

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140916.4 - phase: 0
(250 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
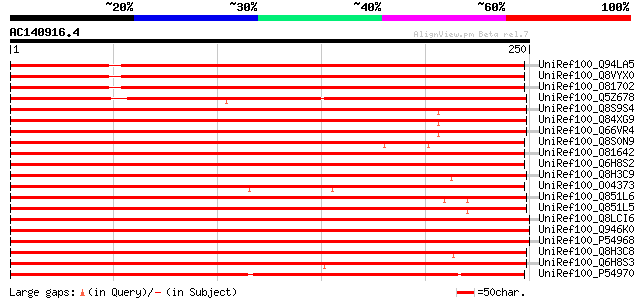
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q94LA5 IAA-amino acid hydrolase, putative [Arabidopsis... 345 5e-94
UniRef100_Q8VYX0 IAA-amino acid conjugate hydrolase-like protein... 345 5e-94
UniRef100_O81702 Gr1-protein [Arabidopsis thaliana] 345 5e-94
UniRef100_Q5Z678 Putative IAA-amino acid hydrolase [Oryza sativa] 316 4e-85
UniRef100_Q8S9S4 Putative IAA-Ala hydrolase [Oryza sativa] 278 1e-73
UniRef100_Q84XG9 IAA-amino acid hydrolase [Oryza sativa] 276 4e-73
UniRef100_Q66VR4 Auxin amidohydrolase [Triticum aestivum] 271 1e-71
UniRef100_Q8S0N9 Putative IAA-Ala hydrolase [Oryza sativa] 269 6e-71
UniRef100_O81642 IAA-Ala hydrolase [Arabidopsis thaliana] 266 3e-70
UniRef100_Q6H8S2 Putative auxin-amidohydrolase precursor [Populu... 266 4e-70
UniRef100_Q8H3C9 Putative IAA amidohydrolase [Oryza sativa] 257 2e-67
UniRef100_O04373 JR3 protein [Arabidopsis thaliana] 254 1e-66
UniRef100_Q851L6 Putative IAA amidohydrolase [Oryza sativa] 248 1e-64
UniRef100_Q851L5 Putative amidohydrolase [Oryza sativa] 247 2e-64
UniRef100_Q8LCI6 IAA-amino acid hydrolase [Arabidopsis thaliana] 244 2e-63
UniRef100_Q946K0 IAA amidohydrolase [Arabidopsis suecica] 243 3e-63
UniRef100_P54968 IAA-amino acid hydrolase 1 [Arabidopsis thaliana] 243 3e-63
UniRef100_Q8H3C8 Putative IAA amidohydrolase [Oryza sativa] 240 2e-62
UniRef100_Q6H8S3 Putative auxin-amidohydrolase precursor [Populu... 238 1e-61
UniRef100_P54970 IAA-amino acid hydrolase homolog 2 precursor [A... 237 2e-61
>UniRef100_Q94LA5 IAA-amino acid hydrolase, putative [Arabidopsis thaliana]
Length = 464
Score = 345 bits (886), Expect = 5e-94
Identities = 166/248 (66%), Positives = 209/248 (83%), Gaps = 5/248 (2%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
MI+DGAL+DVEAIFAVHVSH HPTG+IGSR GPLLAGCG FRAVI+ + + R +A+
Sbjct: 219 MIEDGALDDVEAIFAVHVSHIHPTGVIGSRSGPLLAGCGIFRAVITSE-----DSRGAAN 273
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
+LAAS+AVIS+QGIVSRE++PLDSQVVSVTSF+GG+S D+ PD+VV+GGTFRAFSN+SF
Sbjct: 274 LLLAASSAVISLQGIVSREASPLDSQVVSVTSFDGGHSLDVAPDTVVLGGTFRAFSNSSF 333
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
Y L +RI++V++ Q V+ C A V+FFEK+ IYPPT N+D Y H+KKV+IDLLG +F
Sbjct: 334 YYLKKRIQEVLMDQVGVFGCQATVNFFEKQNAIYPPTTNNDATYNHLKKVTIDLLGDSHF 393
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
+ P MMGAED++FYS++IP+AF++IGIRNE LGS H HSPHF IDED+LP+GAAVHA
Sbjct: 394 TLAPQMMGAEDFAFYSEIIPAAFYFIGIRNEELGSVHIAHSPHFMIDEDSLPVGAAVHAA 453
Query: 241 IAERYLNE 248
+AERYLN+
Sbjct: 454 VAERYLND 461
>UniRef100_Q8VYX0 IAA-amino acid conjugate hydrolase-like protein [Arabidopsis
thaliana]
Length = 441
Score = 345 bits (886), Expect = 5e-94
Identities = 166/248 (66%), Positives = 209/248 (83%), Gaps = 5/248 (2%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
MI+DGAL+DVEAIFAVHVSH HPTG+IGSR GPLLAGCG FRAVI+ + + R +A+
Sbjct: 196 MIEDGALDDVEAIFAVHVSHIHPTGVIGSRSGPLLAGCGIFRAVITSE-----DSRGAAN 250
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
+LAAS+AVIS+QGIVSRE++PLDSQVVSVTSF+GG+S D+ PD+VV+GGTFRAFSN+SF
Sbjct: 251 LLLAASSAVISLQGIVSREASPLDSQVVSVTSFDGGHSLDVAPDTVVLGGTFRAFSNSSF 310
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
Y L +RI++V++ Q V+ C A V+FFEK+ IYPPT N+D Y H+KKV+IDLLG +F
Sbjct: 311 YYLKKRIQEVLMDQVGVFGCQATVNFFEKQNAIYPPTTNNDATYNHLKKVTIDLLGDSHF 370
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
+ P MMGAED++FYS++IP+AF++IGIRNE LGS H HSPHF IDED+LP+GAAVHA
Sbjct: 371 TLAPQMMGAEDFAFYSEIIPAAFYFIGIRNEELGSVHIAHSPHFMIDEDSLPVGAAVHAA 430
Query: 241 IAERYLNE 248
+AERYLN+
Sbjct: 431 VAERYLND 438
>UniRef100_O81702 Gr1-protein [Arabidopsis thaliana]
Length = 464
Score = 345 bits (886), Expect = 5e-94
Identities = 166/248 (66%), Positives = 209/248 (83%), Gaps = 5/248 (2%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
MI+DGAL+DVEAIFAVHVSH HPTG+IGSR GPLLAGCG FRAVI+ + + R +A+
Sbjct: 219 MIEDGALDDVEAIFAVHVSHIHPTGVIGSRSGPLLAGCGIFRAVITSE-----DSRGAAN 273
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
+LAAS+AVIS+QGIVSRE++PLDSQVVSVTSF+GG+S D+ PD+VV+GGTFRAFSN+SF
Sbjct: 274 LLLAASSAVISLQGIVSREASPLDSQVVSVTSFDGGHSLDVAPDTVVLGGTFRAFSNSSF 333
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
Y L +RI++V++ Q V+ C A V+FFEK+ IYPPT N+D Y H+KKV+IDLLG +F
Sbjct: 334 YYLKKRIQEVLMDQVGVFGCQATVNFFEKQNAIYPPTTNNDATYNHLKKVTIDLLGDSHF 393
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
+ P MMGAED++FYS++IP+AF++IGIRNE LGS H HSPHF IDED+LP+GAAVHA
Sbjct: 394 TLAPQMMGAEDFAFYSEIIPAAFYFIGIRNEELGSVHIAHSPHFMIDEDSLPVGAAVHAA 453
Query: 241 IAERYLNE 248
+AERYLN+
Sbjct: 454 VAERYLND 461
>UniRef100_Q5Z678 Putative IAA-amino acid hydrolase [Oryza sativa]
Length = 510
Score = 316 bits (809), Expect = 4e-85
Identities = 163/257 (63%), Positives = 192/257 (74%), Gaps = 16/257 (6%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
MI+ GALEDVEAIFAVHVSH+HPT +IGSR GPLLAGCGFF+AVI G R S D
Sbjct: 242 MIEGGALEDVEAIFAVHVSHQHPTSVIGSRTGPLLAGCGFFKAVIHGGR-------RSGD 294
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIP--------DSVVIGGTF 112
VLAA++ +IS+Q IVSRE++PLDSQVVSV NG + + V+GGTF
Sbjct: 295 AVLAAASTIISLQSIVSREADPLDSQVVSVAMVNGSDHPAATARAAAAEEEEEFVLGGTF 354
Query: 113 RAFSNTSFYQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSI 172
RAFSN SFYQ+ RIE+VI QA V+ C A VDFFE + + YPPTVND +MY HVK V+
Sbjct: 355 RAFSNASFYQVRRRIEEVITAQARVHGCEAAVDFFENQ-SFYPPTVNDARMYAHVKAVAG 413
Query: 173 DLLGQKNFRVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALP 232
+LLG ++R VPPMMGAED+SFYSQV+P+ F+YIG+RNETLGS HTGHSP+F IDED LP
Sbjct: 414 ELLGAGSYRDVPPMMGAEDFSFYSQVVPAGFYYIGVRNETLGSVHTGHSPYFMIDEDVLP 473
Query: 233 IGAAVHATIAERYLNEH 249
GAA HA IAERYL H
Sbjct: 474 TGAAFHAAIAERYLANH 490
>UniRef100_Q8S9S4 Putative IAA-Ala hydrolase [Oryza sativa]
Length = 442
Score = 278 bits (710), Expect = 1e-73
Identities = 137/250 (54%), Positives = 185/250 (73%), Gaps = 1/250 (0%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
MI DGA+E++EAIF VHV+ P G++ SRPGP++AG GFF AVISGK AA P ++ D
Sbjct: 179 MIDDGAVENIEAIFGVHVADVVPIGVVASRPGPVMAGSGFFEAVISGKGGHAALPHHTID 238
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
P+LAAS ++S+Q +VSRE++PLDSQVV+V F GG + ++IPDSV IGGTFRAF SF
Sbjct: 239 PILAASNVIVSLQQLVSREADPLDSQVVTVGKFQGGGAFNVIPDSVTIGGTFRAFLKESF 298
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
QL +RIE+VIV QASV C A VDF +K+ +PPT+N +++ KV+ +++G KN
Sbjct: 299 NQLKQRIEEVIVSQASVQRCNAVVDFLDKDRPFFPPTINSAGLHDFFVKVASEMVGPKNV 358
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFY-IGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHA 239
R P+MGAED++FY+ IP+ ++Y +G+ NET G HSP+FTI+EDALP GAA+ A
Sbjct: 359 RDKQPLMGAEDFAFYADAIPATYYYFLGMYNETRGPQAPHHSPYFTINEDALPYGAALQA 418
Query: 240 TIAERYLNEH 249
++A RYL EH
Sbjct: 419 SLAARYLLEH 428
>UniRef100_Q84XG9 IAA-amino acid hydrolase [Oryza sativa]
Length = 442
Score = 276 bits (706), Expect = 4e-73
Identities = 136/250 (54%), Positives = 184/250 (73%), Gaps = 1/250 (0%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
MI DG +E++EAIF VHV+ P G++ SRPGP++AG GFF AVISGK AA P ++ D
Sbjct: 179 MIDDGTVENIEAIFGVHVADVVPIGVVASRPGPVMAGSGFFEAVISGKGGHAALPHHTID 238
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
P+LAAS ++S+Q +VSRE++PLDSQVV+V F GG + ++IPDSV IGGTFRAF SF
Sbjct: 239 PILAASNVIVSLQQLVSREADPLDSQVVTVGKFQGGGAFNVIPDSVTIGGTFRAFLKESF 298
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
QL +RIE+VIV QASV C A VDF +K+ +PPT+N +++ KV+ +++G KN
Sbjct: 299 NQLKQRIEEVIVSQASVQRCNAVVDFLDKDRPFFPPTINSAGLHDFFVKVASEMVGPKNV 358
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFY-IGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHA 239
R P+MGAED++FY+ IP+ ++Y +G+ NET G HSP+FTI+EDALP GAA+ A
Sbjct: 359 RDKQPLMGAEDFAFYADAIPATYYYFLGMYNETRGPQAPHHSPYFTINEDALPYGAALQA 418
Query: 240 TIAERYLNEH 249
++A RYL EH
Sbjct: 419 SLATRYLLEH 428
>UniRef100_Q66VR4 Auxin amidohydrolase [Triticum aestivum]
Length = 437
Score = 271 bits (693), Expect = 1e-71
Identities = 130/250 (52%), Positives = 183/250 (73%), Gaps = 1/250 (0%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
M++ GA+ ++E +F +HV+ P G++ SRPGP++AG GFF AVISGK AA P ++ D
Sbjct: 174 MVEAGAVVNIEIMFGLHVADSVPIGVLASRPGPIMAGSGFFEAVISGKGGHAALPHHTID 233
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
P+LAAS ++S+Q +VSRE++PLDSQVV+V F GG + ++IPDSV IGGTFRAF SF
Sbjct: 234 PILAASNVIVSLQQLVSREADPLDSQVVTVGKFQGGGAFNVIPDSVTIGGTFRAFMKESF 293
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
QL +RIE+VIV QASV C A VDF +K+ +PPT+N+ ++++ KV +++G N
Sbjct: 294 NQLKQRIEEVIVTQASVQRCSAVVDFLDKDKPFFPPTINNPELHDFFAKVCSEMVGPNNV 353
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFY-IGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHA 239
R P+MGAED+SFY++ +P ++Y +G+ NET G HSP+FTI+EDALP GAA+ A
Sbjct: 354 REKQPLMGAEDFSFYTEAVPKTYYYFVGMLNETRGPQAPHHSPYFTINEDALPYGAAMQA 413
Query: 240 TIAERYLNEH 249
++A RYL EH
Sbjct: 414 SLAARYLLEH 423
>UniRef100_Q8S0N9 Putative IAA-Ala hydrolase [Oryza sativa]
Length = 448
Score = 269 bits (687), Expect = 6e-71
Identities = 130/251 (51%), Positives = 183/251 (72%), Gaps = 3/251 (1%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
M++ GA+++VEAIF HVS E PTG++GSRPGPLLAGCGFF AVI+GK AA+P S D
Sbjct: 185 MVEAGAVDNVEAIFGFHVSVELPTGVVGSRPGPLLAGCGFFEAVITGKGGHAAHPHASVD 244
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
P+LAAS V+++QG+VSRE++PL++QVV+VT F G++ ++IP+S+ IGGTFR FSN F
Sbjct: 245 PILAASTVVLALQGLVSREADPLEAQVVTVTRFLAGDALNVIPESITIGGTFRVFSNEGF 304
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKN- 179
+L RIE+VIV Q++VY C A VDF + PPT+N ++ H + V+ + LG
Sbjct: 305 LRLKRRIEEVIVAQSAVYRCAAAVDFHAGGRPLLPPTINSAALHAHFQAVAAETLGASAA 364
Query: 180 -FRVVPPMMGAEDYSFYSQVIP-SAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAV 237
+ P MG+ED++ +S+ +P S F+++G+RNE G H HSPHF +D+ ALP GAA+
Sbjct: 365 VLGAMEPCMGSEDFAVFSEAVPASHFYFVGVRNEAEGLVHLAHSPHFRVDDAALPYGAAL 424
Query: 238 HATIAERYLNE 248
HA++A RYL+E
Sbjct: 425 HASLAMRYLDE 435
>UniRef100_O81642 IAA-Ala hydrolase [Arabidopsis thaliana]
Length = 440
Score = 266 bits (681), Expect = 3e-70
Identities = 129/248 (52%), Positives = 182/248 (73%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
+++ G LE+V AIF +HV+++ G + SR GP+LAG GFF+A ISGK AA P+++ D
Sbjct: 178 IVEAGVLENVSAIFGLHVTNQLALGQVSSREGPMLAGSGFFKAKISGKGGHAALPQHTID 237
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
P+LAAS ++S+Q +VSRE++PLDSQVV+V F GG + ++IPDSV IGGTFRAFS SF
Sbjct: 238 PILAASNVIVSLQHLVSREADPLDSQVVTVAKFEGGGAFNVIPDSVTIGGTFRAFSTKSF 297
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
QL +RIEQVI +QASV C A VDF E+E +PPTVND +++ K VS D+LG +N+
Sbjct: 298 MQLKKRIEQVITRQASVNMCNATVDFIEEEKPFFPPTVNDKALHQFFKNVSGDMLGIENY 357
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
+ P+MG+ED+SFY Q IP F ++G++N+ + HSP+F ++E+ LP GA++HA+
Sbjct: 358 VEMQPLMGSEDFSFYQQAIPGHFSFVGMQNKARSPMASPHSPYFEVNEELLPYGASLHAS 417
Query: 241 IAERYLNE 248
+A RYL E
Sbjct: 418 MATRYLLE 425
>UniRef100_Q6H8S2 Putative auxin-amidohydrolase precursor [Populus alba x Populus
tremula]
Length = 438
Score = 266 bits (680), Expect = 4e-70
Identities = 128/248 (51%), Positives = 180/248 (71%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
MI +GALE+V AIF +HV+++ P G + SR GPLLAG GFF AVISGK AA P++S D
Sbjct: 175 MIDEGALENVNAIFGLHVANKLPIGEVASRHGPLLAGSGFFEAVISGKGGHAAIPQHSID 234
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
P+LAAS ++S+Q +VSRE++PLDSQVV+V F GG + ++IPDSV GGTFRAF SF
Sbjct: 235 PILAASNVIVSLQHLVSREADPLDSQVVTVAKFQGGGAFNVIPDSVTTGGTFRAFLKESF 294
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
QL +RIE+V+ QA+V C A ++ E E +PPT+ND ++++ + V+ D+LG
Sbjct: 295 MQLRQRIEEVVTGQAAVQRCKAVINLLENEKPFFPPTINDKNLHDYFRVVASDVLGIDKV 354
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
+ + P+MG+ED++FY + IP FF++G++NET + HSP+F I+ED LP GAA+HA+
Sbjct: 355 KDMQPLMGSEDFAFYQEKIPGYFFFVGMQNETRKQLQSPHSPYFEINEDVLPYGAALHAS 414
Query: 241 IAERYLNE 248
+A RYL E
Sbjct: 415 LAARYLLE 422
>UniRef100_Q8H3C9 Putative IAA amidohydrolase [Oryza sativa]
Length = 455
Score = 257 bits (656), Expect = 2e-67
Identities = 120/250 (48%), Positives = 176/250 (70%), Gaps = 1/250 (0%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
++Q+G L+DV AIF +HV G + SRPGP LA G F A I+GK AA P N+ D
Sbjct: 200 VLQEGVLDDVSAIFGLHVDPRIQVGTVTSRPGPFLAASGRFLATITGKGGHAAGPHNAVD 259
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
P+L AS+A++S+Q IV+RE++PL++ V+SVT GG+++++IP+SV GGTFR+ ++
Sbjct: 260 PILTASSAIVSLQQIVARETDPLEAAVISVTFMKGGDAYNVIPESVSFGGTFRSLTSEGL 319
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
L +RI++++ A+V+ C A VDF E+E YP TVND+ MY H + V++D+LG+
Sbjct: 320 SYLKKRIKEIVEAHATVHRCTATVDFMEEERIPYPATVNDEGMYRHARAVAVDVLGEDGV 379
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNE-TLGSTHTGHSPHFTIDEDALPIGAAVHA 239
+V P MG+ED++FY+Q P+AFF IG+ NE T+ + HSPHF +DED LP+GAA+HA
Sbjct: 380 KVGTPFMGSEDFAFYAQRFPAAFFMIGVGNETTMRKVYPLHSPHFVVDEDVLPVGAALHA 439
Query: 240 TIAERYLNEH 249
+A YLN+H
Sbjct: 440 AVAMEYLNKH 449
>UniRef100_O04373 JR3 protein [Arabidopsis thaliana]
Length = 444
Score = 254 bits (649), Expect = 1e-66
Identities = 128/252 (50%), Positives = 181/252 (71%), Gaps = 4/252 (1%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
+++ G LE+V AIF +HV+++ G + SR GP+LAG GFF+A ISGK AA P+++ D
Sbjct: 178 IVEAGVLENVSAIFGLHVTNQLALGQVSSREGPILAGSGFFKAKISGKGGHAALPQHTID 237
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRA---FSN 117
P+LAAS ++S+Q +VSRE++PLDSQVV+V F GG + ++IPDSV IGGTFRA FS
Sbjct: 238 PILAASNVIVSLQHLVSREADPLDSQVVTVAKFEGGGAFNVIPDSVTIGGTFRAFSTFST 297
Query: 118 TSFYQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIY-PPTVNDDQMYEHVKKVSIDLLG 176
SF QL +RIEQVI +QASV C A VDF + T + PPTVND +++ K VS D+LG
Sbjct: 298 KSFMQLKKRIEQVITRQASVNMCNATVDFIARGETFFXPPTVNDKALHQFFKNVSGDMLG 357
Query: 177 QKNFRVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAA 236
+N+ + P+MG+ED+SFY Q IP F ++G++N+ + HSP+F ++E+ LP GA+
Sbjct: 358 IENYVEMQPLMGSEDFSFYQQAIPGHFSFVGMQNKARSPMASPHSPYFEVNEELLPYGAS 417
Query: 237 VHATIAERYLNE 248
+HA++A RYL E
Sbjct: 418 LHASMATRYLLE 429
>UniRef100_Q851L6 Putative IAA amidohydrolase [Oryza sativa]
Length = 414
Score = 248 bits (632), Expect = 1e-64
Identities = 126/256 (49%), Positives = 172/256 (66%), Gaps = 7/256 (2%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
++++G L+D + IFAVHV+ + P G++GSRPGP LAG F A I+GK AA P + D
Sbjct: 153 VLKEGVLDDTQTIFAVHVATDLPAGVVGSRPGPFLAGSARFTATITGKGGHAAEPHLAVD 212
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
P++AAS+AV+S+Q IV+RE+NPL VVSVT+ GG + ++IP+SV +GGT R+ +
Sbjct: 213 PIVAASSAVLSLQQIVARETNPLQGAVVSVTTIKGGEAFNVIPESVTLGGTLRSMTTDGL 272
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
L+ RI +VI QA+V C A VDF E + YP TVND+ MY H K V+ +LG+ N
Sbjct: 273 SYLMNRIREVIEGQAAVNRCTAAVDFMEDKLRPYPATVNDEGMYAHAKAVAESMLGEANV 332
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGI---RNETLGSTHTG----HSPHFTIDEDALPI 233
V P MGAED+ FY+Q IP+AFF IG+ N+ G T HSPHF +DE+ALP+
Sbjct: 333 TVSPMCMGAEDFGFYAQRIPAAFFGIGVGSNGNDGGGMAETTKNQLHSPHFVVDEEALPV 392
Query: 234 GAAVHATIAERYLNEH 249
GAA HA +A YLN++
Sbjct: 393 GAAFHAAVAIEYLNKN 408
>UniRef100_Q851L5 Putative amidohydrolase [Oryza sativa]
Length = 417
Score = 247 bits (630), Expect = 2e-64
Identities = 121/254 (47%), Positives = 173/254 (67%), Gaps = 5/254 (1%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
++++GA++DV+ IF +HV P G++ SRPGP LAG F A I+GK AA P ++ D
Sbjct: 158 VLEEGAVDDVQGIFGMHVDAGLPAGVVASRPGPFLAGSARFTATINGKGGHAAAPHHAVD 217
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
P++A S+AV+S+Q IV+RE++PL VVSVT+ GG + ++IP+SV +GGT R+ +
Sbjct: 218 PIVAVSSAVLSLQQIVARETDPLQGAVVSVTTIKGGEAFNVIPESVTLGGTLRSMTTDGM 277
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
L++RI +VI QA+V C A VDF E + YP TVND++MY H K V+ +LG+ N
Sbjct: 278 SYLMKRIREVIEGQAAVNRCTAAVDFMEDKLPPYPATVNDEEMYAHAKAVAESMLGEANV 337
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTG-----HSPHFTIDEDALPIGA 235
++ P MGAED+ FY+Q IP+AFF IG+ N+ G T HSPHF +DE+ALP+GA
Sbjct: 338 KLSPQGMGAEDFGFYAQRIPAAFFGIGVGNDGGGMAETTTKNQLHSPHFVVDEEALPVGA 397
Query: 236 AVHATIAERYLNEH 249
A HA +A YLN++
Sbjct: 398 AFHAAVAIEYLNKN 411
>UniRef100_Q8LCI6 IAA-amino acid hydrolase [Arabidopsis thaliana]
Length = 442
Score = 244 bits (622), Expect = 2e-63
Identities = 121/250 (48%), Positives = 174/250 (69%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
M++D L+D++ I +VHV P+G IGSRPG +LAG G F + G+ + AA P S D
Sbjct: 182 MLKDEILDDLDGILSVHVFPSIPSGGIGSRPGTVLAGAGLFTVTVHGQGSHAATPHFSKD 241
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
PVLAAS+AV+++Q IVSRE +PL++ VV+V GG++ ++IP S GGTFR+ SN
Sbjct: 242 PVLAASSAVVALQQIVSRELDPLEAGVVTVGYIEGGHAQNVIPQSAKFGGTFRSLSNDGL 301
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
+ RI+++ QASVY C AEV+F EK+ +++P ND+ +YEH KKV+ ++G+ NF
Sbjct: 302 LFIQRRIKEISEAQASVYRCKAEVNFEEKKPSLHPVMNNDEGLYEHGKKVAEAMIGKNNF 361
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
P MG ED+SF++Q +A F +GI+NETLG+ HSP+F +DE+ALP+GAA+HA
Sbjct: 362 HDFPVTMGGEDFSFFTQKTKAAIFVLGIKNETLGAGKPLHSPYFFVDEEALPVGAALHAA 421
Query: 241 IAERYLNEHG 250
+A YL+EHG
Sbjct: 422 MAVSYLDEHG 431
>UniRef100_Q946K0 IAA amidohydrolase [Arabidopsis suecica]
Length = 442
Score = 243 bits (621), Expect = 3e-63
Identities = 121/250 (48%), Positives = 174/250 (69%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
M++D L+D++ I +VHV P+G IGSRPG +LAG G F + G+ + AA P S D
Sbjct: 182 MLKDEILDDLDGILSVHVFPSIPSGGIGSRPGTVLAGAGLFTVTVYGQGSHAATPHFSKD 241
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
PVLAAS+AV+++Q IVSRE +PL++ VV+V GG++ ++IP S GGTFR+ SN
Sbjct: 242 PVLAASSAVVALQQIVSRELDPLEAGVVTVGYIEGGHAQNVIPQSAKFGGTFRSLSNDGL 301
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
+ RI+++ QASVY C AEV+F EK+ +++P ND+ +YEH KKV+ ++G+ NF
Sbjct: 302 LFIQRRIKEISEAQASVYRCKAEVNFEEKKPSLHPVMNNDEGLYEHGKKVAEAMIGKNNF 361
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
P MG ED+SF++Q +A F +GI+NETLG+ HSP+F +DE+ALP+GAA+HA
Sbjct: 362 HDFPVTMGGEDFSFFTQKTKAAIFVLGIKNETLGAGKPLHSPYFFVDEEALPVGAALHAA 421
Query: 241 IAERYLNEHG 250
+A YL+EHG
Sbjct: 422 MAVSYLDEHG 431
>UniRef100_P54968 IAA-amino acid hydrolase 1 [Arabidopsis thaliana]
Length = 442
Score = 243 bits (621), Expect = 3e-63
Identities = 120/250 (48%), Positives = 174/250 (69%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
M++D L+D++ I +VHV P+G IGSRPG +LAG G F + G+ + AA P S D
Sbjct: 182 MLKDEILDDLDGILSVHVFPSIPSGGIGSRPGTVLAGAGLFTVTVHGQGSHAATPHFSKD 241
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
PVLAAS+AV+++Q IVSRE +PL++ VV+V GG++ ++IP S GGTFR+ SN
Sbjct: 242 PVLAASSAVVALQQIVSRELDPLEAGVVTVGYIEGGHAQNVIPQSAKFGGTFRSLSNDGL 301
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
+ RI+++ QASVY C AEV+F EK+ +++P ND+ +YEH KKV+ ++G+ NF
Sbjct: 302 LFIQRRIKEISEAQASVYRCKAEVNFEEKKPSLHPVMNNDEGLYEHGKKVAEAMIGKNNF 361
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
P MG ED+SF++Q +A F +G++NETLG+ HSP+F +DE+ALP+GAA+HA
Sbjct: 362 HDFPVTMGGEDFSFFTQKTKAAIFVLGVKNETLGAGKPLHSPYFFVDEEALPVGAALHAA 421
Query: 241 IAERYLNEHG 250
+A YL+EHG
Sbjct: 422 MAVSYLDEHG 431
>UniRef100_Q8H3C8 Putative IAA amidohydrolase [Oryza sativa]
Length = 444
Score = 240 bits (613), Expect = 2e-62
Identities = 116/253 (45%), Positives = 168/253 (65%), Gaps = 4/253 (1%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
+++ G L+DV IF +HV P G++ SRPGP ++ F A +GK A P ++ D
Sbjct: 189 VLESGLLDDVSVIFGLHVIPNLPVGVVASRPGPFMSAAARFAATFTGKGGHAGVPHDAVD 248
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
PV+A S+AV+S+Q +VSRE++PL++ VVS+T GG+++++IP+S +GGTFR+ ++
Sbjct: 249 PVVAVSSAVLSLQQLVSRETDPLEAAVVSITILKGGDAYNVIPESASLGGTFRSMTDEGL 308
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
L++RI ++I QA V C A VDF E+E YP TVNDD MY H K V+ +LG+ N
Sbjct: 309 AYLMKRIREIIEAQAGVNRCAAAVDFLEEELRPYPATVNDDGMYGHAKAVAEAMLGEANV 368
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNET----LGSTHTGHSPHFTIDEDALPIGAA 236
RV MG ED++FY++ P AFF+IG+ NET + HSPHF +DE ALP+GAA
Sbjct: 369 RVAARSMGGEDFAFYARRSPGAFFFIGVGNETTMGPAAAVRPVHSPHFVLDERALPVGAA 428
Query: 237 VHATIAERYLNEH 249
+HA +A YLN+H
Sbjct: 429 LHAAVAIEYLNKH 441
>UniRef100_Q6H8S3 Putative auxin-amidohydrolase precursor [Populus alba x Populus
tremula]
Length = 432
Score = 238 bits (607), Expect = 1e-61
Identities = 118/250 (47%), Positives = 170/250 (67%), Gaps = 1/250 (0%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
MI+DGAL D EA+F +HV+++ PTG I S GP+ A F I GK AA P N+ D
Sbjct: 177 MIKDGALGDAEAVFGMHVNYKIPTGTIASLSGPVFAAASHFHVKIEGKGGHAAVPHNAVD 236
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
P+LAAS A++++Q ++SRE +PL SQV+S+T GG + ++IP GGT R+ + S
Sbjct: 237 PLLAASFAILALQLLISRELDPLQSQVLSITYVRGGTTLNVIPPYFEFGGTLRSLTTESL 296
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKE-YTIYPPTVNDDQMYEHVKKVSIDLLGQKN 179
+QL R+++V+ QA+V+ C A VD +EKE +YP TVND+++ HV++VS L ++
Sbjct: 297 HQLQRRLKEVVEGQAAVHRCHAHVDMYEKEDVPLYPATVNDEKLNLHVERVSRLLFNPED 356
Query: 180 FRVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHA 239
F++ +M AED+SFY +VIP IGIRNE +G+ H+ HSP+F +DED L IGA++H
Sbjct: 357 FKMGQKVMAAEDFSFYQEVIPGVMLDIGIRNENVGAIHSLHSPYFFLDEDVLSIGASLHT 416
Query: 240 TIAERYLNEH 249
+AE YLNEH
Sbjct: 417 ALAEIYLNEH 426
>UniRef100_P54970 IAA-amino acid hydrolase homolog 2 precursor [Arabidopsis thaliana]
Length = 439
Score = 237 bits (604), Expect = 2e-61
Identities = 122/248 (49%), Positives = 176/248 (70%), Gaps = 3/248 (1%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
M ++GAL++VEAIF +H+S P G SR G LAG G F AVI+GK AA P+++ D
Sbjct: 181 MREEGALKNVEAIFGIHLSARIPFGKAASRAGSFLAGAGVFEAVITGKGGHAAIPQHTID 240
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
PV+AAS+ V+S+Q +VSRE++PLDS+VV+V+ NGGN+ ++IPDS+ IGGT RAF T F
Sbjct: 241 PVVAASSIVLSLQQLVSRETDPLDSKVVTVSKVNGGNAFNVIPDSITIGGTLRAF--TGF 298
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
QL +R+++VI +QA+V+ C A V+ PPTVN+ +Y+ KKV DLLGQ+ F
Sbjct: 299 TQLQQRVKEVITKQAAVHRCNASVNLTPNGREPMPPTVNNKDLYKQFKKVVRDLLGQEAF 358
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
P+MG+ED+S++++ IP F +G+++ET G + HSP + I+ED LP GAA+HA+
Sbjct: 359 VEAAPVMGSEDFSYFAETIPGHFSLLGMQDETNGYA-SSHSPLYRINEDVLPYGAAIHAS 417
Query: 241 IAERYLNE 248
+A +YL E
Sbjct: 418 MAVQYLKE 425
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.135 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 419,588,291
Number of Sequences: 2790947
Number of extensions: 16700238
Number of successful extensions: 37516
Number of sequences better than 10.0: 626
Number of HSP's better than 10.0 without gapping: 519
Number of HSP's successfully gapped in prelim test: 107
Number of HSP's that attempted gapping in prelim test: 36214
Number of HSP's gapped (non-prelim): 633
length of query: 250
length of database: 848,049,833
effective HSP length: 124
effective length of query: 126
effective length of database: 501,972,405
effective search space: 63248523030
effective search space used: 63248523030
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 73 (32.7 bits)
Medicago: description of AC140916.4


