non-LTR retroelement reverse transcrip... 78 4e-13
UniRef100_Q9C697 Reverse transcriptase, putative; 16838-20266 [A... 76 1e-12
UniRef100_Q9SZD8 Hypothetical protein F19B15.120 [Arabidopsis th... 76 2e-12
UniRef100_Q6I5S8 Putative polyprotein [Oryza sativa] 74 8e-12
UniRef100_Q7XMF6 OSJNBa0061G20.3 protein [Oryza sativa] 72 2e-11
UniRef100_Q9FMM8 Non-LTR retroelement reverse transcriptase-like... 72 3e-11
UniRef100_Q9M1F2 Hypothetical protein F9K21.130 [Arabidopsis tha... 71 5e-11
>UniRef100_O80790 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 970
Score = 109 bits (273), Expect = 1e-22
Identities = 77/252 (30%), Positives = 115/252 (45%), Gaps = 17/252 (6%)
Query: 1 QSIFIDESSQDIKIWTHSPTGMYSVKSGYQWLTSNCLS--FAGQTQQETWQWIWKLHLPE 58
+SI D D W + G ++V+S Y+ L G ++ IWKL PE
Sbjct: 599 ESISADALLSDELSWKGTQNGDFTVRSAYELLKPEAEERPLIGSFLKQ----IWKLVAPE 654
Query: 59 NLKHFIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDLLF 118
++ FIWL H + TN R +HL+ ++C C + ES+LH RDCP IW LL
Sbjct: 655 RVRVFIWLVSHMVIMTNVERVRRHLSDIATCSVCNGADESILHVLRDCPAMTPIWQRLLP 714
Query: 119 NKDTNFNSQDFNWWFKHYALALNG--NIFVITCWVLWKARNHKVFNGNDWVC---LSHIY 173
+ N F W F + A +F + W WK R VF G +C L I
Sbjct: 715 QRRQNEFFSQFEWLFTNLDPAKGDWPTLFSMGIWWAWKWRCGDVF-GERKLCRDRLKFIK 773
Query: 174 SLHSDV----VAHISNDVIQKPI-K*VKWIPPLDNIIKVNVDGISLSNHGRSVFGGLIRN 228
+ +V V ++N V + + + ++W P D +K+ DG S + G + G I N
Sbjct: 774 DIAEEVRKAHVGTLNNHVKRARVERMIRWKAPSDRWVKLTTDGASRGHQGLAAASGAILN 833
Query: 229 NNGDWLLGFFMD 240
G+WL GF ++
Sbjct: 834 LQGEWLGGFALN 845
>UniRef100_Q9SHY0 F1E22.12 [Arabidopsis thaliana]
Length = 1055
Score = 109 bits (272), Expect = 1e-22
Identities = 70/252 (27%), Positives = 115/252 (44%), Gaps = 27/252 (10%)
Query: 9 SQDIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQETWQWIWKLHLPENLKHFIWLAM 68
++D W S G +SV+S Y+ LT + + +WK+ +PE +K F+WL
Sbjct: 360 ARDRLSWKFSQDGQFSVRSAYEMLTVD--EVPRPNMASFFNCLWKVRVPERVKTFLWLVG 417
Query: 69 HGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDLLFNKDTN--FNS 126
+ + T R +HL++ + C+ C +ES+LH RDCP L IW ++ + F+
Sbjct: 418 NQAVMTEEERHRRHLSASNVCQVCKGGVESMLHVLRDCPAQLGIWVRVVPQRRQQGFFSK 477
Query: 127 QDFNWWFKHYALALN------GNIFVITCWVLWKARNHKVFNGN-----------DWVCL 169
F W + + IF + W WK R +F N +W
Sbjct: 478 SLFEWLYDNLGDRSGCEDIPWSTIFAVIIWWGWKWRCGNIFGENTKCRDRVKFVKEWAV- 536
Query: 170 SHIYSLHS-DVVAHISNDVIQKPIK*VKWIPPLDNIIKVNVDGISLSNHGRSVFGGLIRN 228
+Y HS +V+ I+ +++ I W+ P +KVN DG S N G + GG++R+
Sbjct: 537 -EVYRAHSGNVLVGITQPRVERMI---GWVSPCVGWVKVNTDGASRGNPGLASAGGVLRD 592
Query: 229 NNGDWLLGFFMD 240
G W GF ++
Sbjct: 593 CTGAWCGGFSLN 604
>UniRef100_O82276 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1231
Score = 108 bits (270), Expect = 2e-22
Identities = 72/240 (30%), Positives = 106/240 (44%), Gaps = 19/240 (7%)
Query: 15 WTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQETWQWIWKLHLPENLKHFIWLAMHGNLPT 74
W + G ++V+S Y L + + IWKL PE ++ FIWL + T
Sbjct: 872 WKGTQDGAFTVRSAYSLLQGDVGD--RPNMGSFFNRIWKLITPERVRVFIWLVSQNVIMT 929
Query: 75 NAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDLL-FNKDTNFNSQDFNWWF 133
N R +HL+ ++ C C + E++LH RDCP IW LL + F SQ W
Sbjct: 930 NVERVRRHLSENAICSVCNGAEETILHVLRDCPAMEPIWRRLLPLRRHHEFFSQSLLEWL 989
Query: 134 KHYALALNG---NIFVITCWVLWKARNHKVFNGNDWVC----------LSHIYSLHSDVV 180
+ G +F + W WK R VF G +C + +H V
Sbjct: 990 FTNMDPVKGIWPTLFGMGIWWAWKWRCCDVF-GERKICRDRLKFIKDMAEEVRRVHVGAV 1048
Query: 181 AHISNDVIQKPIK*VKWIPPLDNIIKVNVDGISLSNHGRSVFGGLIRNNNGDWLLGFFMD 240
+ N V + + ++W P D +K+ DG S NHG + GG IRN G+WL GF ++
Sbjct: 1049 GNRPNGV--RVERMIRWQVPSDGWVKITTDGASRGNHGLAAAGGAIRNGQGEWLGGFALN 1106
>UniRef100_Q9FG26 Non-LTR retroelement reverse transcriptase-like [Arabidopsis
thaliana]
Length = 676
Score = 106 bits (265), Expect = 9e-22
Identities = 69/252 (27%), Positives = 116/252 (45%), Gaps = 23/252 (9%)
Query: 10 QDIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQETWQWIWKLHLPENLKHFIWLAMH 69
+D W + G ++V S Y L + + Q + +W++ +PE + F+WL +
Sbjct: 308 EDKMSWVGTENGRFTVSSAY--LIQSVDEISKQCMSRFFDRVWRVMVPERARIFLWLVGN 365
Query: 70 GNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDL--LFNKDTNFNSQ 127
+ TNA R +H+ C C + ESL+H RDCP + IW + + + F +
Sbjct: 366 QVVLTNAERVRRHMADSDVCPLCKGASESLIHVLRDCPAMMGIWMRVVPVMEQRRFFETS 425
Query: 128 DFNWWFKHYALALNG------NIFVITCWVLWKARNHKVFNGNDWVCLSHIYSLHSDV-- 179
W + + + +F +T W WK R VF G D C + L S V
Sbjct: 426 LLEWMYGNLKERSDSERRSWPTLFALTVWWGWKWRCGYVF-GEDSRCRDRVKFLKSAVAE 484
Query: 180 --VAHIS------NDVIQKPIK*VKWIPPLDNIIKVNVDGISLSNHGRSVFGGLIRNNNG 231
AH++ DV+ + + + W P + + +N DG S N G++ GG+IR+ +G
Sbjct: 485 VEAAHLAANGDAREDVLVERM--IAWRKPAEGWVTMNTDGASHGNPGQATAGGVIRDEHG 542
Query: 232 DWLLGFFMDFVV 243
WL+GF ++ V
Sbjct: 543 SWLVGFALNIGV 554
>UniRef100_Q9SJZ8 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 321
Score = 95.9 bits (237), Expect = 2e-18
Identities = 60/201 (29%), Positives = 98/201 (47%), Gaps = 18/201 (8%)
Query: 57 PENLKHFIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDL 116
PE ++ F+WL + + TN R +HL+ C+ C E++LH RDCP IW L
Sbjct: 3 PERVRVFLWLVVQQVIITNVERYRRHLSDTRVCQICQGGEETILHVLRDCPAMAGIWSRL 62
Query: 117 LFNKDTN--FNSQDFNWWFKHYALALNGN---IFVITCWVLWKARNHKVFNGNDWVCLSH 171
+ F + W +K+ L G+ +FV+ W WK R +F GN C
Sbjct: 63 VPRDQIRQFFTASLLEWIYKN--LRERGSWPTVFVMAVWWGWKWRCGNIFGGNG-KCRDR 119
Query: 172 IYSLHSDVVAHIS--------NDV-IQKPIK*VKWIPPLDNIIKVNVDGISLSNHGRSVF 222
+ + D+ ++ N+V + + + V W+ P D +K+N DG S N G +
Sbjct: 120 VKFI-KDLAEEVAIANAFVKGNEVRVSRVERLVSWVSPEDGWVKLNTDGASRGNPGFATA 178
Query: 223 GGLIRNNNGDWLLGFFMDFVV 243
GG++R++NG W+ GF ++ V
Sbjct: 179 GGVLRDHNGAWIGGFAVNIGV 199
>UniRef100_Q7XJS2 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1344
Score = 92.4 bits (228), Expect = 2e-17
Identities = 65/246 (26%), Positives = 106/246 (42%), Gaps = 22/246 (8%)
Query: 10 QDIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQ------ETWQWIWKLHLPENLKHF 63
+D W HS G+YSVK+GY++L+ Q + + IW LH ++ F
Sbjct: 977 EDSFCWLHSHNGLYSVKTGYEFLSKQVHHRLYQEAKVKPSVNSLFDKIWNLHTAPKIRIF 1036
Query: 64 IWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWD-DLLFNKDT 122
+W A+HG +P + + SD C C E++ H +CPL+ +W L + +
Sbjct: 1037 LWKALHGAIPVEDRLRTRGIRSDDGCLMCDTENETINHILFECPLARQVWAITHLSSAGS 1096
Query: 123 NFNSQDFNWWFKHYALALNGNI-----FVI--TCWVLWKARNHKVFNGNDWVCLSHI--- 172
F++ + + L ++ FV W LWK RN +F G + + +
Sbjct: 1097 EFSNSVYTNMSRLIDLTQQNDLPHHLRFVSPWILWFLWKNRNALLFEGKGSITTTLVDKA 1156
Query: 173 ---YSLHSDVVAHISNDVIQKPIK*VKWIPPLDNIIKVNVDGISLSNHGRSVFGGLIRNN 229
Y H+ ND +K +K KW PPL +K N+ H S ++R++
Sbjct: 1157 YEAYHEWFSAQTHMQND--EKHLKITKWCPPLPGELKCNIGFAWSKQHHFSGASWVVRDS 1214
Query: 230 NGDWLL 235
G LL
Sbjct: 1215 QGKVLL 1220
>UniRef100_O22148 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1374
Score = 90.9 bits (224), Expect = 5e-17
Identities = 64/247 (25%), Positives = 106/247 (42%), Gaps = 23/247 (9%)
Query: 7 ESSQDIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQ-------ETWQWIWKLHLPEN 59
+ ++D W +S +G YSVKSGY W+ + ++ Q+ +Q IWKL +P
Sbjct: 1004 KETRDRFTWEYSRSGHYSVKSGY-WVMTEIINQRNNPQEVLQPSLDPIFQQIWKLDVPPK 1062
Query: 60 LKHFIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDLLFN 119
+ HF+W ++ L + A +HL + SC RC + E++ H CP + W
Sbjct: 1063 IHHFLWRCVNNCLSVASNLAYRHLAREKSCVRCPSHGETVNHLLFKCPFARLTWAISPLP 1122
Query: 120 KDTNFNSQDFNWWFKHYALALNGN---------IFVITCWVLWKARNHKVFNGNDWVCLS 170
+ + H+ L+++ + + W LWK RN VF G ++
Sbjct: 1123 APPGGEWAESLFRNMHHVLSVHKSQPEESDHHALIPWILWRLWKNRNDLVFKGREFTAPQ 1182
Query: 171 HIYSLHSDVVAHISNDVIQKPI------K*VKWIPPLDNIIKVNVDGISLSNHGRSVFGG 224
I D+ A + Q + + VKW PP +K N DG + G G
Sbjct: 1183 VILKATEDMDAWNNRKEPQPQVTSSTRDRCVKWQPPSHGWVKCNTDGAWSKDLGNCGVGW 1242
Query: 225 LIRNNNG 231
++RN+ G
Sbjct: 1243 VLRNHTG 1249
>UniRef100_Q9LDL8 Hypothetical protein AT4g08830 [Arabidopsis thaliana]
Length = 947
Score = 85.5 bits (210), Expect = 2e-15
Identities = 68/256 (26%), Positives = 109/256 (42%), Gaps = 45/256 (17%)
Query: 9 SQDIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQETWQWIWKLHLPENLKHFIWLAM 68
++D W +S G+++VKS Y+ LT + + +W++ E +K F+W
Sbjct: 656 ARDRLSWGYSADGVFTVKSAYRLLTED--HDPRPNMAAFFDRLWRVVALERVKTFLW--- 710
Query: 69 HGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDLLFNKDTN--FNS 126
H+ S C+ C E++LH +DCP IW L+ + + FN
Sbjct: 711 -------------HIGDTSVCQVCKGGDETILHVLKDCPSIAGIWRRLVQVQRSYDFFNG 757
Query: 127 QDFNWWFKHYAL--ALNG----NIFVITCWVLWKARNHKVFNGNDWVC------------ 168
F W + + + A G +F I W WK R VF G C
Sbjct: 758 SLFGWLYVNLGMKNAETGYAWATLFAIVVWWSWKWRCGYVF-GEVGKCRDRVKFFRDLAA 816
Query: 169 -LSHIYSLHSDVVAHISNDVIQKPIK*VKWIPPLDNIIKVNVDGISLSNHGRSVFGGLIR 227
+SH +++HS + + + + V W PP +K+N DG S N G + GG++R
Sbjct: 817 EVSHAHAIHSQ-----NGGLRTRVERLVAWKPPDGEWVKLNTDGASRGNLGLATTGGVLR 871
Query: 228 NNNGDWLLGFFMDFVV 243
+ G W GF +D V
Sbjct: 872 DGIGHWCGGFALDIGV 887
>UniRef100_Q9SZ87 RNA-directed DNA polymerase-like protein [Arabidopsis thaliana]
Length = 1274
Score = 81.3 bits (199), Expect = 4e-14
Identities = 52/165 (31%), Positives = 80/165 (47%), Gaps = 13/165 (7%)
Query: 10 QDIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQETWQW-IWKLHLPENLKHFIWLAM 68
QD +W +G Y+ K+GY N SF WQ IWK+H +KHF+W AM
Sbjct: 934 QDSLVWLPVKSGEYTTKTGYALAKLN--SFPASQLDFNWQKNIWKIHTSPKVKHFLWKAM 991
Query: 69 HGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWD--DLLFN-KDTNFN 125
G LP + +++ ++ +CKRCG + ES LH CP + +W+ +LFN + +
Sbjct: 992 KGALPVGEALSRRNIEAEVTCKRCGQT-ESSLHLMLLCPYAKKVWELAPVLFNPSEATHS 1050
Query: 126 SQDFNWWFKHYALAL------NGNIFVITCWVLWKARNHKVFNGN 164
S +AL + ++ W LWKARN +F+ +
Sbjct: 1051 SVALLLVDAKRMVALPPTGLGSAPLYPWLLWHLWKARNRLIFDNH 1095
>UniRef100_Q9SHP2 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 773
Score = 80.9 bits (198), Expect = 5e-14
Identities = 50/174 (28%), Positives = 77/174 (43%), Gaps = 18/174 (10%)
Query: 9 SQDIKIWTHSPTGMYSVKSGYQWLT-------SNCLSFAGQTQQETWQWIWKLHLPENLK 61
+ D W ++ G YSVKSGY L ++ S + Q + IWK + P +K
Sbjct: 452 ANDAITWIYTHDGNYSVKSGYHLLRKLSQQQHASLPSPNEVSAQTVFTNIWKQNAPPKIK 511
Query: 62 HFIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWD------- 114
HF W + H LPT + L +D +C+RCG + E + H C +S IW+
Sbjct: 512 HFWWRSAHNALPTAGNLKRRRLITDDTCQRCGEASEDVNHLLFQCRVSKEIWEQAHIKLC 571
Query: 115 --DLLFNKDTNFNSQDFNWWFKHYALALNGNIFVITCWVLWKARNHKVFNGNDW 166
D L + N N + + + + ++F W +WK RN +FN W
Sbjct: 572 PGDSLMSNSFNQNLESIQ--KLNQSARKDVSLFPFIGWRIWKMRNDLIFNNKRW 623
>UniRef100_Q9FW98 Putative non-LTR retroelement reverse transcriptase [Oryza sativa]
Length = 1382
Score = 79.0 bits (193), Expect = 2e-13
Identities = 61/251 (24%), Positives = 98/251 (38%), Gaps = 29/251 (11%)
Query: 11 DIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQ------------QETWQWIWKLHLPE 58
D W H G+YSV+S Y S FA Q+ Q+ W+ +WK++ P
Sbjct: 1015 DFASWPHDKLGLYSVRSAYNLARSEAF-FADQSNSGRGMASRLLESQKDWKGLWKINAPG 1073
Query: 59 NLKHFIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDLLF 118
+K +W A H L T +H+ S C C +++ H F CP + IW+++
Sbjct: 1074 KMKITLWRAAHECLATGFQLRRRHIPSTDGCVFCNRD-DTVEHVFLFCPFAAQIWEEIKG 1132
Query: 119 NKDTNFNSQDFN----WWFKHYALALN--GNIFVITCWVLWKARNHKVFNGNDWV----- 167
F+ W F + + +T W +W+ARN+ N N V
Sbjct: 1133 KCAVKLGRNGFSTMRQWIFDFLKRGSSHANTLLAVTFWHIWEARNN-TKNNNGTVHPQRV 1191
Query: 168 ---CLSHIYSLHSDVVAHISNDVIQKPIK*VKWIPPLDNIIKVNVDGISLSNHGRSVFGG 224
LS++ + + +W PP ++ +N D S+ G
Sbjct: 1192 VIKILSYVDMILKHNTKTVDGQRGGNTQAIPRWQPPPASVWMINSDAAIFSSSRTMGVGA 1251
Query: 225 LIRNNNGDWLL 235
LIR+N G L+
Sbjct: 1252 LIRDNTGKCLV 1262
>UniRef100_Q60DM2 F-box domain containing protein [Oryza sativa]
Length = 855
Score = 78.2 bits (191), Expect = 3e-13
Identities = 60/251 (23%), Positives = 98/251 (38%), Gaps = 29/251 (11%)
Query: 11 DIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQ------------QETWQWIWKLHLPE 58
D W H G+YSV+S Y + S FA Q+ Q+ W+ +WK++ P
Sbjct: 488 DFASWPHEKLGLYSVRSAYNMVRSEAF-FADQSNSGRGMASRLLESQKDWKGLWKINAPG 546
Query: 59 NLKHFIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDLLF 118
+K +W A H L T +H+ S C C +++ H F CP + IW+++
Sbjct: 547 KMKIILWRAAHECLATGFQLPRRHIPSTDGCVFCNRD-DTVEHVFLFCPFAAQIWEEIKR 605
Query: 119 NKDTNFNSQDF----NWWFKHY--ALALNGNIFVITCWVLWKARNHKVFNGNDWV----- 167
F +W F + + +T W +W+ARN+ N N V
Sbjct: 606 KCAVKLGRNGFSTMRHWIFDFLKRGSSHTNTLLAVTFWHIWEARNN-TKNNNGTVHPQRV 664
Query: 168 ---CLSHIYSLHSDVVAHISNDVIQKPIK*VKWIPPLDNIIKVNVDGISLSNHGRSVFGG 224
LS++ + + + +W P + +N D S+ G
Sbjct: 665 VIKILSYVDMILKHNTKTVDGQRGENTQAIPRWQPLPAGVWMINSDAAIFSSSRTMGVGA 724
Query: 225 LIRNNNGDWLL 235
LIR+N G L+
Sbjct: 725 LIRDNTGKCLV 735
>UniRef100_Q7Y193 Putative reverse transcriptase [Oryza sativa]
Length = 404
Score = 78.2 bits (191), Expect = 3e-13
Identities = 60/251 (23%), Positives = 98/251 (38%), Gaps = 29/251 (11%)
Query: 11 DIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQ------------QETWQWIWKLHLPE 58
D W H G+YSV+S Y + S FA Q+ Q+ W+ +WK++ P
Sbjct: 37 DFASWPHEKLGLYSVRSAYNMVRSEAF-FADQSNSGRGMASRLLESQKDWKGLWKINAPG 95
Query: 59 NLKHFIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDLLF 118
+K +W A H L T +H+ S C C +++ H F CP + IW+++
Sbjct: 96 KMKIILWRAAHECLATGFQLPRRHIPSTDGCVFCNRD-DTVEHVFLFCPFAAQIWEEIKR 154
Query: 119 NKDTNFNSQDF----NWWFKHY--ALALNGNIFVITCWVLWKARNHKVFNGNDWV----- 167
F +W F + + +T W +W+ARN+ N N V
Sbjct: 155 KCAVKLGRNGFSTMRHWIFDFLKRGSSHTNTLLAVTFWHIWEARNN-TKNNNGTVHPQRV 213
Query: 168 ---CLSHIYSLHSDVVAHISNDVIQKPIK*VKWIPPLDNIIKVNVDGISLSNHGRSVFGG 224
LS++ + + + +W P + +N D S+ G
Sbjct: 214 VIKILSYVDMILKHNTKTVDGQRGENTQAIPRWQPLPAGVWMINSDAAIFSSSRTMGVGA 273
Query: 225 LIRNNNGDWLL 235
LIR+N G L+
Sbjct: 274 LIRDNTGKCLV 284
>UniRef100_Q9ZQF8 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1225
Score = 77.8 bits (190), Expect = 4e-13
Identities = 50/162 (30%), Positives = 76/162 (46%), Gaps = 16/162 (9%)
Query: 15 WTHSPTGMYSVKSGYQWLTSNC------LSFAGQTQQETWQWIWKLHLPENLKHFIWLAM 68
W+ + +G Y+VKSGY W + L F G + +WK+ KHF W +
Sbjct: 885 WSFTKSGNYTVKSGY-WAARDLSRPTCDLPFQGPSVSALQAQVWKIKTTRKFKHFEWQCL 943
Query: 69 HGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIW-------DDLLFNKD 121
G L TN +H+ ++ C RCGA ES+ H CP S IW + +F ++
Sbjct: 944 SGCLATNQRLFSRHIGTEKVCPRCGAEEESINHLLFLCPPSRQIWALSPIPSSEYIFPRN 1003
Query: 122 TNFNSQDFNW-WFKHYALALN-GNIFVITCWVLWKARNHKVF 161
+ F + DF K + +A + IF W +WK+RN +F
Sbjct: 1004 SLFYNFDFLLSRGKEFDIAEDIMEIFPWILWYIWKSRNRFIF 1045
>UniRef100_Q9C697 Reverse transcriptase, putative; 16838-20266 [Arabidopsis thaliana]
Length = 1142
Score = 76.3 bits (186), Expect = 1e-12
Identities = 47/166 (28%), Positives = 72/166 (43%), Gaps = 12/166 (7%)
Query: 10 QDIKIWTHSPTGMYSVKSGYQWLTSNC---LSFAGQTQQETWQWIWKLHLPENLKHFIWL 66
+D W + G Y+VKSGY + + G +IWK+ P L+HF+W
Sbjct: 784 EDTLGWHFTKAGNYTVKSGYHTARLDLNEGTTLIGPDLTTLKAYIWKVQCPPKLRHFLWQ 843
Query: 67 AMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIW-------DDLLFN 119
+ G +P + + + D C CGAS ES+ HT C + IW +F
Sbjct: 844 ILSGCVPVSENLRKRGILCDKGCVSCGASEESINHTLFQCHPARQIWALSQIPTAPGIFP 903
Query: 120 KDTNFNSQDFNWWFKHYALALNGNIFVITCWVLWKARNHKVFNGND 165
++ F + D +W ++ + W +WKARN KVF D
Sbjct: 904 SNSIFTNLDHLFW--RIPSGVDSAPYPWIIWYIWKARNEKVFENVD 947
>UniRef100_Q9SZD8 Hypothetical protein F19B15.120 [Arabidopsis thaliana]
Length = 575
Score = 75.9 bits (185), Expect = 2e-12
Identities = 61/252 (24%), Positives = 101/252 (39%), Gaps = 37/252 (14%)
Query: 11 DIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQET-------WQWIWKLHLPENLKHF 63
D W ++ +G Y+VKSGY W+ + ++ Q+ + +Q IWK ++HF
Sbjct: 211 DSYTWDYTSSGDYTVKSGY-WVLTQIINKRSSPQEVSEPSLNPIYQKIWKSQTSPKIQHF 269
Query: 64 IWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIW---------- 113
+W + +LP A +HL+ +S+C RC + E++ H C + W
Sbjct: 270 LWKCLSNSLPVAGALAYRHLSKESACIRCPSCKETVNHLLFKCTFARLTWAISSIPIPLG 329
Query: 114 ----DDLLFNKDTNFNSQDFN-WWFKHYALALNGNIFVITCWVLWKARNHKVFNGNDWVC 168
D + N FN + N W K + W LWK RN VF G ++
Sbjct: 330 GEWADSIYVNLYWVFNLGNGNPQWEK------ASQLVPWLLWRLWKNRNELVFRGREFNA 383
Query: 169 LSHIYSLHSDV----VAHISNDVIQKP----IK*VKWIPPLDNIIKVNVDGISLSNHGRS 220
+ D+ + + KP +W PP +K N D ++ R
Sbjct: 384 QEVLRRAEDDLEEWRIRTEAESCGTKPQVNRSSCGRWRPPPHQWVKCNTDATWNRDNERC 443
Query: 221 VFGGLIRNNNGD 232
G ++RN G+
Sbjct: 444 GIGWVLRNEKGE 455
>UniRef100_Q6I5S8 Putative polyprotein [Oryza sativa]
Length = 1419
Score = 73.6 bits (179), Expect = 8e-12
Identities = 44/177 (24%), Positives = 77/177 (42%), Gaps = 26/177 (14%)
Query: 5 IDESSQDIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQETWQWIWKLHLPENLKHFI 64
I E ++D +W S +G+Y+ KS Y + F G+T ++ + +W+ P K F
Sbjct: 247 IHEGAEDTILWRWSASGLYTAKSAY------VMQFMGKTFSDSTELVWETWAPGKHKFFA 300
Query: 65 WLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDLLFNKDTNF 124
WL + T + ++ C+ C ++E++ H F +CP + +W D+ F
Sbjct: 301 WLLAQNRIWTADHLQCRGWPNNYFCQLCFRNLETVQHLFMECPFTQQVWRDI----GRKF 356
Query: 125 NSQDF----------NWWFKH----YALALNG--NIFVITCWVLWKARNHKVFNGND 165
Q F NWW + L G + + T W +W RN ++F G +
Sbjct: 357 RPQRFLHVTTDNTLLNWWREQTDEAEKLQQKGLRSAVLTTLWEIWMERNDRIFRGKE 413
>UniRef100_Q7XMF6 OSJNBa0061G20.3 protein [Oryza sativa]
Length = 1463
Score = 72.0 bits (175), Expect = 2e-11
Identities = 42/174 (24%), Positives = 72/174 (41%), Gaps = 19/174 (10%)
Query: 5 IDESSQDIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQETWQWIWKLHLPENLKHFI 64
++E +D W + TG Y+ KS Y FAG+ IWK P K F
Sbjct: 1266 LEEGVEDTITWRWTNTGKYTAKSAY------LAQFAGREDSRAATLIWKTWAPSKCKKFA 1319
Query: 65 WLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDLLFNKDTN- 123
WL + + T+ + ++ C+ C ++E H F+DCP + IWD ++ + N
Sbjct: 1320 WLVLQNRIWTSDRLQLRQWPNNYFCQLCYRNLEMAQHLFKDCPFTREIWDKIMARMNHNQ 1379
Query: 124 ----FNSQD--FNWWFKHYALALN------GNIFVITCWVLWKARNHKVFNGND 165
N+ + +WW ++ ++ W +W RN +VF +
Sbjct: 1380 PFPVHNTDESLVDWWESRTKQQSKCQSKGMQSLHILLSWEIWCERNRRVFKDKE 1433
>UniRef100_Q9FMM8 Non-LTR retroelement reverse transcriptase-like protein
[Arabidopsis thaliana]
Length = 308
Score = 71.6 bits (174), Expect = 3e-11
Identities = 50/183 (27%), Positives = 83/183 (45%), Gaps = 17/183 (9%)
Query: 74 TNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIWDDLLFNKDTN--FNSQDFNW 131
T+ R +HL++ + + +ES+LH FRDCP L IW + + F+ F W
Sbjct: 2 TDEERHRRHLSASNVSQVYIGGVESVLHVFRDCPAQLGIWVRFVPRRRQQGFFSKSLFEW 61
Query: 132 WFKHYALALN------GNIFVITCWVLWKARNHKVFNGNDWVCLSHIYSLHSDVV----A 181
+ + + IF + W WK R +F G + C + + VV A
Sbjct: 62 LYDNLCDRSSCEDIPWSTIFAVIIWWGWKWRCSNIF-GENTKCRDRVKFVKEWVVEVYRA 120
Query: 182 HISNDVI----QKPIK*VKWIPPLDNIIKVNVDGISLSNHGRSVFGGLIRNNNGDWLLGF 237
H+ N ++ + + + W+ P +KVN DG S N G + GG++R+ G W GF
Sbjct: 121 HLGNALVGSTQPRVERLIGWVLPCVGWVKVNTDGASRGNPGLASAGGVLRDCEGAWCGGF 180
Query: 238 FMD 240
++
Sbjct: 181 SLN 183
>UniRef100_Q9M1F2 Hypothetical protein F9K21.130 [Arabidopsis thaliana]
Length = 851
Score = 70.9 bits (172), Expect = 5e-11
Identities = 51/172 (29%), Positives = 74/172 (42%), Gaps = 20/172 (11%)
Query: 8 SSQDIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQETW------QWIWKLHLPENLK 61
S D IW+++ TG Y+V+SGY WL+++ S T + IW L + LK
Sbjct: 619 SKTDRLIWSYNSTGDYTVRSGY-WLSTHDPSNTIPTMAKPHGSVDLKTKIWNLPIMPKLK 677
Query: 62 HFIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTFRDCPLSLAIW---DDLLF 118
HF+W + LPT + + D C RC ES+ H CP + W D L+
Sbjct: 678 HFLWRILSKALPTTDRLTTRGMRIDPGCPRCRRENESINHALFTCPFATMAWRLSDTPLY 737
Query: 119 NKDTNFNSQDFNWWFKHYALALNGNIFVIT--------CWVLWKARNHKVFN 162
N+ + N + L L + W +WKARN+ VFN
Sbjct: 738 RSSILSNNIEDN--ISNILLLLQNTTITDSQKLIPFWLLWRIWKARNNVVFN 787
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.337 0.147 0.506
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 608,875,567
Number of Sequences: 2790947
Number of extensions: 25138376
Number of successful extensions: 87925
Number of sequences better than 10.0: 188
Number of HSP's better than 10.0 without gapping: 108
Number of HSP's successfully gapped in prelim test: 80
Number of HSP's that attempted gapping in prelim test: 87586
Number of HSP's gapped (non-prelim): 235
length of query: 364
length of database: 848,049,833
effective HSP length: 129
effective length of query: 235
effective length of database: 488,017,670
effective search space: 114684152450
effective search space used: 114684152450
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 75 (33.5 bits)
Medicago: description of AC141436.5


