

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141114.8 - phase: 0
(239 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
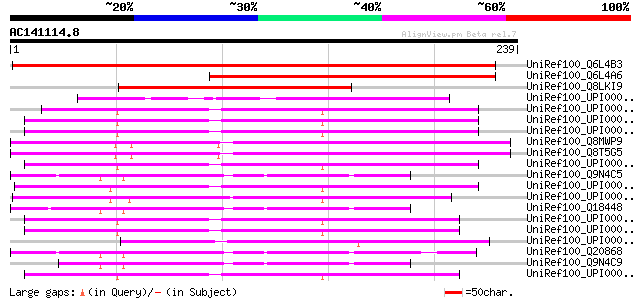
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6L4B3 Putative polyprotein [Solanum demissum] 282 6e-75
UniRef100_Q6L4A6 Putative reverse transcriptase [Solanum demissum] 171 1e-41
UniRef100_Q8LKI9 NBS/LRR resistance protein-like protein [Capsic... 130 2e-29
UniRef100_UPI000034DCE8 UPI000034DCE8 UniRef100 entry 104 2e-21
UniRef100_UPI000024BB57 UPI000024BB57 UniRef100 entry 86 6e-16
UniRef100_UPI000024D3A0 UPI000024D3A0 UniRef100 entry 86 8e-16
UniRef100_UPI000024BEDA UPI000024BEDA UniRef100 entry 86 8e-16
UniRef100_Q8MWP9 Polyprotein [Schistosoma japonicum] 85 1e-15
UniRef100_Q8T5G5 Polyprotein [Schistosoma japonicum] 85 1e-15
UniRef100_UPI0000247D3B UPI0000247D3B UniRef100 entry 84 2e-15
UniRef100_Q9N4C5 Hypothetical protein Y76B12C.5 [Caenorhabditis ... 82 2e-14
UniRef100_UPI000024A3B3 UPI000024A3B3 UniRef100 entry 81 2e-14
UniRef100_UPI00003604D1 UPI00003604D1 UniRef100 entry 81 2e-14
UniRef100_Q18448 Hypothetical protein C34D4.5 [Caenorhabditis el... 81 3e-14
UniRef100_UPI000024D3A8 UPI000024D3A8 UniRef100 entry 78 2e-13
UniRef100_UPI000024BEE4 UPI000024BEE4 UniRef100 entry 78 2e-13
UniRef100_UPI0000018E3B UPI0000018E3B UniRef100 entry 78 2e-13
UniRef100_Q20868 Hypothetical protein F56C9.2 [Caenorhabditis el... 78 2e-13
UniRef100_Q9N4C9 Hypothetical protein Y75D11A.4 [Caenorhabditis ... 77 3e-13
UniRef100_UPI000024B6D5 UPI000024B6D5 UniRef100 entry 77 5e-13
>UniRef100_Q6L4B3 Putative polyprotein [Solanum demissum]
Length = 832
Score = 282 bits (721), Expect = 6e-75
Identities = 133/228 (58%), Positives = 166/228 (72%)
Query: 2 YEGVSTSVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQELAPRCMLFADDV 61
Y T VRT G +E FP+ IGLHQGS LSP+LF LV+D LT IQE P CMLFADD+
Sbjct: 370 YYTAKTRVRTVGGDSEHFPVEIGLHQGSVLSPFLFALVMDELTRSIQETVPWCMLFADDI 429
Query: 62 VLVGESREEVNGRLETWRQALEAYGFRLSRSKTEYMEWNFSGRRSRSTLEVKVGDHIIPQ 121
VL+ E+R+ VN RLE WRQ LE+ GFRLSR+KTEY+ FS + +EV++ +IP+
Sbjct: 430 VLIDETRDRVNARLEVWRQTLESKGFRLSRTKTEYLGCKFSDGLDETDVEVRLAAQVIPK 489
Query: 122 VTRFKYLGSFVQNDGEIEADVSHRIQAGWLKWRRASGVLCDKKVPLKLKGKFYRTAIRPA 181
F+YLG+ +Q G+I+ DV+HR+ A W+KWR ASGVLCDKK+ KLKGKFYR +RPA
Sbjct: 490 KESFRYLGAVIQGSGDIDDDVTHRVGAAWMKWRLASGVLCDKKISPKLKGKFYRVVVRPA 549
Query: 182 LLYGTECWAVKSQHENQVSVTEMRMLRWMSGKTRQDRIRNDTIREGRG 229
LLYG ECW VK+ H +++ V EMRMLRWM G TR D+IRN+ IRE G
Sbjct: 550 LLYGAECWPVKNAHVHKMHVAEMRMLRWMCGHTRSDKIRNEVIREKVG 597
>UniRef100_Q6L4A6 Putative reverse transcriptase [Solanum demissum]
Length = 213
Score = 171 bits (433), Expect = 1e-41
Identities = 77/135 (57%), Positives = 99/135 (73%)
Query: 95 EYMEWNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFVQNDGEIEADVSHRIQAGWLKWR 154
EY+ FS + +EV++ +IP+ FKYLG+ +Q G+I+ DV+HR+ A W+KWR
Sbjct: 7 EYLGCKFSDVLDETDVEVRLAAQVIPKKESFKYLGAVIQGSGDIDDDVTHRVGAAWMKWR 66
Query: 155 RASGVLCDKKVPLKLKGKFYRTAIRPALLYGTECWAVKSQHENQVSVTEMRMLRWMSGKT 214
ASGVLCDKK+PLKLKGKFYR +RPALLYG ECW VK+ H +++ V EMRMLRWM G T
Sbjct: 67 LASGVLCDKKIPLKLKGKFYRVVVRPALLYGAECWPVKNAHVHKMHVAEMRMLRWMCGHT 126
Query: 215 RQDRIRNDTIREGRG 229
R D+IRN+ IRE G
Sbjct: 127 RSDKIRNEVIREKVG 141
>UniRef100_Q8LKI9 NBS/LRR resistance protein-like protein [Capsicum annuum]
Length = 122
Score = 130 bits (328), Expect = 2e-29
Identities = 61/110 (55%), Positives = 83/110 (75%)
Query: 52 PRCMLFADDVVLVGESREEVNGRLETWRQALEAYGFRLSRSKTEYMEWNFSGRRSRSTLE 111
P CMLFADDVVL+ E+R VN +LE WRQ LE+ GFR+SR+KTEY+E F+ R + +
Sbjct: 10 PWCMLFADDVVLIDETRGGVNDKLELWRQTLESKGFRVSRTKTEYVECKFNDVRRENEVV 69
Query: 112 VKVGDHIIPQVTRFKYLGSFVQNDGEIEADVSHRIQAGWLKWRRASGVLC 161
V++ + + +FKYLGS +Q++GEI+ DVSHRI AGW+KW+ ASGV+C
Sbjct: 70 VRLEAQEVKKRDKFKYLGSVIQSNGEIDEDVSHRIGAGWMKWKLASGVMC 119
>UniRef100_UPI000034DCE8 UPI000034DCE8 UniRef100 entry
Length = 183
Score = 104 bits (259), Expect = 2e-21
Identities = 65/175 (37%), Positives = 99/175 (56%), Gaps = 17/175 (9%)
Query: 33 PYLFTLVLDVLTEHIQELAPRCMLFADDVVLVGESREEVNGRLETWRQALEAYGFRLSRS 92
P L +V+D LT+ +++ +P +FA D+V+ SRE+V +LE R ALE+ S
Sbjct: 19 PLLVAMVMDRLTDEVRQESPWTTMFAGDIVMC--SREQVEEKLEERRFALES-------S 69
Query: 93 KTEYMEWNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFVQNDGEIEADVSHRIQAGWLK 152
KTE E + SG E+K G+ + K LGS VQ GE +V R+QAGW
Sbjct: 70 KTEN-ERDLSGSVRLQGEEIKKGEDL-------KNLGSTVQTSGECGKEVKKRVQAGWNW 121
Query: 153 WRRASGVLCDKKVPLKLKGKFYRTAIRPALLYGTECWAVKSQHENQVSVTEMRML 207
W + SGV+CD+ V K+K K +T RPA+++G E ++ + E ++ + EM+ L
Sbjct: 122 WGKVSGVMCDRGVSAKIKRKVDKTVARPAIIFGLETVPLRKRQEAELELAEMKAL 176
>UniRef100_UPI000024BB57 UPI000024BB57 UniRef100 entry
Length = 312
Score = 86.3 bits (212), Expect = 6e-16
Identities = 59/214 (27%), Positives = 105/214 (48%), Gaps = 13/214 (6%)
Query: 16 TEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQE-------LAPRCMLFADDVVLVGESR 68
+E F I+ G+ QG L+P LF + + +L H L R MLFADD + ++
Sbjct: 8 SEPFSISSGIKQGCVLAPTLFGIFIALLLRHAFGSATEGIYLRTRDMLFADDAAVTTHTQ 67
Query: 69 EEVNGRLETWRQALEAYGFRLSRSKTEYMEWNFSGRRSRSTLEVKVGDHIIPQVTRFKYL 128
EE+ + + A + +G +S KT N + + + + V D+ + V +F YL
Sbjct: 68 EELQSLMTHFSMACKDFGLTISLKKT-----NVLSQDTATPPTITVDDYQLDVVHQFTYL 122
Query: 129 GSFVQNDGEIEADVSHRI-QAGWLKWRRASGVLCDKKVPLKLKGKFYRTAIRPALLYGTE 187
GS + ++ ++A++ RI +A R + V + ++ K Y + LLYG+E
Sbjct: 123 GSTITDNLSLDAELDRRIGKAASTLARLTTRVWTNHRLTTATKMAVYNACVISTLLYGSE 182
Query: 188 CWAVKSQHENQVSVTEMRMLRWMSGKTRQDRIRN 221
W ++ E +++ +R LR + G T QD++ N
Sbjct: 183 TWTTYARQERRLNTFHLRGLRRILGITWQDKVSN 216
>UniRef100_UPI000024D3A0 UPI000024D3A0 UniRef100 entry
Length = 823
Score = 85.9 bits (211), Expect = 8e-16
Identities = 59/222 (26%), Positives = 107/222 (47%), Gaps = 13/222 (5%)
Query: 8 SVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQE-------LAPRCMLFADD 60
+++ +E F I+ G+ QG L+P LF + +L H L R MLFADD
Sbjct: 570 TIQFNGSCSEPFSISSGVKQGCVLAPTLFGIFFALLLRHAFGSATEGIYLRTRDMLFADD 629
Query: 61 VVLVGESREEVNGRLETWRQALEAYGFRLSRSKTEYMEWNFSGRRSRSTLEVKVGDHIIP 120
+ ++EE+ + + A + +G +S KT N + + + + V D+ +
Sbjct: 630 AAVTTHTQEELQSLMTRFSMACKDFGLTISLKKT-----NVLSQDTATPPTITVDDYQLD 684
Query: 121 QVTRFKYLGSFVQNDGEIEADVSHRI-QAGWLKWRRASGVLCDKKVPLKLKGKFYRTAIR 179
V +F YLGS + ++ ++A++ RI +A R + V + ++ K Y +
Sbjct: 685 VVHQFTYLGSTITDNLSLDAELDRRIGKAASTLARLTTRVWTNHRLTTATKMAVYNACVI 744
Query: 180 PALLYGTECWAVKSQHENQVSVTEMRMLRWMSGKTRQDRIRN 221
LLYG+E W ++ E +++ +R LR + G T QD++ N
Sbjct: 745 STLLYGSETWTTYARQERRLNTFHLRGLRRILGITWQDKVSN 786
>UniRef100_UPI000024BEDA UPI000024BEDA UniRef100 entry
Length = 821
Score = 85.9 bits (211), Expect = 8e-16
Identities = 59/222 (26%), Positives = 107/222 (47%), Gaps = 13/222 (5%)
Query: 8 SVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQE-------LAPRCMLFADD 60
+++ +E F I+ G+ QG L+P LF + +L H L R MLFADD
Sbjct: 568 TIQFNGSCSEPFSISSGVKQGCVLAPTLFGIFFALLLRHAFGSATEGIYLRTRDMLFADD 627
Query: 61 VVLVGESREEVNGRLETWRQALEAYGFRLSRSKTEYMEWNFSGRRSRSTLEVKVGDHIIP 120
+ ++EE+ + + A + +G +S KT N + + + + V D+ +
Sbjct: 628 AAVTTHTQEELQSLMTRFSMACKDFGLTISLKKT-----NVLSQDTATPPTITVDDYQLD 682
Query: 121 QVTRFKYLGSFVQNDGEIEADVSHRI-QAGWLKWRRASGVLCDKKVPLKLKGKFYRTAIR 179
V +F YLGS + ++ ++A++ RI +A R + V + ++ K Y +
Sbjct: 683 VVHQFTYLGSTITDNLSLDAELDRRIGKAASTLARLTTRVWTNHRLTTATKMAVYNACVI 742
Query: 180 PALLYGTECWAVKSQHENQVSVTEMRMLRWMSGKTRQDRIRN 221
LLYG+E W ++ E +++ +R LR + G T QD++ N
Sbjct: 743 STLLYGSETWTTYARQERRLNTFHLRGLRRILGITWQDKVSN 784
>UniRef100_Q8MWP9 Polyprotein [Schistosoma japonicum]
Length = 976
Score = 85.1 bits (209), Expect = 1e-15
Identities = 63/250 (25%), Positives = 111/250 (44%), Gaps = 20/250 (8%)
Query: 1 MYEGVSTSVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQ--------ELAP 52
+Y + VR + + G+ Q LSP+LF ++D+L E +L P
Sbjct: 629 LYSNTTCRVRAYGRLSSELTTSSGVRQACPLSPFLFNFIIDILLELTLSSSDFPGVDLFP 688
Query: 53 RCML----FADDVVLVGESREEVNGRLETWRQALEAYGFRLSRSKTEYM--EWNFSGRRS 106
L +ADD+VL+ E +++ L T + G R S SK + + +W
Sbjct: 689 GDKLTDLEYADDIVLLSEDADKMQDFLTTLNMNVSMLGMRFSPSKCKMLLQDW------L 742
Query: 107 RSTLEVKVGDHIIPQVTRFKYLGSFVQNDGEIEADVSHRIQAGWLKWRRASGVLCDKKVP 166
S ++ +G I V RF YLGS + +G + ++S RI + + + +
Sbjct: 743 NSAPKLVIGRETIECVNRFTYLGSLISPNGLVSDEISARIHKARSAFANLRHLWRRRDIR 802
Query: 167 LKLKGKFYRTAIRPALLYGTECWAVKSQHENQVSVTEMRMLRWMSGKTRQDRIRNDTIRE 226
L KG+ Y A+R L YG E W ++ + ++ V + R LR ++ +R+ N +R
Sbjct: 803 LMTKGRVYCAAVRSVLPYGCETWPLRVEDIRRILVFDHRCLRNIARVCWDNRVSNAWVRN 862
Query: 227 GRGGIHSRKV 236
G + + +
Sbjct: 863 RVLGKYGKSI 872
>UniRef100_Q8T5G5 Polyprotein [Schistosoma japonicum]
Length = 1091
Score = 85.1 bits (209), Expect = 1e-15
Identities = 63/250 (25%), Positives = 111/250 (44%), Gaps = 20/250 (8%)
Query: 1 MYEGVSTSVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQ--------ELAP 52
+Y + VR + + G+ Q LSP+LF ++D+L E +L P
Sbjct: 744 LYSNTTCRVRAYGRLSSELTTSSGVRQACPLSPFLFNFIIDILLELTLSSSDFPGVDLFP 803
Query: 53 RCML----FADDVVLVGESREEVNGRLETWRQALEAYGFRLSRSKTEYM--EWNFSGRRS 106
L +ADD+VL+ E +++ L T + G R S SK + + +W
Sbjct: 804 GDKLTDLEYADDIVLLSEDADKMQDFLTTLNMNVSMLGMRFSPSKCKMLLQDW------L 857
Query: 107 RSTLEVKVGDHIIPQVTRFKYLGSFVQNDGEIEADVSHRIQAGWLKWRRASGVLCDKKVP 166
S ++ +G I V RF YLGS + +G + ++S RI + + + +
Sbjct: 858 NSAPKLVIGRETIECVNRFTYLGSLISPNGLVSDEISARIHKARSAFANLRHLWRRRDIR 917
Query: 167 LKLKGKFYRTAIRPALLYGTECWAVKSQHENQVSVTEMRMLRWMSGKTRQDRIRNDTIRE 226
L KG+ Y A+R L YG E W ++ + ++ V + R LR ++ +R+ N +R
Sbjct: 918 LMTKGRVYCAAVRSVLPYGCETWPLRVEDIRRILVFDHRCLRNIARVCWDNRVSNAWVRN 977
Query: 227 GRGGIHSRKV 236
G + + +
Sbjct: 978 RVLGKYGKSI 987
>UniRef100_UPI0000247D3B UPI0000247D3B UniRef100 entry
Length = 789
Score = 84.3 bits (207), Expect = 2e-15
Identities = 59/222 (26%), Positives = 106/222 (47%), Gaps = 13/222 (5%)
Query: 8 SVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQE-------LAPRCMLFADD 60
+V+ +E F I+ G+ QG L+P LF + +L H L R MLFADD
Sbjct: 536 TVQFNGSCSEPFSISSGVKQGCVLAPTLFGIFFALLLRHAFGSATEGIYLRTRDMLFADD 595
Query: 61 VVLVGESREEVNGRLETWRQALEAYGFRLSRSKTEYMEWNFSGRRSRSTLEVKVGDHIIP 120
+ ++EE+ + + A + +G +S KT N + + + + V D+ +
Sbjct: 596 AAVTTHTQEELQSLMTRFSMACKDFGLTISLKKT-----NVLSQDTATPPTITVDDYQLD 650
Query: 121 QVTRFKYLGSFVQNDGEIEADVSHRI-QAGWLKWRRASGVLCDKKVPLKLKGKFYRTAIR 179
V + YLGS + ++ ++A++ RI +A R + V + ++ K Y +
Sbjct: 651 VVHQLTYLGSTITDNLSLDAELDRRIGKAASTLARLTTRVWTNHRLTTATKMALYNACVI 710
Query: 180 PALLYGTECWAVKSQHENQVSVTEMRMLRWMSGKTRQDRIRN 221
LLYG+E W ++ E +++ +R LR + G T QD++ N
Sbjct: 711 STLLYGSETWTTYARQERRLNTFHLRGLRRILGITWQDKVSN 752
>UniRef100_Q9N4C5 Hypothetical protein Y76B12C.5 [Caenorhabditis elegans]
Length = 938
Score = 81.6 bits (200), Expect = 2e-14
Identities = 67/200 (33%), Positives = 100/200 (49%), Gaps = 19/200 (9%)
Query: 1 MYEGVSTSVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLD-VLTEHIQELAP------- 52
+ EG + D +V T G+ QG + SP LF+ L +LT+ ELA
Sbjct: 696 LMEGGQAEITVHDKKLKVNLCT-GIRQGDSASPALFSAALQAILTDCDNELAGVGISVEG 754
Query: 53 ---RCMLFADDVVLVGESREEVNGRLETWRQALEAYGFRLSRSKTEYMEWNFSGRRSRST 109
R + FADDVVL+ + EEV RLE + YG ++++SKT ++ F RS
Sbjct: 755 RHIRRLEFADDVVLICSTPEEVQERLEILDRISSNYGLKINQSKTVLLKNKF----CRSQ 810
Query: 110 LEVKVGDHIIPQVTRFKYLGSFVQNDGEIEADVSHRIQAGWLKWRRASGVLCDKKVPLKL 169
+ G IIP V +YLG ++ G I+ ++S RI+AGW VL + +P K
Sbjct: 811 DILFNGSPIIP-VPGCRYLGRWIDISGSIDEEISRRIRAGWGALVGIKEVL--RIMPNKE 867
Query: 170 KGKFYRTAIRPALLYGTECW 189
+ ++ + PALLY +E W
Sbjct: 868 RIILFKQNVLPALLYASETW 887
>UniRef100_UPI000024A3B3 UPI000024A3B3 UniRef100 entry
Length = 851
Score = 81.3 bits (199), Expect = 2e-14
Identities = 59/250 (23%), Positives = 112/250 (44%), Gaps = 36/250 (14%)
Query: 3 EGVSTSVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEH---------------- 46
+ + +++ +E F I+ G+ QG L+P LF + + +L H
Sbjct: 515 QNMKGTIQFNGSCSEPFSISSGIKQGCVLAPTLFGIFIALLLRHAFGSATEGIYLRTRSD 574
Query: 47 --------------IQELAPRCMLFADDVVLVGESREEVNGRLETWRQALEAYGFRLSRS 92
++E R MLFADD + ++EE+ + + A + +G +S
Sbjct: 575 GKLFNLSRLKAKTKVRETLIRDMLFADDAAVTTHTQEELQSLMTHFSMACKDFGLTISLK 634
Query: 93 KTEYMEWNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFVQNDGEIEADVSHRI-QAGWL 151
KT N + + + + V D+ + V +F YLGS + ++ ++A++ RI +A
Sbjct: 635 KT-----NVLSQDTATPPTITVDDYQLDVVHQFTYLGSTITDNLSLDAELDRRIGKAAST 689
Query: 152 KWRRASGVLCDKKVPLKLKGKFYRTAIRPALLYGTECWAVKSQHENQVSVTEMRMLRWMS 211
R + V + ++ K Y + LLYG+E W ++ E +++ +R LR +
Sbjct: 690 LARLTTRVWTNHRLTTATKMAVYNACVISTLLYGSETWTTYARQERRLNTFHLRGLRRIL 749
Query: 212 GKTRQDRIRN 221
G T QD++ N
Sbjct: 750 GITWQDKVSN 759
>UniRef100_UPI00003604D1 UPI00003604D1 UniRef100 entry
Length = 649
Score = 81.3 bits (199), Expect = 2e-14
Identities = 63/238 (26%), Positives = 105/238 (43%), Gaps = 32/238 (13%)
Query: 2 YEGVSTSVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEH--------------- 46
++G+ V + T+ F I+ G+ QG L+P LF L + H
Sbjct: 410 HDGMQGLVLSNGDTSAPFDISNGVKQGCVLAPVLFNLFFTCVLNHALRDLDRGIYLRYRL 469
Query: 47 ------IQELAPRCM---------LFADDVVLVGESREEVNGRLETWRQALEAYGFRLSR 91
++ L R LFADD L+ + ++ ++ + +A +G +S
Sbjct: 470 DGSLFDLRRLNARTKTIERLVHDALFADDCALMAHTEPDLQIIVDRFAEAARLFGLMISL 529
Query: 92 SKTEYMEWNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFVQNDGEIEADVSHRI-QAGW 150
SKTE M G S V + + V +FKYLGS + DG ++ +++ RI +A
Sbjct: 530 SKTEVMHQPAPGT-SAPPPNVSIEGTELKVVEQFKYLGSTISCDGSLDKEIAQRISKASQ 588
Query: 151 LKWRRASGVLCDKKVPLKLKGKFYRTAIRPALLYGTECWAVKSQHENQVSVTEMRMLR 208
R + V+ +K + L+ K K Y+ + +LLYG E W ++ Q+ MR LR
Sbjct: 589 SLGRLRTRVINNKNIKLETKIKVYKAVVLTSLLYGCETWTTYRKYIKQLERFHMRSLR 646
>UniRef100_Q18448 Hypothetical protein C34D4.5 [Caenorhabditis elegans]
Length = 624
Score = 80.9 bits (198), Expect = 3e-14
Identities = 66/200 (33%), Positives = 98/200 (49%), Gaps = 19/200 (9%)
Query: 1 MYEGVSTSVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLD-VLTEHIQELAP------- 52
M EG + D +V + G+ QG + SP LF+ L +LT+ E A
Sbjct: 296 MMEGGQAEISVHDKKLKV-NLRTGVRQGDSASPALFSAALQAILTDCDNEFAGVGIKVEG 354
Query: 53 ---RCMLFADDVVLVGESREEVNGRLETWRQALEAYGFRLSRSKTEYMEWNFSGRRSRST 109
R + FADDVVL+ + EEV RLE + YG ++++SKT ++ F RS
Sbjct: 355 RHIRRLEFADDVVLICSTPEEVQERLEILDRISSIYGLKINQSKTVLLKNKF----CRSQ 410
Query: 110 LEVKVGDHIIPQVTRFKYLGSFVQNDGEIEADVSHRIQAGWLKWRRASGVLCDKKVPLKL 169
G IIP V +YLG ++ G I+ ++S RI+AGW VL + +P K
Sbjct: 411 DVFFNGSPIIP-VPGCRYLGRWIDISGSIDEEISRRIRAGWGALVGIKEVL--RIMPNKE 467
Query: 170 KGKFYRTAIRPALLYGTECW 189
+ ++ + PALLY +E W
Sbjct: 468 RIILFKQNVLPALLYASETW 487
>UniRef100_UPI000024D3A8 UPI000024D3A8 UniRef100 entry
Length = 729
Score = 78.2 bits (191), Expect = 2e-13
Identities = 55/213 (25%), Positives = 101/213 (46%), Gaps = 13/213 (6%)
Query: 8 SVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQE-------LAPRCMLFADD 60
+++ +E F I+ G+ QG L+P LF + +L H L R MLFADD
Sbjct: 522 TIQFNGSCSEPFSISSGVKQGCVLAPTLFGIFFALLLRHAFGSATEGIYLRTRDMLFADD 581
Query: 61 VVLVGESREEVNGRLETWRQALEAYGFRLSRSKTEYMEWNFSGRRSRSTLEVKVGDHIIP 120
+ ++EE+ + + A + +G +S KT N + + + + V D+ +
Sbjct: 582 AAVTTHTQEELQSLMTRFSMACKDFGLTISLKKT-----NVLSQDTATPPTITVDDYQLD 636
Query: 121 QVTRFKYLGSFVQNDGEIEADVSHRI-QAGWLKWRRASGVLCDKKVPLKLKGKFYRTAIR 179
V +F YLGS + ++ ++A++ RI +A R + V + ++ K Y +
Sbjct: 637 VVHQFTYLGSTITDNLSLDAELDRRIGKAASTLARLTTRVWTNHRLTTATKMAVYNACVI 696
Query: 180 PALLYGTECWAVKSQHENQVSVTEMRMLRWMSG 212
LLYG+E W ++ E +++ +R LR + G
Sbjct: 697 STLLYGSETWTTYARQERRLNTFHLRGLRRILG 729
>UniRef100_UPI000024BEE4 UPI000024BEE4 UniRef100 entry
Length = 725
Score = 78.2 bits (191), Expect = 2e-13
Identities = 55/213 (25%), Positives = 101/213 (46%), Gaps = 13/213 (6%)
Query: 8 SVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQE-------LAPRCMLFADD 60
+++ +E F I+ G+ QG L+P LF + +L H L R MLFADD
Sbjct: 518 TIQFNGSCSEPFSISSGVKQGCVLAPTLFGIFFALLLRHAFGSATEGIYLRTRDMLFADD 577
Query: 61 VVLVGESREEVNGRLETWRQALEAYGFRLSRSKTEYMEWNFSGRRSRSTLEVKVGDHIIP 120
+ ++EE+ + + A + +G +S KT N + + + + V D+ +
Sbjct: 578 AAVTTHTQEELQSLMTRFSMACKDFGLTISLKKT-----NVLSQDTATPPTITVDDYQLD 632
Query: 121 QVTRFKYLGSFVQNDGEIEADVSHRI-QAGWLKWRRASGVLCDKKVPLKLKGKFYRTAIR 179
V +F YLGS + ++ ++A++ RI +A R + V + ++ K Y +
Sbjct: 633 VVHQFTYLGSTITDNLSLDAELDRRIGKAASTLARLTTRVWTNHRLTTATKMAVYNACVI 692
Query: 180 PALLYGTECWAVKSQHENQVSVTEMRMLRWMSG 212
LLYG+E W ++ E +++ +R LR + G
Sbjct: 693 STLLYGSETWTTYARQERRLNTFHLRGLRRILG 725
>UniRef100_UPI0000018E3B UPI0000018E3B UniRef100 entry
Length = 262
Score = 78.2 bits (191), Expect = 2e-13
Identities = 48/175 (27%), Positives = 91/175 (51%), Gaps = 6/175 (3%)
Query: 53 RCMLFADDVVLVGESREEVNGRLETWRQALEAYGFRLSRSKTEYMEWNFSGRRSRSTLEV 112
R MLFADD +V S+E + ++ + QA + + +S +KT+ M G+ + + +
Sbjct: 2 RDMLFADDAAVVAHSQEHLQTLMDHFSQACQDFSLVISLNKTKTM-----GQGTENPPSI 56
Query: 113 KVGDHIIPQVTRFKYLGSFVQNDGEIEADVSHRIQAGWLKWRRASGVLCDK-KVPLKLKG 171
+ ++ + V F YLGS + ++ ++ ++S RI +R+ + + D K+ K
Sbjct: 57 TINNYTLEAVNTFTYLGSTITDNLSLDVELSRRIGKAASTFRKLTERVWDNGKLTTHTKV 116
Query: 172 KFYRTAIRPALLYGTECWAVKSQHENQVSVTEMRMLRWMSGKTRQDRIRNDTIRE 226
+ YR + LLYG+E W + S E +++ +R LR + G T +D + + I E
Sbjct: 117 QVYRACVISTLLYGSETWTLYSHQERRLNSFYLRCLRRILGVTWKDMVTHTAILE 171
>UniRef100_Q20868 Hypothetical protein F56C9.2 [Caenorhabditis elegans]
Length = 772
Score = 78.2 bits (191), Expect = 2e-13
Identities = 71/231 (30%), Positives = 109/231 (46%), Gaps = 26/231 (11%)
Query: 1 MYEGVSTSVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLD-VLTEHIQELAP------- 52
M +G + D +V T G+ QG + SP LF+ L +LT+ E A
Sbjct: 444 MMDGGQAEITVHDKKLKVNLCT-GVRQGDSASPALFSAALQAILTDCDNEFAGVGINVEG 502
Query: 53 ---RCMLFADDVVLVGESREEVNGRLETWRQALEAYGFRLSRSKTEYMEWNFSGRRSRST 109
R + FADDVVL+ + EV RLE + YG ++++SKT ++ F RS
Sbjct: 503 RHIRRLEFADDVVLICSTPGEVQERLEILDRISSNYGLKINQSKTVLLKNKF----CRSQ 558
Query: 110 LEVKVGDHIIPQVTRFKYLGSFVQNDGEIEADVSHRIQAGWLKWRRASGVLCDKKVPLKL 169
+ G IIP V +YLG ++ G I+ ++S RI+AGW VL + +P K
Sbjct: 559 DVLFNGSPIIP-VPGCRYLGRWIDISGSIDEEISRRIRAGWGALVGIKEVL--RIMPNKE 615
Query: 170 KGKFYRTAIRPALLYGTECWAVKSQHENQVSVTEMRMLRWMSGKTRQDRIR 220
+ ++ + PALLY +E W + + +R+ R +SG IR
Sbjct: 616 RIILFKQNVLPALLYASETWTCNAG-------STLRLKRTVSGLIDAAEIR 659
>UniRef100_Q9N4C9 Hypothetical protein Y75D11A.4 [Caenorhabditis elegans]
Length = 480
Score = 77.4 bits (189), Expect = 3e-13
Identities = 61/177 (34%), Positives = 90/177 (50%), Gaps = 18/177 (10%)
Query: 24 GLHQGSTLSPYLFTLVLD-VLTEHIQELAP----------RCMLFADDVVLVGESREEVN 72
G+ QG + SP LF+ L +LT+ ELA R + FADDVVL+ + EEV
Sbjct: 174 GVRQGDSASPALFSAALQAILTDCDNELAGVGISVEGRHIRRLEFADDVVLICSTPEEVQ 233
Query: 73 GRLETWRQALEAYGFRLSRSKTEYMEWNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFV 132
RLE + YG ++ +SKT ++ F RS + G IIP V +YLG ++
Sbjct: 234 ERLEILDRISSYYGLKIDQSKTVLLKNKF----CRSQDVLFNGSPIIP-VPGCRYLGRWI 288
Query: 133 QNDGEIEADVSHRIQAGWLKWRRASGVLCDKKVPLKLKGKFYRTAIRPALLYGTECW 189
G I+ ++S RI+AGW VL + +P K ++ + PALLY ++ W
Sbjct: 289 DISGSIDEEISRRIRAGWGALVGIKEVL--RIMPNKENIILFKQNVLPALLYASKTW 343
>UniRef100_UPI000024B6D5 UPI000024B6D5 UniRef100 entry
Length = 728
Score = 76.6 bits (187), Expect = 5e-13
Identities = 55/213 (25%), Positives = 100/213 (46%), Gaps = 13/213 (6%)
Query: 8 SVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQE-------LAPRCMLFADD 60
+V+ +E F I+ G+ QG L+P LF + +L H L R MLFADD
Sbjct: 521 TVQFNGSCSEPFSISSGVKQGCVLAPTLFGIFFALLLRHAFGSATEGIYLRTRDMLFADD 580
Query: 61 VVLVGESREEVNGRLETWRQALEAYGFRLSRSKTEYMEWNFSGRRSRSTLEVKVGDHIIP 120
+ ++EE+ + + A + +G +S KT N + + + + V D+ +
Sbjct: 581 AAVTTHTQEELQSLMTRFSMACKDFGLTISLKKT-----NVLSQDTATPPTITVDDYQLD 635
Query: 121 QVTRFKYLGSFVQNDGEIEADVSHRI-QAGWLKWRRASGVLCDKKVPLKLKGKFYRTAIR 179
V + YLGS + ++ ++A++ RI +A R + V + ++ K Y +
Sbjct: 636 VVHQLTYLGSTITDNLSLDAELDRRIGKAASTLARLTTRVWTNHRLTTATKMALYNACVI 695
Query: 180 PALLYGTECWAVKSQHENQVSVTEMRMLRWMSG 212
LLYG+E W ++ E +++ +R LR + G
Sbjct: 696 STLLYGSETWTTYARQERRLNTFHLRGLRRILG 728
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.136 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 388,374,485
Number of Sequences: 2790947
Number of extensions: 15269523
Number of successful extensions: 32284
Number of sequences better than 10.0: 1170
Number of HSP's better than 10.0 without gapping: 283
Number of HSP's successfully gapped in prelim test: 887
Number of HSP's that attempted gapping in prelim test: 31482
Number of HSP's gapped (non-prelim): 1220
length of query: 239
length of database: 848,049,833
effective HSP length: 124
effective length of query: 115
effective length of database: 501,972,405
effective search space: 57726826575
effective search space used: 57726826575
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 73 (32.7 bits)
Medicago: description of AC141114.8


