

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141111.3 + phase: 0 /pseudo
(64 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
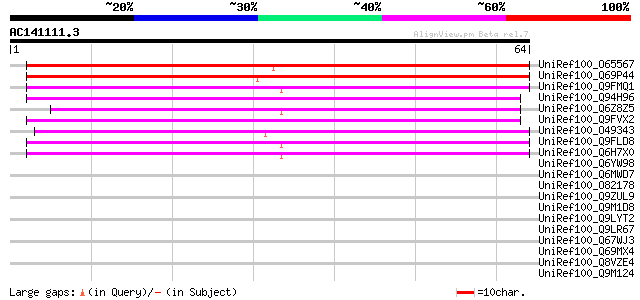
Sequences producing significant alignments: (bits) Value
UniRef100_O65567 Puative protein [Arabidopsis thaliana] 87 1e-16
UniRef100_Q69P44 Putative fertility restorer [Oryza sativa] 69 3e-11
UniRef100_Q9FMQ1 Gb|AAF19552.1 [Arabidopsis thaliana] 48 7e-05
UniRef100_Q94H96 Hypothetical protein OSJNBb0048A17.10 [Oryza sa... 48 7e-05
UniRef100_Q6Z8Z5 Putative pentatricopeptide (PPR) repeat-contain... 47 2e-04
UniRef100_Q9FVX2 Hypothetical protein F2P24.7 [Arabidopsis thali... 47 2e-04
UniRef100_O49343 Hypothetical protein At2g30780 [Arabidopsis tha... 45 4e-04
UniRef100_Q9FLD8 Gb|AAD55291.1 [Arabidopsis thaliana] 45 5e-04
UniRef100_Q6H7X0 Putative pentatricopeptide (PPR) repeat-contain... 45 5e-04
UniRef100_Q6YW98 PPR protein-like protein [Oryza sativa] 44 0.001
UniRef100_Q6MWD7 B1358B12.18 protein [Oryza sativa] 44 0.001
UniRef100_O82178 Hypothetical protein At2g35130 [Arabidopsis tha... 44 0.001
UniRef100_Q9ZUL9 Hypothetical protein At2g02150 [Arabidopsis tha... 43 0.002
UniRef100_Q9M1D8 Hypothetical protein T2O9.30 [Arabidopsis thali... 43 0.002
UniRef100_Q9LYT2 Hypothetical protein F17J16_90 [Arabidopsis tha... 43 0.002
UniRef100_Q9LR67 F21B7.18 [Arabidopsis thaliana] 43 0.002
UniRef100_Q67WJ3 Pentatricopeptide (PPR) repeat-containing prote... 42 0.003
UniRef100_Q69MX4 Pentatricopeptide (PPR) repeat-containing prote... 42 0.003
UniRef100_Q8VZE4 AT4g01570/T15B16_21 [Arabidopsis thaliana] 42 0.003
UniRef100_Q9M124 Hypothetical protein AT4g01570 [Arabidopsis tha... 42 0.003
>UniRef100_O65567 Puative protein [Arabidopsis thaliana]
Length = 1075
Score = 87.0 bits (214), Expect = 1e-16
Identities = 41/63 (65%), Positives = 50/63 (79%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGG-RIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
I+ YNT+ITGYGK M+ A+G+F L +EPDETSYRSMIEGWGRA NYE+A+ YY+
Sbjct: 520 IIAYNTLITGYGKIFKMEAAQGLFHRLCNIGLEPDETSYRSMIEGWGRADNYEEAKHYYQ 579
Query: 62 ELK 64
ELK
Sbjct: 580 ELK 582
Score = 30.8 bits (68), Expect = 9.4
Identities = 17/55 (30%), Positives = 29/55 (51%), Gaps = 2/55 (3%)
Query: 6 YNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNY-EKARW 58
YNT+I YG ++ A G+ + GR I PD+ +Y +++ R + E +W
Sbjct: 1012 YNTLIKAYGIGGMVEEAVGLVKEMRGRNIIPDKVTYTNLVTALRRNDEFLEAIKW 1066
>UniRef100_Q69P44 Putative fertility restorer [Oryza sativa]
Length = 962
Score = 68.9 bits (167), Expect = 3e-11
Identities = 35/63 (55%), Positives = 45/63 (70%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+V YNT+ITGYGK S+M A VF L + PDET+YRSMIEG+GRA Y++A YY
Sbjct: 402 VVAYNTVITGYGKVSDMQKAMEVFDRLKSAGLAPDETTYRSMIEGFGRADKYKQAILYYR 461
Query: 62 ELK 64
+L+
Sbjct: 462 KLR 464
Score = 35.8 bits (81), Expect = 0.29
Identities = 22/60 (36%), Positives = 32/60 (52%), Gaps = 1/60 (1%)
Query: 6 YNTMITGYGKASNMDGAEGVFLTLGGRI-EPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
YN MI YG+ ++G V L R EPD SY ++I+ +G AG E A +E++
Sbjct: 859 YNIMINIYGRKGWIEGVANVLAELKSRGGEPDLYSYNTLIKAYGIAGMPEDAVKLMQEMR 918
>UniRef100_Q9FMQ1 Gb|AAF19552.1 [Arabidopsis thaliana]
Length = 816
Score = 47.8 bits (112), Expect = 7e-05
Identities = 23/63 (36%), Positives = 37/63 (58%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+V YNT+I G + AE + L + + ++PD +Y S+I G+G AGN ++ YE
Sbjct: 564 LVTYNTLIDGLSMTGKLSEAEDLLLEISRKGLKPDVFTYNSLISGYGFAGNVQRCIALYE 623
Query: 62 ELK 64
E+K
Sbjct: 624 EMK 626
Score = 40.4 bits (93), Expect = 0.012
Identities = 22/63 (34%), Positives = 36/63 (56%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVF-LTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+ +YN +I G K M+ AE +F L R+ P +Y ++I+G+ +AGN EK+ E
Sbjct: 214 VFIYNVLIDGLCKGKRMNDAEQLFDEMLARRLLPSLITYNTLIDGYCKAGNPEKSFKVRE 273
Query: 62 ELK 64
+K
Sbjct: 274 RMK 276
Score = 35.4 bits (80), Expect = 0.38
Identities = 20/63 (31%), Positives = 35/63 (54%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
++ YNT+I GY KA N + + V + IEP ++ ++++G +AG E A +
Sbjct: 249 LITYNTLIDGYCKAGNPEKSFKVRERMKADHIEPSLITFNTLLKGLFKAGMVEDAENVLK 308
Query: 62 ELK 64
E+K
Sbjct: 309 EMK 311
Score = 32.0 bits (71), Expect = 4.2
Identities = 18/62 (29%), Positives = 31/62 (49%), Gaps = 1/62 (1%)
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
V+YNTMI GY + ++ GA + + ++PD +Y +I + G E A +
Sbjct: 390 VIYNTMIDGYCRKGDLVGARMKIEAMEKQGMKPDHLAYNCLIRRFCELGEMENAEKEVNK 449
Query: 63 LK 64
+K
Sbjct: 450 MK 451
>UniRef100_Q94H96 Hypothetical protein OSJNBb0048A17.10 [Oryza sativa]
Length = 484
Score = 47.8 bits (112), Expect = 7e-05
Identities = 20/61 (32%), Positives = 34/61 (54%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
+ +N+++ K+ N+ A+ +F + R PD +Y ++EGWGRA N K R Y E
Sbjct: 170 LAAFNSLLGALCKSKNVRKAQEIFDKMNSRFSPDAKTYSILLEGWGRAPNLPKMREVYSE 229
Query: 63 L 63
+
Sbjct: 230 M 230
>UniRef100_Q6Z8Z5 Putative pentatricopeptide (PPR) repeat-containing protein [Oryza
sativa]
Length = 747
Score = 46.6 bits (109), Expect = 2e-04
Identities = 22/59 (37%), Positives = 35/59 (59%), Gaps = 1/59 (1%)
Query: 6 YNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYEEL 63
YNT+I G+G M+ A VF + + PD T+Y +++ W RAG+ E AR ++E+
Sbjct: 245 YNTLIWGFGLCKKMEAALRVFGDMKDHGVTPDVTTYNTLLNAWVRAGDLESARKVFDEM 303
Score = 36.2 bits (82), Expect = 0.22
Identities = 18/62 (29%), Positives = 33/62 (53%), Gaps = 1/62 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+ YNT++ + +A +++ A VF + G + SY MI+G+ AG E+A +
Sbjct: 277 VTTYNTLLNAWVRAGDLESARKVFDEMPGAGFAQNSVSYNVMIKGYVEAGKVEEAVGLFS 336
Query: 62 EL 63
E+
Sbjct: 337 EM 338
>UniRef100_Q9FVX2 Hypothetical protein F2P24.7 [Arabidopsis thaliana]
Length = 481
Score = 46.6 bits (109), Expect = 2e-04
Identities = 20/61 (32%), Positives = 35/61 (56%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
+V +N +++ K+ N+ A+ VF + R PD +Y ++EGWG+ N KAR + E
Sbjct: 167 LVAFNGLLSALCKSKNVRKAQEVFENMRDRFTPDSKTYSILLEGWGKEPNLPKAREVFRE 226
Query: 63 L 63
+
Sbjct: 227 M 227
>UniRef100_O49343 Hypothetical protein At2g30780 [Arabidopsis thaliana]
Length = 452
Score = 45.4 bits (106), Expect = 4e-04
Identities = 23/62 (37%), Positives = 33/62 (53%), Gaps = 1/62 (1%)
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTLG-GRIEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
V YN +I GY A N D E F + G +EPD +Y+ M+ G+ +GN + YE
Sbjct: 210 VTYNFLIAGYMTAWNWDKMEATFQEMKRGPVEPDTDTYQLMLRGYANSGNLNRMEEMYEV 269
Query: 63 LK 64
+K
Sbjct: 270 IK 271
Score = 35.4 bits (80), Expect = 0.38
Identities = 17/63 (26%), Positives = 34/63 (52%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGG-RIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+V YN +++ YG+ + E F L ++ P+ +Y +I G+ A N++K ++
Sbjct: 174 VVTYNILVSVYGRLLMVKNMEAAFEELQKVKLPPNSVTYNFLIAGYMTAWNWDKMEATFQ 233
Query: 62 ELK 64
E+K
Sbjct: 234 EMK 236
>UniRef100_Q9FLD8 Gb|AAD55291.1 [Arabidopsis thaliana]
Length = 678
Score = 45.1 bits (105), Expect = 5e-04
Identities = 22/63 (34%), Positives = 37/63 (57%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+V YNTMI YGK + A + + R IEP+ +Y ++I WG+AG ++A ++
Sbjct: 400 VVTYNTMIKIYGKTMEHEKATNLVQEMQSRGIEPNAITYSTIISIWGKAGKLDRAATLFQ 459
Query: 62 ELK 64
+L+
Sbjct: 460 KLR 462
Score = 36.2 bits (82), Expect = 0.22
Identities = 21/62 (33%), Positives = 34/62 (53%), Gaps = 1/62 (1%)
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
+ Y+T+I+ +GKA +D A +F L +E D+ Y++MI + R G A+ E
Sbjct: 436 ITYSTIISIWGKAGKLDRAATLFQKLRSSGVEIDQVLYQTMIVAYERVGLMGHAKRLLHE 495
Query: 63 LK 64
LK
Sbjct: 496 LK 497
>UniRef100_Q6H7X0 Putative pentatricopeptide (PPR) repeat-containing protein [Oryza
sativa]
Length = 397
Score = 45.1 bits (105), Expect = 5e-04
Identities = 22/63 (34%), Positives = 34/63 (53%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+ YN ++ Y +A GA +F + EPD SY +++ +GRAG +E A +E
Sbjct: 152 VYAYNALMEAYSRAGLPQGASEIFSLMQHMGCEPDRASYNILVDAYGRAGLHEDAEAVFE 211
Query: 62 ELK 64
ELK
Sbjct: 212 ELK 214
>UniRef100_Q6YW98 PPR protein-like protein [Oryza sativa]
Length = 794
Score = 43.9 bits (102), Expect = 0.001
Identities = 26/63 (41%), Positives = 37/63 (58%), Gaps = 3/63 (4%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVF--LTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYY 60
+V YNT+I G KA N+DGA +F L L G I PDE +Y ++I+G RA A +
Sbjct: 535 VVTYNTLINGLCKARNLDGAVRLFKELQLKG-ISPDEITYGTLIDGLLRAHRENDAMMLF 593
Query: 61 EEL 63
+ +
Sbjct: 594 QNI 596
>UniRef100_Q6MWD7 B1358B12.18 protein [Oryza sativa]
Length = 609
Score = 43.9 bits (102), Expect = 0.001
Identities = 22/62 (35%), Positives = 38/62 (60%), Gaps = 1/62 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLG-GRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+V +NTMI+G +A ++DGAE + + + PD +Y ++I+G R G E AR +E
Sbjct: 275 VVSFNTMISGMCRAGDLDGAETLHRRMSEAGVTPDVYTYGALIQGLCRVGRIEDARGVFE 334
Query: 62 EL 63
++
Sbjct: 335 KM 336
Score = 31.2 bits (69), Expect = 7.2
Identities = 18/62 (29%), Positives = 31/62 (49%), Gaps = 1/62 (1%)
Query: 4 VVYNTMITGYGKASNMDGAEGVF-LTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
V Y +I+G KA AE V + +EPD T+Y +I+ + R G+ + +E
Sbjct: 451 VTYTALISGLSKAGRSADAERVLGEMMEAGLEPDNTTYTMVIDAFCRKGDVKTGLRLLKE 510
Query: 63 LK 64
++
Sbjct: 511 MQ 512
>UniRef100_O82178 Hypothetical protein At2g35130 [Arabidopsis thaliana]
Length = 591
Score = 43.9 bits (102), Expect = 0.001
Identities = 22/63 (34%), Positives = 34/63 (53%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+ VYN ++ Y +A GA +F + EPD SY M++ +GRAG + A +E
Sbjct: 334 VYVYNALMESYSRAGYPYGAAEIFSLMQHMGCEPDRASYNIMVDAYGRAGLHSDAEAVFE 393
Query: 62 ELK 64
E+K
Sbjct: 394 EMK 396
Score = 33.1 bits (74), Expect = 1.9
Identities = 20/62 (32%), Positives = 31/62 (49%), Gaps = 1/62 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYE 61
I YN +I YGKA ++ E +F+ L + PD ++ S I + R Y K +E
Sbjct: 474 ISTYNILINIYGKAGFLERIEELFVELKEKNFRPDVVTWTSRIGAYSRKKLYVKCLEVFE 533
Query: 62 EL 63
E+
Sbjct: 534 EM 535
Score = 32.0 bits (71), Expect = 4.2
Identities = 18/60 (30%), Positives = 33/60 (55%), Gaps = 1/60 (1%)
Query: 6 YNTMITGYGKASNMDGAEGVFLTLGG-RIEPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
YN MI YGKAS + ++ + + +P+ +Y +++ + R G EKA +E+L+
Sbjct: 267 YNLMINLYGKASKSYMSWKLYCEMRSHQCKPNICTYTALVNAFAREGLCEKAEEIFEQLQ 326
Score = 30.8 bits (68), Expect = 9.4
Identities = 17/63 (26%), Positives = 32/63 (49%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
++ +N +I YG+ AE +++ L R P E +Y +I+ + AG E+A
Sbjct: 155 VICFNLLIDAYGQKFQYKEAESLYVQLLESRYVPTEDTYALLIKAYCMAGLIERAEVVLV 214
Query: 62 ELK 64
E++
Sbjct: 215 EMQ 217
>UniRef100_Q9ZUL9 Hypothetical protein At2g02150 [Arabidopsis thaliana]
Length = 1107
Score = 43.1 bits (100), Expect = 0.002
Identities = 21/63 (33%), Positives = 36/63 (56%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+ YN MI K +++ A G+F + R + PD +Y SMI+G+G+ G + ++E
Sbjct: 130 VFTYNIMIDCMCKEGDVEAARGLFEEMKFRGLVPDTVTYNSMIDGFGKVGRLDDTVCFFE 189
Query: 62 ELK 64
E+K
Sbjct: 190 EMK 192
Score = 34.3 bits (77), Expect = 0.85
Identities = 19/62 (30%), Positives = 30/62 (47%), Gaps = 1/62 (1%)
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTLGGRI-EPDETSYRSMIEGWGRAGNYEKARWYYEE 62
V YN+MI G+GK +D F + EPD +Y ++I + + G +Y E
Sbjct: 166 VTYNSMIDGFGKVGRLDDTVCFFEEMKDMCCEPDVITYNALINCFCKFGKLPIGLEFYRE 225
Query: 63 LK 64
+K
Sbjct: 226 MK 227
Score = 33.9 bits (76), Expect = 1.1
Identities = 23/60 (38%), Positives = 30/60 (49%), Gaps = 1/60 (1%)
Query: 6 YNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
YN +I G+ KA NMD A + L GR I+PD Y + I G E A+ E+K
Sbjct: 343 YNALIHGFVKAKNMDRALELLNELKGRGIKPDLLLYGTFIWGLCSLEKIEAAKVVMNEMK 402
Score = 32.7 bits (73), Expect = 2.5
Identities = 20/63 (31%), Positives = 32/63 (50%), Gaps = 1/63 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLG-GRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+V Y +I G A M AE +F + + P+ SY ++I G+ +A N ++A
Sbjct: 305 VVTYTALIDGLCDAERMKEAEELFGKMDTAGVIPNLASYNALIHGFVKAKNMDRALELLN 364
Query: 62 ELK 64
ELK
Sbjct: 365 ELK 367
>UniRef100_Q9M1D8 Hypothetical protein T2O9.30 [Arabidopsis thaliana]
Length = 473
Score = 43.1 bits (100), Expect = 0.002
Identities = 23/64 (35%), Positives = 39/64 (60%), Gaps = 3/64 (4%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVF--LTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYY 60
+V Y MITGY + +D A+ +F +T+ G++ P+ +Y SMI G AG + +A W
Sbjct: 359 VVCYTVMITGYVVSGELDKAKEMFREMTVKGQL-PNVFTYNSMIRGLCMAGEFREACWLL 417
Query: 61 EELK 64
+E++
Sbjct: 418 KEME 421
Score = 30.8 bits (68), Expect = 9.4
Identities = 18/63 (28%), Positives = 32/63 (50%), Gaps = 3/63 (4%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLT--LGGRIEPDETSYRSMIEGWGRAGNYEKARWYY 60
++ Y T+I G +A N++ + FL + PD Y MI G+ +G +KA+ +
Sbjct: 324 VLHYTTLIDGLSRAGNLEACK-YFLDEMVKAGCRPDVVCYTVMITGYVVSGELDKAKEMF 382
Query: 61 EEL 63
E+
Sbjct: 383 REM 385
>UniRef100_Q9LYT2 Hypothetical protein F17J16_90 [Arabidopsis thaliana]
Length = 526
Score = 42.7 bits (99), Expect = 0.002
Identities = 21/60 (35%), Positives = 37/60 (61%), Gaps = 1/60 (1%)
Query: 6 YNTMITGYGKASNMDGAEGVFLTLG-GRIEPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
Y TM++ Y AS+M+GAE F + EP+ +Y ++I+G+ +A + EK YE+++
Sbjct: 330 YTTMLSAYVNASDMEGAEKFFKRIKVDGFEPNIVTYGTLIKGYAKANDVEKMMEVYEKMR 389
Score = 36.6 bits (83), Expect = 0.17
Identities = 19/57 (33%), Positives = 29/57 (50%), Gaps = 1/57 (1%)
Query: 9 MITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
+IT YGK N +GAE V L P+ SY +++E +GR G A + ++
Sbjct: 88 LITAYGKLGNFNGAERVLSVLSKMGSTPNVISYTALMESYGRGGKCNNAEAIFRRMQ 144
Score = 36.2 bits (82), Expect = 0.22
Identities = 20/64 (31%), Positives = 39/64 (60%), Gaps = 3/64 (4%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVF--LTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYY 60
IV Y T+I GY KA++++ V+ + L G I+ ++T ++++ GR N+ A +Y
Sbjct: 362 IVTYGTLIKGYAKANDVEKMMEVYEKMRLSG-IKANQTILTTIMDASGRCKNFGSALGWY 420
Query: 61 EELK 64
+E++
Sbjct: 421 KEME 424
Score = 36.2 bits (82), Expect = 0.22
Identities = 20/64 (31%), Positives = 32/64 (49%), Gaps = 4/64 (6%)
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTL----GGRIEPDETSYRSMIEGWGRAGNYEKARWY 59
+ Y ++ + + AE VF TL ++PD+ Y MI + +AGNYEKAR
Sbjct: 153 ITYQIILKTFVEGDKFKEAEEVFETLLDEKKSPLKPDQKMYHMMIYMYKKAGNYEKARKV 212
Query: 60 YEEL 63
+ +
Sbjct: 213 FSSM 216
>UniRef100_Q9LR67 F21B7.18 [Arabidopsis thaliana]
Length = 660
Score = 42.7 bits (99), Expect = 0.002
Identities = 22/55 (40%), Positives = 34/55 (61%), Gaps = 1/55 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKA 56
+ YN ++ G A +D AE VF + GRI+PD +Y +MI+G+ +AG +KA
Sbjct: 222 LYTYNFLMNGLVSAMFVDSAERVFEVMESGRIKPDIVTYNTMIKGYCKAGQTQKA 276
>UniRef100_Q67WJ3 Pentatricopeptide (PPR) repeat-containing protein-like [Oryza
sativa]
Length = 908
Score = 42.4 bits (98), Expect = 0.003
Identities = 21/61 (34%), Positives = 39/61 (63%), Gaps = 3/61 (4%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
+V +NTMI+GY +AS+M +E +F + +PD S+ +I+G+ + G E AR +++
Sbjct: 306 LVSWNTMISGYTQASDMKESEKLFWEMP---DPDTVSWNLIIQGFMQKGEAEHARGFFDR 362
Query: 63 L 63
+
Sbjct: 363 M 363
>UniRef100_Q69MX4 Pentatricopeptide (PPR) repeat-containing protein-like [Oryza
sativa]
Length = 612
Score = 42.4 bits (98), Expect = 0.003
Identities = 20/59 (33%), Positives = 37/59 (61%), Gaps = 3/59 (5%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+V +N+MI+GY + +M+ A +F + R D S+ SM++G+ +AG+ E AR ++
Sbjct: 219 VVTWNSMISGYSRHGDMENARKMFDAMPER---DVVSWNSMLDGYAQAGDVEMARLVFD 274
Score = 40.4 bits (93), Expect = 0.012
Identities = 20/61 (32%), Positives = 33/61 (53%), Gaps = 3/61 (4%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
+V YN+M+ GY ++ GA +F G D ++ SMI G+ R G+ E AR ++
Sbjct: 188 VVSYNSMVAGYVAEGDLAGARNLF---DGMARRDVVTWNSMISGYSRHGDMENARKMFDA 244
Query: 63 L 63
+
Sbjct: 245 M 245
Score = 35.8 bits (81), Expect = 0.29
Identities = 20/62 (32%), Positives = 35/62 (56%), Gaps = 3/62 (4%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
+++ T++T Y K M+ A +F ++G + P S+ SMI G+G G EKA + E
Sbjct: 354 VLLLTTLLTMYAKCGVMETAREIFNSMGEKSVP---SWNSMIIGYGLHGQSEKALELFLE 410
Query: 63 LK 64
++
Sbjct: 411 ME 412
Score = 30.8 bits (68), Expect = 9.4
Identities = 16/61 (26%), Positives = 30/61 (48%), Gaps = 2/61 (3%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVF--LTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYY 60
IV +N ++ Y K + G+F + G P+E ++ S++ G+ EK RW +
Sbjct: 281 IVSWNVILALYAKLRDWRECLGLFDVMIAEGNTVPNEKTFVSVLTACANLGDLEKGRWVH 340
Query: 61 E 61
+
Sbjct: 341 D 341
>UniRef100_Q8VZE4 AT4g01570/T15B16_21 [Arabidopsis thaliana]
Length = 805
Score = 42.4 bits (98), Expect = 0.003
Identities = 24/62 (38%), Positives = 36/62 (57%), Gaps = 1/62 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
IV+YNT+I GKA+ +D A +F + I PD SY +MIE +AG ++A Y +
Sbjct: 700 IVMYNTLINALGKATRLDEATQLFDHMKSNGINPDVVSYNTMIEVNSKAGKLKEAYKYLK 759
Query: 62 EL 63
+
Sbjct: 760 AM 761
>UniRef100_Q9M124 Hypothetical protein AT4g01570 [Arabidopsis thaliana]
Length = 454
Score = 42.4 bits (98), Expect = 0.003
Identities = 24/62 (38%), Positives = 36/62 (57%), Gaps = 1/62 (1%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
IV+YNT+I GKA+ +D A +F + I PD SY +MIE +AG ++A Y +
Sbjct: 246 IVMYNTLINALGKATRLDEATQLFDHMKSNGINPDVVSYNTMIEVNSKAGKLKEAYKYLK 305
Query: 62 EL 63
+
Sbjct: 306 AM 307
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.137 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 112,926,584
Number of Sequences: 2790947
Number of extensions: 3451186
Number of successful extensions: 10064
Number of sequences better than 10.0: 391
Number of HSP's better than 10.0 without gapping: 59
Number of HSP's successfully gapped in prelim test: 333
Number of HSP's that attempted gapping in prelim test: 8885
Number of HSP's gapped (non-prelim): 1388
length of query: 64
length of database: 848,049,833
effective HSP length: 40
effective length of query: 24
effective length of database: 736,411,953
effective search space: 17673886872
effective search space used: 17673886872
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC141111.3


