

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141322.2 - phase: 0
(233 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
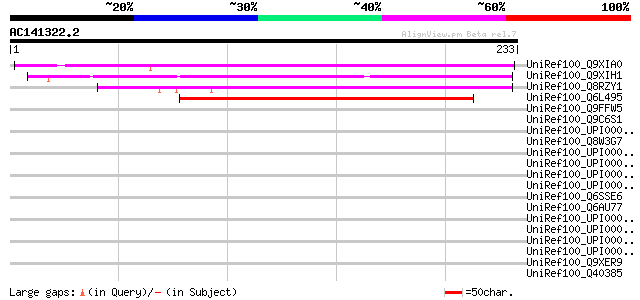
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9XIA0 F13F21.24 protein [Arabidopsis thaliana] 191 2e-47
UniRef100_Q9XIH1 Hypothetical protein At2g16190 [Arabidopsis tha... 168 1e-40
UniRef100_Q8RZY1 P0034C09.31 protein [Oryza sativa] 152 5e-36
UniRef100_Q6L495 Hypothetical protein P0554F08.11 [Oryza sativa] 139 6e-32
UniRef100_Q9FFW5 Similarity to protein kinase [Arabidopsis thali... 45 0.002
UniRef100_Q9C6S1 Hypothetical protein F5M6.18 [Arabidopsis thali... 44 0.003
UniRef100_UPI0000222289 UPI0000222289 UniRef100 entry 44 0.004
UniRef100_Q8W3G7 Putative hydroxyproline-rich glycoprotein [Oryz... 44 0.004
UniRef100_UPI00003617D5 UPI00003617D5 UniRef100 entry 43 0.006
UniRef100_UPI0000289AD4 UPI0000289AD4 UniRef100 entry 43 0.006
UniRef100_UPI0000361A97 UPI0000361A97 UniRef100 entry 43 0.008
UniRef100_UPI0000361A96 UPI0000361A96 UniRef100 entry 43 0.008
UniRef100_Q6SSE6 Plus agglutinin [Chlamydomonas reinhardtii] 43 0.008
UniRef100_Q6AU77 Hypothetical protein OSJNBa0014C03.6 [Oryza sat... 43 0.008
UniRef100_UPI00003617D6 UPI00003617D6 UniRef100 entry 42 0.010
UniRef100_UPI000024B6BB UPI000024B6BB UniRef100 entry 42 0.017
UniRef100_UPI00002B1219 UPI00002B1219 UniRef100 entry 42 0.017
UniRef100_UPI0000265027 UPI0000265027 UniRef100 entry 42 0.017
UniRef100_Q9XER9 Transcription factor-like protein [Arabidopsis ... 42 0.017
UniRef100_Q40385 Pistil extensin-like protein [Nicotiana alata] 42 0.017
>UniRef100_Q9XIA0 F13F21.24 protein [Arabidopsis thaliana]
Length = 331
Score = 191 bits (484), Expect = 2e-47
Identities = 103/231 (44%), Positives = 138/231 (59%), Gaps = 4/231 (1%)
Query: 3 DSEQPTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAP 62
D P+ +PL M FQ+ PN P L P + P S+ PPP + V
Sbjct: 96 DPPPPSHQIPLWMSNYFQQT---PNPPQLVTHFFPPSGLAPPSSNLTPPPVKRPVTGSVR 152
Query: 63 V-RSRRRGPPRNGPIPTPFIWATDRRAKIHSLNHLLQNRIFNITGDVKCKSCQTKFQMSF 121
+ RSR ++ I PF WAT+RR +I SL +L N+I ITG+V+C+ C+ +Q+S+
Sbjct: 153 IYRSRSTVSKKSDTISPPFPWATNRRGEIQSLEYLESNQITTITGEVQCRHCEKVYQVSY 212
Query: 122 DLVAKFDVISQYLVTNFNTMHDRAPETLMYPRLMKCVHCNQENCVKPVIAEKKKNINWLF 181
+L +F + ++ +T M DRA + YP +C C +E VKPVIAE+K INWLF
Sbjct: 213 NLRERFAEVVKFYLTEKRKMRDRAHKDWAYPEQRRCELCGREKAVKPVIAERKSQINWLF 272
Query: 182 LLLGQMLGCCKLEQLKYFCKYNSHHRTGAKNRVLYLTYLELCKQLDPSLPL 232
LLLGQ LG C LEQLK FCK++ +HRTGAK+RVLYLTY+ LCK L P L
Sbjct: 273 LLLGQTLGFCTLEQLKNFCKHSKNHRTGAKDRVLYLTYMGLCKMLQPKSDL 323
>UniRef100_Q9XIH1 Hypothetical protein At2g16190 [Arabidopsis thaliana]
Length = 303
Score = 168 bits (425), Expect = 1e-40
Identities = 94/226 (41%), Positives = 129/226 (56%), Gaps = 7/226 (3%)
Query: 9 QSLPLPML---TPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRS 65
Q++P P + TP + +PP Q + T+ + P PP Q ++ PV
Sbjct: 79 QAVPPPNVSVRTPLPYQPSEEVLPPPQLNQVA-TVALATPRRGRPPGGQARRNSKRPVAG 137
Query: 66 RRRGPPRNGPIPTPFIWATDRRAKIHSLNHLLQNRIFNITGDVKCKSCQTKFQMSFDLVA 125
R +P P+ WAT + KI S L N I I+G V CK+C + ++L
Sbjct: 138 VERNVGDREIVP-PYPWATKKPGKIQSFRDLSSNNINVISGQVHCKTCDRTDTVEYNLEE 196
Query: 126 KFDVISQYLVTNFNTMHDRAPETLMYPRLMKCVHCNQENCVKPVIAEKKKNINWLFLLLG 185
KF + Y+ N M RAP + P+L+ C C E +KPV++E+K+ INWLFLLLG
Sbjct: 197 KFSELYGYIKVNKEEMRHRAPGSWSTPKLIPCRTCKSE--MKPVMSERKEEINWLFLLLG 254
Query: 186 QMLGCCKLEQLKYFCKYNSHHRTGAKNRVLYLTYLELCKQLDPSLP 231
QMLGCC L+QL+YFC+ NS HRTG+K+RV+Y+TYL LCKQLDP P
Sbjct: 255 QMLGCCTLDQLRYFCQLNSKHRTGSKDRVVYITYLSLCKQLDPEGP 300
>UniRef100_Q8RZY1 P0034C09.31 protein [Oryza sativa]
Length = 316
Score = 152 bits (385), Expect = 5e-36
Identities = 83/199 (41%), Positives = 111/199 (55%), Gaps = 8/199 (4%)
Query: 41 NIPQPSHPPPPPPQDVVDNHAPVRSRR-RGPPRNGP-----IPTPFIWATDRRAKIH--S 92
N P+ S PP + ++N +S R RG NG + T F W T + +
Sbjct: 115 NSPRSSLLAPPSNRRRLNNPDEGQSPRGRGEEANGDNGVLVMATSFPWVTSADLPVLHCT 174
Query: 93 LNHLLQNRIFNITGDVKCKSCQTKFQMSFDLVAKFDVISQYLVTNFNTMHDRAPETLMYP 152
L +L I ++ G C C + +++DL AKF + Y+ N + M DRAPE M P
Sbjct: 175 LESMLLKGITSVEGKATCNRCSAEVPIAYDLDAKFREVRDYVAANIHIMDDRAPEHWMCP 234
Query: 153 RLMKCVHCNQENCVKPVIAEKKKNINWLFLLLGQMLGCCKLEQLKYFCKYNSHHRTGAKN 212
RL C C ++ C+ P I +K+ INWLFL LGQMLGCC LE LK+FCK +H TGAK
Sbjct: 235 RLPDCGSCGKKACMWPQIPNEKREINWLFLFLGQMLGCCTLEGLKFFCKNTKNHCTGAKT 294
Query: 213 RVLYLTYLELCKQLDPSLP 231
RVLY Y+E+C+QLDP P
Sbjct: 295 RVLYYAYIEMCRQLDPQGP 313
>UniRef100_Q6L495 Hypothetical protein P0554F08.11 [Oryza sativa]
Length = 315
Score = 139 bits (350), Expect = 6e-32
Identities = 63/135 (46%), Positives = 87/135 (63%)
Query: 79 PFIWATDRRAKIHSLNHLLQNRIFNITGDVKCKSCQTKFQMSFDLVAKFDVISQYLVTNF 138
P+ WAT+ AK HSL L + I NI G+ +C+ C T+ + +++ KF +S Y N+
Sbjct: 172 PYPWATNEVAKHHSLVELARRDIININGEARCRRCDTRKMIVYNIATKFREVSDYFRQNY 231
Query: 139 NTMHDRAPETLMYPRLMKCVHCNQENCVKPVIAEKKKNINWLFLLLGQMLGCCKLEQLKY 198
M+DRA M P + C C E C++PVIA +K+ INWLFLLLG+ LG C L+QLKY
Sbjct: 232 QHMNDRAQARWMNPVVPNCDSCGHERCMRPVIAAEKERINWLFLLLGETLGLCTLDQLKY 291
Query: 199 FCKYNSHHRTGAKNR 213
FC + + HRTGAK+R
Sbjct: 292 FCAHTNRHRTGAKDR 306
>UniRef100_Q9FFW5 Similarity to protein kinase [Arabidopsis thaliana]
Length = 681
Score = 45.1 bits (105), Expect = 0.002
Identities = 28/71 (39%), Positives = 35/71 (48%), Gaps = 5/71 (7%)
Query: 12 PLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQD---VVDNHAPVRSRRR 68
PLP+L+P + PPLQ QP T + P P PPP PPQ VV + P
Sbjct: 6 PLPILSPPSSNSSTTAPPPLQTQPT--TPSAPPPVTPPPSPPQSPPPVVSSSPPPPVVSS 63
Query: 69 GPPRNGPIPTP 79
PP + P P+P
Sbjct: 64 PPPSSSPPPSP 74
>UniRef100_Q9C6S1 Hypothetical protein F5M6.18 [Arabidopsis thaliana]
Length = 1201
Score = 43.9 bits (102), Expect = 0.003
Identities = 25/73 (34%), Positives = 35/73 (47%), Gaps = 8/73 (10%)
Query: 7 PTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRSR 66
P++S+P P+ P P PP P P + +IP PS PPPPPP + ++
Sbjct: 583 PSRSIPPPLAQP------PPPRPPPPPPPPPSSRSIPSPSAPPPPPPPP--PSFGSTGNK 634
Query: 67 RRGPPRNGPIPTP 79
R+ P P P P
Sbjct: 635 RQAQPPPPPPPPP 647
Score = 37.7 bits (86), Expect = 0.24
Identities = 21/54 (38%), Positives = 24/54 (43%), Gaps = 8/54 (14%)
Query: 26 PNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRSRRRGPPRNGPIPTP 79
P PPL + +P L P P PPPPPP P S R P + P P P
Sbjct: 577 PPPPPLPSRSIPPPLAQPPPPRPPPPPP--------PPPSSRSIPSPSAPPPPP 622
Score = 37.0 bits (84), Expect = 0.41
Identities = 25/76 (32%), Positives = 32/76 (41%), Gaps = 3/76 (3%)
Query: 6 QPTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQ--PSHPPPPPPQDVVDNHAPV 63
QP P P L F+ P+ PP P PL + PS PPPPPP N P+
Sbjct: 503 QPPPPPPPPPLFMSTTSFS-PSQPPPPPPPPPLFTSTTSFSPSQPPPPPPLPSFSNRDPL 561
Query: 64 RSRRRGPPRNGPIPTP 79
+ + + P P P
Sbjct: 562 TTLHQPINKTPPPPPP 577
Score = 34.3 bits (77), Expect = 2.7
Identities = 25/76 (32%), Positives = 29/76 (37%), Gaps = 10/76 (13%)
Query: 4 SEQPTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPV 63
S P+Q P P L F N PL P+ P PPPPPP + P
Sbjct: 540 SFSPSQPPPPPPLPSFS------NRDPLTTLHQPINKTPP----PPPPPPPPLPSRSIPP 589
Query: 64 RSRRRGPPRNGPIPTP 79
+ PPR P P P
Sbjct: 590 PLAQPPPPRPPPPPPP 605
Score = 33.5 bits (75), Expect = 4.6
Identities = 24/85 (28%), Positives = 30/85 (35%), Gaps = 3/85 (3%)
Query: 7 PTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRSR 66
P P P+ T PP PL ++ PS PPPPPP + S
Sbjct: 484 PPPPPPPPLFTSTTSFSPSQPPPPPPPPPLFMSTTSFSPSQPPPPPPPPPLFTSTTSFSP 543
Query: 67 RRGPPRNGPIPTPFIWATDRRAKIH 91
+ PP P P P D +H
Sbjct: 544 SQPPP---PPPLPSFSNRDPLTTLH 565
>UniRef100_UPI0000222289 UPI0000222289 UniRef100 entry
Length = 198
Score = 43.5 bits (101), Expect = 0.004
Identities = 27/74 (36%), Positives = 34/74 (45%), Gaps = 7/74 (9%)
Query: 7 PTQSLPLPMLTPFQRMFAHPN-IPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRS 65
P SLP P+ P + P IPP LP + IP P P PPPP ++ +P+
Sbjct: 57 PIPSLPAPIPPPPAMILLPPAPIPPPPAPILPPSSPIPPPPAPIPPPPAPILPPSSPI-- 114
Query: 66 RRRGPPRNGPIPTP 79
PP PIP P
Sbjct: 115 ----PPPPAPIPPP 124
Score = 35.8 bits (81), Expect = 0.92
Identities = 25/71 (35%), Positives = 31/71 (43%), Gaps = 7/71 (9%)
Query: 7 PTQSLPLPMLTPFQRMFAHPN-IPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRS 65
P P P+L P + P IPP LP + IP P P PPPP + AP+
Sbjct: 78 PIPPPPAPILPPSSPIPPPPAPIPPPPAPILPPSSPIPPPPAPIPPPPAMISPPPAPI-- 135
Query: 66 RRRGPPRNGPI 76
PP+ PI
Sbjct: 136 ----PPQPAPI 142
>UniRef100_Q8W3G7 Putative hydroxyproline-rich glycoprotein [Oryza sativa]
Length = 464
Score = 43.5 bits (101), Expect = 0.004
Identities = 35/108 (32%), Positives = 50/108 (45%), Gaps = 18/108 (16%)
Query: 12 PLPMLTPFQRMFAHPN--IPPLQQQPLPLTLNIPQPSHPPP-------PPP----QDVVD 58
PLP P + HP+ +PP QQP P ++ + Q PPP PPP Q
Sbjct: 237 PLPASPPQPQHQHHPHRPLPPQHQQPKPPSMRL-QKIRPPPISTPVARPPPVHNHQIPNP 295
Query: 59 NHAPVRSRRRGP-PRNGPIPTPFIWATDRRAKIHSLNHLLQNRIFNIT 105
NH P R P P P+P P +WA + + + +L+N +F+ T
Sbjct: 296 NHNPAFHRPPPPQPMPMPMPGPPVWAD---SPVTAYMRILENSLFSAT 340
>UniRef100_UPI00003617D5 UPI00003617D5 UniRef100 entry
Length = 707
Score = 43.1 bits (100), Expect = 0.006
Identities = 25/61 (40%), Positives = 32/61 (51%), Gaps = 9/61 (14%)
Query: 20 QRMFAHPNIPPLQQQPLPLTLN-IP--QPSHPPPPPPQDVVDNHAPVRSRRRGPPRNGPI 76
Q++ P IPP QQQP+P +P QP PPPPPPQ PV ++ PP +
Sbjct: 325 QQIMPQPQIPPSQQQPVPPQAQPVPSQQPPPPPPPPPQQ------PVPTQPAVPPAQPNM 378
Query: 77 P 77
P
Sbjct: 379 P 379
>UniRef100_UPI0000289AD4 UPI0000289AD4 UniRef100 entry
Length = 300
Score = 43.1 bits (100), Expect = 0.006
Identities = 21/48 (43%), Positives = 23/48 (47%)
Query: 32 QQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRSRRRGPPRNGPIPTP 79
Q PLP TL+ +P PPPPP P SR R P PIP P
Sbjct: 25 QSSPLPRTLSTSRPPEPPPPPSGRPKGEDFPPASRARAPSTPTPIPPP 72
>UniRef100_UPI0000361A97 UPI0000361A97 UniRef100 entry
Length = 411
Score = 42.7 bits (99), Expect = 0.008
Identities = 28/76 (36%), Positives = 31/76 (39%), Gaps = 5/76 (6%)
Query: 4 SEQPTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPV 63
S P LPLP+ P P PPL P P P PS PPPPPP +P
Sbjct: 21 SPPPPLPLPLPLPLPLPLPPPPPPPPPLPPPPPPPPPPPPSPSPPPPPPPPP-----SPP 75
Query: 64 RSRRRGPPRNGPIPTP 79
PP + P P P
Sbjct: 76 PPPPPPPPPSPPPPPP 91
Score = 39.7 bits (91), Expect = 0.064
Identities = 22/54 (40%), Positives = 24/54 (43%), Gaps = 7/54 (12%)
Query: 26 PNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRSRRRGPPRNGPIPTP 79
P+ PP PLPL L +P P PPPPPP P PP P P P
Sbjct: 20 PSPPPPLPLPLPLPLPLPLPPPPPPPPPLPPPPPPPP-------PPPPSPSPPP 66
Score = 37.7 bits (86), Expect = 0.24
Identities = 26/76 (34%), Positives = 28/76 (36%), Gaps = 8/76 (10%)
Query: 4 SEQPTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPV 63
S P LPLP+ P P PP P P P PS PPPPP P
Sbjct: 19 SPSPPPPLPLPLPLPLPLPLPPPPPPPPPLPPPPPPPPPPPPSPSPPPPP--------PP 70
Query: 64 RSRRRGPPRNGPIPTP 79
PP P P+P
Sbjct: 71 PPSPPPPPPPPPPPSP 86
Score = 34.7 bits (78), Expect = 2.1
Identities = 25/75 (33%), Positives = 28/75 (37%), Gaps = 2/75 (2%)
Query: 7 PTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHP--PPPPPQDVVDNHAPVR 64
P P P P P +P PLPL L P P P PPPPP +P
Sbjct: 5 PPPPAPAPPPPPPPSPSPPPPLPLPLPLPLPLPLPPPPPPPPPLPPPPPPPPPPPPSPSP 64
Query: 65 SRRRGPPRNGPIPTP 79
PP + P P P
Sbjct: 65 PPPPPPPPSPPPPPP 79
Score = 32.7 bits (73), Expect = 7.8
Identities = 19/52 (36%), Positives = 19/52 (36%)
Query: 26 PNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRSRRRGPPRNGPIP 77
P PP P P P PS PPPPPP P PP P P
Sbjct: 65 PPPPPPPPSPPPPPPPPPPPSPPPPPPPSSSPPPPPPPSPPPPPPPPPAPPP 116
>UniRef100_UPI0000361A96 UPI0000361A96 UniRef100 entry
Length = 248
Score = 42.7 bits (99), Expect = 0.008
Identities = 28/76 (36%), Positives = 31/76 (39%), Gaps = 5/76 (6%)
Query: 4 SEQPTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPV 63
S P LPLP+ P P PPL P P P PS PPPPPP +P
Sbjct: 30 SPPPPLPLPLPLPLPLPLPPPPPPPPPLPPPPPPPPPPPPSPSPPPPPPPPP-----SPP 84
Query: 64 RSRRRGPPRNGPIPTP 79
PP + P P P
Sbjct: 85 PPPPPPPPPSPPPPPP 100
Score = 39.7 bits (91), Expect = 0.064
Identities = 22/54 (40%), Positives = 24/54 (43%), Gaps = 7/54 (12%)
Query: 26 PNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRSRRRGPPRNGPIPTP 79
P+ PP PLPL L +P P PPPPPP P PP P P P
Sbjct: 29 PSPPPPLPLPLPLPLPLPLPPPPPPPPPLPPPPPPPP-------PPPPSPSPPP 75
Score = 37.7 bits (86), Expect = 0.24
Identities = 26/76 (34%), Positives = 28/76 (36%), Gaps = 8/76 (10%)
Query: 4 SEQPTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPV 63
S P LPLP+ P P PP P P P PS PPPPP P
Sbjct: 28 SPSPPPPLPLPLPLPLPLPLPPPPPPPPPLPPPPPPPPPPPPSPSPPPPP--------PP 79
Query: 64 RSRRRGPPRNGPIPTP 79
PP P P+P
Sbjct: 80 PPSPPPPPPPPPPPSP 95
Score = 34.7 bits (78), Expect = 2.1
Identities = 27/75 (36%), Positives = 29/75 (38%), Gaps = 5/75 (6%)
Query: 7 PTQSLPLPMLTPFQRMFAHPNIP-PLQQQPLPLTLNIPQPSHPPPP-PPQDVVDNHAPVR 64
P P P P P +P PL PLPL L +P P PPPP PP P
Sbjct: 14 PPPPAPAPPPPPPPSPSPPPPLPLPL---PLPLPLPLPPPPPPPPPLPPPPPPPPPPPPS 70
Query: 65 SRRRGPPRNGPIPTP 79
PP P P P
Sbjct: 71 PSPPPPPPPPPSPPP 85
Score = 32.7 bits (73), Expect = 7.8
Identities = 19/52 (36%), Positives = 19/52 (36%)
Query: 26 PNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRSRRRGPPRNGPIP 77
P PP P P P PS PPPPPP P PP P P
Sbjct: 74 PPPPPPPPSPPPPPPPPPPPSPPPPPPPSSSPPPPPPPSPPPPPPPPPAPPP 125
>UniRef100_Q6SSE6 Plus agglutinin [Chlamydomonas reinhardtii]
Length = 3409
Score = 42.7 bits (99), Expect = 0.008
Identities = 27/77 (35%), Positives = 29/77 (37%), Gaps = 1/77 (1%)
Query: 4 SEQPTQSLPLPMLTPFQRMFAHP-NIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAP 62
S P P L P A P +PP P P L PQP P PPP + AP
Sbjct: 1320 SPTPPAPAPAAALPPLPPSPAPPLPVPPASPAPSPSPLRPPQPQTPAMPPPSPAPPSPAP 1379
Query: 63 VRSRRRGPPRNGPIPTP 79
G P P PTP
Sbjct: 1380 PSPAPPGVPPPPPTPTP 1396
Score = 37.0 bits (84), Expect = 0.41
Identities = 18/43 (41%), Positives = 23/43 (52%), Gaps = 6/43 (13%)
Query: 35 PLPLTLNIPQPSHPPPPPPQDVVDNHAPVRSRRRGPPRNGPIP 77
P+P + PQP PPPPPP+ P R+ R PP + P P
Sbjct: 551 PMPPSPRPPQPPSPPPPPPR------PPPRAPRPSPPFHPPSP 587
Score = 34.3 bits (77), Expect = 2.7
Identities = 21/55 (38%), Positives = 24/55 (43%), Gaps = 5/55 (9%)
Query: 7 PTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHA 61
P + L ML P A P PP L L QPS PPPPPP ++ A
Sbjct: 1961 PRRHLMQQMLQPPAAAVAAPPPPPASSSALVL-----QPSPPPPPPPSQLLIQQA 2010
Score = 33.1 bits (74), Expect = 6.0
Identities = 21/71 (29%), Positives = 23/71 (31%), Gaps = 5/71 (7%)
Query: 7 PTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRSR 66
P P+P P P PP P P L P P P P PP + AP
Sbjct: 1155 PPSPEPVPPTPP-----PSPPAPPSPAPPSPQPLEPPSPEPPSPAPPSPAPPSPAPPSPE 1209
Query: 67 RRGPPRNGPIP 77
P P P
Sbjct: 1210 PPSPEPPSPEP 1220
Score = 32.7 bits (73), Expect = 7.8
Identities = 23/73 (31%), Positives = 29/73 (39%), Gaps = 2/73 (2%)
Query: 7 PTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRSR 66
P P P+L P P P PLP + P+P P PPP + AP +
Sbjct: 1123 PEPPSPAPLLPPSPDP-PSPAPPSPMPPPLPTSPPSPEPVPPTPPPSPPAPPSPAPPSPQ 1181
Query: 67 RRGPPRNGPIPTP 79
PP P P+P
Sbjct: 1182 PLEPPSPEP-PSP 1193
>UniRef100_Q6AU77 Hypothetical protein OSJNBa0014C03.6 [Oryza sativa]
Length = 847
Score = 42.7 bits (99), Expect = 0.008
Identities = 18/36 (50%), Positives = 21/36 (58%)
Query: 20 QRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQD 55
+R+ P +PPL P P T PQP PPPPPP D
Sbjct: 193 RRLRHRPQVPPLVSSPAPGTPVTPQPPPPPPPPPPD 228
>UniRef100_UPI00003617D6 UPI00003617D6 UniRef100 entry
Length = 669
Score = 42.4 bits (98), Expect = 0.010
Identities = 22/47 (46%), Positives = 26/47 (54%), Gaps = 3/47 (6%)
Query: 20 QRMFAHPNIPPLQQQPLPLTLN-IP--QPSHPPPPPPQDVVDNHAPV 63
Q++ P IPP QQQP+P +P QP PPPPPPQ V V
Sbjct: 247 QQIMPQPQIPPSQQQPVPPQAQPVPSQQPPPPPPPPPQQPVPTQPAV 293
>UniRef100_UPI000024B6BB UPI000024B6BB UniRef100 entry
Length = 519
Score = 41.6 bits (96), Expect = 0.017
Identities = 31/85 (36%), Positives = 40/85 (46%), Gaps = 17/85 (20%)
Query: 7 PTQSLPLPMLTPFQRMFAH-----PNIPPLQQQPLPLTLNI-PQPSHPPPPPPQDVVDNH 60
P QS LP P M +H P I P+Q +PLP I P SHP PPPP V
Sbjct: 355 PMQSHALPPPPPITPMQSHALPPPPPIMPMQSEPLPPPAQIMPMQSHPLPPPPPLV---- 410
Query: 61 APVRSRRRGPP------RNGPIPTP 79
++S+ PP ++ P+P P
Sbjct: 411 -QMQSQPMPPPPPTVSFQSQPLPPP 434
Score = 41.6 bits (96), Expect = 0.017
Identities = 30/80 (37%), Positives = 37/80 (45%), Gaps = 7/80 (8%)
Query: 7 PTQSLPLPMLTPFQRMFAH-----PNIPPLQQQPLPLTLNI-PQPSHP-PPPPPQDVVDN 59
P QS PLP P M +H P I P+Q LP I P SH PPPPP + +
Sbjct: 313 PIQSQPLPPPPPITPMQSHALPPPPPITPMQSHALPPPPPITPMQSHALPPPPPITPMQS 372
Query: 60 HAPVRSRRRGPPRNGPIPTP 79
HA P ++ P+P P
Sbjct: 373 HALPPPPPIMPMQSEPLPPP 392
Score = 37.7 bits (86), Expect = 0.24
Identities = 22/55 (40%), Positives = 27/55 (49%), Gaps = 8/55 (14%)
Query: 7 PTQSLPLPMLTPFQRMFAHPNIPP-----LQQQPLPL---TLNIPQPSHPPPPPP 53
P QS PLP M +HP PP +Q QP+P T++ PPPPPP
Sbjct: 383 PMQSEPLPPPAQIMPMQSHPLPPPPPLVQMQSQPMPPPPPTVSFQSQPLPPPPPP 437
Score = 35.8 bits (81), Expect = 0.92
Identities = 22/52 (42%), Positives = 25/52 (47%), Gaps = 5/52 (9%)
Query: 7 PTQSLPLPMLTPFQRMFAHPNIPP-----LQQQPLPLTLNIPQPSHPPPPPP 53
P QS PLP P +M + P PP Q QPLP Q + PPPPP
Sbjct: 397 PMQSHPLPPPPPLVQMQSQPMPPPPPTVSFQSQPLPPPPPPLQANGLPPPPP 448
Score = 34.7 bits (78), Expect = 2.1
Identities = 25/76 (32%), Positives = 33/76 (42%), Gaps = 13/76 (17%)
Query: 12 PLPMLTP-FQRMFAH--PN-----IPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPV 63
P P++T ++ FAH P+ +PP Q P P P PPPPP + + V
Sbjct: 446 PPPLITNGLKQAFAHKFPSRPQDVLPPANQNNFP-----PPPPPPPPPPLAPTLPPYQHV 500
Query: 64 RSRRRGPPRNGPIPTP 79
S PP P P P
Sbjct: 501 TSTSNPPPPPPPPPLP 516
>UniRef100_UPI00002B1219 UPI00002B1219 UniRef100 entry
Length = 247
Score = 41.6 bits (96), Expect = 0.017
Identities = 27/74 (36%), Positives = 32/74 (42%), Gaps = 5/74 (6%)
Query: 7 PTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRSR 66
P S+P P TP P+ PP P+ +P PSHPPPP P + P S
Sbjct: 18 PPPSIPPP--TPPPPSPPPPSPPPPSPPPISPPPALPPPSHPPPPSPSPPPPSLPPPPS- 74
Query: 67 RRGPPRNGPIPTPF 80
PP PI PF
Sbjct: 75 --SPPHPPPITPPF 86
Score = 34.7 bits (78), Expect = 2.1
Identities = 21/56 (37%), Positives = 26/56 (45%), Gaps = 4/56 (7%)
Query: 24 AHPNIPPLQQQPLPLTLNIPQPSHPP--PPPPQDVVDNHAPVRSRRRGPPRNGPIP 77
AHP PP P P +IP P+ PP PPPP + P+ PP + P P
Sbjct: 6 AHP--PPSLPPPSPPPPSIPPPTPPPPSPPPPSPPPPSPPPISPPPALPPPSHPPP 59
>UniRef100_UPI0000265027 UPI0000265027 UniRef100 entry
Length = 243
Score = 41.6 bits (96), Expect = 0.017
Identities = 24/73 (32%), Positives = 31/73 (41%), Gaps = 5/73 (6%)
Query: 7 PTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRSR 66
P + PL P A P+ PP + +P +P P PPPPPP AP +
Sbjct: 81 PLATPPLSADEPSDEPSAPPSKPPARPEPGAAKAPVPPPPRPPPPPPM-----AAPKLEK 135
Query: 67 RRGPPRNGPIPTP 79
PPR G +P
Sbjct: 136 EPRPPRPGWTSSP 148
>UniRef100_Q9XER9 Transcription factor-like protein [Arabidopsis thaliana]
Length = 1392
Score = 41.6 bits (96), Expect = 0.017
Identities = 25/73 (34%), Positives = 32/73 (43%), Gaps = 1/73 (1%)
Query: 7 PTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPVRSR 66
P+ LP L P P +PP QP P L+ P PPPPPP + + S
Sbjct: 1082 PSPPLPPSSLPPPPPAALFPPLPPPPSQPPPPPLSPPPSPPPPPPPPSQSLTTQLSIASH 1141
Query: 67 RRGPPRNG-PIPT 78
+ P + G P PT
Sbjct: 1142 HQIPFQPGFPPPT 1154
Score = 41.2 bits (95), Expect = 0.022
Identities = 27/76 (35%), Positives = 39/76 (50%), Gaps = 2/76 (2%)
Query: 4 SEQPTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPV 63
+E+ T+ PLP +P + P +PPL P P + +P PS PPPPP + P
Sbjct: 1048 AEKSTEFNPLPEDSPPLPQESPPPLPPLPPSPPPPSPPLP-PSSLPPPPPAALFPPLPPP 1106
Query: 64 RSRRRGPPRNGPIPTP 79
S+ PP + P P+P
Sbjct: 1107 PSQPPPPPLSPP-PSP 1121
>UniRef100_Q40385 Pistil extensin-like protein [Nicotiana alata]
Length = 430
Score = 41.6 bits (96), Expect = 0.017
Identities = 25/77 (32%), Positives = 34/77 (43%), Gaps = 10/77 (12%)
Query: 6 QPTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQP---SHPPPPPPQDVVDNHAP 62
+P S P P + P P PP +P P + QP PPPPPP V +P
Sbjct: 146 RPKPSAPSPPVKP-------PPPPPSPCKPSPPDQSAKQPPPAKQPPPPPPPPPVKAPSP 198
Query: 63 VRSRRRGPPRNGPIPTP 79
+++ PP P P+P
Sbjct: 199 SPAKQSPPPPRAPSPSP 215
Score = 34.7 bits (78), Expect = 2.1
Identities = 25/76 (32%), Positives = 28/76 (35%), Gaps = 5/76 (6%)
Query: 4 SEQPTQSLPLPMLTPFQRMFAHPNIPPLQQQPLPLTLNIPQPSHPPPPPPQDVVDNHAPV 63
S P + P P P P P +Q P P P PPPPPP AP
Sbjct: 197 SPSPAKQSPPPPRAPSPSPATQP---PTKQPPPPPRAKKSPP--PPPPPPVAYPPVMAPS 251
Query: 64 RSRRRGPPRNGPIPTP 79
S PP P P+P
Sbjct: 252 PSPAAEPPIVAPFPSP 267
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.323 0.139 0.450
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 449,002,207
Number of Sequences: 2790947
Number of extensions: 21833532
Number of successful extensions: 172044
Number of sequences better than 10.0: 1352
Number of HSP's better than 10.0 without gapping: 325
Number of HSP's successfully gapped in prelim test: 1189
Number of HSP's that attempted gapping in prelim test: 149256
Number of HSP's gapped (non-prelim): 8848
length of query: 233
length of database: 848,049,833
effective HSP length: 123
effective length of query: 110
effective length of database: 504,763,352
effective search space: 55523968720
effective search space used: 55523968720
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 73 (32.7 bits)
Medicago: description of AC141322.2


