

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
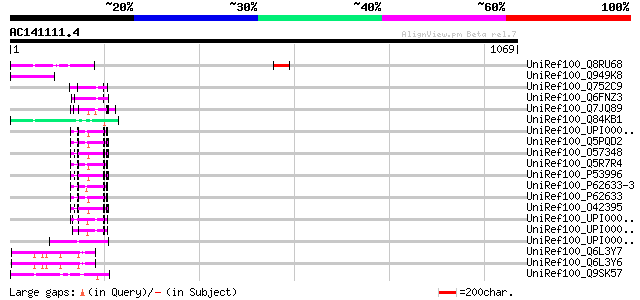
Query= AC141111.4 - phase: 0 /pseudo
(1069 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8RU68 Putative 22 kDa kafirin cluster; Ty3-Gypsy type... 89 5e-16
UniRef100_Q949K8 Gag polyprotein [Cicer arietinum] 67 3e-09
UniRef100_Q752C9 AFR646Wp [Ashbya gossypii] 67 4e-09
UniRef100_Q6FNZ3 Similar to sp|P53849 Saccharomyces cerevisiae Y... 65 8e-09
UniRef100_Q7JQ89 CnjB protein [Tetrahymena thermophila] 65 1e-08
UniRef100_Q84KB1 Gag-protease polyprotein [Cucumis melo] 63 4e-08
UniRef100_UPI00003AA82A UPI00003AA82A UniRef100 entry 63 5e-08
UniRef100_Q5PQD2 Hypothetical protein [Xenopus laevis] 63 5e-08
UniRef100_O57348 Cellular nucleic acid binding protein [Gallus g... 63 5e-08
UniRef100_Q5R7R4 Hypothetical protein DKFZp470E2414 [Pongo pygma... 63 5e-08
UniRef100_P53996 Cellular nucleic acid binding protein [Mus musc... 63 5e-08
UniRef100_P62633-3 Splice isoform 3 of P62633 [Homo sapiens] 63 5e-08
UniRef100_P62633 Cellular nucleic acid binding protein [Homo sap... 63 5e-08
UniRef100_O42395 Cellular nucleic acid binding protein [Gallus g... 63 5e-08
UniRef100_UPI000036346C UPI000036346C UniRef100 entry 62 7e-08
UniRef100_UPI0000318F83 UPI0000318F83 UniRef100 entry 62 7e-08
UniRef100_UPI000049964B UPI000049964B UniRef100 entry 62 1e-07
UniRef100_Q6L3Y7 Putative gag-pol polyprotein [Solanum demissum] 62 1e-07
UniRef100_Q6L3Y6 Putative gag-pol polyprotein [Solanum demissum] 62 1e-07
UniRef100_Q9SK57 Putative retroelement pol polyprotein [Arabidop... 62 1e-07
>UniRef100_Q8RU68 Putative 22 kDa kafirin cluster; Ty3-Gypsy type [Oryza sativa]
Length = 1230
Score = 89.4 bits (220), Expect = 5e-16
Identities = 58/180 (32%), Positives = 93/180 (51%), Gaps = 12/180 (6%)
Query: 1 RREFLNRYFPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELAKFYPHYTAETAEFSKCIK 60
+ F +YFPE V+ KE EFLELKQG+ SV EY +F LA+F P + + SK +
Sbjct: 147 KEAFYKKYFPESVKRMKEKEFLELKQGNKSVAEYEIEFSRLARFAPEFV--QTDGSKARR 204
Query: 61 IENGLRADIKRAIGYQKIRNFSELVSSCRIYEEDTKAHYKVMSERRGKGQLSRPKPYSAP 120
E+GLR +KR + ++ F E+VS ++ E K +++ +R GQ + + P
Sbjct: 205 FESGLRQPLKRRVEAFELTIFREVVSKAQLLE---KGYHE---QRIEHGQPQKKFKTNNP 258
Query: 121 PDKGK*RLKDERRPKMRDAPTDIVCFKCG--EKGHKSNVCDRDEKKCFRCGKKGHTLADC 178
++G R + +M+ ++ KC + H ++C +CF CG+ GHT C
Sbjct: 259 QNQG--RFRGNYSGQMQRKSSENQGRKCPICQGSHVPSICPNCWGRCFECGEAGHTRYQC 316
Score = 49.3 bits (116), Expect = 6e-04
Identities = 24/34 (70%), Positives = 30/34 (87%), Gaps = 1/34 (2%)
Query: 557 LDKFVVVFIDDILIYSETEEEHAEHLK*VFLATL 590
LD+FVVVFIDDILIYS+T+E+HA HL+ + L TL
Sbjct: 641 LDQFVVVFIDDILIYSKTKEDHANHLR-IVLQTL 673
>UniRef100_Q949K8 Gag polyprotein [Cicer arietinum]
Length = 277
Score = 67.0 bits (162), Expect = 3e-09
Identities = 34/93 (36%), Positives = 56/93 (59%)
Query: 1 RREFLNRYFPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELAKFYPHYTAETAEFSKCIK 60
+R+FL++YFP R K +FL+L QG ++V EYAAKF L++ + + E E C +
Sbjct: 133 KRKFLDKYFPMSTRTKLGDDFLKLHQGSLTVGEYAAKFESLSRHFRFFREEIDEPFMCHR 192
Query: 61 IENGLRADIKRAIGYQKIRNFSELVSSCRIYEE 93
++GL+ +I+ ++ I+ F LV CR E+
Sbjct: 193 FQDGLKYEIQDSVLPLGIQRFQALVEKCREVED 225
>UniRef100_Q752C9 AFR646Wp [Ashbya gossypii]
Length = 163
Score = 66.6 bits (161), Expect = 4e-09
Identities = 29/74 (39%), Positives = 37/74 (49%), Gaps = 2/74 (2%)
Query: 127 RLKDERRPKMRDAPTDIVCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCKRG--DIV 184
R + R + C++CG H + C +DE KC+ CGK GH DC G +
Sbjct: 87 RFSNSSRSSFSGRLNKVSCYRCGGPNHMAKDCLQDETKCYSCGKSGHISRDCPSGPSEKT 146
Query: 185 CYNFNEEGHISLQC 198
CYN NE GHIS C
Sbjct: 147 CYNCNESGHISRDC 160
Score = 53.9 bits (128), Expect = 3e-05
Identities = 21/66 (31%), Positives = 36/66 (53%), Gaps = 6/66 (9%)
Query: 144 VCFKCGEKGHKSNVC----DRDEKKCFRCGKKGHTLADCKRGDIVCYNFNEEGHISLQCT 199
+C+ C GH + C + K+C+ CG+ GH +C C+N ++ GH+S CT
Sbjct: 24 LCYNCNMPGHIQSECTLPRSAEHKQCYNCGETGHVRGECNIQK--CFNCSQAGHVSRDCT 81
Query: 200 QPKKVR 205
+P++ R
Sbjct: 82 EPRRSR 87
Score = 47.0 bits (110), Expect = 0.003
Identities = 22/80 (27%), Positives = 30/80 (37%), Gaps = 18/80 (22%)
Query: 145 CFKCGEKGHKSNVCDRDEKK------------------CFRCGKKGHTLADCKRGDIVCY 186
CF C + GH S C + C+RCG H DC + + CY
Sbjct: 67 CFNCSQAGHVSRDCTEPRRSRFSNSSRSSFSGRLNKVSCYRCGGPNHMAKDCLQDETKCY 126
Query: 187 NFNEEGHISLQCTQPKKVRT 206
+ + GHIS C +T
Sbjct: 127 SCGKSGHISRDCPSGPSEKT 146
Score = 44.3 bits (103), Expect = 0.020
Identities = 20/41 (48%), Positives = 26/41 (62%), Gaps = 1/41 (2%)
Query: 162 EKKCFRCGKKGHTLADCKRGDIVCYNFNEEGHISLQCTQPK 202
+K C+ CGK GH LAD + +CYN N GHI +CT P+
Sbjct: 3 QKACYVCGKLGH-LADNCDSERLCYNCNMPGHIQSECTLPR 42
>UniRef100_Q6FNZ3 Similar to sp|P53849 Saccharomyces cerevisiae YNL255c GIS2 [Candida
glabrata]
Length = 155
Score = 65.5 bits (158), Expect = 8e-09
Identities = 28/71 (39%), Positives = 40/71 (55%), Gaps = 2/71 (2%)
Query: 130 DERRPKMRDAPTDIVCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCKRG--DIVCYN 187
+ R+ + A ++ C+KCG H + C + + KC+ CG+ GH DC G + VCYN
Sbjct: 82 EPRKGRFGAASKNVSCYKCGGPNHVARDCMQTDTKCYSCGRFGHVSRDCPNGPNEKVCYN 141
Query: 188 FNEEGHISLQC 198
NE GHIS C
Sbjct: 142 CNETGHISRDC 152
Score = 60.1 bits (144), Expect = 4e-07
Identities = 24/74 (32%), Positives = 42/74 (56%), Gaps = 6/74 (8%)
Query: 138 DAPTDIVCFKCGEKGHKSNVCDR----DEKKCFRCGKKGHTLADCKRGDIVCYNFNEEGH 193
D ++ +C+ C + GH + C + K+C+ CG+ GH ++C C+N N+ GH
Sbjct: 18 DCDSERLCYNCNQPGHVQSECTMPRTVEHKQCYNCGETGHVKSECSIQR--CFNCNQTGH 75
Query: 194 ISLQCTQPKKVRTG 207
+S +C +P+K R G
Sbjct: 76 VSRECPEPRKGRFG 89
Score = 48.5 bits (114), Expect = 0.001
Identities = 19/43 (44%), Positives = 27/43 (62%), Gaps = 1/43 (2%)
Query: 162 EKKCFRCGKKGHTLADCKRGDIVCYNFNEEGHISLQCTQPKKV 204
+K C+ CGK GH DC + +CYN N+ GH+ +CT P+ V
Sbjct: 3 QKACYVCGKIGHLADDCD-SERLCYNCNQPGHVQSECTMPRTV 44
Score = 42.4 bits (98), Expect = 0.076
Identities = 19/66 (28%), Positives = 31/66 (46%), Gaps = 12/66 (18%)
Query: 145 CFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADC---KRG-------DIVCYNFNEEGHI 194
C+ CGE GH + C ++CF C + GH +C ++G ++ CY H+
Sbjct: 49 CYNCGETGHVKSECSI--QRCFNCNQTGHVSRECPEPRKGRFGAASKNVSCYKCGGPNHV 106
Query: 195 SLQCTQ 200
+ C Q
Sbjct: 107 ARDCMQ 112
>UniRef100_Q7JQ89 CnjB protein [Tetrahymena thermophila]
Length = 1748
Score = 64.7 bits (156), Expect = 1e-08
Identities = 30/82 (36%), Positives = 43/82 (51%), Gaps = 9/82 (10%)
Query: 135 KMRDAPTDIVCFKCGEKGHKSNVCDRDEKK-----CFRCGKKGHTLADCKR----GDIVC 185
+ + P CFKCGE+GH S C +K+ CF+C ++GH DC G C
Sbjct: 1520 QQQQKPRGGACFKCGEEGHISKDCPNPQKQQQKNTCFKCKQEGHISKDCPNSQNSGGNKC 1579
Query: 186 YNFNEEGHISLQCTQPKKVRTG 207
+N N+EGH+S C P + + G
Sbjct: 1580 FNCNQEGHMSKDCPNPSQKKKG 1601
Score = 62.8 bits (151), Expect = 5e-08
Identities = 33/106 (31%), Positives = 48/106 (45%), Gaps = 12/106 (11%)
Query: 129 KDERRPKMRDAPTDIVCFKCGEKGHKSNVCDRDEKK---CFRCGKKGHTLADC------- 178
KD P+ + CFKC ++GH S C ++K CF+CG++GH DC
Sbjct: 1462 KDCTEPQQQGRKQSGACFKCNQEGHMSKDCPNQQQKKSGCFKCGEEGHFSKDCPNPQKQQ 1521
Query: 179 --KRGDIVCYNFNEEGHISLQCTQPKKVRTGGKVFALTGTQTVNED 222
K C+ EEGHIS C P+K + F +++D
Sbjct: 1522 QQKPRGGACFKCGEEGHISKDCPNPQKQQQKNTCFKCKQEGHISKD 1567
Score = 58.5 bits (140), Expect = 1e-06
Identities = 29/71 (40%), Positives = 39/71 (54%), Gaps = 13/71 (18%)
Query: 145 CFKCGEKGHKSNVCDRDEK----KCFRCGKKGHTLADC------KRGDIVCYNFNEEGHI 194
CFKC ++GH S C + KCF C ++GH DC K+G C+N EEGH
Sbjct: 1555 CFKCKQEGHISKDCPNSQNSGGNKCFNCNQEGHMSKDCPNPSQKKKG---CFNCGEEGHQ 1611
Query: 195 SLQCTQPKKVR 205
S +CT+ +K R
Sbjct: 1612 SRECTKERKER 1622
Score = 53.9 bits (128), Expect = 3e-05
Identities = 24/69 (34%), Positives = 36/69 (51%), Gaps = 10/69 (14%)
Query: 145 CFKCGEKGHKSNVCDRDEKK-------CFRCGKKGHTLADC---KRGDIVCYNFNEEGHI 194
CFKCG+ GH + C +++ CF+C ++GH DC ++ C+ EEGH
Sbjct: 1451 CFKCGKVGHMAKDCTEPQQQGRKQSGACFKCNQEGHMSKDCPNQQQKKSGCFKCGEEGHF 1510
Query: 195 SLQCTQPKK 203
S C P+K
Sbjct: 1511 SKDCPNPQK 1519
Score = 43.1 bits (100), Expect = 0.045
Identities = 18/52 (34%), Positives = 28/52 (53%), Gaps = 7/52 (13%)
Query: 163 KKCFRCGKKGHTLADC-------KRGDIVCYNFNEEGHISLQCTQPKKVRTG 207
K CF+CGK GH DC ++ C+ N+EGH+S C ++ ++G
Sbjct: 1449 KGCFKCGKVGHMAKDCTEPQQQGRKQSGACFKCNQEGHMSKDCPNQQQKKSG 1500
>UniRef100_Q84KB1 Gag-protease polyprotein [Cucumis melo]
Length = 429
Score = 63.2 bits (152), Expect = 4e-08
Identities = 60/238 (25%), Positives = 95/238 (39%), Gaps = 27/238 (11%)
Query: 1 RREFLNRYFPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELAKFYPHYTAETAEFSKCIK 60
+ F ++F +R K EFL L+QGDM+V +Y A+F L++F P A E ++ K
Sbjct: 138 KESFYAKFFSASLRDAKRQEFLNLEQGDMTVEQYDAEFDMLSRFAPEMIA--TEAARADK 195
Query: 61 IENGLRADIKRAI-GYQKIRNFSELVSSCRIYEEDTKAHYKVMSERRGKGQLSRPKPYSA 119
GLR DI+ + ++ + L + + ++ K GQ + +
Sbjct: 196 FVRGLRLDIQGLVRAFRPATHADALRLAVDLSLQERANSSKTAGRGSTSGQKRKAEQQPV 255
Query: 120 PPDKGK*RLKDERRPKMRDAPTDIVCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCK 179
P + R E R + P F+ GE R + C CGK H L C
Sbjct: 256 PVPQRNFRPGGEFR-SFQQKP-----FEAGEAA-------RGKPLCTTCGK--HHLGRCL 300
Query: 180 RGDIVCYNFNEEGHISLQC---------TQPKKVRTGGKVFALTGTQTVNED*LIRGT 228
G C+ +EGH + +C Q G+ FA T+ ++ GT
Sbjct: 301 FGTRTCFKCRQEGHTADRCPLRVTGIAQNQGAGAPHQGRAFATNRTEAEKAGTVVTGT 358
>UniRef100_UPI00003AA82A UPI00003AA82A UniRef100 entry
Length = 171
Score = 62.8 bits (151), Expect = 5e-08
Identities = 28/77 (36%), Positives = 44/77 (56%), Gaps = 6/77 (7%)
Query: 129 KDERRPKMRDAPTDIVCFKCGEKGHKSNVCDR-DEKKCFRCGKKGHTLADCKRGDIVCYN 187
KD + PK + C+ CG+ GH + CD DE+KC+ CG+ GH DC + + CY
Sbjct: 79 KDCKEPKRE---REQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTK--VKCYR 133
Query: 188 FNEEGHISLQCTQPKKV 204
E GH+++ C++ +V
Sbjct: 134 CGETGHVAINCSKTSEV 150
Score = 60.8 bits (146), Expect = 2e-07
Identities = 23/59 (38%), Positives = 33/59 (54%), Gaps = 4/59 (6%)
Query: 144 VCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCK----RGDIVCYNFNEEGHISLQC 198
+C++CGE GH + CD E C+ CG+ GH DCK + CYN + GH++ C
Sbjct: 47 ICYRCGESGHLAKDCDLQEDACYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDC 105
Score = 56.6 bits (135), Expect = 4e-06
Identities = 23/56 (41%), Positives = 33/56 (58%), Gaps = 3/56 (5%)
Query: 145 CFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADC-KRGDIVCYNFNEEGHISLQCT 199
C+ CGE GH C + KC+RCG+ GH +C K ++ CY E GH++ +CT
Sbjct: 113 CYSCGEFGHIQKDCTK--VKCYRCGETGHVAINCSKTSEVNCYRCGESGHLARECT 166
Score = 54.3 bits (129), Expect = 2e-05
Identities = 26/83 (31%), Positives = 34/83 (40%), Gaps = 22/83 (26%)
Query: 145 CFKCGEKGHKSNVCDRDEKK----------------------CFRCGKKGHTLADCKRGD 182
CFKCG GH + C + C+RCG+ GH DC +
Sbjct: 6 CFKCGRTGHWARECPTGIGRGRGMRSRGRAGFQFMSSSLPDICYRCGESGHLAKDCDLQE 65
Query: 183 IVCYNFNEEGHISLQCTQPKKVR 205
CYN GHI+ C +PK+ R
Sbjct: 66 DACYNCGRGGHIAKDCKEPKRER 88
Score = 43.9 bits (102), Expect = 0.026
Identities = 16/39 (41%), Positives = 24/39 (61%), Gaps = 1/39 (2%)
Query: 141 TDIVCFKCGEKGHKSNVCDR-DEKKCFRCGKKGHTLADC 178
T + C++CGE GH + C + E C+RCG+ GH +C
Sbjct: 127 TKVKCYRCGETGHVAINCSKTSEVNCYRCGESGHLAREC 165
>UniRef100_Q5PQD2 Hypothetical protein [Xenopus laevis]
Length = 179
Score = 62.8 bits (151), Expect = 5e-08
Identities = 28/77 (36%), Positives = 44/77 (56%), Gaps = 6/77 (7%)
Query: 129 KDERRPKMRDAPTDIVCFKCGEKGHKSNVCDR-DEKKCFRCGKKGHTLADCKRGDIVCYN 187
KD + PK + C+ CG+ GH + CD DE+KC+ CG+ GH DC + + CY
Sbjct: 87 KDCKEPKRE---REQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTK--VKCYR 141
Query: 188 FNEEGHISLQCTQPKKV 204
E GH+++ C++ +V
Sbjct: 142 CGETGHVAINCSKTSEV 158
Score = 57.0 bits (136), Expect = 3e-06
Identities = 23/61 (37%), Positives = 34/61 (55%), Gaps = 6/61 (9%)
Query: 144 VCFKCGEKGHKSNVCD--RDEKKCFRCGKKGHTLADCK----RGDIVCYNFNEEGHISLQ 197
+C++CGE GH + CD D + C+ CG+ GH DCK + CYN + GH++
Sbjct: 53 ICYRCGESGHLAKDCDLQEDVEACYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARD 112
Query: 198 C 198
C
Sbjct: 113 C 113
Score = 56.6 bits (135), Expect = 4e-06
Identities = 23/56 (41%), Positives = 33/56 (58%), Gaps = 3/56 (5%)
Query: 145 CFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADC-KRGDIVCYNFNEEGHISLQCT 199
C+ CGE GH C + KC+RCG+ GH +C K ++ CY E GH++ +CT
Sbjct: 121 CYSCGEFGHIQKDCTK--VKCYRCGETGHVAINCSKTSEVNCYRCGESGHLARECT 174
Score = 55.5 bits (132), Expect = 9e-06
Identities = 26/68 (38%), Positives = 34/68 (49%), Gaps = 5/68 (7%)
Query: 145 CFKCGEKGHKSNVCDRDEKK----CFRCGKKGHTLADCKRGDIV-CYNFNEEGHISLQCT 199
C+ CG GH + C +++ C+ CGK GH DC D CY+ E GHI CT
Sbjct: 76 CYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCT 135
Query: 200 QPKKVRTG 207
+ K R G
Sbjct: 136 KVKCYRCG 143
Score = 49.3 bits (116), Expect = 6e-04
Identities = 27/91 (29%), Positives = 35/91 (37%), Gaps = 30/91 (32%)
Query: 145 CFKCGEKGHKSNVCDRDEKK----------------------------CFRCGKKGHTLA 176
CFKCG GH + C + C+RCG+ GH
Sbjct: 6 CFKCGRSGHWARECPTGGGRGRGMRSRGRGGFTSDRGFQFVSSSLPDICYRCGESGHLAK 65
Query: 177 DCKRGDIV--CYNFNEEGHISLQCTQPKKVR 205
DC + V CYN GHI+ C +PK+ R
Sbjct: 66 DCDLQEDVEACYNCGRGGHIAKDCKEPKRER 96
Score = 43.9 bits (102), Expect = 0.026
Identities = 16/39 (41%), Positives = 24/39 (61%), Gaps = 1/39 (2%)
Query: 141 TDIVCFKCGEKGHKSNVCDR-DEKKCFRCGKKGHTLADC 178
T + C++CGE GH + C + E C+RCG+ GH +C
Sbjct: 135 TKVKCYRCGETGHVAINCSKTSEVNCYRCGESGHLAREC 173
>UniRef100_O57348 Cellular nucleic acid binding protein [Gallus gallus]
Length = 172
Score = 62.8 bits (151), Expect = 5e-08
Identities = 28/77 (36%), Positives = 44/77 (56%), Gaps = 6/77 (7%)
Query: 129 KDERRPKMRDAPTDIVCFKCGEKGHKSNVCDR-DEKKCFRCGKKGHTLADCKRGDIVCYN 187
KD + PK + C+ CG+ GH + CD DE+KC+ CG+ GH DC + + CY
Sbjct: 80 KDCKEPKRE---REQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTK--VKCYR 134
Query: 188 FNEEGHISLQCTQPKKV 204
E GH+++ C++ +V
Sbjct: 135 CGETGHVAINCSKTSEV 151
Score = 57.8 bits (138), Expect = 2e-06
Identities = 28/75 (37%), Positives = 36/75 (47%), Gaps = 5/75 (6%)
Query: 138 DAPTDIVCFKCGEKGHKSNVCDRDEKK----CFRCGKKGHTLADCKRGDIV-CYNFNEEG 192
D D C+ CG GH + C +++ C+ CGK GH DC D CY+ E G
Sbjct: 62 DLQEDEACYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDCDHADEQKCYSCGEFG 121
Query: 193 HISLQCTQPKKVRTG 207
HI CT+ K R G
Sbjct: 122 HIQKDCTKVKCYRCG 136
Score = 57.8 bits (138), Expect = 2e-06
Identities = 22/60 (36%), Positives = 36/60 (59%), Gaps = 5/60 (8%)
Query: 144 VCFKCGEKGHKSNVCD-RDEKKCFRCGKKGHTLADCK----RGDIVCYNFNEEGHISLQC 198
+C++CGE GH + CD ++++ C+ CG+ GH DCK + CYN + GH++ C
Sbjct: 47 ICYRCGESGHLAKDCDLQEDEACYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDC 106
Score = 56.6 bits (135), Expect = 4e-06
Identities = 23/56 (41%), Positives = 33/56 (58%), Gaps = 3/56 (5%)
Query: 145 CFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADC-KRGDIVCYNFNEEGHISLQCT 199
C+ CGE GH C + KC+RCG+ GH +C K ++ CY E GH++ +CT
Sbjct: 114 CYSCGEFGHIQKDCTK--VKCYRCGETGHVAINCSKTSEVNCYRCGESGHLARECT 167
Score = 52.4 bits (124), Expect = 7e-05
Identities = 27/84 (32%), Positives = 35/84 (41%), Gaps = 23/84 (27%)
Query: 145 CFKCGEKGHKSNVCDRDEKK----------------------CFRCGKKGHTLADCK-RG 181
CFKCG GH + C + C+RCG+ GH DC +
Sbjct: 6 CFKCGRTGHWARECPTGIGRGRGMRSRGRAGFQFMSSSLPDICYRCGESGHLAKDCDLQE 65
Query: 182 DIVCYNFNEEGHISLQCTQPKKVR 205
D CYN GHI+ C +PK+ R
Sbjct: 66 DEACYNCGRGGHIAKDCKEPKRER 89
Score = 43.9 bits (102), Expect = 0.026
Identities = 16/39 (41%), Positives = 24/39 (61%), Gaps = 1/39 (2%)
Query: 141 TDIVCFKCGEKGHKSNVCDR-DEKKCFRCGKKGHTLADC 178
T + C++CGE GH + C + E C+RCG+ GH +C
Sbjct: 128 TKVKCYRCGETGHVAINCSKTSEVNCYRCGESGHLAREC 166
>UniRef100_Q5R7R4 Hypothetical protein DKFZp470E2414 [Pongo pygmaeus]
Length = 170
Score = 62.8 bits (151), Expect = 5e-08
Identities = 28/77 (36%), Positives = 44/77 (56%), Gaps = 6/77 (7%)
Query: 129 KDERRPKMRDAPTDIVCFKCGEKGHKSNVCDR-DEKKCFRCGKKGHTLADCKRGDIVCYN 187
KD + PK + C+ CG+ GH + CD DE+KC+ CG+ GH DC + + CY
Sbjct: 78 KDCKEPKRE---REQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTK--VKCYR 132
Query: 188 FNEEGHISLQCTQPKKV 204
E GH+++ C++ +V
Sbjct: 133 CGETGHVAINCSKTSEV 149
Score = 60.8 bits (146), Expect = 2e-07
Identities = 23/59 (38%), Positives = 33/59 (54%), Gaps = 4/59 (6%)
Query: 144 VCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCK----RGDIVCYNFNEEGHISLQC 198
+C++CGE GH + CD E C+ CG+ GH DCK + CYN + GH++ C
Sbjct: 46 ICYRCGESGHLAKDCDLQEDACYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDC 104
Score = 56.6 bits (135), Expect = 4e-06
Identities = 23/56 (41%), Positives = 33/56 (58%), Gaps = 3/56 (5%)
Query: 145 CFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADC-KRGDIVCYNFNEEGHISLQCT 199
C+ CGE GH C + KC+RCG+ GH +C K ++ CY E GH++ +CT
Sbjct: 112 CYSCGEFGHIQKDCTK--VKCYRCGETGHVAINCSKTSEVNCYRCGESGHLARECT 165
Score = 55.5 bits (132), Expect = 9e-06
Identities = 26/82 (31%), Positives = 34/82 (40%), Gaps = 21/82 (25%)
Query: 145 CFKCGEKGHKSNVCDRDEKK---------------------CFRCGKKGHTLADCKRGDI 183
CFKCG GH + C + C+RCG+ GH DC +
Sbjct: 6 CFKCGRSGHWARECPTGGGRGRGMRSRGRGFQFVSSSLPDICYRCGESGHLAKDCDLQED 65
Query: 184 VCYNFNEEGHISLQCTQPKKVR 205
CYN GHI+ C +PK+ R
Sbjct: 66 ACYNCGRGGHIAKDCKEPKRER 87
Score = 43.9 bits (102), Expect = 0.026
Identities = 16/39 (41%), Positives = 24/39 (61%), Gaps = 1/39 (2%)
Query: 141 TDIVCFKCGEKGHKSNVCDR-DEKKCFRCGKKGHTLADC 178
T + C++CGE GH + C + E C+RCG+ GH +C
Sbjct: 126 TKVKCYRCGETGHVAINCSKTSEVNCYRCGESGHLAREC 164
>UniRef100_P53996 Cellular nucleic acid binding protein [Mus musculus]
Length = 178
Score = 62.8 bits (151), Expect = 5e-08
Identities = 28/77 (36%), Positives = 44/77 (56%), Gaps = 6/77 (7%)
Query: 129 KDERRPKMRDAPTDIVCFKCGEKGHKSNVCDR-DEKKCFRCGKKGHTLADCKRGDIVCYN 187
KD + PK + C+ CG+ GH + CD DE+KC+ CG+ GH DC + + CY
Sbjct: 86 KDCKEPKRE---REQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTK--VKCYR 140
Query: 188 FNEEGHISLQCTQPKKV 204
E GH+++ C++ +V
Sbjct: 141 CGETGHVAINCSKTSEV 157
Score = 57.8 bits (138), Expect = 2e-06
Identities = 28/75 (37%), Positives = 36/75 (47%), Gaps = 5/75 (6%)
Query: 138 DAPTDIVCFKCGEKGHKSNVCDRDEKK----CFRCGKKGHTLADCKRGDIV-CYNFNEEG 192
D D C+ CG GH + C +++ C+ CGK GH DC D CY+ E G
Sbjct: 68 DLQEDEACYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDCDHADEQKCYSCGEFG 127
Query: 193 HISLQCTQPKKVRTG 207
HI CT+ K R G
Sbjct: 128 HIQKDCTKVKCYRCG 142
Score = 57.8 bits (138), Expect = 2e-06
Identities = 22/60 (36%), Positives = 36/60 (59%), Gaps = 5/60 (8%)
Query: 144 VCFKCGEKGHKSNVCD-RDEKKCFRCGKKGHTLADCK----RGDIVCYNFNEEGHISLQC 198
+C++CGE GH + CD ++++ C+ CG+ GH DCK + CYN + GH++ C
Sbjct: 53 ICYRCGESGHLAKDCDLQEDEACYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDC 112
Score = 56.6 bits (135), Expect = 4e-06
Identities = 23/56 (41%), Positives = 33/56 (58%), Gaps = 3/56 (5%)
Query: 145 CFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADC-KRGDIVCYNFNEEGHISLQCT 199
C+ CGE GH C + KC+RCG+ GH +C K ++ CY E GH++ +CT
Sbjct: 120 CYSCGEFGHIQKDCTK--VKCYRCGETGHVAINCSKTSEVNCYRCGESGHLARECT 173
Score = 50.8 bits (120), Expect = 2e-04
Identities = 27/90 (30%), Positives = 35/90 (38%), Gaps = 29/90 (32%)
Query: 145 CFKCGEKGHKSNVCDRDEKK----------------------------CFRCGKKGHTLA 176
CFKCG GH + C + C+RCG+ GH
Sbjct: 6 CFKCGRSGHWARECPTGGGRGRGMRSRGRGGFTSDRGFQFVSSSLPDICYRCGESGHLAK 65
Query: 177 DCK-RGDIVCYNFNEEGHISLQCTQPKKVR 205
DC + D CYN GHI+ C +PK+ R
Sbjct: 66 DCDLQEDEACYNCGRGGHIAKDCKEPKRER 95
Score = 43.9 bits (102), Expect = 0.026
Identities = 16/39 (41%), Positives = 24/39 (61%), Gaps = 1/39 (2%)
Query: 141 TDIVCFKCGEKGHKSNVCDR-DEKKCFRCGKKGHTLADC 178
T + C++CGE GH + C + E C+RCG+ GH +C
Sbjct: 134 TKVKCYRCGETGHVAINCSKTSEVNCYRCGESGHLAREC 172
>UniRef100_P62633-3 Splice isoform 3 of P62633 [Homo sapiens]
Length = 167
Score = 62.8 bits (151), Expect = 5e-08
Identities = 28/77 (36%), Positives = 44/77 (56%), Gaps = 6/77 (7%)
Query: 129 KDERRPKMRDAPTDIVCFKCGEKGHKSNVCDR-DEKKCFRCGKKGHTLADCKRGDIVCYN 187
KD + PK + C+ CG+ GH + CD DE+KC+ CG+ GH DC + + CY
Sbjct: 75 KDCKEPKRE---REQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTK--VKCYR 129
Query: 188 FNEEGHISLQCTQPKKV 204
E GH+++ C++ +V
Sbjct: 130 CGETGHVAINCSKTSEV 146
Score = 60.8 bits (146), Expect = 2e-07
Identities = 23/59 (38%), Positives = 33/59 (54%), Gaps = 4/59 (6%)
Query: 144 VCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCK----RGDIVCYNFNEEGHISLQC 198
+C++CGE GH + CD E C+ CG+ GH DCK + CYN + GH++ C
Sbjct: 43 ICYRCGESGHLAKDCDLQEDACYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDC 101
Score = 58.5 bits (140), Expect = 1e-06
Identities = 27/79 (34%), Positives = 34/79 (42%), Gaps = 18/79 (22%)
Query: 145 CFKCGEKGHKSNVCDR------------------DEKKCFRCGKKGHTLADCKRGDIVCY 186
CFKCG GH + C D C+RCG+ GH DC + CY
Sbjct: 6 CFKCGRSGHWARECPTGGGRGRGMRSRGRGGFTSDRDICYRCGESGHLAKDCDLQEDACY 65
Query: 187 NFNEEGHISLQCTQPKKVR 205
N GHI+ C +PK+ R
Sbjct: 66 NCGRGGHIAKDCKEPKRER 84
Score = 56.6 bits (135), Expect = 4e-06
Identities = 23/56 (41%), Positives = 33/56 (58%), Gaps = 3/56 (5%)
Query: 145 CFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADC-KRGDIVCYNFNEEGHISLQCT 199
C+ CGE GH C + KC+RCG+ GH +C K ++ CY E GH++ +CT
Sbjct: 109 CYSCGEFGHIQKDCTK--VKCYRCGETGHVAINCSKTSEVNCYRCGESGHLARECT 162
Score = 43.9 bits (102), Expect = 0.026
Identities = 16/39 (41%), Positives = 24/39 (61%), Gaps = 1/39 (2%)
Query: 141 TDIVCFKCGEKGHKSNVCDR-DEKKCFRCGKKGHTLADC 178
T + C++CGE GH + C + E C+RCG+ GH +C
Sbjct: 123 TKVKCYRCGETGHVAINCSKTSEVNCYRCGESGHLAREC 161
>UniRef100_P62633 Cellular nucleic acid binding protein [Homo sapiens]
Length = 177
Score = 62.8 bits (151), Expect = 5e-08
Identities = 28/77 (36%), Positives = 44/77 (56%), Gaps = 6/77 (7%)
Query: 129 KDERRPKMRDAPTDIVCFKCGEKGHKSNVCDR-DEKKCFRCGKKGHTLADCKRGDIVCYN 187
KD + PK + C+ CG+ GH + CD DE+KC+ CG+ GH DC + + CY
Sbjct: 85 KDCKEPKRE---REQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTK--VKCYR 139
Query: 188 FNEEGHISLQCTQPKKV 204
E GH+++ C++ +V
Sbjct: 140 CGETGHVAINCSKTSEV 156
Score = 60.8 bits (146), Expect = 2e-07
Identities = 23/59 (38%), Positives = 33/59 (54%), Gaps = 4/59 (6%)
Query: 144 VCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCK----RGDIVCYNFNEEGHISLQC 198
+C++CGE GH + CD E C+ CG+ GH DCK + CYN + GH++ C
Sbjct: 53 ICYRCGESGHLAKDCDLQEDACYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDC 111
Score = 56.6 bits (135), Expect = 4e-06
Identities = 23/56 (41%), Positives = 33/56 (58%), Gaps = 3/56 (5%)
Query: 145 CFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADC-KRGDIVCYNFNEEGHISLQCT 199
C+ CGE GH C + KC+RCG+ GH +C K ++ CY E GH++ +CT
Sbjct: 119 CYSCGEFGHIQKDCTK--VKCYRCGETGHVAINCSKTSEVNCYRCGESGHLARECT 172
Score = 52.8 bits (125), Expect = 6e-05
Identities = 26/89 (29%), Positives = 34/89 (37%), Gaps = 28/89 (31%)
Query: 145 CFKCGEKGHKSNVCDRDEKK----------------------------CFRCGKKGHTLA 176
CFKCG GH + C + C+RCG+ GH
Sbjct: 6 CFKCGRSGHWARECPTGGGRGRGMRSRGRGGFTSDRGFQFVSSSLPDICYRCGESGHLAK 65
Query: 177 DCKRGDIVCYNFNEEGHISLQCTQPKKVR 205
DC + CYN GHI+ C +PK+ R
Sbjct: 66 DCDLQEDACYNCGRGGHIAKDCKEPKRER 94
Score = 43.9 bits (102), Expect = 0.026
Identities = 16/39 (41%), Positives = 24/39 (61%), Gaps = 1/39 (2%)
Query: 141 TDIVCFKCGEKGHKSNVCDR-DEKKCFRCGKKGHTLADC 178
T + C++CGE GH + C + E C+RCG+ GH +C
Sbjct: 133 TKVKCYRCGETGHVAINCSKTSEVNCYRCGESGHLAREC 171
>UniRef100_O42395 Cellular nucleic acid binding protein [Gallus gallus]
Length = 172
Score = 62.8 bits (151), Expect = 5e-08
Identities = 28/77 (36%), Positives = 44/77 (56%), Gaps = 6/77 (7%)
Query: 129 KDERRPKMRDAPTDIVCFKCGEKGHKSNVCDR-DEKKCFRCGKKGHTLADCKRGDIVCYN 187
KD + PK + C+ CG+ GH + CD DE+KC+ CG+ GH DC + + CY
Sbjct: 80 KDCKEPKRE---REQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTK--VKCYR 134
Query: 188 FNEEGHISLQCTQPKKV 204
E GH+++ C++ +V
Sbjct: 135 CGETGHVAINCSKTSEV 151
Score = 59.3 bits (142), Expect = 6e-07
Identities = 23/60 (38%), Positives = 36/60 (59%), Gaps = 5/60 (8%)
Query: 144 VCFKCGEKGHKSNVCD-RDEKKCFRCGKKGHTLADCK----RGDIVCYNFNEEGHISLQC 198
+C++CGE GH + CD +++K C+ CG+ GH DCK + CYN + GH++ C
Sbjct: 47 ICYRCGESGHLAKDCDLQEDKACYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDC 106
Score = 57.8 bits (138), Expect = 2e-06
Identities = 28/75 (37%), Positives = 36/75 (47%), Gaps = 5/75 (6%)
Query: 138 DAPTDIVCFKCGEKGHKSNVCDRDEKK----CFRCGKKGHTLADCKRGDIV-CYNFNEEG 192
D D C+ CG GH + C +++ C+ CGK GH DC D CY+ E G
Sbjct: 62 DLQEDKACYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDCDHADEQKCYSCGEFG 121
Query: 193 HISLQCTQPKKVRTG 207
HI CT+ K R G
Sbjct: 122 HIQKDCTKVKCYRCG 136
Score = 56.6 bits (135), Expect = 4e-06
Identities = 23/56 (41%), Positives = 33/56 (58%), Gaps = 3/56 (5%)
Query: 145 CFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADC-KRGDIVCYNFNEEGHISLQCT 199
C+ CGE GH C + KC+RCG+ GH +C K ++ CY E GH++ +CT
Sbjct: 114 CYSCGEFGHIQKDCTK--VKCYRCGETGHVAINCSKTSEVNCYRCGESGHLARECT 167
Score = 52.4 bits (124), Expect = 7e-05
Identities = 27/84 (32%), Positives = 35/84 (41%), Gaps = 23/84 (27%)
Query: 145 CFKCGEKGHKSNVCDRDEKK----------------------CFRCGKKGHTLADCK-RG 181
CFKCG GH + C + C+RCG+ GH DC +
Sbjct: 6 CFKCGRTGHWARECPTGIGRGRGMRSRGRAGFQFMSSSLPDICYRCGESGHLAKDCDLQE 65
Query: 182 DIVCYNFNEEGHISLQCTQPKKVR 205
D CYN GHI+ C +PK+ R
Sbjct: 66 DKACYNCGRGGHIAKDCKEPKRER 89
Score = 43.9 bits (102), Expect = 0.026
Identities = 16/39 (41%), Positives = 24/39 (61%), Gaps = 1/39 (2%)
Query: 141 TDIVCFKCGEKGHKSNVCDR-DEKKCFRCGKKGHTLADC 178
T + C++CGE GH + C + E C+RCG+ GH +C
Sbjct: 128 TKVKCYRCGETGHVAINCSKTSEVNCYRCGESGHLAREC 166
>UniRef100_UPI000036346C UPI000036346C UniRef100 entry
Length = 167
Score = 62.4 bits (150), Expect = 7e-08
Identities = 26/71 (36%), Positives = 38/71 (52%), Gaps = 4/71 (5%)
Query: 132 RRPKMRDAPTDIVCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCK----RGDIVCYN 187
R P+ R + C++CGE GH + CD+ E C+ C K GH DCK + +CYN
Sbjct: 31 RGPRGRGRVKEQFCYRCGEHGHIARDCDQPEDSCYNCHKSGHISRDCKEPKREREHLCYN 90
Query: 188 FNEEGHISLQC 198
+ GH++ C
Sbjct: 91 CGKAGHVARDC 101
Score = 59.3 bits (142), Expect = 6e-07
Identities = 27/76 (35%), Positives = 37/76 (48%), Gaps = 15/76 (19%)
Query: 145 CFKCGEKGHKSNVCDRD---------------EKKCFRCGKKGHTLADCKRGDIVCYNFN 189
CF CG GH C + E+ C+RCG+ GH DC + + CYN +
Sbjct: 9 CFGCGRTGHWIKDCPKSSGPRGRGPRGRGRVKEQFCYRCGEHGHIARDCDQPEDSCYNCH 68
Query: 190 EEGHISLQCTQPKKVR 205
+ GHIS C +PK+ R
Sbjct: 69 KSGHISRDCKEPKRER 84
Score = 58.5 bits (140), Expect = 1e-06
Identities = 24/56 (42%), Positives = 33/56 (58%), Gaps = 3/56 (5%)
Query: 145 CFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADC-KRGDIVCYNFNEEGHISLQCT 199
C+ CGE GH +CD+ KC+RCG+ GH C K + CYN + GH++ CT
Sbjct: 109 CYSCGEFGHIQKLCDK--VKCYRCGEIGHVAVQCSKASETNCYNCGKAGHVARDCT 162
Score = 54.3 bits (129), Expect = 2e-05
Identities = 24/73 (32%), Positives = 43/73 (58%), Gaps = 6/73 (8%)
Query: 129 KDERRPKMRDAPTDIVCFKCGEKGHKSNVCDR-DEKKCFRCGKKGHTLADCKRGDIVCYN 187
+D + PK + +C+ CG+ GH + C+ +E+KC+ CG+ GH C + + CY
Sbjct: 75 RDCKEPKRE---REHLCYNCGKAGHVARDCEHANEQKCYSCGEFGHIQKLCDK--VKCYR 129
Query: 188 FNEEGHISLQCTQ 200
E GH+++QC++
Sbjct: 130 CGEIGHVAVQCSK 142
Score = 38.5 bits (88), Expect = 1.1
Identities = 17/54 (31%), Positives = 24/54 (43%), Gaps = 15/54 (27%)
Query: 164 KCFRCGKKGHTLADCKRG---------------DIVCYNFNEEGHISLQCTQPK 202
+CF CG+ GH + DC + + CY E GHI+ C QP+
Sbjct: 8 ECFGCGRTGHWIKDCPKSSGPRGRGPRGRGRVKEQFCYRCGEHGHIARDCDQPE 61
>UniRef100_UPI0000318F83 UPI0000318F83 UniRef100 entry
Length = 167
Score = 62.4 bits (150), Expect = 7e-08
Identities = 25/71 (35%), Positives = 39/71 (54%), Gaps = 4/71 (5%)
Query: 132 RRPKMRDAPTDIVCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCK----RGDIVCYN 187
R P+ R ++ C++CG++GH CD+ E C+ C K GH DCK + CYN
Sbjct: 31 RGPRGRGRGKELFCYRCGDQGHMVKDCDQTEDSCYNCHKSGHISRDCKEPKREREQQCYN 90
Query: 188 FNEEGHISLQC 198
+ GH++ +C
Sbjct: 91 CGKAGHMAREC 101
Score = 57.8 bits (138), Expect = 2e-06
Identities = 27/76 (35%), Positives = 36/76 (46%), Gaps = 15/76 (19%)
Query: 145 CFKCGEKGHKSNVCDRD---------------EKKCFRCGKKGHTLADCKRGDIVCYNFN 189
CF CG GH C E C+RCG +GH + DC + + CYN +
Sbjct: 9 CFGCGRPGHWVKNCPTSSGLRGRGPRGRGRGKELFCYRCGDQGHMVKDCDQTEDSCYNCH 68
Query: 190 EEGHISLQCTQPKKVR 205
+ GHIS C +PK+ R
Sbjct: 69 KSGHISRDCKEPKRER 84
Score = 57.0 bits (136), Expect = 3e-06
Identities = 24/56 (42%), Positives = 31/56 (54%), Gaps = 3/56 (5%)
Query: 145 CFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADC-KRGDIVCYNFNEEGHISLQCT 199
CF CG GH +CD+ KC+RCG GH C K + CYN + GH++ CT
Sbjct: 109 CFTCGTLGHIQKLCDK--VKCYRCGGIGHVALQCSKASETTCYNCGKAGHVAKDCT 162
Score = 55.1 bits (131), Expect = 1e-05
Identities = 23/57 (40%), Positives = 34/57 (59%), Gaps = 3/57 (5%)
Query: 145 CFKCGEKGHKSNVCDR-DEKKCFRCGKKGHTLADCKRGDIVCYNFNEEGHISLQCTQ 200
C+ CG+ GH + CD +E+KCF CG GH C + + CY GH++LQC++
Sbjct: 88 CYNCGKAGHMARECDHANEQKCFTCGTLGHIQKLCDK--VKCYRCGGIGHVALQCSK 142
Score = 40.4 bits (93), Expect = 0.29
Identities = 15/37 (40%), Positives = 21/37 (56%), Gaps = 1/37 (2%)
Query: 143 IVCFKCGEKGHKSNVCDR-DEKKCFRCGKKGHTLADC 178
+ C++CG GH + C + E C+ CGK GH DC
Sbjct: 125 VKCYRCGGIGHVALQCSKASETTCYNCGKAGHVAKDC 161
>UniRef100_UPI000049964B UPI000049964B UniRef100 entry
Length = 389
Score = 61.6 bits (148), Expect = 1e-07
Identities = 43/129 (33%), Positives = 58/129 (44%), Gaps = 6/129 (4%)
Query: 85 VSSCRIYEEDTKAHYKVMSERRGKGQLSRPKPYSAPPDKGK*RLKDERRPKMRDAPTDIV 144
VS +I E KA K + ++ K + + K GK + P+ + +D
Sbjct: 234 VSMAKISERGQKAMTKKVKKQLEKEEKEKLKALKKCIICGKIGHTSKDCPQNENKGSDC- 292
Query: 145 CFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADC---KRGDIVCYNFNEEGHISLQC--T 199
CF CGE GH S C E+KCF CGK GH DC K + C+ E GH+ C
Sbjct: 293 CFICGETGHISKDCPNAERKCFVCGKTGHKSRDCPKAKGNNRPCFICGEIGHLDRDCPNK 352
Query: 200 QPKKVRTGG 208
KK + GG
Sbjct: 353 NEKKEKKGG 361
>UniRef100_Q6L3Y7 Putative gag-pol polyprotein [Solanum demissum]
Length = 1351
Score = 61.6 bits (148), Expect = 1e-07
Identities = 56/212 (26%), Positives = 95/212 (44%), Gaps = 39/212 (18%)
Query: 4 FLNRYFPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELAKFYPHYTAE--------TAEF 55
FL +FP +++ K EFL LKQ +SV EY+ KF +L+++ P A+ A
Sbjct: 106 FLGHFFPRELKEVKVREFLTLKQESLSVHEYSLKFTQLSRYAPEMVADMRNRMSLFVAGL 165
Query: 56 SKCIKIENGLRA------DIKRAIGY------QKIRNFSELVSSCRIYEEDTKAHYKVMS 103
S+ + + G A DI R + Y +K+R+ E + R+ + + S
Sbjct: 166 SR-LSSKEGRAAMLIGDMDISRLMVYVQQVEEEKLRDREEFRNK-RVKTRNESGQQRGNS 223
Query: 104 ER-----RGKGQLSRPKPYSAPPDKGK*RLKDERRPKMRDAPTD----------IVCFKC 148
R R KG + AP +G+ +++ + K+ A + C KC
Sbjct: 224 NRSSFQQRQKGPATSSARAPAPRYRGEHNVQNSKDFKVTPAQSSGSVVRGGSSFPACAKC 283
Query: 149 GEKGHKSNVCDRDEKKCFRCGKKGHTLADCKR 180
G + H C + + CFRCG++GH + +C +
Sbjct: 284 G-RVHPGK-CRQGQTCCFRCGQEGHFMKECPK 313
Score = 43.9 bits (102), Expect = 0.026
Identities = 19/27 (70%), Positives = 22/27 (81%)
Query: 557 LDKFVVVFIDDILIYSETEEEHAEHLK 583
LD FV+VFIDDILIYS E +HA HL+
Sbjct: 585 LDMFVIVFIDDILIYSRNEGDHASHLR 611
>UniRef100_Q6L3Y6 Putative gag-pol polyprotein [Solanum demissum]
Length = 1351
Score = 61.6 bits (148), Expect = 1e-07
Identities = 56/212 (26%), Positives = 95/212 (44%), Gaps = 39/212 (18%)
Query: 4 FLNRYFPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELAKFYPHYTAE--------TAEF 55
FL +FP +++ K EFL LKQ +SV EY+ KF +L+++ P A+ A
Sbjct: 106 FLGHFFPRELKEVKVREFLTLKQESLSVHEYSLKFTQLSRYAPEMVADMRNRMSLFVAGL 165
Query: 56 SKCIKIENGLRA------DIKRAIGY------QKIRNFSELVSSCRIYEEDTKAHYKVMS 103
S+ + + G A DI R + Y +K+R+ E + R+ + + S
Sbjct: 166 SR-LSSKEGRAAMLIGDMDISRLMVYVQQVEEEKLRDREEFRNK-RVKTRNESGQQRGNS 223
Query: 104 ER-----RGKGQLSRPKPYSAPPDKGK*RLKDERRPKMRDAPTD----------IVCFKC 148
R R KG + AP +G+ +++ + K+ A + C KC
Sbjct: 224 NRSSFQQRQKGPATSSARAPAPRYRGEHNVQNSKDFKVTPAQSSGSVVRGGSSFPACAKC 283
Query: 149 GEKGHKSNVCDRDEKKCFRCGKKGHTLADCKR 180
G + H C + + CFRCG++GH + +C +
Sbjct: 284 G-RVHPGK-CRQGQTCCFRCGQEGHFMKECPK 313
Score = 43.9 bits (102), Expect = 0.026
Identities = 19/27 (70%), Positives = 22/27 (81%)
Query: 557 LDKFVVVFIDDILIYSETEEEHAEHLK 583
LD FV+VFIDDILIYS E +HA HL+
Sbjct: 585 LDMFVIVFIDDILIYSRNEGDHASHLR 611
>UniRef100_Q9SK57 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1611
Score = 61.6 bits (148), Expect = 1e-07
Identities = 56/218 (25%), Positives = 90/218 (40%), Gaps = 22/218 (10%)
Query: 3 EFLNRYFPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELAKFYPHYTAETAEFSKCIKIE 62
EF +YFP + + E +L+L QG+ +V EY +F L ++ E E ++ +
Sbjct: 248 EFNKKYFPPEAWDRLECAYLDLVQGNRTVREYDEEFNRLRRYVGRELEE--EQAQLRRFI 305
Query: 63 NGLRADIKRAIGYQKIRNFSELVSSCRIYEEDTKAHYKVMSERRGKGQLSRPKPYSAPPD 122
GLR +I+ + + SELV + EE + + E+ R + P D
Sbjct: 306 RGLRIEIRNHCLVRTFNSVSELVERAAMIEEGIEEERYLNREKAP----IRNNQSTKPAD 361
Query: 123 KGK*RLKDERRPKMRDAPTDIVCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCKRGD 182
K R D+ DA T C CG K H S C + C RCG K H + C + +
Sbjct: 362 KK--RKFDKVDNTKSDAKTG-ECVTCG-KNH-SGTCWKAIGACGRCGSKDHAIQSCPKME 416
Query: 183 -----------IVCYNFNEEGHISLQCTQPKKVRTGGK 209
C+ + GH+ +C + + G+
Sbjct: 417 PGQSKVLGEETRTCFYCGKTGHLKRECPKLTAEKQAGQ 454
Score = 41.2 bits (95), Expect = 0.17
Identities = 16/27 (59%), Positives = 24/27 (88%)
Query: 557 LDKFVVVFIDDILIYSETEEEHAEHLK 583
LD+FV++FI+DIL+YS++ E H EHL+
Sbjct: 804 LDEFVIIFINDILVYSKSWEAHQEHLR 830
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.369 0.167 0.628
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,349,396,461
Number of Sequences: 2790947
Number of extensions: 46157742
Number of successful extensions: 260690
Number of sequences better than 10.0: 3658
Number of HSP's better than 10.0 without gapping: 790
Number of HSP's successfully gapped in prelim test: 2878
Number of HSP's that attempted gapping in prelim test: 243217
Number of HSP's gapped (non-prelim): 12791
length of query: 1069
length of database: 848,049,833
effective HSP length: 138
effective length of query: 931
effective length of database: 462,899,147
effective search space: 430959105857
effective search space used: 430959105857
T: 11
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 80 (35.4 bits)
Medicago: description of AC141111.4


