

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141115.10 + phase: 0 /pseudo
(173 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
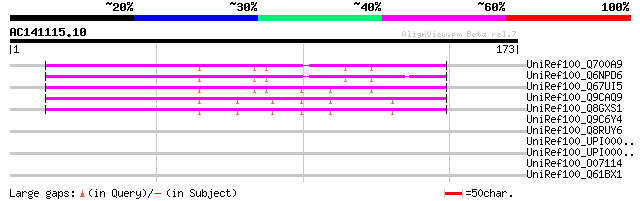
UniRef100_Q700A9 C2 domain-containing protein [Cicer arietinum] 120 1e-26
UniRef100_Q6NPD6 At2g22125 [Arabidopsis thaliana] 115 5e-25
UniRef100_Q67UI5 C2 domain-containing protein-like [Oryza sativa] 100 2e-20
UniRef100_Q9CAQ9 Hypothetical protein T5M16.5 [Arabidopsis thali... 51 1e-05
UniRef100_Q8GXS1 Hypothetical protein At1g77460/T5M16_5 [Arabido... 51 1e-05
UniRef100_Q9C6Y4 Hypothetical protein T7O23.25 [Arabidopsis thal... 44 0.002
UniRef100_Q8RUY6 Hypothetical protein At2g22125 [Arabidopsis tha... 41 0.015
UniRef100_UPI0000316B45 UPI0000316B45 UniRef100 entry 32 8.9
UniRef100_UPI000023D3DB UPI000023D3DB UniRef100 entry 32 8.9
UniRef100_O07114 Sensory transduction histidine kinase [Mastigoc... 32 8.9
UniRef100_Q61BX1 Hypothetical protein CBG13182 [Caenorhabditis b... 32 8.9
>UniRef100_Q700A9 C2 domain-containing protein [Cicer arietinum]
Length = 248
Score = 120 bits (301), Expect = 1e-26
Identities = 82/180 (45%), Positives = 100/180 (55%), Gaps = 45/180 (25%)
Query: 13 MDALFLLIQGWSACPVEVSRDQSNAAAYAIPLLQNLIQFGPVLFFEKAEFI-------LV 65
+D+LFLL Q WSACP EVSR QS AAA AIP LQ LIQ GP F EKAEF+ LV
Sbjct: 62 LDSLFLLRQAWSACPAEVSRAQSIAAADAIPFLQYLIQSGPPRFQEKAEFLLQCLPGTLV 121
Query: 66 MIVKRGNNMRQCVGNQG---KITL-------------EANQEWDERFTYMVL*ECSSRTE 109
+I+KRGNNM+Q VGN KITL N EWDE F++ E + +
Sbjct: 122 VIIKRGNNMKQSVGNPSVYCKITLGNNPPRLTKVVSTGPNPEWDESFSWSF--ESPPKGQ 179
Query: 110 ASYL-LQKQA*SGK-------------------ADEHTLLPTSKSGQPRNLEVELKWSNK 149
++ + ++ GK A E+TLLP SKSG PRNLE+E +WSNK
Sbjct: 180 KLHISCKNKSKVGKSKFGKVTIQIDRVVMLGAVAGEYTLLPASKSGPPRNLEIEFQWSNK 239
>UniRef100_Q6NPD6 At2g22125 [Arabidopsis thaliana]
Length = 309
Score = 115 bits (288), Expect = 5e-25
Identities = 82/180 (45%), Positives = 100/180 (55%), Gaps = 46/180 (25%)
Query: 13 MDALFLLIQGWSACPVEVSRDQSNAAAYAIPLLQNLIQFGPVLFFEKAEFI-------LV 65
+DALFLL Q WSACP EVSR QS AAA AIPLLQ LIQ GP F EKAEF+ LV
Sbjct: 133 LDALFLLRQAWSACPAEVSRAQSVAAADAIPLLQYLIQSGPPRFQEKAEFLLQCLPGTLV 192
Query: 66 MIVKRGNNMRQCVGNQG---KITL-------------EANQEWDERFTYMVL*ECSSRTE 109
+ +KRGNNM+Q VGN KITL N EWDE F++ E + +
Sbjct: 193 VTIKRGNNMKQSVGNPSVFCKITLGNNPPRQTKVISTGPNPEWDESFSWSF--ESPPKGQ 250
Query: 110 ASYL-LQKQA*SGK-------------------ADEHTLLPTSKSGQPRNLEVELKWSNK 149
++ + ++ GK A E++LLP SKSG PRNLE+E +WSNK
Sbjct: 251 KLHISCKNKSKMGKSSFGKVTIQIDRVVMLGAVAGEYSLLPESKSG-PRNLEIEFQWSNK 309
>UniRef100_Q67UI5 C2 domain-containing protein-like [Oryza sativa]
Length = 983
Score = 100 bits (248), Expect = 2e-20
Identities = 70/178 (39%), Positives = 92/178 (51%), Gaps = 41/178 (23%)
Query: 13 MDALFLLIQGWSACPVEVSRDQSNAAAYAIPLLQNLIQFGPVLFFEKAEFI-------LV 65
+D+L+LL Q W AC E+ + QS AA+ AIPLLQ LIQ GP F EKAE + L
Sbjct: 806 LDSLYLLRQAWGACAAEIFKAQSVAASEAIPLLQYLIQSGPPRFQEKAELLLQCLPGTLT 865
Query: 66 MIVKRGNNMRQCVGNQG---KITL-------------EANQEWDERFTY---------MV 100
+ +KRGNN+RQ VGN K+TL A EWDE F + +
Sbjct: 866 VTIKRGNNLRQSVGNPSAFCKLTLGNNPPRLTKIVSTGATPEWDEAFAWAFDSPPKGQKL 925
Query: 101 L*ECSSRT--------EASYLLQKQA*SGK-ADEHTLLPTSKSGQPRNLEVELKWSNK 149
C + + + + + + G A E+TLLP SKSG RNLE+E +WSNK
Sbjct: 926 HISCKNNSKFGKKSFGKVTIQIDRVVMLGSVAGEYTLLPESKSGPNRNLEIEFQWSNK 983
>UniRef100_Q9CAQ9 Hypothetical protein T5M16.5 [Arabidopsis thaliana]
Length = 2110
Score = 50.8 bits (120), Expect = 1e-05
Identities = 46/179 (25%), Positives = 77/179 (42%), Gaps = 42/179 (23%)
Query: 13 MDALFLLIQGWSACPVEVSRDQSNAAAYAIPLLQNLIQFGPVLFFEKAEFI-------LV 65
+D L+LL W+ ++V++ Q+ AA AIP+LQ L++ P F +KA+ + L
Sbjct: 1926 LDILYLLRHSWTNMSIDVAKSQAMIAAEAIPVLQMLMKTCPPRFHDKADSLLHCLPGCLT 1985
Query: 66 MIVKRGNNMRQ---------------CVGNQGKITLEA-NQEWDERFTY---------MV 100
+ V R NN++Q C Q K+ + EW E FT+ +
Sbjct: 1986 VNVMRANNLKQSMATTNAFCQLTIGNCPPRQTKVVSNSTTPEWKEGFTWAFDVPPKGQKL 2045
Query: 101 L*ECSSRT--------EASYLLQKQA*SGKADEHTLL--PTSKSGQPRNLEVELKWSNK 149
C S++ + + K G+ L SK R+L++E+ WSN+
Sbjct: 2046 HIICKSKSTFGKTTLGRVTIQIDKVVTEGEYSGSLSLNHENSKDASSRSLDIEIAWSNR 2104
>UniRef100_Q8GXS1 Hypothetical protein At1g77460/T5M16_5 [Arabidopsis thaliana]
Length = 434
Score = 50.8 bits (120), Expect = 1e-05
Identities = 46/179 (25%), Positives = 77/179 (42%), Gaps = 42/179 (23%)
Query: 13 MDALFLLIQGWSACPVEVSRDQSNAAAYAIPLLQNLIQFGPVLFFEKAEFI-------LV 65
+D L+LL W+ ++V++ Q+ AA AIP+LQ L++ P F +KA+ + L
Sbjct: 250 LDILYLLRHSWTNMSIDVAKSQAMIAAEAIPVLQMLMKTCPPRFHDKADSLLHCLPGCLT 309
Query: 66 MIVKRGNNMRQ---------------CVGNQGKITLEA-NQEWDERFTY---------MV 100
+ V R NN++Q C Q K+ + EW E FT+ +
Sbjct: 310 VNVMRANNLKQSMATTNAFCQLTIGNCPPRQTKVVSNSTTPEWKEGFTWAFDVPPKGQKL 369
Query: 101 L*ECSSRT--------EASYLLQKQA*SGKADEHTLL--PTSKSGQPRNLEVELKWSNK 149
C S++ + + K G+ L SK R+L++E+ WSN+
Sbjct: 370 HIICKSKSTFGKTTLGRVTIQIDKVVTEGEYSGSLSLNHENSKDASSRSLDIEIAWSNR 428
>UniRef100_Q9C6Y4 Hypothetical protein T7O23.25 [Arabidopsis thaliana]
Length = 2114
Score = 43.9 bits (102), Expect = 0.002
Identities = 31/112 (27%), Positives = 53/112 (46%), Gaps = 24/112 (21%)
Query: 11 SWMDALFLLIQGWSACPVEVSRDQSNAAAYAIPLLQNLIQF-----GPVLFFEKAEFI-- 63
S MD ++ L Q W+ P E +R Q+ AA AIP+LQ +++ P F E+ +
Sbjct: 1930 SAMDTIYTLRQSWTTMPTETARSQAVLAADAIPVLQLMMKSKLKSPAPSSFHERGNSLLN 1989
Query: 64 -----LVMIVKRGNNMRQ-----------CVGNQGKITLEANQE-WDERFTY 98
L + +KRG+N+++ C + K+ ++ W E FT+
Sbjct: 1990 CLPGSLTVAIKRGDNLKRSNAFCRLIIDNCPTKKTKVVKRSSSPVWKESFTW 2041
>UniRef100_Q8RUY6 Hypothetical protein At2g22125 [Arabidopsis thaliana]
Length = 109
Score = 40.8 bits (94), Expect = 0.015
Identities = 19/27 (70%), Positives = 23/27 (84%), Gaps = 1/27 (3%)
Query: 123 ADEHTLLPTSKSGQPRNLEVELKWSNK 149
A E++LLP SKSG PRNLE+E +WSNK
Sbjct: 84 AGEYSLLPESKSG-PRNLEIEFQWSNK 109
>UniRef100_UPI0000316B45 UPI0000316B45 UniRef100 entry
Length = 252
Score = 31.6 bits (70), Expect = 8.9
Identities = 20/42 (47%), Positives = 25/42 (58%), Gaps = 4/42 (9%)
Query: 35 SNAAAYAIPLLQNLIQFGP-VLFFEKAEFILVMIVKRGNNMR 75
S A AY+ N IQF P VL FEK +FIL +KR N++
Sbjct: 205 SYAGAYSFSQSWNFIQFSPPVLAFEKKKFIL---IKRKQNLK 243
>UniRef100_UPI000023D3DB UPI000023D3DB UniRef100 entry
Length = 3213
Score = 31.6 bits (70), Expect = 8.9
Identities = 17/63 (26%), Positives = 34/63 (52%)
Query: 64 LVMIVKRGNNMRQCVGNQGKITLEANQEWDERFTYMVL*ECSSRTEASYLLQKQA*SGKA 123
L ++ R N+ +G+ T+E ++ + FTY++L + S + S+L++ A A
Sbjct: 1518 LASLIHRTGNIASELGHVFSQTMEKHENRYQYFTYLLLEQGESINDPSFLVRPSAVLRSA 1577
Query: 124 DEH 126
D+H
Sbjct: 1578 DDH 1580
>UniRef100_O07114 Sensory transduction histidine kinase [Mastigocladus laminosus]
Length = 308
Score = 31.6 bits (70), Expect = 8.9
Identities = 22/92 (23%), Positives = 43/92 (45%), Gaps = 10/92 (10%)
Query: 2 QIQWHVKKLSWMDALFLLIQGWSACPVEVSRDQSNAAAYAIPLLQNLIQFGPVLFFEKAE 61
+++WH + LS + + L + SR ++ A +P ++ LI L +
Sbjct: 135 RVEWHPESLSLQECVDLAL----------SRIRTRVATDPLPQIKTLISPNLPLVKADGD 184
Query: 62 FILVMIVKRGNNMRQCVGNQGKITLEANQEWD 93
+++ +I K +N + QG+IT+ AN D
Sbjct: 185 WLVEVIAKLVDNACKFTPPQGQITITANPNSD 216
>UniRef100_Q61BX1 Hypothetical protein CBG13182 [Caenorhabditis briggsae]
Length = 700
Score = 31.6 bits (70), Expect = 8.9
Identities = 14/43 (32%), Positives = 23/43 (52%)
Query: 43 PLLQNLIQFGPVLFFEKAEFILVMIVKRGNNMRQCVGNQGKIT 85
P +I FG FF FI++ I GN+ +Q + ++G+ T
Sbjct: 20 PSCNKIISFGVYSFFIVVNFIVLCIPNTGNHYQQMIDDEGRTT 62
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.328 0.139 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 271,198,532
Number of Sequences: 2790947
Number of extensions: 9669363
Number of successful extensions: 21113
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 21092
Number of HSP's gapped (non-prelim): 19
length of query: 173
length of database: 848,049,833
effective HSP length: 118
effective length of query: 55
effective length of database: 518,718,087
effective search space: 28529494785
effective search space used: 28529494785
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 70 (31.6 bits)
Medicago: description of AC141115.10


